Abstract
Causality has been a burgeoning field of research leading to the point where the literature abounds with different components addressing distinct parts of causality. For researchers, it has been increasingly difficult to discern the assumptions they have to abide by in order to glean sound conclusions from causal concepts or methods. This paper aims to disambiguate the different causal concepts that have emerged in causal inference and causal discovery from observational data by attributing them to different levels of Pearl’s Causal Hierarchy. We will provide the reader with a comprehensive arrangement of assumptions necessary to engage in causal reasoning at the desired level of the hierarchy. Therefore, the assumptions underlying each of these causal concepts will be emphasized and their concomitant graphical components will be examined. We show which assumptions are necessary to bridge the gaps between causal discovery, causal identification and causal inference from a parametric and a non-parametric perspective. Finally, this paper points to further research areas related to the strong assumptions that researchers have glibly adopted to take part in causal discovery, causal identification and causal inference.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
Causality is a field that has percolated multiple research areas such as medical treatment (Shalit 2020), policy-making (Kreif and DiazOrdaz 2019), social science (Sobel and Legare 2014) epidemiology (Halloran and Struchiner 1995) and cybersecurity (Andrew et al. 2022; Dhir et al. 2021). Historically, the fundamental problem of causality, the fact that we cannot observe the outcome under treatment as well as control in a single unit of observation, has long precluded researchers from making causal claims (Holland 1986). Therefore, the earliest methods for drawing causal conclusions from data were the randomized controlled trials (RCTs), where units of analysis were randomly assigned treatment or control, eliminating any confounding relation between assignment and outcome. However, in many cases randomized control trials are unethical or impractical. This has set the stage for causal research with observational data.
While research on causality with observational data is burgeoning, more specific subfields of causality are starting to emerge. Nowadays, the number of different causal concepts is increasing exponentially. The reason for this is twofold. First, different causal concepts correspond to different subtasks of causality. One can be interested in exploring the causalFootnote 1 dynamics by exploiting statistical properties, which is known as causal discovery. Alternatively, one can engage in estimation of an outcome variable under possible alternations, known as causal inference. The latter can also be further differentiated into three levels of increasingly complex queries known as Pearl’s Causal Hierarchy (Bareinboim et al. 2022). Some approaches to the two more complex levels of the hierarchy require the use of different calculi to reduce the query of interest to known quantities that provide a unique solution to the query, which is called causal identification (Pearl 1997). Second, causal concepts are merely ramifications of the assumptions a researcher is willing to adopt. For every departure of putative assumptions, new causal concepts emerge that account for that deviation. For example, in epidemiology patients that take a vaccine might not only protect themselves but also those they come in contact with. Therefore, epidemiologists probably would like to work with causal concepts that account for interference.
The most straightforward way of disentangling causal concepts is by starting to examine what different queries the field of causal inference is able to address. Hence, we start by elucidating the three different levels over which causal questions can be formulated, namely the associational, interventional and counterfactual level corresponding to the action of seeing, doing and imagining respectively (Pearl and Mackenzie 2018). See Table 1 for an overview of the different queries on the hierarchy.
Specifically, we will highlight causal concepts that help addressing corresponding queries at each level of the hierarchy. Because the hierarchy is of increasing complexity, there exists a causal object that can be utilized to address queries at all three levels of the hierarchy, which is the Structural Causal Model (SCM). This object will be formally introduced in Sect. 2. Often, the full specifications of the SCM are empirically unattainable, but one is able to use surrogate causal objects to still address queries at lower levels of the hierarchy. It has been proven by the Causal Hierarchy Theorem (CHT) (Bareinboim et al. 2022) that queries at higher levels of the hierarchy can generally not be addressed with information of lower levels only. Fig. 1 is adopted from Bareinboim et al. (2022) and shows how the different levels of the hierarchy relate to each other. It also outlines the scope of this paper.
In this paper, we try to disentangle different causal concepts based on the query it is supposed to address as well as its assumptions lying underneath. Therefore, we give an overview of reasoning with causality from observational data to (causal) estimands and estimatesFootnote 2 at each level of the hierarchy where we take the flow of causality as illustrated by Fig. 2. At the first two levels of the hierarchy, one starts with observational data and some assumptions based on the data generating process. Applying algorithms that recover the (causal) structure of the variables will generate graphical objects such as causal diagrams. These graphical components can be subjected to additional assumptions that enable identification of a causal query. During the identification process, the causal query is rewritten to contain only the observational information necessary to address the query. Inference is then concerned with the process of identifying the relevant observational information for a specific question of interest. We will unify the non-parametric approach to causal inference with the parametric approach and describe how they have emerged from a different appreciation of the fundamental problem of causality. Specifically, the different assumptions both approaches start from will be highlighted and their meeting point will be identified.
The flow illustrated by Fig. 2 is not binding. Some scholars might solely be interested in the causal structure among the variables while other scholars rely on domain experts to design the graphical structure and focus only on causal identification and inference. This paper does not primarily focus on the different methods and algorithms available to do structure learning, identification and inference, but mainly tries to delineate the different causal concepts as well as their underlying assumptions to engage in structure learning, identification and inference. However, when the assumptions are contingent on the algorithms used, we will include them in the discussion although the algorithms’ inner workings will generally not be emphasized. By illuminating the causal concepts and making explicit concomitant assumptions we hope to encourage non-experimental causal research that social scientists sometimes eschew (Grosz et al. 2020).
Flow from observational data to estimands and estimates at all levels of Pearl’s Causal Hierarchy. The center part contains the input (observational data), the intermediate object (graphical component) and the output (estimands and estimates) of a potential flow of causal reasoning. While the light blue part indicates at what stage assumptions have to be taken into account, the purple part indicates which task is involved at what stage
There have been many survey papers written on causality, but, to the best of our knowledge, none of them have tried to bridge the gap between causal discovery and causal inference from the point of view of necessary assumptions. One of the first causal inference papers that can be considered a survey paper is the work of Holland (1986) summarizing the research of Rubin (1974, 1978) as well as uniting it with philosophical and statistical authors and the well-known Granger Causality (Granger 1969). The subsequent survey that picked up the advances made in non-experimental estimation techniques is of Nichols (2007) followed by a survey paper on the specific matching technique (Stuart 2010). Finally, the most recent paper of causal inference under the potential outcome model as well as its underlying assumptions and various departures from those assumptions has been written by Yao et al. (2021).
As regards surveys on causal discovery, there are general methodological surveys (Glymour et al. 2019; Nogueira et al. 2021, 2022), surveys that focus on continuous optimization (Vowels et al. 2022), constraint-based methods (Yu et al. 2016), time series methods (Assaad et al. 2022) and various aspects related to assumptions and practical use of the methods (Eberhardt 2009, 2016; Malinsky and Danks 2018). For a general survey on causal learning with different sorts of data, we refer the reader to (Guo et al. 2020a).
Central to this paper are the different levels of the hierarchy and therefore we draw the relation between causal inference and Bayesian Network inference. There is much (summarizing) research on discrete Bayesian Network inference (Friedman 2009), while non-parametric (Hanea et al. 2015) and hybrid (Salmerón et al. 2018; Shenoy and West 2011; Langseth et al. 2009) Bayesian Network inference are less developed.
Concretely, this paper is structured as follows. After a brief introduction of preliminaries and Structural Causal Models (SCMs) in Sect. 2, this paper starts with an incipient concept of causality in Sect. 3, the Potential Outcome Framework, and continues to examine the assumptions inherent to this concept. Then, we introduce Bayesian Networks, d-separation and some equivalent Markov assumptions at the associational level of the hierarchy in Sect. 4. We will show how the latter contributes to Bayesian Network inference and highlight the available tools and contingent assumptions available to conduct inference at the first level of the hierarchy in Sect. 4.2. Concepts and assumptions at the interventional level of the hierarchy will be introduced in Sect. 5. In Sect. 5.1 different sets of assumptions that allow non-parametric as well as parametric structure learning are introduced. Subsequent Sect. 5.2 delineates different assumptions and concepts for non-parametric as well as parametric inference approaches while enunciating the meeting point between the two approaches. Some possible deviations from putative assumptions at the interventional level together with relevant references are mentioned in Sect. 5.3. We continue by the introduction of various counterfactual models and inference techniques to reason with causality at the counterfactual level of the hierarchy at Sect. 6. Finally, in Sect. 7 we summarize the results and propose future research directions based on the articulated assumptions.
2 Preliminaries
In this section, we discuss general preliminaries and notation conventions we will follow throughout the paper. Random variables are denoted by capital letters and unless specified otherwise, Y denotes the outcome variable, T the treatment variable and Z possible confounding variables. Generally, \(X=\{X_1,\ldots ,X_n\}\) denotes the set containing random variables \(X_i\) that take values \(x_i\) in corresponding state space \(\Omega _{X_i}\).
A graph is denoted by \(G=(V,E)\) where \(V=\{V_1,\ldots ,V_n\}\) is the set of vertices (or nodes) and E the set of edges. A graph can be directed when every edge has a direction, undirected when no edge has a direction or partially directed when some but not all edges have a direction. A graph can contain a cycle when there exists a directed path from a node to itself. When there is no such path and the graph is directed, we call this a directed acyclic graph (DAG). Edges can also be bidirected. A graph containing only directed and bidirected arrows without directed cycles is called an acyclic directed mixed graph (ADMG)
When there is a directed edge from node T to node Y (or \(T\xrightarrow []{}Y\), in short), we say that T is a parent of Y and Y a child of T. The set of parents of Y is denoted by \(\text {pa}(Y)\) and the set of children of T is denoted by \(\text {ch}(T)\). An ancestor of Y is a node with a directed path to Y, including Y itself. The set of ancestors is denoted by \(\text {an}(Y)\). Similarly, a descendent of Y is a node with a directed path from Y, including Y itself, and the set is denoted by \(\text {de}(Y)\). The set of non-descendants of Y is denoted by \(\text {nonde}(Y)\). Note that due to the exclusion of the node itself, this is not the same as \(\text {an}(Y)\). Finally, we distinguish a couple of different directed graph structures: We call the structure \(T\xrightarrow []{}Z\xrightarrow []{}Y\) a chain, \(T\xleftarrow []{}Z\xrightarrow []{}Y\) a fork and \(T\xrightarrow []{}Z\xleftarrow []{}Y\) a v-structure. In the latter case, Z is called the collider.
A topological sort < is any linear ordering of the nodes for which \(T\xrightarrow []{}Z\) implies \(T<Z\) in the ordering. Note that this can only exist when the corresponding graph is acyclic. A subgraph \(G_i\) is defined to be the graph restricted to the nodes that precede and include \(V_i\) in the topological sort and a mutilated graph \(G_{\overline{W}}\) is the graph for which all arrows to \(W\subseteq V\) are deleted.
When graphs are endowed with probabilistic meaning, the random variables \(X=\{X_1,\ldots ,X_n\}\) will correspond to nodes of the graph \(V=\{V_1,\ldots ,V_n\}\) and therefore V will inherit the probability distributions and state spaces from X (meaning P(V) and \(v_i\) will correspond to P(X) and \(x_i\), respectively). In this case, \(\text {pa}(V_i)\) refers to the random variables that are associated with the parents of \(V_i\). The assignment of random variables \(\text {pa}(V_i)\) is denoted by \(pa_i\), which is an element of state space \(\Omega _{\text {pa}(V_i)}\).
2.1 Structural causal models
The true subject of our investigation to address all levels of the hierarchy is the Structural Causal Model. We start by formally introducing it:
Definition 1
(Structural Causal Models (SCM)) A Structural Causal Model M consists of an ordered set of endogenous variables \(V=\{V_1,\ldots ,V_n\}\), exogenous variables U and a set of functions \(F=\{f_1,\ldots ,f_n\}\) such that:
-
1.
