Abstract
Allometric equations were developed for estimating aboveground biomass carbon (AGBC) in five tree species grown in a tree-based intercropping system at the University of Guelph Agroforestry Research Station, Guelph, Ontario, Canada. A total of 66 representative trees from five species: red oak (Quercus rubra) [n = 12], black walnut (Juglans nigra) [n = 16], black locust (Robinia pseudoacacia) [n = 10], white ash (Fraxinus americana) [n = 15], Norway spruce (Picea abies) [n = 13] were selected, harvested and their aboveground biomass and carbon content were quantified. Three commonly used allometric models were used to develop predictive equations. Regression models were developed and parameterized for each tree species and the best are presented based on information criteria (AIC, AICc, and BIC), mean absolute percentage error (MAPE), over/under estimation (MOUE), root mean square error (RMSE), R2, and regression coefficients (a, b) of the observed/predicted (OP) linear regression analysis. All equations with diameter at breast height (D) only and D and tree height (H) as the predictor variables fitted the AGBC data well, with R2 > 97% and RMSE < 40. However, a power model using D as the only predictor is recommended as the best model for black walnut, black locust, white ash, and Norway spruce. The models presented are the best fitted allometric equations for the indicated species and are recommended for these species, growing on similar soils under the same temperate conditions at densities of < 125 tree per hectare.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The integration of agroforestry systems into climate change strategies is a promising approach to address environmental and societal challenges while promoting sustainable development. The sequestration of carbon in aboveground biomass (AGBC) through woody cropping systems in Canada is seen as a capable land-use strategy to achieve the country's climate change mitigation goals, (Environment and Climate Change Canada, 2020). Agroforestry systems can also benefit growers economically by providing valuable feedstock to support green technologies in Canada (Bazrgar et al. 2022). Understanding the contribution of agroforestry systems to offset Green House Gases (GHG) emissions requires accurate quantification of AGBC at the individual tree species level (Altanzagas et al. 2019), especially since the growth and biomass of trees grown under agroforestry cropping systems can vary significantly depending on species and agroclimatic conditions (Jassal et al. 2013; Fortier et al. 2010; Pinno et al. 2010; Truax et al. 2012). It is therefore critical to understand the long-term biomass accumulation of promising tree species under agroforestry land-use systems in different regions of Canada and their adaptability to diverse agroclimatic conditions. This understanding is essential for assessing biomass productivity at a system level and to monitor carbon stocks on a spatial and temporal scale across the country (Cooper 1983; Chambers et al. 2001; Specht and West 2003; Kebede and Soromessa 2018).
There are currently no predictive models available to estimate the aboveground biomass for tree species under tree-based intercropping systems on marginal lands in southern Ontario, Canada. Regression models are commonly used and cost-effective methods for predicting aboveground biomass and carbon stocks in various woody crop production systems (Ravindranath and Ostwald 2008), and researchers have employed resultant allometric equations for biomass estimation due to their accuracy and efficiency (Rathore et al. 2021; Lambert et al. 2005). These equations primarily utilize tree growth characteristics such as diameter at breast height (D), height, and collar diameter as regressors (Guiabao 2016; Naik et al. 2019). This study aimed to evaluate species-specific nonlinear regression equations for predicting carbon sequestration in aboveground biomass of five common tree species suitable for use in an intercropping agroforestry land-use systems in southern Ontario, Canada.
Materials and methods
Site descriptions
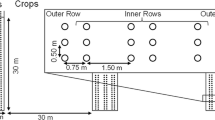
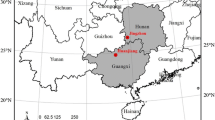
The study was conducted in a 30-ha tree-based intercropping system established in 1987 at the University of Guelph Agroforestry Research Station (GARS), Guelph, Ontario, Canada (latitude 430 32′ 28″ N, 800 12′ 32″ W) (Fig. 1). The average annual precipitation at GARS is 904 mm of which 338 mm is received during the growing season (May to September) of 144 frost-free days. Most of the land at GARS is Class 3–4 agricultural land, based on the Canadian Land Inventory (CLI) classification, and soils are classified as Gray Brown Luvisols, with fine sandy loam texture (Bazrgar et al. 2022). The tree density is 111 trees ha−1 with a within row spacing of 6 m and between row spacing of 6 and 15 m. The tree row width is 2 m, where no crops are grown. Tree alleys are intercropped with a crop rotation of corn (Zea mays), and soybean (Glycine max).