For all \(V_i\in V\), there exist a corresponding subset of exogenous variables \(U_i\subseteq U\) and a mapping \(f_i:\Omega _{\text {pa}(V_i)\cup U_i}\xrightarrow []{}\Omega _{V_i}\) that maps the state space of parents of \(V_i\) together with \(U_i\) to the state space of \(V_i\):
$$\begin{aligned} v_i = f_i(pa_i,u_i). \end{aligned}$$ -
2.
The error terms u are drawn from a probability distribution P(U) over exogenous variables with state space \(\Omega _{U}\).
The SCMs are also known as the Structural Equation Models (SEMs). They can be either parametric or non-parametric. Non-parametric structural equation models are sometimes invoked because assumptions about functional forms between respective exogenous and endogenous variables are costly. It is important to note that the SCM does not assume the independence of exogenous variables.Footnote 3 However, when this additional property is satisfied, the models are known as non-parametric structural equation models with independent errors (NPSEM-ie) as will be illustrated in Sect. 6. Since the NPSEM-ie has been subject to criticism about their implicit assumptions and unification with graphical components (Richardson and Robins 2013b), we will consider the SCM to be the true object of investigation in the rest of the paper. Henceforth we will not assume the independence of errors unless explicitly stated. The SCMs are assumed to be acyclic, also called recursive. Recursiveness allows a topological sort to exist over the endogenous variables.
Frequently, the true SCM is unattainable due to a limited ability to observe a system (Rubenstein et al. 2017), and one has to settle for surrogate models that do not have equal expressive power, but can be sufficient to answer queries of the first two levels of the hierarchy, see Fig. 1. Before introducing these surrogate models in later sections, we will first introduce the core assumptions and some targets of interest via the so-called Potential Outcome Framework in the next section.
3 Potential outcome framework
In this section, the Potential Outcome Framework (or Neyman-Rubin Causal Model) as developed by Rubin (1974) is introduced. The potential outcomes ground the most granular sort of queries of the causal hierarchy, the counterfactual, and the framework incorporates the core assumptions of causal inference. That means that claims about potential outcomes are equivalent to counterfactual claims. Therefore, we will regularly draw comparisons between different levels of the hierarchy and the Potential Outcome Framework. The framework is logically equivalent to the Structural Causal Model framework (Pearl 2009), which was introduced in the previous section. The necessary methods and targets of interest will be defined along with accessory assumptions. For a full picture of these methods and assumptions, we refer the reader to Fig. 3. This section naturally revolves around the concept of potential outcomes.
Methods for inferring causal claims under different assumptions. The ignorability assumption and stable unit-treatment value assumption (SUTVA) are implicit in randomized controlled trials for which we can draw causal claims. When strong ignorability holds together with SUTVA, the adjustment formula should be invoked to calculate causal estimates
3.1 Potential outcomes
Before the potential outcome is introduced, first the treatment will be defined:
Definition 2
(Treatment variable) The treatment variable T is a random variable that takes on different values for treatment t.
Definition 3
(Potential outcome) The potential outcome random variables are denoted by \(Y(T=t)\) (or Y(t) in short) for different treatment values \(T=t\). For a unit of observation i (or unit in short) and treatment value t, we denote the potential outcome realizations \(y_{i}^t\) to be the outcome that would have been observed if unit i had been exposed to treatment t.
Classically, t has been considered to take on binary values corresponding to treatment (1) and control (0) (Rubin 1974). The first target of interest emerges naturally from this definition and is called the unit-level causal effect.
Definition 4
(Unit-level causal effect) Considering binary treatment t, the unit-level causal effect for unit i is defined as \(\tau _i= y_{i}^1 - y_{i}^0\).
The potential outcome of unit i cannot be observed for treatment \(t=1\) and control \(t=0\) in a single observation leading to the fundamental problem of causal inference (Holland 1986). This means that the unit-level causal effect cannot be calculated exactly but only estimated. We call \(y_i^t\) counterfactual when unit i has not been exposed to treatment t but to another treatment value \(t' \ne t\). The unit-level causal effect also has its statistical population counterpart, the average causal effect.
Definition 5
(Average causal effect) Considering binary treatment t, the average causal effect for a population \(i=1,\ldots , n\) is defined as
3.2 Randomized control trials
Randomized controlled trials are widely considered to be the golden standard to retrieve average causal effects. That is because inherent to the randomized controlled trials are three assumptions. The first assumption is called consistency:
Assumption 1
(Consistency) Let \(T = t\) be the treatment assignment. Let Y be the observed outcome. Then, consistency is satisfied if
Informally, the assumption forces one to unambiguously define treatment and tie the potential outcomes to the observed variables. Earliest claims for the use of this assumptions to ’simplify matters’ date back to the seventies (Gibbard and Harper 1978), but have been formalized later by Robins (1986). Despite the fact that consistency can be derived from the definition of potential outcome variables (Malinsky et al. 2019) (which will be discussed in Sect. 6), scholars (VanderWeele 2009) propound the view that consistency is an assumption rather than a definition or axiom. Although this assumption is sometimes known as the no-multiple-treatment assumption, some researchers draw a firm distinction between the two (VanderWeele and Hernan 2013). Consistency can be a strong assumption in the observational setting, but it is implicit in randomized controlled trials, because exposure to treatment is a result of experimental design (Cole and Frangakis 2009).
The second assumption is known as the no-interference assumption (Cox 1958). It explicitly states that a potential outcome of a unit is not dependent on treatment received by other units. More formally,
Assumption 2
(No-interference) Let \(t_i\) be the treatment assignment of unit i for \(i=1,\ldots ,n\). Then no-interference is satisfied if
Interference is also known as spillover. In a randomized controlled trial the investigator can prevent causal spillover by designing the experiment such that different units do not interact.
A combination of both consistency and no-interference leads to the stable unit-treatment value assumption (SUTVA) (Rubin 1980). While interference is hard to restrain in the observational setting, in many causal inference applications the stable unit-treatment value assumption is implicitly adopted. Although a randomized control trial poses limitations on SUTVA violations, the strength of the randomized control trial lies in its implication of the ignorability assumption:
Assumption 3
(Ignorability/exchangeability) Consider binary treatment assignment random variable T and potential outcome under treatment Y(1) and control Y(0). Then, ignorability is satisfied if
where \(\perp \!\!\! \perp _P\) means independence in probability.
In words, the potential outcomes under treatment are independent of treatment assignment. In this case, we can ignore how units ended up in the treatment or control group. Equivalently, the group that received treatment could have been exchanged with the group receiving control resulting in the same potential outcome.
The three assumptions together constitute the randomized controlled trial (as illustrated in Fig. 3) and make calculation of the average causal effect possible by means of reasoning with potential outcomes. Besides the use of potential outcomes, the Potential Outcome Framework contains one additional element that enables one to bypass the fundamental problem of causality beyond randomized controlled trials, which is the assignment mechanism (Imbens and Rubin 2010).
3.3 Beyond randomized control trials
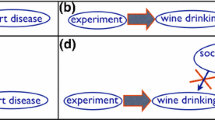
Unlike for randomized controlled trials, the ignorability assumption is easily violated when dealing with observational data, because the treatment and control group are rarely truly exchangeable. A confounder can causally influence the treatment variable as well as the outcome variable as illustrated in Fig. 4. Therefore, more lenient assumptions can be adopted to render the calculation of causal effects under the Potential Outcome Framework still possible in the presence of confounders.
Assumption 4
(Conditional inorability) Let Z denote confounding variables. Consider binary treatment assignment random variable T. Then conditional ignorability is satisfied if
That means that treatment and control group are generally not exchangeable, but they become exchangeable when we condition on the confounding set. For that reason, conditonal ignorability is also known as the unconfoundedness assumption. It is useful to adjust for confounding to reach conditional ignorability as long as the probability of receiving treatment and control remains strictly positive in each of the created subgroups. The positivity assumption guarantees this is the case.
Assumption 5
(Positivity) Let Z denote confounding variables. Then positivity is satisfied if
There is a tradeoff between conditional ignorability and positivity by virtue of adjusting for covariates (D’Amour et al. 2021), which is the process of conditioning on subgroups of the data that share similar covariate values. Intuitively, the more covariates are adjusted for, the smaller the subgroups become. This can lead to subgroups being entirely assigned to either treatment or control, which is a violation of the positivity assumption. Contrary, not sufficiently adjusting for high dimensional covariates may lead to violations of conditional ignorability assumptions. In Sect. 5.2.2 we explain how this problem gives rise to the use of parametric approaches as opposed to non-parametric approaches. Both conditional ignorability and positivity together are called strong ignorability (Rosenbaum and Rubin 1983; Imbens and Rubin 2015).
Vested with all of the above assumptions, one is able to calculate the average causal effect. Assume binary treatment assignment variable T and confounding set Z:
While the first two equalities follow from the laws of probability and expectation, the third equality is a result of conditional ignorability and positivity and the fourth equality a result of consistency. This result is also called the adjustment formula and the underlying assumptions are summarized in Fig. 3. The formula requires one to have insight into the assignment mechanism: the conditional probabilities of treatment given covariates and potential outcomes. This is the second element that constitutes the potential outcome framework.
When conditional ignorability does not apply, causal inference becomes significantly harder. In some cases instrumental variables, those that causally influence the treatment but not the outcome variable, can be utilized (Hartford et al. 2017), the joint distribution of latent and observed confounders can be extracted from variational auto-encoders (Louizos et al. 2017) and network data as a proxy for latent confounders can still be used to substantiate causal effects (Guo et al. 2020b).
Consistency follows from the definitions of the Structural Causal Models and hence the literature rejecting this assumption is not rich (Pearl 2009). SUTVA can easily be violated by departures from the no-interference assumption. Concepts that emerge from this departure at the second level of the hierarchy will be discussed in Sect. 5.3.
4 Associational level of the hierarchy
In this section we introduce concepts and associated assumptions that are necessary to address questions at the first level of Pearl’s Causal Hierarchy, the associational level (see Fig. 1). The chapter starts with some preliminaries on the relation between probability functions and graphical models. We continue with explaining the features of Bayesian Networks (BNs) and introduce Markov Random Fields (MRFs). Because structure learning of Bayesian Networks has much resemblance with causal discovery, we refer to Sect. 5.1 for information about structure learning. Finally, different inference methods are discussed for different types of Bayesian Networks. For an overview of items that are covered in this section, we refer the reader to Fig. 5.
Assumptions and concepts discussed at the associational level of the hierarchy. Probability distributions and graphical models can be tied together by means of the Markov assumptions. The minimality assumption can be adopted optionally for a parsimonious encoding of the joint distribution. The resulting object can either be a Bayesian Network or a Markov Random Field. Because Bayesian Networks can be endowed with causal meaning as well, inference methods for various sorts of Bayesian Networks are discussed
4.1 Bayesian networks
In order to address queries at the first level, we need to tie the random variables to the graphical components introduced. This is only possible when we invoke additional assumptions. Let \(X_1, \ldots , X_n\) be random variables with joint probability distribution \(P(x_1, \ldots , x_n)\). According to the chain rule of probability, this can be factorized as
In Bayesian Networks, the random variables are represented by the nodes of a directed acyclic graph and the probabilistic dependencies are represented by the edges via the local Markov assumption:
Assumption 6
(Local Markov) Let \(P(x_1, \ldots , x_n)\) be the joint probability distribution of random variables \(X_i\) corresponding to nodes \(V_i\in V\) in the directed acyclic graph \(G=(V,E)\). Then the local Markov assumption holds, if for every \(X_i\) the following holds in the graph:
Since the local Markov assumption ties the random variables together with the graphical structure, V is assumed to inherit all the probabilistic properties from X. Henceforth, we will use \(P(v_1, \ldots , v_n)\) instead of \(P(x_1, \ldots , x_n)\) to denote the probability distribution of the random variables. The use of the underscore P to imply independence in probability is not superfluous as there also exists independence in the graph defined by d-separation and denoted by symbol \(\perp \!\!\! \perp _G\).