Aboveground biomass sampling and carbon quantification
Tree harvest occurred during the winter of 2020 using a commercial harvester. Representative trees (n = 66) of red oak (Quercus rubra) [n = 12], black walnut (Juglans nigra) [n = 16], black locust (Robinia pseudoacacia) [n = 10], white ash (Fraxinus americana) [n = 15], Norway spruce (Picea abies) [n = 13] were selected. For this purpose, we used stratified random sampling. For this we first divide the tree population into strata based on the diameter classes. Then, within each stratum, we randomly selected trees to form the sample. The proportion of each stratum in the final sample was equal to the proportion of the stratum in the population. This method was employed to ensure that our sample represents the entire range of diameter classes in the population. Trunk, branches, and needles (for Norway spruce) for each tree were separated and weighed in the field using a compact truck loader, Kubota CTL Model SVL75-2, and a digital crane hanging scale (VEVOR), with a 1000 kg capacity, following the protocols of Zhou et al. (2011), Battulga et al. (2013), and Bazrgar et al. (2020). The moisture content (MC) at the time of harvest for each component of a tree was quantified using sub-samples and MC (%) was used to convert whole-tree green weights to oven-dry weights. C concentration for each separated component in each tree sample was quantified using the combustion method (LECO CR412) (Table 1), and actual C content in aboveground biomass (AGB) was calculated using (1):
where AGBC refers to aboveground biomass carbon (kg C), AGB refers to aboveground dry biomass (kg) and MCC refers to the mean carbon concentration (%) in the tree.
Tree heights were measured using a normal measuring tape (Keson, Tape measure 1/2 IN X 100 FT/30M) and diameter at breast height (1.3 m aboveground) were measured using a diameter tape (Friedrich Richter Steel Diameter Tape). As the trees were already cut down, the above tapes were used on trees on the ground.
Developing regression equations
In order to develop allometric equations, three commonly used non-linear regression models were evaluated for estimating the AGBC of trees (Table 2). These models were selected by reviewing existing literature on biomass estimation equations, especially those relevant to our studied tree species and similar regions. Regression models were developed and parameterized for each tree species and the best of them were used as candidate models for predicting AGBC. For this purpose, the measured tree data were divided into two portions, the first of which was used for model fitting (Training set) and the second of which was used for model validation (Test set). Tree height (H, in m) and diameter at breast height (D, in cm) were used as regressors. The predicted C sequestration in AGB was calculated by multiplying the predicted AGB by the respective C concentration in AGB associated with each species.
Models were fitted to the data of H, D and AGBC using SAS 9.4 (SAS Institute Inc. Cary, NC), and parameterization in each model was done separately for each species using the Iterative Optimization Method ( Chen et al. 2012; Bazrgar et al. 2022). In this method, starter values are repeatedly entered to estimate the best model parameters through the least squares method.
Final values for parameters were estimated based on statistical significance, the model R2 and the Root Mean Square Error (RMSE) of non-linear regression analysis. The model R2 (pseudo R2 for non-linear regression) was calculated based on the below (2):
The best-fitted equation for each tree species was selected using the following standard criteria:
-
(1)
The scatter plot of observed and predicted AGBC values (OP) were plotted against the 1:1 line using the test set of the tree data (Pineiro et al. 2008; Bazrgar et al 2022). Post analysis of the OP linear regression was conducted using SAS 9.4 and goodness of fit was evaluated based on: (a) RMSE, which indicates the bias in the model (deviation of the actual slope value from the 1:1 line) (Savage 1993), (b) the Coefficient of Determination (adjusted-R2), which is a measure of the proportion of the variance in observed values explained by the predicted values, and (c) the OP linear regression coefficients (significance of slope = 1 and intercept = 0 which show the least deviation from 1:1 line).
-
(2)
Mean absolute percentage error (MAPE) given below (3):
$$MAPE= 1/n \sum_{i=1}^{n}| \frac{Observed i-Predicted i}{Observed i }|\times 100$$(3) -
(3)
Over/under estimation (MOUE) (%) was also calculated based on the formula below (4) and used to evaluate the predictive equations.
$$MOUE=\frac{Mean observed-Mean predicted}{Mean observed}\times 100$$(4) -
(4)
Fit statistics of nonlinear regression analysis including AIC (Akaike information criteria), AICc (Akaike's Corrected Information Criterion), and BIC (Bayesian Information Criterion). Information criteria are important measures for selecting non-linear models because they provide a quantitative measure of how well a particular model fits the data while considering model complexity. These criteria help to strike a balance between model fit and model complexity, allowing for the selection of the most appropriate non-linear model. Therefore, this study deployed these criteria for further validation of the selected model. The lower fit statistic values indicate a better-fit model.
Statistical analysis
All regression analyses were carried out using SAS 9.4 applying the NLIN, NLMIXED and REG procedure to perform statistical analyses and model fitting and evaluation. All linear and non-linear assumptions including normality of the standardized residuals and variance homogeneity were verified with plots of standardized residuals for all equations using SAS 9.4.
Results
Model parameterization
Table 3 shows the parameters developed for the regression equations using diameter (D) at breast height and height (H) data. The results indicated that all equations with D only and D and H as the predictor variables fitted the AGBC data well, with an R2 > 97% and RMSE < 40.
Model validation—Predicted versus observed regression of fitted models
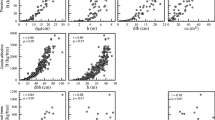
Results of the validation of allometric equations (goodness of fit) for each tree species using the observed/predicted (OP) linear regression are presented in Table 4. The results indicated that for red oak, Eqs. 1 and 2 showed acceptable prediction of AGBC and could be considered as candidate models (b ± SE = 1.08 ± 0.078, and 0.92 ± 0.074, respectively). For black walnut, all equations predicted acceptable values based on the evaluation statistics; however, Eq. 1 had the lowest RMSE (16.9 kg C tree−1) and could be the best candidate for predicting AGBC. Equation 2 gave an acceptable prediction of AGBC in Norway spruce based on the OP analysis. The coefficient of linear regression in this equation was the closest to 1 (b ± SE = 0.97 ± 0.053). However, the RMSE (4.7 kg C tree−1) was lowest in Eq. 1 suggesting that both Eqs. 1, and 2 were candidate equations for Norway spruce. OP analysis showed all three equations were good for black locust (Fig. 2); however, Eq. 1 had the highest R2 (0.93), lowest RMSE (17.6 kg C tree−1), and the closest coefficient of regression to 1 (b ± SE = 0.79 ± 0.125). Therefore, Eq. 1 could be considered the best candidate based on the OP analysis results. All equations predicted AGBC satisfactorily in white ash based on the OP analysis; however, for this species the R2 for the OP linear regression was lower than other spices (0.64, 0.65, 0.65, for Eq. 1, 2, and 3, respectively). The R2 values associated with all three predictive equations for white ash were higher than 0.60, which can be accepted given the statistically significant (p > 0.0311, 0.0281, 0.0278, for Eq. 1, 2, and 3, respectively). All equations had similar RMSE, although Eq. 1 showed the closest coefficient regression to 1 (b ± SE = 0.94 ± 0.32).
Model validation – Information criteria, mean absolute percentage error (MAPE) and over/under estimation (MOUE)
Results showed that for red oak, information criteria statistics, similar to the MAPE (10%) and MOUE (1.8%) support the results of OP analysis and indicated Eq. 2 as the best fitted model for predicting AGBC for this species under agroforestry systems. For black walnut, AIC, AICc, and BIC were lowest in Eq. 2 (84.2, 89, and 84.8, respectively), but were not considerably lower than the Eq. 1 fit statistics (86.4, 91.2, 87 respectively) (Table 5). MAPE (11.6%) and MOUE (3.6%) were also the least in Eq. 1 for black walnut. Findings from this study indicated that Eq. 1 showed the lowest AIC, AICc, BIC and MAPE (18.8%) for Norway spruce. However, the MOUE suggested Eq. 2 (− 15.6%) as the best equation for this species. For black locust, supporting the results from OP analysis, information criteria suggested Eq. 1 as the best predictor model for AGBC. Prediction with Eq. 1 also had the lowest MAPE (32.6%) for this species; however, the lowest MOUE (− 1%) was associated with Eq. 2 for this species. For white ash, the results showed that Eq. 2 had the lowest AIC, AICc, and BIC (71, 77, 71.3, respectively), which were not extensively different from the Eq. 1 statistics (73.8, 79.8, 74.1, respectively). MAPE (25.91 kg C tree−1) and MOUE (− 0.71 kg C tree−1) were both lowest in Eq. 1 (Table 6).