Definition 6
d-separation A path p between T and Y is d-connected in the directed acyclic graph \(G=(V,E)\) by a set of nodes \(C\subseteq V\backslash \{T,Y\}\) if
-
1.
p does not contain a chain \(\cdots \xrightarrow []{}Z\xrightarrow []{}\cdots\) or fork \(\cdots \xleftarrow []{}Z\xrightarrow []{}\cdots\), where Z is contained in C.
-
2.
all colliders of the path p are in C or have a descendant in C.
If there are no d-connecting paths between T and Y given C, then T and Y are d-separated by C which is denoted by \(T \perp \!\!\! \perp _G Y \mid C\).
The concept of graph independencies gives rise to a reformulation of the local Markov assumption to the global Markov assumption:
Assumption 7
(Global markov) Let \(P(v_1, \ldots , v_n)\) be the joint probability distribution of random variables corresponding to the nodes \(V_i\in V\). Let \(\perp \!\!\! \perp _G\) denote d-separation in the directed acyclic graph \(G=(V,E)\) and \(\perp \!\!\! \perp _P\) independence in distribution. Then the global Markov assumption holds if for all \(T, Y, Z \subseteq V\)
By relating the independencies of the graph to the independencies of the distribution, one can leverage the graphical structure for a parsimonious factorization of the joint probability distribution. This can also be directly assumed.
Assumption 8
(Bayesian network factorization) Let \(P(v_1, \ldots , v_n)\) be the joint probability distribution of random variables corresponding to the nodes \(V_i\in V\) in the directed acyclic graph \(G=(V,E)\). Then the Bayesian Network Factorization assumption holds if we can factorize the distribution according to the corresponding graphical structure:
Example 1
Consider the Bayesian Network displayed by Fig. 4. According to the Bayesian Network Factorization assumption, the joint probability distribution P(Z, Y, T) can be factorized to \(P(Z)P(T\mid Z) P(Y \mid Z, T)\).
It has been shown that the local Markov assumption, the global Markov assumption and the Bayesian Network Factorization are equivalent when positivity is assumed (Koller D, Friedman 2009). A probability distribution P is said to be Markov relative (or Markov in short) to \(G=(V,E)\)Footnote 4 if the MarkovFootnote 5 assumption holds.
While the Markov assumption imposes restrictions on the probability distribution via the graphical structure, an additional assumption is necessary to enforce limitations on the graphical structure by means of the probability distribution dependencies. This assumption comes in various forms of increasing strength: SGS-minimality, P-minimality and faithfulness (Zhang 2013). We discuss P-minimality here (Pearl 2009), but before introducing this assumption, the concept of a preferred graph needs to be introduced:
Definition 7
(Preferred graph) Let P be the set of distributions that is Markov relative to \(G=(V,E)\) and \(G'=(V,E')\). Then \(G'\) is (strictly) preferred to G if the conditional independence relations of G are a (proper) subset of the conditional independence relations of \(G'\).
Assumption 9
(Minimality) Let P be the set of distributions that is Markov relative to \(G=(V,E)\). We assume that minimality is satisfied with respect to G if P is not Markov relative to a strictly preferred graph \(G'=(V,E')\) to G.
Although minimality is a desirable assumption because it allows one to encode the joint distribution in the most parsimonious graphical structure possible, it is not required to answer queries at the first level of the hierarchy.
In concluding this section, it is worth emphasizing that not all independence relations can be encoded by a Bayesian Network, as exemplified by the following counterexample:
Example 2
Let \(X_1, X_2, X_3, X_4\) be random variables. Then there does not exist a Bayesian Network satisfying conditional independence relations \(X_1\perp \!\!\! \perp _P X_2 \mid \{X_3, X_4\}\) and \(X_3\perp \!\!\! \perp _P X_4 \mid \{X_1, X_2\}\).
Therefore, there is another graphical structure that can represent conditional independencies, which is the Markov Random Field (MRF). Markov Random Fields can account for cyclic probability relations and work with potential functions, but they cannot account for directionality. For more information about Markov Random Fields, we refer the reader to the work by Koller D, Friedman (2009). Both Bayesian Networks and Markov Random Fields are probabilistic graphical models as they unify joint probability distributions with graphical structures. Because the lack of directionality excludes Markov Random Fields from causal reasoning at the second and the third level of the hierarchy, only inference on Bayesian Networks will be examined at this stage.
4.2 Bayesian network inference
The emphasis so far has been on the concepts and assumptions necessary to address different kinds of (associational) queries. We will now delve into the identification of the relevant components to obtain answers of interest, known as inference. Since associational queries require estimation, there are no exact solutions to associational queries. However, the inference algorithms that identify the relevant components for these queries can be of exact or approximate nature. Because many queries of interest are NP-hard, we will refer the reader to the appropriate literature for the corresponding exact or approximate algorithms. In this section, the role of the previously introduced material and assumptions is emphasized and their necessity at inference at the first level of the hierarchy is explained.
As illustrated in the previous section, at the first level of the hierarchy, there are two components tied to each other by the Markov assumption: the independencies implied by the graphical structure and the independencies implied by the probability distribution. Let \(P(v_1,\ldots ,v_n)\) be a probability distribution that is Markov to a directed acyclic graph \(G=(V,E)\). It should be noted that such a Bayesian Network is not unique, since a probability distribution can be Markov to multiple Bayesian Networks.
We focus specifically on marginal inference, that is the probability of a random variable \(v_n\) when marginalizing the rest of the variables out:
The Bayesian Network Factorization assumption allows to rewrite this in a more efficient way:
By leveraging the independencies implied by the Bayesian Network, the sums can be evaluated more efficiently, leading to a less expensive way to compute queries of interest. Naturally, efficiency increases as the Bayesian Network becomes more minimal.
Exact methods in discrete Bayesian Networks that exploit the Bayesian Network structure are variable elimination and message passing. When the structure of the Bayesian Network is not sufficient to reach the desired computational results, approximate methods can be used. Among them are sampling methods and variational inference. For a full overview of these various methods, we refer the reader to the work by Koller D, Friedman (2009) and Salmerón et al. (2018).
Inference on hybrid (combination of discrete and continuous) Bayesian Networks is much less developed. Obviously, continuous variables can be effectively discretized such that well-established discrete methods can be used (Beuzen et al. 2018). When the continuous variables are assumed to have conditional Gaussian distribution, other well-established inference methods based on the joint tree methods exist (Koller D, Friedman 2009). However, in this case, the discrete variables cannot be dependent on continuous parents.
Another powerful method for dealing with hybrid variables in Bayesian Networks uses the mixtures of truncated exponentials (MTE) to approximate distributions, because they are closed under marginalization (Langseth et al. 2009). A similar technique approximates mixture of polynomials (MOP) for which closed inference techniques exist (Shenoy and West 2011).
However, some methods do not assume any distribution, such as a method based on importance sampling by Yuan and Druzdzel (2007). There is an entire field of Bayesian Network inference without parametric assumptions. An extensive survey paper on existing methods and applications is written by Hanea et al. (2015). Finally, some best practices for working with Bayesian Network as a modeling technique are described by Chen and Pollino (2012).
5 Interventional level of the hierarchy
This section discusses the causal assumptions and components necessary to address queries at the second level of the hierarchy. We start with the various sets of assumptions necessary to conduct parametric as well as non-parametric causal discovery in Sect. 5.1, specified in Fig. 6. In Sect. 5.2 we show how the output of causal discovery, a causal diagram, forms the basis of a non-parametric approach as well as a parametric approach, where the approaches differ based on a different appreciation of the fundamental problem of causality. The non-parametric approach adopts assumptions inherent to Causal Bayesian Networks that enable inference, while the parametric approach emerges by observing that the fundamental problem of causality requires estimation by definition. Fig. 7 shows the specifications of the different concepts and assumptions necessary for causal inference for each of the two approaches. Finally, we discuss causal concepts that emerge when deviating from putative assumptions in Sect. 5.3.
5.1 Causal discovery
This section will discuss causal discovery from the point of view of necessary assumption expanding on previous assumptive approaches (Eberhardt 2009). Technical details will be discussed when they are contingent on the introduced assumptions, but for a broader account of why causal discovery methods fail in the absence of assumptions, we refer to a survey paper by Runge (2018). Although this section can serve as a blueprint for which method to use when certain assumptions are adopted, a more practical guide about the application of causal discovery methods can be found in the work by Malinsky and Danks (2018). While using interventional data can lead to significant improvements to causal structure learning (Hauser and Bühlmann 2015; Silva 2016), in this survey we restrict ourselves to recovering the structure with observational data alone. Because we consider observational data to be the only source of information at both the first and the second level of the hierarchy, structure learning at the first two levels of the hierarchy collapse (Mahmood 2011). Additionally, in this survey we limit ourselves to static causal discovery methods, which are causal discovery methods that do not account for the passage of time. There is a body of survey papers on causal discovery methods for longitudinal data and the additional assumptions necessary (Assaad et al. 2022; Runge 2018).
An assumption most causal discovery methods revolve around is the i.i.d. assumption.
Assumption 10
(Independent and identically distributed (i.i.d.)) The observational data is independent and identically distributed.
We first discuss structure learning when the causal sufficiency, Markov, faithfulness, acyclicity and i.i.d. assumptions are satisfied. We then move on to causal discovery with violations of the causal sufficiency assumption and subsequently discuss relaxations of the faithfulness assumption. Some of these approaches are summarized in Fig. 6.Footnote 6 However, there are assumption sets that allow conducting causal discovery beyond the assumption sets in Fig. 6. Concepts that emerge when the Markov or the i.i.d. assumptions are violated are discussed in Sect. 5.3.
Because the goal of causal discovery is to recover the graphical structure from observational data, the core assumption within causal discovery should imply features of this underlying structure from the probability distributions (that are learned from the data). The strongest form of that assumption was already touched upon in Sect. 4.1 and is called faithfulness:
Assumption 11
(Faithfulness) Let \(P(v_1, \ldots , v_n)\) be the joint probability distribution of random variables \(V_i \in V\) corresponding to the nodes in the graph \(G=(V,E)\). Let \(\perp \!\!\! \perp _G\) denote d-separation in a graph \(G=(V,E)\) and \(\perp \!\!\! \perp _P\) be the independencies in distribution. Then the probability distribution P is faithful to G if for all \(T, Y, Z \subseteq V\):
A probability distribution can be faithful to a graph that is acyclic. If this is the case then the acyclicity assumption holds in addition to faithfulness. Practitioners that adopt faithfulness are not necessarily expected to have access to the full probability distributions but are equipped with appropriate independence tests to find (conditional) independencies in the data. In order to complete the first collection of assumptions necessary to conduct causal discovery, we highlight the causal sufficiency assumption:
Assumption 12
(Causal sufficiency) A set of variables V is assumed to be causal sufficient if and only if V contains all common causes of two or more variables in V.
When causal sufficiency is assumed, the subject of investigation is the directed acyclic graph that best fits the data generating process of the observational data. As most causal discovery methods do not uniquely determine the entire directed acyclic graph, one additional definition should be introduced:
Definition 8
(Completed partially directed acyclic graph) Directed acyclic graphs that entail the same conditional independencies are said to be in the same Markov Equivalence Class (MEC) for DAGs. The MEC for DAGs is represented by a Completed Partially Directed Acyclic Graph (CPDAG) for which an edge is directed if all directed acyclic graphs in the MEC agree on the direction of the edge and undirected otherwise.
The causal sufficiency, Markov, faithfulness, acyclicity and i.i.d. assumptions make up the first assumption set that allow causal discovery.
Causal discovery assumption sets: the different purple circles represent possible sets of assumptions described in Sects. 5.1.1–5.1.3 under which causal discovery can be conducted. The boxes represent the assumptions necessary for causal discovery, which may have overlap with multiple assumption sets. The color of the boxes indicate the nature of the assumptions: while light blue represents sampling assumptions, ivory blue indicates assumptions on the data generating process and darker blue is used for causal assumptions. (Color figure online)
5.1.1 Causal discovery with causal sufficiency
Vested with this collection of assumptions as illustrated in the top circle of Fig. 6, the structure of the underlying data generating process could be investigated with observational data alone. The first algorithm was the Spirtes, Glymour and Scheines algorithm (SGS) (Spirtes et al. 1990) closely followed by the Peter-Clarke algorithm (PC) (Spirtes and Glymour 1991). Both are constraint-based methods, meaning they aim to exploit the conditional independencies to inform the structure of the graph. This means that they require the use of reliable conditional independence testing methods. The algorithms output the CPDAG based on observational data.
Besides constraint-based methods, there are also score-based methods. Score-based methods employ the same assumptions, take in the same input and generate the same output as constraint-based methods, but work fundamentally differently. The methods start with a specific CPDAG and fit it to the data. The fit is scored based on a scoring system and compared to the score of a slightly different CPDAG. The best fit is kept and the algorithm continues in the same way. In order to restrain the enormous search space they often have a forward and a backward phase. The forward phase keeps adding edges which improves the score the most. When no edges can be added that can improve the score, the backward phase starts removing edges that improve the score the most. If there is no edge that can be removed to improve the score, the algorithm ends (Chickering 2003). Score-based methods require the use of the appropriate score based on the nature of the data.
5.1.2 Causal discovery without causal sufficiency
The causal sufficiency assumption can be relaxed. In this case, we acknowledge that there can be missing common causes in the observational data and the target of interest should be able to account for unobserved confounders. The smallest superclass of DAGs that accounts for the presence of unobserved confounders and is closed under marginalization is a Maximal Ancestral Graph (MAG) (Richardson and Spirtes 2002). Similar to how multiple DAGs can encode the same independence constraints, multiple MAGs can also represent the same conditional independencies. This gives rise to the Partial Ancestral Graph (PAG) that represents the Markov Equivalence Class of MAGs with the same independence constraints.
It is important to note that the existence of unobserved confounding also leads to a slightly modified version of d-separation that represents conditional independencies with respect to the MAG, called m-separation. This leads to natural extensions of the Markov assumption and the faithfulness assumption that go by the semi-Markov assumption and m-faithfulness.
Algorithms that can extract the PAG from observational data such as Fast Causal Inference (FCI) (Spirtes et al. 2000), Greedy Fast Causal Inference (GFCI) (Ogarrio et al. 2016) and Really Fast Causal Inference (RFCI) (Colombo et al. 2012) rely on the i.i.d. assumption, the semi-Markov assumption and the m-faithfulness assumption to an acyclic system as illustrated in the bottom right circle of Fig. 6.
There are two main drawbacks with the algorithms introduced so far. First, either traditional faithfulness or its extension to unobserved confounder models (m-faithfulness) is assumed. Faithfulness is a strong assumption and it can be easy to find examples where faithfulness is violated (Andersen 2013). Second, the output of all introduced algorithms entails a representation of a Markov Equivalence Class. In order to exploit the obtained graphical structure for inference purposes, additional assumptions on the data generating process should be adopted to direct the edges in the graphical structure, which the algorithm could not provide. Both drawbacks can be skirted by assuming restrictions on the data generating process beforehand. This will be discussed in the next section.
5.1.3 Parametric causal discovery and relaxations of faithfulness
In Pearl’s Causal Hierarchy the true subject of investigation is the Structural Causal Model (SCM). Because the true SCM is almost always unattainable, one is forced to settle for a surrogate model for which at least questions of lower levels of the hierarchy can be addressed. However, by taking parametric assumptions on the distribution of the underlying SCM, other assumptions can be bypassed.
These methods are based on Functional Causal Models (FCMs), which are equivalent (Goudet et al. 2019) to earlier introduced SCMs, where one writes the dependent variable as a function of its parents and a noise term. A special case of a FCM is Linear Non-Gaussian Acyclic Model (LiNGAM) and is defined as follows:
Assumption 13
(LiNGAM) A SCM M with an ordered set of endogenous variables \(V=\{V_1,\ldots ,V_n\}\), exogenous variables \(U=\{U_1,\ldots ,U_n\}\) and a set of functions \(F=\{f_1,\ldots ,f_n\}\) is assumed to be a Linear Non-Gaussian Acyclic Model if:
-
1.
Every function \(f_i\) is a linear function of its parents in the topological sort and exogenous variable term \(u_i\):
$$\begin{aligned} v_i = f_i(pa_i,u_i) = \sum _{j:V_j\in \text {pa}(V_i)}b_{ij}v_j+u_i. \end{aligned}$$ -
2.
The error terms \(u_i\) are drawn from exogenous variables \(U_i\in U\), which are continuous, mutually independent and follow a non-Gaussian distribution.
When LiNGAM is assumed, methods exist to fully recover the DAG (Shimizu et al. 2006) based on independent component analysis (ICA-LiNGAM). Faithfulness can be dropped, but causal sufficiency, acyclity and the i.i.d. assumptions should be adopted. The assumption set has been summarized in Fig. 6. Complementary LiNGAM discovery methods were further developed to account for the violation of causal sufficiency (Hoyer et al. 2008). In addition, there are also variants that allow for a violation of the acyclicity assumption (Lacerda et al. 2012).
There are also alternative assumptions (to LiNGAM) on the data generating process that can be used to sideline the faithfulness assumptions and retrieve the full DAG. Some of those assume an additive noise data generating process (Hoyer et al. 2008a; Peters et al. 2014). More general methods assume a post-linear form (Zhang and Hyvärinen 2009), where it has been proven that in all but 5 model specification cases the causal direction is identifiable. Even though faithfulness does not have to be assumed in some cases, less restrictive assumptions do have to be adopted (Peters et al. 2014).
If one is not willing to commit to additional assumptions about the data generating process, but still considers faithfulness too strong of an assumption, one can adopt one of the many weaker versions of faithfulness (Zhang and Spirtes 2015), such as adjacency faithfulness (Spirtes et al. 2000; Ramsey et al. 2017), 2-adjacency faithfulness (Marx et al. 2021) and frugality (Forster et al. 2018) for which causal discovery algorithms exist or could be developed.
5.2 Identification and inference
In this section, we discuss how the concepts from causal discovery can be used for parametric as well as non-parametric inference. While we acknowledge the discussion about to what degree the result of causal discovery can be called ’causal’ (Dawid 2010), in this section we assume that the ADMGs and DAGs convey causal meaning, making them causal diagrams. We first discuss how non-parametric causal inference contributed to causal inference and emphasize its assumptions. Next, we describe what assumptions the parametric approach to causal inference adopts and where both approaches meet. Both approaches can be summarized by Fig. 7.
Causal diagrams are the basis for causal inference. They can be endowed with assumptions from Sect. 3 to allow inferring causal statements under the g-formula. Alternatively, the diagrams can be subjected to non-parametric assumptions to obtain Causal Bayesian Networks, which can be leveraged for inference with the truncated factorization formula
5.2.1 Non-parametric causal inference
In order to be able to infer causal statements, it should be specified how the causal meaning is conveyed on top of the earlier introduced Bayesian Networks. This leads to the definition of Causal Bayesian Networks. We adopt the ’missing link’ definition as described by Bareinboim et al. (2012) among multiple equivalent definitions of Causal Bayesian Networks because its definition intuitively implicates the (SGS-)minimality assumption. We try to dissect the assumptions inherent to the definitions. Central to this notation are (atomic) interventions and therefore we need to introduce the do-operator and the accessory interventional distribution.
Definition 9
(Interventional distribution) Let Y and S be random variables. The interventional distribution \(P(y\mid do(S=s))\) encodes the probability that \(Y=y\) given that S is forced to take value s (denoted by the do-operator \(do(S=s)\), or do(s) in short) with probability 1.
We first look at Bayesian Networks that do not contain latent variables, which we call Markovian.
Markovian causal Bayesian networks
The behavior of the do-operator within a Bayesian Network can be assumed by the modularity assumption:
Assumption 14
(Modularity) Let P be a probability distribution Markov relative to Bayesian Network \(G=(V,E)\) and let \(S\subseteq V\). Then we say an intervention \(do(S=s)\) is modular if:
-
1.
For every \(V_i\in V\backslash S\), where S and \(\text {pa}(V_i)\) are disjoint in G, the interventional distribution by intervening on the parents of \(V_i\) is invariant to other interventions in the graph:
$$\begin{aligned} P(v_i\mid do(S=s), do(\text {pa}(V_i)=pa_i))= P(v_i\mid do(\text {pa}(V_i)=pa_i)). \end{aligned}$$ -
2.
For every \(V_i\in V\), the interventional distribution by intervening on the parents of \(V_i\) yields the same distribution as observing the parents of \(V_i\):
$$\begin{aligned} P(v_i\mid do(S=s), do(\text {pa}(V_i)=pa_i))= P(v_i\mid do(S=s), \text {pa}(V_i)=pa_i). \end{aligned}$$
Modularity specifies how the interventional distributions operate within the context of a Bayesian Network. We can now define a Causal Bayesian Network:
Definition 10
((Markovian) Causal Bayesian Network) Let P be a probability distribution Markov relative to Bayesian Network \(G=(V,E)\). Then \(G=(V,E)\) is said to be a Causal Bayesian Network (CBN) if for all \(S\subseteq V\) and \(V_i\in V\backslash S\):
-
1.
\(P(v_i\mid do(S=s))\) is Markov relative to G.
-
2.
The intervention \(do(S=s)\) is modular.
The assumptions of the interventional distributions implicit in the definition of Causal Bayesian Networks immediately imply (SGS-)minimality in case the conditional probability distributions are strictly positive. In case they are deterministic, there still is good reason to assume (SGS-)minimality (Zhang and Spirtes 2011).
As the Markov assumption implies a factorization of a Bayesian Network, in a similar way the modularity assumption implicit in the Causal Bayesian Networks enforces the truncated factorization for interventional distributions (Bareinboim et al. 2012):
Assumption 15
(Truncated factorization) Let P be a probability distribution Markov relative to Bayesian Network \(G=(V,E)\). Let \(S\subseteq V\) be the set random variables where is intervened upon. Then we assume that the truncated factorization holds if:
and 0 otherwise.
The truncated factorization property implicit in Markovian Causal Bayesian Networks reduces marginal inference in Markovian Causal Bayesian Networks to marginal inference in the mutilated Bayesian Networks. These are the networks that are obtained when removing all the arrows to these nodes where is intervened upon. Inference techniques discussed in the previous section can be used accordingly. Although the truncated factorization property is sometimes known as the g-formula (Perkovic 2020), it will be emphasized in Sect. 5.2.2 that the g-formula is derived from a different appreciation of the fundamental problem of causality as shown in Fig. 7.