Selected models
The best-fitted equation for each tree species was selected using the above standard criteria and are presented in presented in Table 7.
Discussion
Selecting the best-fit model
All of the evaluation statistics (OP analysis, MAPE and MOUE and information criteria statistics) confirmed that Eq. 2 (Table 7) was the best-fit model for predicting AGBC in red oak grown under an intercropping scenario. For black walnut, OP analysis supported the results from MAPE and MOUE and suggested Eq. 1 (Table 7) as the best fitted equation. Fit statistics were lowest in Eq. 2, but they are not considerably less than Eq. 1 fit statistics. Therefore, we selected Eq. 1 as the best-fit equation for black walnut, intercropped. For Norway spruce, OP analysis indicated both Eqs. 1 and 2 were the best candidate equations. Therefore, based on the lower AIC, AICC and BIC (Table 5), we recommend Eq. 1 (Table 7) as the best predictor of AGBC for Norway spruce, grown in an intercropped scenario. Findings from this study also suggested Eq. 1 as the best predictor model for AGBC for black locust cultivated under an intercropping agroforestry scenario based on the results from the OP analysis, information criteria and MAPE. For white ash, notwithstanding the fact that fit statistics were lowest in Eq. 2, the current study suggests that Eq. 1 (Table 7) is the best-fit model for this species when intercropped, due to the indicated results from OP analysis, MOUE and MAPE.
For most of the tested tree species (red oak excepted) the power function using D as the only predictor was sufficient to provide the best model. Other studies have also indicated that the power equation is easy to fit to biomass data, requires only basic system inventory data to apply in practice, and usually provides reasonably accurate predictions (Ter-Mikaelian and Korzukhin 1997; Jenkins et al. 2003; Wang 2006; Sierra et al. 2007; Basuki et al. 2009). However, the results of this study demonstrate that adding tree height or height classes as an additional predictor into biomass equations can significantly improve the model fitting and performance for red oak.
Conclusion
This study parameterized, evaluated, and suggested species-specific nonlinear regression equations for predicting carbon sequestration in aboveground biomass of five common tree species grown in an intercropping scenario in southern Ontario, Canada. The suggested models were selected based on OP analysis, information criteria, MAPE and MOUE statistics. Many scientists have recommended linear regressing OP values for evaluating quantitative models and post analyzing the regression coefficients (Monte et al. 1996; Loehle 1997; Kobayashi and Salam 2000; Knightes and Cyterski 2005; Pineiro et al. 2008, Altanzagas et al. 2019). Information criteria are also crucial in selecting non-linear models due to their ease of assessing the fit of the model to the data, penalize model complexity, striking a balance between fit and complexity, and facilitating model comparison.
Different regression models were found as best fits for AGBC sequestration in five tree species under an intercropping system. Results indicate that all equations with D only and D and H as the predictor variables fitted the AGBC data well, with R2 > 97% and RMSE < 40. However, a power function using D as the only predictor was selected as the best model for black walnut, black locust, white ash, and Norway spruce.
Based on the results of this study, the listed equations in Table 7, for the respective tree species, are the best fitted allometric equations that should be used for open-grown and low tress density (< 125 tree per hectare) tree-based intercropping systems in the temperate regions. Compared to trees grown in urban settings or in the forest, trees grown in intercropping systems differ significantly in the following ways: (a) Intercropping systems provide trees with leached nutrients from fertilizers applied annually to adjacent agricultural crops. The growth rate of these trees can therefore be greater than that of trees in a forest or in an urban area. (b) Intercropping trees are heavily pruned (branches) periodically to ensure light penetration and to reduce competition for light. Hence, the aboveground biomass derived from the developed equations for the tested tree species indicates the actual standing biomass excluding the pruned biomass. This is important to calculate the carbon in the standing biomass for carbon trading purposes in temperate tree-based intercropping systems. (c) There is a significant difference between tree densities in an intercropping system (around 100 to 125 trees per ha) and those reported for forests (over 2000 trees per ha). It is, therefore, not reasonable to expect that the allometric equations developed from this study can be applied in urban, forest or parkland settings due to the specific management aspects of temperate tree-based intercropping systems as indicated above. However, any researcher / landowner / municipalities can now take our developed allometric equations and apply to different tree-based land-use systems (urban plantings, forestry setting, roof top trees etc.) and test for the best-fit and can report deviations, if any.