Semi-Markovian Causal Bayesian Networks
The concepts and assumptions introduced in this section do naturally extend to the case when the models allow for unobserved confounding variables as is the case in semi-Markovian models. Naturally, the Markov assumption cannot be adopted but is replaced by a semi-Markov assumption. Although the full specifications of the semi-Markovian Causal Bayesian Network have been detailed by Bareinboim et al. (2022), we would like to emphasize that inherent to that definition is an adjusted version of the Markov assumption and Modularity assumption tailor-made to account for the complexities when latent variables are involved.
As we described in Sect. 5.1, the object that emerges when unobserved confounding random variables are at play is an ADMG. Naturally, the Markov assumption as defined above does not hold when unobserved confounders are involved, because the latent confounders cannot be conditioned on. By generalizing d-separation to m-separation, we can extend the Markov assumption to ADMGs (Richardson 2014) resulting in the semi-Markov assumption. Similarly, as in the original Markov assumption, the semi-Markov assumption can also be expressed in terms of m-separation or in terms of truncated factorization of the distribution. It has been shown that both definitions are equivalent (Richardson 2014), but for specifications of the semi-Markov assumption or the associated semi-modularity assumption, we refer the reader to the article by Bareinboim et al. (2022). These assumptions together give rise to the semi-Markovian Causal Bayesian Network
Definition 11
(Semi-Markovian Causal Bayesian Network) Let P be a probability distribution Markov relative to an ADMG \(G=(V,E)\). Then \(G=(V,E)\) is said to be a Causal Bayesian Network if for all \(S\subseteq V\) and \(V_i\in V\backslash S\):
-
1.
\(P(v_i\mid do(S=s))\) is semi-Markov relative to G.
-
2.
The intervention \(do(S=s)\) is semi-modular.
Obviously, the factorization implied by the semi-Markov assumption also leads to a form of truncated factorization of interventional distributions. For a full overview of this factorization and subsequent ways to marginalize out variables, we refer the reader to (the appendix of) Bareinboim et al. (2022). It has been proven that the do-calculus provides a complete toolkit necessary to rewrite interventional distributions to observational distribution and the rules of do-calculus are implied by the assumptions implicit in the definition of the semi-Markovian Bayesian Network (Shpitser and Pearl 2006). Completeness of the do-calculus means that the do-calculus will provide an observational distribution for each interventional distribution if it exists. When the interventional distributions cannot be written in observational terms, the distribution is called unidentifiable. Identification is a necessary condition for both non-parametric and parametric causal inference approaches
5.2.2 Parametric causal inference
Apart from some causal discovery methods, most of the concepts discussed so far are non-parametric concepts. Since potential outcomes by nature imply missing values, the fundamental problem of causality is essentially an estimation problem. That is why substantial contributions to causal inference also involve estimation. We briefly discuss the motivation of parametric causal inference and we then address the parametric counterpart of the truncated factorization (parametric g-formula) based on assumptions introduced in Sect. 3. At the third level of the hierarchy, these concepts will be extended (see Sect. 6).
The following example motivates the use of parametric methods as a result of estimation problems: according to the truncated factorization, the interventional probability \(P(y\mid do(T=t))\) corresponding to the DAG of Fig. 4 can be converted to observation probabilities:
This is also known as the back-door adjustment (Pearl 2009). Although using parametric methods would require additional assumptions on the functional form, there are two main benefits to using parametric approaches. First, when considering continuous treatment variables, the query of interest \(P(y\mid do(T=t))\) might not be available from data for the intervention \(do(T=t)\) of interest. Second, taking into account high-dimensional covariates Z, summing over all the strata z could be intractable. Both estimation problems can be circumvented by assuming the functional form (Hernan and Robins 2020).
When returning to the fundamental problem of causality and the adjustment formula as a result of various assumptions in Sect. 3, calculating the conditional expectation \(\mathbb {E}[Y\mid do(T=t)]\) of Fig. 4 can be reduced to evaluating \(\mathbb {E}_Z[\mathbb {E}[Y\mid T,Z]]\). This would require the evaluation of \(\mathbb {E}[Y\mid T,Z]\) adjusted for the probability P(z). However, a non-parametric evaluation of \(\mathbb {E}[Y\mid T,Z]\) is impossible when Z is high-dimensional. Therefore, one can fit a linear regression model to the data to receive the estimates for \(\mathbb {E}[Y\mid T,Z]\) for each combination of (t, z) and only estimate the P(z) for the z that are present in the data. This is called standardization based on parametric models, or in more general form, the parametric g-formula.
Alternatively, \(\mathbb {E}(Y\mid do(T=t))\) can be further reduced to
meaning we can marginalize out z from the joint probability if we account for the conditional probability \(P(t\mid z)\) for ending up in the treatment group \(T=t\). When Z is high-dimensional, this cannot be completed with non-parametric methods, but parametric model specifications need to be assumed. Logistic regression would be a straightforward choice in case of binary treatment. This is an example of inverse probability weighting (IPW).
Together with g-estimation methods, inverse probability weighting and the parametric g-formula belong to the family of g-methods, a class of methods that allows the computation of the average causal effects under time-varying treatments (Naimi et al. 2016). All these methods rely on the availability of a causal diagram and on assumptions that have been described in Sect. 3. These assumptions include consistency, positivity, (conditional) ignorability and no-interference as illustrated by Fig. 7. The connection between the g-formula and the truncated factorization formula looms large because the latter stems from the non-parametric causality research while the former originates in its parametric counterpart, both being derived from different assumptions.
In a similar way, expressions with the do-operator, such as \(\mathbb {E}(Y\mid do(T=t))\), can be formulated as expressions containing potential outcomes, \(\mathbb {E}(Y(t))\). Nonetheless, identifiable potential outcomes queries cannot always be reduced to observational queries via the do-calculus, as nested counterfactuals require more refined tooling for reduction. In Sect. 6 we explain that some properties of the do-calculus can be extended to account for the reduction of nested counterfactuals to observational queries as well (Malinsky et al. 2019).
5.3 Discovery, identification and inference with more relaxations
There are also many more departures from traditional assumptions in causal discovery and inference that we have omitted so far and will be discussed here. Deviations that we henceforth consider are departures from the no-interference assumption, departures that allow for context-specific independence and departures that consider a different kind of intervention.
All of the discussed causal discovery methods in Sect. 5.1 are based on the i.i.d. assumption as illustrated by Fig. 6. There is also an entire body of work in terms of causal discovery and inference when this assumption is violated (Arbour et al. 2016; Maier et al. 2013b, a; Lauritzen and Richardson 2002; Hudgens and Halloran 2008; Tchetgen and VanderWeele 2012; Ogburn and VanderWeele 2014; Peña 2016; Bhattacharya et al. 2020; Sherman and Shpitser 2018; Aronow and Samii 2017). As has been tenaciously demonstrated, the causal graphs that emerge as a result of causal discovery under interference, depend on the different kinds of causal interference present (Ogburn and VanderWeele 2014). Causal research under interference has been bifurcating.
On the one hand, graphs with violations of the i.i.d. assumption allow directed edges for causal relationships as well as undirected edges for stable symmetric relationships. These can consequently be accounted for by either Laurritzen-Wermuth-Frydenberg chain graphs (Lauritzen 1996; Lauritzen and Richardson 2002; Bhattacharya et al. 2020) or Andersson-Madigan-Perlman chain graphs (Andersson et al. 2001) depending on the Markov property interpreted. Generalization of the former by relaxing causal sufficiency leads to segregated graphs (Shpitser 2015; Sherman and Shpitser 2018). Complete identification and inference methods for segregated graphs with stable symmetric relationships are established (Sherman and Shpitser 2018). Alternatively, an absorption of the Andersson-Madigan-Perlman chain graphs in combination with ADMGs (Richardson and Spirtes 2002) leads to a new family of causal graphs for which causal discovery methods exist for observational and interventional data (Peña 2016).
On the other hand, extending the rules of d-separation to relational d-separation, a criterion for conditional independence in case of relational data, has given rise to an alternative representation, that enables the existence of independencies of relational data, called abstract ground graphs (Maier et al. 2013a). With an extension of the Peter-Clark algorithm, the Relational Causal Discovery (RCD) algorithm (Maier et al. 2013b) makes it possible to extract the true relational causal structure in case of violations of the no-interference assumption. For every perspective, the relational causal model corresponds to an abstract ground graph. Inference is also possible under abstract ground graphs (Arbour et al. 2016).
Because the Markov assumption has occasionally been criticized (Cartwright 1999) and defended (Hausman and Woodward 1999), there have also been attempts to relax the Markov assumption. Claiming that any variable is independent of its non-descendants given its parents excludes the possibility of conditional independence relations that only hold for a subset of realizations of conditioning variables (Duarte and Solus 2021). Relaxing the Markov property to a kind of Markov property that allows for context-specific independence (CSI) relations calls for different causal concepts that can account for this such as Bayesian Multinets (Geiger and Heckerman 1996), conditional probability tables (CPTs) with regularity structure (Boutilier et al. 1996), Staged Trees and Chain Event Graphs (Smith and Anderson 2008) and labeled directed acyclic graphs (LDAGs) (Pensar et al. 2015). Various algorithms for causal discovery exist for Staged Trees (Carli et al. 2020; Leonelli and Varando 2021) as well as for LDAGs (Hyttinen et al. 2018) (with a slightly adapted version of faithfulness). There are also inference methods available when context-specific independence is involved (Tikka et al. 2019).
Besides the atomic or hard interventions discussed in Sect. 5.2, there are also stochastic or soft interventions. These interventions do not force the intervened variable to take on a fixed value, but merely replace the underlying causal mechanism by a known function (Correa and Bareinboim 2020; Eberhardt and Scheines 2007). The do-calculus falls short in converting causal queries with soft interventions or conditional interventions. For that we need a more general calculus, called \(\sigma\)-calculus (Correa and Bareinboim 2020) that can account for stochastic interventions and comes with a concomitant inference algorithm.
6 Counterfactuals
The components introduced in the previous two sections are not sufficient to address queries at the third level of the hierarchy. While the second level represents interventions on conditioning variables, the third level corresponds to interventions on conditioned variables. As mentioned in Sect. 3, the object necessary to reason at all levels of the hierarchy, including the counterfactual level, is the SCM. Next is an example of how a SCM can be utilized to reason at the counterfactual level of the hierarchy when the Causal Bayesian Network falls short:
Example 3
Assume the Linear Gaussian (Markovian) Causal Bayesian Network corresponding to the graph \(X\xrightarrow []{} Y\) with
The intervention distribution \(P(Y\mid do(X=1))\) can be computed via the truncated factorization formula and results in \(\mathcal {N}(2.5,1)\). However, the counterfactual distribution \(P(Y(X=0) \mid X=1, Y=4)\), meaning the probability of Y had X been set to 0 given that \(X=1\) and \(Y=4\), cannot be computed with a Causal Bayesian Network alone. In order to compute this counterfactual query, access to the SCM is required.
Therefore, assume the following structural equations in the SCM:
The evidence of the counterfactual query, \(X=1\) and \(Y=4\), can be used to update the distribution of the exogenous variables in the SCM to \(u_1\sim \delta (1)\) and \(u_2\sim \delta (4)\), with \(\delta ()\) being the Dirac delta measure. Ingesting the intervention \(X=0\) into the updated structural equations leads to a complete evaluation of the counterfactual query: \(P(Y(X=0) \mid X=1, Y=4)=f_2(X=0,u_2)=\delta (4)\).
One of the reasons much research has been dedicated to the first two levels of the hierarchy is that access to the fully specified SCM is considered to be implausible. While the above Linear Gaussian (Markovian) Bayesian Network gives rise to a natural separation between the endogenous and exogenous variables, the interaction between the observed and latent variables is often unknown, rendering access to the fully specified SCM ’hopeless’ (Bareinboim et al. 2022). Despite the inaccessibility of the fully specified SCM, scholars have painstakingly reasoned with counterfactual models, because it plays an essential role in mediation analysis (Robins et al. 2022; Robins and Richardson 2011). Some counterfactual models have antagonized scholars that have argued that the introduced assumptions are not scientific because they lack the possibility of empirical validation (Dawid 2000).
The definition of the one-step-ahead potential outcomes and recursive substitution imply counterfactual consistency and causal irrelevance. Additional independence assumptions need to be adopted to yield a counterfactual model, which can either be a FFRCISTG or a NPSEM-ie. The SWIG unifies these models with graphical approaches and features a factorization and modularity property. Together with the positivity assumption, inference can be conducted via the extended g-formula or the edge g-formula.
In this section we will generalize the Potential Outcome Framework as introduced in Sect. 3, which is equivalent to the Structural Causal Model framework introduced in Sect. 2, shedding new light on the assumptions involved at the third level of the hierarchy (see Fig. 8).Footnote 7 We emphasize the different counterfactual models emerging from assumptions and highlight the inference tools available for each model. Throughout this section, we assume the existence of a topological sort on the random variables.
6.1 One-step-ahead potential outcomes
The very definition of counterfactuals entails the existence of a hypothetical world that may not be empirically verifiable. Therefore, we start by assuming the existence of one-step-ahead potential outcomes.
Assumption 16
(One-step-ahead potential outcomes) Let \(X_1, \ldots , X_n\) be random variables corresponding to nodes \(V_1, \ldots , V_n\). Then for all \(V_i\in V\) and possible assignments of parents \(pa_i\in \Omega _{\text {pa}(V_i)}\), we assume the existence of one-step-ahead potential outcomes \(V_i(\text {pa}(V_i)=pa_i)\).
Note that \(V_i(\text {pa}(V_i)=pa_i)\) corresponds to the notation introduced in the Potential Outcome Framework of Sect. 3. Intuitively, the one-step-ahead potential outcome corresponds to the response \(V_i\) had the parents of \(V_i\) been set to \(pa_i\). This is emphasized as an assumption because the assumed potential outcomes could possibly be counterfactual and therefore presuming the existence of a hypothetical world. Since not all potential outcomes naturally depend on possible assignments of parent nodes in the topological sort, it is necessary to extend the definition of potential outcomes via recursive substitution.
Assumption 17
(Recursive substitution) Let \(X=\{X_1, \ldots , X_n\}\) be random variables corresponding to nodes \(V=\{V_1, \ldots , V_n\}\). Assume the existence of one-step-ahead potential outcomes \(V_i(\text {pa}(V_i)=pa_i)\) for all \(V_i\in V\) and possible assignments of parents \(pa_i\in \Omega _{\text {pa}(V_i)}\). Then for all \(S\subset V\) and \(s\in \Omega _S\) we assume that \(V_i(s)\) can be expressed recursively:
\(V_i(s)\) is thus the potential outcome where the parents of \(V_i\) that are in S had been set to s and variables for which \(V_j\in \text {pa}(V_i)\setminus S\) are set to the values these potential outcomes would have had had S been set to s, denoted by \(V_j(s)\).
Example 4
Assume the topological sort over the random variables Z, T, Y as implied by Fig. 4. Then, we assume the one-step-ahead potential outcome Y(z) is defined recursively as
Expressing potential outcomes recursively brings along desirable properties as illustrated by Fig. 8. First of all, it directly implies the consistency assumption introduced in Sect. 3 (Malinsky et al. 2019). Second, it proves the so-called causal irrelevance: every potential outcome derived from recursive substitution \(V_i(s)\) can be expressed as a unique minimally causal relevant subset of \(W\subseteq S\): \(V_i(s)=V_i(w)\). The reader can find the specifications of a minimally causal relevant subset and the proof in the work of Malinsky et al. (2019). Equivalence between the Structural Causal Model and the Potential Outcome Framework follows from the equivalent representation of the one-step-ahead counterfactual \(V_i(pa_i)\) as the output of the structural equation \(f_i(pa_i, u_i)\) (by letting \(\vec {u_i}=\{V_i(pa_i)\mid pa_i\in \Omega _{\text {pa}(V_i)}\}\) and setting \(f_i(pa_i, u_i)=(\vec {u_i})_{pa_i}=V_i(pa_i)\)).
6.2 Counterfactual models
In addition to consistency and causal irrelevance, independence relations are assumed to reason about counterfactuals. The literature splits along the lines of which independence assumptions exactly to adopt. There is the more conservative finest fully randomized causally interpretable structured tree graph (FFRCISTG) and the more restrictive non-parametric structural equation model with independent errors (NPSEM-ie). We start by introducing the FFRCISTG independencies.
Assumption 18
(FFRCISTGS independencies) Assume one-step-ahead counterfactuals by recursive substitution. Let v be an assignment for random variables V and let \(pa_i\) be the restriction of that assignment to parents variables of \(V_i\). Then for each assignment v, the corresponding one-step-ahead counterfactuals consistent with v are mutually independent:
where \(V_{i}< V_{i+1}\) in the topological sort.
It is important to note that all counterfactual random variables are consistent with each other in the sense that there is no contrary assignment among them. Extra independencies across contradicting assignments are imposed by assuming independencies of the error terms in the non-parametric structural equation models. Formally, the counterfactual random variables that are independent in the NPSEM-ie model are:
Assumption 19
(NPSEM-ie independencies) Assume one-step-ahead counterfactuals by recursive substitution. Then the set of one-step-ahead counterfactuals across possibly contradictory interventions are mutually independent:
where \(V_{i}< V_{i+1}\) by the topological sort.
Because the NPSEM-ie independencies also contain the FFRCISTGS independencies, the NPSEM-ie model is strictly stronger than the FFRCISTGS model. Consistency and causal irrelevance are implicit in the NPSEM-ie as well as the FFRCISTGS model.
Example 5
Assume one-step-ahead potential outcome random variables corresponding to the nodes Z, T, Y respecting the topological sort of Fig. 4. Then, following the FFRCISTGS model, for assignment \(z_1,t\) we have independencies:
In addition to the previous independencies, according to the NPSEM-ie model, other independencies across contradictory assignments \(z_1\) and \(z_2\) are implied, such as:
While DAGs and ADMGs are not expressive enough to account for reasoning with one-step-ahead potential outcomes with either NPSEM-ie independencies or FFRCISTGS independencies, a more refined graphical construction called a Single World Intervention Graph (SWIG) was introduced via a node-splitting operation based on causal irrelevance. The SWIG can encode the independence relations of either the NPSEM-ie or the FFRCISTGS. Similarly to how the Causal Bayesian Networks assume a factorization property of the interventional distributions and modularity property about the nature of interventions, the SWIGs obey properties that specify the behavior of counterfactual distributions. Both the NPSEM-ie model and the FFRCISTGS model together with consistency imply these factorization and modularity properties for SWIGs (Richardson and Robins 2013a) as illustrated by Fig. 8.
6.3 Inference
Inference on the counterfactual level is concerned with the identification of the relevant components necessary to address counterfactual queries. In order to calculate the distribution of counterfactuals under different interventions, we can use the g-formula, which we have introduced in Sect. 5.2.2. This formula can be extended to account for unit-specific interventions and the distribution of that intervention (Young et al. 2014) resulting in the extended g-formula (Robins et al. 2004; Richardson and Robins 2013b):
Proposition 20
(Extended G-formula) Let \(S\subset V\) and V(s) be the one-step-ahead counterfactual defined by recursive substitution. Then given positivity, the joint distribution can be written as:
The formula is equivalent to the factorization and modularity property implicit in SWIGs and has been proven by Richardson and Robins (2013b) and Shpitser et al. (2022). The strength of the formula is that it rewrites counterfactual distributions in terms of observational distributions, but unlike the g-formula, the extended g-formula also accounts for nested counterfactuals by having a term for every \(V_i\in V\). Analogously, the do-calculus can be extended to rewrite nested counterfactuals such as dynamic treatment regimes or path-specific interventions. For that reason, po-calculus has been introduced by Malinsky et al. (2019) as a generalization of the do-calculus resulting from consistency, causal irrelevance and factorization on SWIGs. Although the po-calculus implies the do-calculus for interventional queries (Malinsky et al. 2019), it has been shown that additional identification results consisting of nested counterfactuals follow from the po-calculus (Shpitser et al. 2022).
As there exists a hierarchy of causal queries, there is also a hierarchy of interventions. The most granular form of interventions are node interventions according to the hierarchy of interventions of Shpitser and Tchetgen (2016). Node interventions are a specific form of edge interventions which in turn are a specific form of path interventions. Multiple targets of interest in mediation analysis are defined as edge interventions and for this reason, the extended g-formula has also been extended to the edge g-formula (Shpitser and Tchetgen 2016). While node interventions are associated with the FFRCISTG model and require the extended g-formula for identification, edge interventions correspond to the NPSEM-ie model and require the edge g-formula for identification as shown in Fig. 8.
7 Discussion and future directions
In this paper, we have coalesced the scattered research on causality into a whole by pinning different research areas to a place on Pearl’s Causal Hierarchy. The concepts and contingent assumptions necessary to address different queries of the hierarchy have been emphasized. There remains a body of directions open for future research.
Because Pearl’s Causal Hierarchy rests upon the existence of the Structural Causal Models and recursiveness is often assumed for these models, most of the research in causality has been operating under the assumption of acyclicity. However, cyclical causal relations are assumed to be common in real-world applications such as climatological phenomena (Cox et al. 2000). Therefore research is now emerging that accounts for the existence of feedback loops in Structural Causal Models (Bongers et al. 2021), but more research is necessary.
In order to bring the concepts retrieved from causal discovery to the realm of inference, some sort of assumption about the nature of interventions should be included. As discussed by Dawid (2010), the Bayesian Network that constitutes the independence relations is an entirely different object than the Causal Bayesian Network. Both objects can be related by assuming, among other assumptions, that edges convey some causal meaning. Research could point out how the relation between causal discovery and causal inference can be strengthened even more.
Because multiple agents can be at play at a causal model conducting interventions, it makes sense to use causal models as a basis of strategic interactions and take causal models to the game theoretical realm. There has not been much research in this field yet (Grimbly et al. 2021), apart from a few exceptions (Soto et al. 2020; Maes et al. 2007). Further research can point out how additional assumptions can bolster the connection between game theoretical concepts and causality.
While causality has gained more traction from multiple disciplines, we hope this paper helps promulgate causal concepts across different research areas. We encourage researchers to examine the causal question they aim to address, assess the validity of the assumptions they are abiding by and employ the suitable causal concepts.
Notes
The word ‘causal’ is considered to be ambiguous by some scholars (Dawid 2010). Therefore we will also refer to this process as structure learning.
Estimands are the targets of inference and estimates are targets of estimation. Because restricting the scope to estimands would dismiss (semi)-parametric causal inference methods that contributed to the second and the third level of the hierarchy, we decided to include parametric inference methods only in regard to causality.
In Definition 1 this can be observed from the fact that for \(V_i, V_j \in V\), the corresponding \(U_i\) and \(U_j\) can overlap.
Sometimes a probability distribution is said to be Markov to the Bayesian Network corresponding to the graph \(G=(V,E)\).
The Markov assumption is in specific cases known as the causal Markov assumption. Technically, the assumption is only causal when the concomitant graphical component has causal meaning (which will be introduced in Sect. 5).