References
Altanzagas B, Luo YK, Altansukh B, Dorjsuren C, Fang JY, Hu HF (2019) Allometric equations for estimating the aboveground biomass of five forest tree species in Khangai Mongolia. Forests 10:661. https://doi.org/10.3390/f10080661
Basuki TM, van Laake PE, Skidmore AK, Hussin YA (2009) Allometric equations for estimating the aboveground biomass in tropical lowland Dipterocarp forests. For Ecol Manage 257:1684–1694
Battulga P, Tsogtbaatar J, Dulamsuren C, Hauck M (2013) Equations for estimating the aboveground biomass of Larix sibirica in the forest-steppe of Mongolia. Journal of Forestry Research 24:431–437
Bazrgar AB, Ng A, Coleman B, Ashiq MW, Gordon A, Thevathasan N (2020) Long-term monitoring of soil carbon sequestration in woody and herbaceous bioenergy crop production systems on marginal lands in southern Ontario, Canada. Sustainability 12(9):3901
Bazrgar AB, Sidders D, Thevathasan N (2022) Predicting aboveground biomass carbon sequestration potential in hybrid poplar clones under afforestation plantation management in southern Ontario. Canada Forest Chronicle 98(1):89–102. https://doi.org/10.5558/tfc2022-011
Chambers PA, Guy M, Roberts ES, Charlton MN, Kent R, Gagnon C, Grove G, Foster N (2001) Nutrients and their impact on the Canadian environment. Agriculture and Agri-Food Canada, Environment Canada, Fisheries and Oceans Canada, Health Canada, and Natural Resources Canada, Ottawa, ON.
Chen Y, Fang S, Huang Y, Eeckhout L, Fursin G, Temam O, Wu C (2012) Deconstructing iterative optimization. Trans Architect Code Optimiz. https://doi.org/10.1145/2355585.2355594
Cooper C (1983) Carbon storage in managed forests. Can J for Res 13:155–166. https://doi.org/10.1139/x83-022
Dong L, Zhang L, Li F (2015) Developing additive systems of biomass equations for nine hardwood species in Northeast China. Trees 29:1149–1163
Environment and Climate Change Canada (2020) National Inventory Report 1990–2018: Greenhouse gas sources and sinks in Canada: Executive summary. Available from: https://www.canada.ca/en/environment-climate-change/services/climate-change/greenhouse-gas-emissions.html [accessed 6 June 2023].
Fortier J, Gagnon D, Truax B, Lambert F (2010) Biomass and volume yield after 6 years in multiclonal hybrid poplar riparian buffer strips. Biomass Bioenerg 34:1028–1040
Guiabao EG (2016) Aboveground carbon stock assessment of mango-based agroforestry in Bulbul, Rizal, Kalinga, Philippines. Int J Int Res Innovat 4(2):19–25
Hosoda K, Iehara T (2010) Aboveground biomass equations for individual trees of Cryptomeria japonica, Chamaecyparis obtusa and Larix kaempferi in Japan. J for Res Jpn 15:299–306
Jassal RS, Black TA, Arevalo C, Jones H, Bhatti JS, Sidders D (2013) Carbon sequestration and water use of a young hybrid poplar plantation in north-central Alberta. Biomass Bioenergy 56:323–333. https://doi.org/10.1016/j.biombioe.2013.05.023
Jenkins JC, Chojnacky DC, Heath LS, Birdsey RA (2003) National-scale biomass estimators for United States tree species. For Sci 49:12–35
Kebede B, Soromessa T (2018) Allometric equations for aboveground biomass estimation of Olea europaea L. subsp. cuspidata in Mana Angetu Forest. Ecosyst Health Sustain 4:1–12. https://doi.org/10.1080/20964129.2018.1433951
Knightes CD, Cyterski M (2005) Evaluating predictive errors of a complex environmental model using a general linear model and least square means. Ecol Model 186:366–374
Kobayashi K, Salam MU (2000) Comparing simulated and measured values using mean squared deviation and its components. Agron J 92:345–352
Lambert MC, Ung CH, Raulier F (2005) Canadian national tree aboveground biomass equations. Can J for Res 35:1996–2018. https://doi.org/10.1139/X05-112
Loehle C (1997) A hypothesis testing framework for evaluating ecosystem model performance. Ecol Model 97:153–165
Monte L, Hakanson L, Bergstrom U, Brittain J, Heling R (1996) Uncertainty analysis and validation of environmental models: the empirically based uncertainty analysis. Ecol Model 91:139–152
Naik SK, Sarkar PK, Das B, Singh AK, Bhatt BP (2019) Biomass production and carbon stocks estimate in mango orchards of hot and subhumid climate in eastern region. India Carbon Manag 10(5):477–487
Piñeiro G, Perelman S, Guerschman JP, Paruelo JM (2008) How to evaluate models: observed versus predicted or predicted vs. observed? Ecol Model 216:316–322
Pinno B, Thomas B, Belanger N (2010) Predicting the productivity of a young hybrid poplar clone under intensive plantation management in northern Alberta, Canada using soil and site characteristics. New For 39:89–103
Rathore AC, Mehta H, Islam S, Saroj PL, Sharma NK, Jayaprakash J, Gupta AK, Dubey RK, Ghosh BN, Prasad R, Kumar D, Raizada A (2021) Biomass, carbon stocks estimation and predictive modeling in mango-based land uses on degraded lands in Indian Sub-Himalayas. Agroforest Syst. https://doi.org/10.1007/s10457-021-00660-4
Ravindranath NH, Ostwald M (2008) Carbon inventory methods: Handbook for greenhouse gas inventory. Carbon Mitig Round Wood Prod Projects, Springer Science, Delft: Advances in Global Change Research, Springer.
Savage MJ (1993) Statistical aspects of model validation. In: Workshop on the field water balance in the modelling of cropping systems. University of Pretoria, South Africa.
Sierra CA, del Valle JI, Orrego SA, Moreno FH, Harmon MA, Zapata M, Colorado GJ, Herrera MA, Lara W, Restrepo DE, Berrouet LM, Loaiza LM, Benjumea JF (2007) Total carbon stocks in a tropical forest landscape of the Porce region, Colombia. For Ecol Manage 243:299–309
Specht A, West PW (2003) Estimation of biomass and sequestered carbon on farm forest plantations in northern New South Wales, Australia. Biomass Bioenergy 25:363–379. https://doi.org/10.1016/S0961-9534(03)00050-3
Ter-Mikaelian MT, Korzukhin MD (1997) Biomass equations for sixty-five North American tree species. For Ecol Manag 97:1–24. https://doi.org/10.1016/S0378-1127(97)00019-4
Truax B, Gagnon D, Fortier J, Lambert F (2012) Yield in 8-year-old hybrid poplar plantations on abandoned farmland along climatic and soil fertility gradients. For Ecol Manag 267:228–239
Wang C (2006) Biomass allometric equations for 10 co-occurring tree species in Chinese temperate forests. For Ecol Manage 222:9–16
Xue Y, Yang Z, Wang X, Lin Z, Li D, Su S (2016) Tree biomass allocation and its model Additivity for Casuarina equisetifolia in a tropical forest of Hainan Island, China. PLoS ONE 11:1–20
Zhou X, Brandle JR, Awada TN, Schoeneberger MM, Martin DL, Xin Y, Tang Z (2011) The use of forest-derived specific gravity for the conversion of volume to biomass for open-grown trees on agricultural land. Biomass Bioenergy 35:1721–1731
Acknowledgements
The authors are thankful to the Ontario Ministry of Agriculture, Food and Rural Affairs (OMAFRA) for funding this project under the Special Initiative program. The authors are also thankful to all historical contributions made by previous graduate students and research field assistants which made this study possible.
Author information
Authors and Affiliations
Contributions
A.B. wrote the main manuscript text, data collection and statistical analysis. N.T. PI for the project and reviewed the manuscript. A.M. and J.S. reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bazrgar, A.B., Thevathasan, N., Gordon, A. et al. Allometric equations for estimating aboveground biomass carbon in five tree species grown in an intercropping agroforestry system in southern Ontario, Canada. Agroforest Syst 98, 739–749 (2024). https://doi.org/10.1007/s10457-023-00942-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10457-023-00942-z