The list of assumption sets is not exhaustive as more possible assumption sets will be described that allow conducting causal discovery. Although constraint-based and score-based causal discovery algorithms require the use of appropriate conditional independence tests and scoring methods respectively, these are not mentioned as assumptions because they are algorithm-specific.
The SWIG does not immediately apply the edge g-formula, but the graphical structure of the SWIG can be generalized to allow edge interventions as has been shown by Shpitser and Tchetgen (2016).
References
Andersen H (2013) When to expect violations of causal faithfulness and why it matters. Philos Sci 80(5):672–683. https://doi.org/10.1086/673937
Andersson SA, Madigan D, Perlman MD (2001) Alternative markov properties for chain graphs. Scand J Stat 28(1):33–85. https://doi.org/10.1111/1467-9469.00224
Andrew A, Spillard S, Collyer J, et al (2022) Developing optimal causal cyber-defence agents via cyber security simulation. https://doi.org/10.48550/arXiv.2207.12355
Arbour D, Garant D, Jensen D (2016) Inferring network effects from observational data. In: Proceedings of the 22nd ACM SIGKDD international conference on knowledge discovery and data mining. Association for Computing Machinery, New York, KDD ’16, pp 715–724, https://doi.org/10.1145/2939672.2939791
Aronow P, Samii C (2017) Estimating average causal effects under general interference, with application to a social network experiment. Ann Appl Stat 11(4):1912–1947. https://doi.org/10.1214/16-AOAS1005
Assaad CK, Devijver E, Gaussier E (2022) Survey and evaluation of causal discovery methods for time series. J Artif Intell Res 73:767–819. https://doi.org/10.1613/jair.1.13428
Bareinboim E, Brito C, Pearl J (2012) Local characterizations of causal bayesian networks. In: Graph structures for knowledge representation and reasoning. Springer, Berlin, pp 1–17, https://doi.org/10.1007/978-3-642-29449-5_1
Bareinboim E, Correa JD, Ibeling D et al (2022) On pearl’s hierarchy and the foundations of causal inference. Probab Causal Inference 10(1145/3501714):3501743
Beuzen T, Marshall L, Splinter KD (2018) A comparison of methods for discretizing continuous variables in bayesian networks. Environ Modell Softw 108:61–66. https://doi.org/10.1016/j.envsoft.2018.07.007
Bhattacharya R, Malinsky D, Shpitser I (2020) Causal inference under interference and network uncertainty. In: Adams RP, Gogate V (eds) In: Proceedings of The 35th Uncertainty in Artificial Intelligence Conference, PMLR, pp 1028–1038
Bongers S, Forré P, Peters J et al (2021) Foundations of structural causal models with cycles and latent variables. Ann Stat 49(5):2885–2915
Boutilier C, Friedman N, Goldszmidt M, et al (1996) Context-specific independence in bayesian networks. In: Proceedings of the twelfth international conference on uncertainty in artificial intelligence. Morgan Kaufmann Publishers Inc., San Francisco, UAI’96, pp 115–123
Carli F, Leonelli M, Riccomagno E, et al (2020) The R package stagedtrees for structural learning of stratified staged trees. https://doi.org/10.48550/arXiv.2004.06459
Cartwright N (1999) Causal diversity and the markov condition. Synthese 121(1/2):3–27 (http://www.jstor.org/stable/20118219)
Chen SH, Pollino CA (2012) Good practice in bayesian network modelling. Environ Modell Softw 37:134–145. https://doi.org/10.1016/j.envsoft.2012.03.012
Chickering DM (2003) Optimal structure identification with greedy search. J Mach Learn Res 3:507–554
Cole SR, Frangakis CE (2009) The consistency statement in causal inference: a definition or an assumption? Epidemiology 20(1):3–5. https://doi.org/10.1097/EDE.0b013e31818ef366
Colombo D, Maathuis MH, Kalisch M, et al (2012) Learning high-dimensional directed acyclic graphs with latent and selection variables. Ann Stat pp 294–321. https://doi.org/10.1214/11-AOS940
Correa J, Bareinboim E (2020) A calculus for stochastic interventions: causal effect identification and surrogate experiments. In: Proceedings of the AAAI conference on artificial intelligence 06:10093–10100. https://doi.org/10.1609/aaai.v34i06.6567
Cox DR (1958) Planning of experiments. Wiley, New York
Cox PM, Betts RA, Jones CD et al (2000) Acceleration of global warming due to carbon-cycle feedbacks in a coupled climate model. Nature 408(6809):184–187. https://doi.org/10.1038/35041539
D’Amour A, Ding P, Feller A et al (2021) Overlap in observational studies with high-dimensional covariates. J Econ 221(2):644–654. https://doi.org/10.1016/j.jeconom.2019.10.014
Dawid AP (2000) Causal inference without counterfactuals. J Am Stat Assoc 95(450):407–424. https://doi.org/10.1080/01621459.2000.10474210
Dawid AP (2010) Beware of the dag! In: Proceedings of workshop on Ccausality: objectives and assessment at NIPS 2008, vol 6. PMLR, Whistler, pp 59–86, https://proceedings.mlr.press/v6/dawid10a.html
Dhir N, Hoeltgebaum H, Adams N, et al (2021) Prospective artificial intelligence approaches for active cyber defence. https://doi.org/10.48550/arXiv.2104.09981
Duarte E, Solus L (2021) Representation of context-specific causal models with observational and interventional data. https://doi.org/10.48550/arXiv.2101.09271
Eberhardt F (2009) Introduction to the epistemology of causation. Philos Compass 4(6):913–925. https://doi.org/10.1111/j.1747-9991.2009.00243.x
Eberhardt F (2016) Introduction to the foundations of causal discovery. Int J Data Sci Anal 3:81–91. https://doi.org/10.1007/s41060-016-0038-6
Eberhardt F, Scheines R (2007) Interventions and causal inference. Philos Sci 74(5):981–995. https://doi.org/10.1086/525638
Forster M, Raskutti G, Stern R et al (2018) The frugal inference of causal relations. Br J Philos Sci 69(3):821–848. https://doi.org/10.1093/bjps/axw033
Geiger D, Heckerman D (1996) Knowledge representation and inference in similarity networks and bayesian multinets. Artif Intell 82(1):45–74. https://doi.org/10.1016/0004-3702(95)00014-3
Gibbard A, Harper WL (1978) Counterfactuals and two kinds of expected utility. In: Ifs. Springer, pp 153–190, https://doi.org/10.1007/978-94-009-9117-0_8
Glymour C, Zhang K, Spirtes P (2019) Review of causal discovery methods based on graphical models. Front Genet 10:524. https://doi.org/10.3389/fgene.2019.00524
Goudet O, Kalainathan D, Sebag M, et al (2019) Learning bivariate functional causal models. In: Cause effect pairs in machine learning. Springer, pp 101–153
Granger CW (1969) Investigating causal relations by econometric models and cross-spectral methods. Econometrica: J Econ Soc pp 424–438
Grimbly SJ, Shock J, Pretorius A (2021) Causal multi-agent reinforcement learning: review and open problems. https://doi.org/10.48550/arXiv.2111.06721
Grosz MP, Rohrer JM, Thoemmes F (2020) The taboo against explicit causal inference in nonexperimental psychology. Perspect Psychol Sci 15(5):1243–1255. https://doi.org/10.1177/1745691620921521
Guo R, Cheng L, Li J et al (2020) A survey of learning causality with data: Problems and methods. ACM Comput Surv (CSUR) 53(4):1–37. https://doi.org/10.1145/3397269
Guo R, Li J, Liu H (2020b) Learning individual causal effects from networked observational data. In: Proceedings of the 13th international conference on web search and data mining, pp 232–240, https://doi.org/10.1145/3336191.3371816
Halloran ME, Struchiner CJ (1995) Causal inference in infectious diseases. Epidemiology pp 142–151. https://doi.org/10.1097/00001648-199503000-00010
Hanea A, Napoles OM, Ababei D (2015) Non-parametric bayesian networks: Improving theory and reviewing applications. Reliabil Eng Syst Saf 144:265–284. https://doi.org/10.1016/j.ress.2015.07.027
Hartford J, Lewis G, Leyton-Brown K, et al (2017) Deep iv: a flexible approach for counterfactual prediction. In: International conference on machine learning, PMLR, proceedings of machine learning research, pp 1414–1423
Hauser A, Bühlmann P (2015) Jointly interventional and observational data: estimation of interventional markov equivalence classes of directed acyclic graphs. J R Stat Soc Ser B (Stat Methodol) 77(1):291–318. https://doi.org/10.1111/rssb.12071
Hausman DM, Woodward J (1999) Independence, invariance and the causal markov condition. Br J Philos Sci 50(4):521–583. https://doi.org/10.1093/bjps/50.4.521
Hernan M, Robins J (2020) Causal inference: what if. CRC Press, Boca Raton
Holland PW (1986) Statistics and causal inference. J Am Stat Assoc 81(396):945–960
Hoyer P, Janzing D, Mooij JM, et al (2008a) Nonlinear causal discovery with additive noise models. Adv Neural Inform Process Syst 21
Hoyer PO, Shimizu S, Kerminen AJ et al (2008) Estimation of causal effects using linear non-gaussian causal models with hidden variables. Int J Approx Reason 49(2):362–378. https://doi.org/10.1016/j.ijar.2008.02.006
Hudgens MG, Halloran ME (2008) Toward causal inference with interference. J Am Stat Assoc 103(482):832–842. https://doi.org/10.1198/016214508000000292
Hyttinen A, Pensar J, Kontinen J, et al (2018) Structure learning for bayesian networks over labeled dags. In: Proceedings of the ninth international conference on probabilistic graphical models, proceedings of machine learning research, vol 72. PMLR, pp 133–144
Imbens GW, Rubin DB (2010) Rubin causal model. Palgrave Macmillan UK, London, pp 229–241
Imbens GW, Rubin DB (2015) Causal inference for statistics, social, and biomedical sciences: an introduction. Cambridge University Press, Cambridge https://doi.org/10.1017/CBO9781139025751
Koller D, Friedman N (2009) Probabilistic graphical models: principles and techniques. MIT Press, Cambridge
Kreif N, DiazOrdaz K (2019) Machine learning in policy evaluation: new tools for causal inference. https://doi.org/10.48550/arXiv.1903.00402
Lacerda G, Spirtes PL, Ramsey J, et al (2012) Discovering cyclic causal models by independent components analysis. https://doi.org/10.48550/arXiv.1206.3273
Langseth H, Nielsen TD, Rumí R et al (2009) Inference in hybrid bayesian networks. Reliabil Eng Syst Saf 94(10):1499–1509. https://doi.org/10.1016/j.ress.2009.02.027
Lauritzen SL (1996) Graphical models. Oxford University Press, Oxford
Lauritzen SL, Richardson TS (2002) Chain graph models and their causal interpretations. J R Stat Soc Ser B (Stat Methodol) 64(3):321–348. https://doi.org/10.1111/1467-9868.00340
Leonelli M, Varando G (2021) Context-specific causal discovery for categorical data using staged trees. https://doi.org/10.48550/arXiv.2106.04416
Louizos C, Shalit U, Mooij JM, et al (2017) Causal effect inference with deep latent-variable models. Adv Neural Inform Process Syst 30
Maes S, Meganck S, Manderick B (2007) Inference in multi-agent causal models. Int J Approx Reason 46(2):274–299. https://doi.org/10.1016/j.ijar.2006.09.005
Mahmood A (2011). Structure learning of causal bayesian networks: a survey. https://doi.org/10.7939/R35717N51
Maier M, Marazopoulou K, Arbour D, et al (2013a) A sound and complete algorithm for learning causal models from relational data. https://doi.org/10.48550/arXiv.1309.6843
Maier M, Marazopoulou K, Jensen D (2013b) Reasoning about independence in probabilistic models of relational data. https://doi.org/10.48550/arXiv.1302.4381
Malinsky D, Danks D (2018) Causal discovery algorithms: a practical guide. Philos Compass 13(1):e12470. https://doi.org/10.1111/phc3.12470
Malinsky D, Shpitser I, Richardson T (2019) A potential outcomes calculus for identifying conditional path-specific effects. In: Proceedings of the twenty-second international conference on artificial intelligence and statistics, PMLR, pp 3080–3088
Marx A, Gretton A, Mooij JM (2021) A weaker faithfulness assumption based on triple interactions. In: Proceedings of the thirty-seventh conference on uncertainty in artificial intelligence, PMLR, Proceedings of machine learning research, pp 451–460
Naimi AI, Cole SR, Kennedy EH (2016) An introduction to g methods. Int J Epidemiol 46(2):756–762. https://doi.org/10.1093/ije/dyw323
Nichols A (2007) Causal inference with observational data. Stata J 7(4):507–541. https://doi.org/10.1177/1536867X0800700403
Nogueira AR, Gama J, Ferreira CA (2021) Causal discovery in machine learning: theories and applications. J Dyn Games 8(3):203. https://doi.org/10.3934/jdg.2021008
Nogueira AR, Pugnana A, Ruggieri S et al (2022) Methods and tools for causal discovery and causal inference. Wiley Interdiscip Rev 12(2):e1449. https://doi.org/10.1002/widm.1449
Ogarrio JM, Spirtes P, Ramsey J (2016) A hybrid causal search algorithm for latent variable models. In: Proceedings of the eighth international conference on probabilistic graphical models, PMLR, pp 368–379
Ogburn EL, VanderWeele TJ (2014) Causal diagrams for interference. Stat Sci 29(4):559–578. https://doi.org/10.1214/14-STS501
Pearl J (1997) On the identification of nonparametric structural models. In: Berkane M (ed) Latent variable modeling and applications to causality. Springer, New York, pp 29–68, https://doi.org/10.1007/978-1-4612-1842-5_3
Pearl J (2009) Causality. Cambridge University Press. https://doi.org/10.1017/CBO9780511803161
Pearl J, Mackenzie D (2018) The book of why: the new science of cause and effect. Basic books
Peña JM (2016) Learning acyclic directed mixed graphs from observations and interventions. In: Conference on probabilistic graphical models, PMLR, pp 392–402
Pensar J, Nyman H, Koski T et al (2015) Labeled directed acyclic graphs: a generalization of context-specific independence in directed graphical models. Data Mining Knowledge Discov 29(2):503–533. https://doi.org/10.1007/s10618-014-0355-0
Perkovic E (2020) Identifying causal effects in maximally oriented partially directed acyclic graphs. In: Proceedings of the 36th conference on uncertainty in artificial intelligence (UAI), proceedings of machine learning research, vol 124. PMLR, pp 530–539
Peters J, Mooij JM, Janzing D et al (2014) Causal discovery with continuous additive noise models. J Mach Learn Res 15(58):2009–2053
Ramsey J, Glymour M, Sanchez-Romero R et al (2017) A million variables and more: the fast greedy equivalence search algorithm for learning high-dimensional graphical causal models, with an application to functional magnetic resonance images. Int J Data Sci Anal 3(2):121–129. https://doi.org/10.1007/s41060-016-0032-z
Richardson T, Spirtes P (2002) Ancestral graph markov models. Ann Stat 30(4):962–1030.
Richardson TS (2014) A factorization criterion for acyclic directed mixed graphs. https://doi.org/10.48550/arXiv.1406.6764
Richardson TS, Robins JM (2013a) Single world intervention graphs: a primer
Richardson TS, Robins JM (2013b) Single world intervention graphs (swigs): aunification of the counterfactual and graphical approaches to causality. Center for the Statistics and the Social Sciences, University of Washington Series Working Paper 128(30)
Robins J (1986) A new approach to causal inference in mortality studies with a sustained exposure period-application to control of the healthy worker survivor effect. Math Modell 7(9–12):1393–1512
Robins J, Hernán M, Siebert U (2004) Effects of multiple interventions. Comparat Quant Health Risks 1:2191–2230
Robins JM, Richardson TS (2011) Alternative graphical causal models and the identification of direct effects. In: Causality and psychopathology: finding the determinants of disorders and their cures, vol 84. Oxford University Press, pp 103–158, https://doi.org/10.1093/oso/9780199754649.003.0011
Robins JM, Richardson TS, Shpitser I (2022) An interventionist approach to mediation analysis. In: Probabilistic and causal inference: the works of Judea Pearl, pp 713–764, https://doi.org/10.1145/3501714.3501754
Rosenbaum PR, Rubin DB (1983) The central role of the propensity score in observational studies for causal effects. Biometrika 70(1):41–55. https://doi.org/10.1093/biomet/70.1.41
Rubenstein PK, Weichwald S, Bongers S, et al (2017) Causal consistency of structural equation models. https://doi.org/10.48550/arXiv.1707.00819
Rubin DB (1974) Estimating causal effects of treatments in randomized and nonrandomized studies. J Eucat Psychol 66(5):688–701. https://doi.org/10.1037/h0037350
Rubin DB (1978) Bayesian inference for causal effects: the role of randomization. Ann Stat 6(1):34–58. https://doi.org/10.1214/aos/1176344064
Rubin DB (1980) Randomization analysis of experimental data: the fisher randomization test comment. J Am Stat Assoc 75(371):591–593. https://doi.org/10.2307/2287653
Runge J (2018) Causal network reconstruction from time series: from theoretical assumptions to practical estimation. Chaos 28(7):075310. https://doi.org/10.1063/1.5025050
Salmerón A, Rumí R, Langseth H et al (2018) A review of inference algorithms for hybrid bayesian networks. J Artif Intell Res 62:799–828. https://doi.org/10.1613/jair.1.11228
Shalit U (2020) Can we learn individual-level treatment policies from clinical data? Biostatistics 21(2):359–362. https://doi.org/10.1093/biostatistics/kxz043
Shenoy PP, West JC (2011) Inference in hybrid bayesian networks using mixtures of polynomials. Int J Approx Reason 52(5):641–657. https://doi.org/10.1016/j.ijar.2010.09.003
Sherman E, Shpitser I (2018) Identification and estimation of causal effects from dependent data. Adv Neural Inform Process Syst 31
Shimizu S, Hoyer PO, Hyvärinen A et al (2006) A linear non-gaussian acyclic model for causal discovery. J Mach Learn Res 7(72):2003–2030
Shpitser I (2015) Segregated graphs and marginals of chain graph models. Adv Neural Inform Process Syst 28
Shpitser I, Pearl J (2006) Identification of joint interventional distributions in recursive semi-markovian causal models. In: Proceedings of the 21st national conference on artificial intelligence-volume 2. AAAI Press, AAAI’06, pp 1219–1226
Shpitser I, Tchetgen ET (2016) Causal inference with a graphical hierarchy of interventions. Ann Stat 44(6):2433–2466. https://doi.org/10.1214/15-AOS1411
Shpitser I, Richardson TS, Robins JM (2022) Multivariate counterfactual systems and causal graphical models, 1st edn., Association for computing machinery, New York, pp 813–852. https://doi.org/10.1145/3501714.3501757
Silva R (2016) Observational-interventional priors for dose-response learning. Adv Neural Inform Process Syst 29
Smith JQ, Anderson PE (2008) Conditional independence and chain event graphs. Artif Intell 172(1):42–68. https://doi.org/10.1016/j.artint.2007.05.004
Sobel DM, Legare CH (2014) Causal learning in children. WIREs Cognit Sci 5(4):413–427. https://doi.org/10.1002/wcs.1291
Soto MG, Sucar LE, Escalante HJ (2020) Causal games and causal nash equilibrium. Res Comput Sci 149:123–133
Spirtes P, Glymour C (1991) An algorithm for fast recovery of sparse causal graphs. Soc Sci Comput Rev 9(1):62–72. https://doi.org/10.1177/089443939100900106
Spirtes P, Glymour CN, Scheines R (1990) Causality from probability. In: Conference proceedings: advanced computing for the social sciences
Spirtes P, Glymour CN, Scheines R, et al (2000) Causation, prediction, and search, 2nd edn. MIT Press, Cambridge
Stuart EA (2010) Matching methods for causal inference: a review and a look forward. Stat Sci 25(1):1–21. https://doi.org/10.1214/09-STS313
Tchetgen EJT, VanderWeele TJ (2012) On causal inference in the presence of interference. Stat Methods Med Res 21(1):55–75. https://doi.org/10.1177/0962280210386779
Tikka S, Hyttinen A, Karvanen J (2019) Identifying causal effects via context-specific independence relations. Adv Neural Inform Process Syst 32:15
VanderWeele TJ (2009) Concerning the consistency assumption in causal inference. Epidemiology 20(6):880–883. https://doi.org/10.1097/EDE.0b013e3181bd5638
VanderWeele TJ, Hernan MA (2013) Causal inference under multiple versions of treatment. J Causal Inference 1(1):1–20. https://doi.org/10.1515/jci-2012-0002
Vowels MJ, Camgoz NC, Bowden R (2022) D’ya like dags? A survey on structure learning and causal discovery. ACM Comput Surv (CSUR). https://doi.org/10.1145/3527154
Yao L, Chu Z, Li S et al (2021) A survey on causal inference. ACM Trans Knowl Disco Data (TKDD) 15(5):1–46. https://doi.org/10.1145/3444944
Young JG, Hernán MA, Robins JM (2014) Identification, estimation and approximation of risk under interventions that depend on the natural value of treatment using observational data. Epidemiol Methods 3(1):1–19. https://doi.org/10.1515/em-2012-0001
Yu K, Li J, Liu L (2016) A review on algorithms for constraint-based causal discovery.
Yuan C, Druzdzel MJ (2007) Importance sampling for general hybrid bayesian networks. In: Artificial intelligence and statistics, proceedings of machine learning research, vol 2. PMLR, pp 652–659
Zhang J (2013) A comparison of three occam’s razors for markovian causal models. Br J Philos Sci 64:423–448. https://doi.org/10.1093/bjps/axs005
Zhang J, Spirtes P (2011) Intervention, determinism, and the causal minimality condition. Synthese 182(3):335–347. https://doi.org/10.1007/s11229-010-9751-1
Zhang J, Spirtes P (2015) The three faces of faithfulness. Synthese 193(4):1011–1027. https://doi.org/10.1007/s11229-015-0673-9
Zhang K, Hyvärinen A (2009) On the identifiability of the post-nonlinear causal model. In: Proceedings of the twenty-fifth conference on uncertainty in artificial intelligence, pp 647–655
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Appendix A: Explanation figures
Appendix A: Explanation figures
All but two pictures are based on this color/shape scheme. The text next to the arrows indicates what meaning the arrows generally convey with one exception: when assumptions have a directed arrow to another assumption outside its group of assumption, it means ’can be combined with’. (Color figure online)
Second exception: the different purple circles refer to different tasks. The boxes represent the assumptions necessary for this task, which may have overlap among tasks. The color of the boxes indicate the nature of the assumptions: while light blue represents sampling assumptions, ivory blue indicates assumptions on the data generating process (DGP) and darker blue is used for causal assumptions. (Color figure online)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Vonk, M.C., Malekovic, N., Bäck, T. et al. Disentangling causality: assumptions in causal discovery and inference. Artif Intell Rev 56, 10613–10649 (2023). https://doi.org/10.1007/s10462-023-10411-9
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10462-023-10411-9