Abstract
Spatial capture–recapture (SCR) using search–encounter methods estimate population abundance from encounters of individually recognisable animals along an a priori-designated search path. We applied search–encounter SCR methods and photographic sampling to estimate the abundance of plains zebras (Equus quagga) at Telperion and Ezemvelo nature reserves, South Africa. We analysed encounter data by comparing four hazard-function models for the detection process. The abundance estimate under three models was just above 1000 animals (95% credible intervals c. 960, 1220) versus 811 (719, 917) for the remaining model. The former estimates were broadly similar to aerial counts conducted around the same time. Standard deviation in locations around individual activity centres (\(\sigma _{move}\)) was c. 0.8 km, with little difference between models. In situations where structured surveys are not possible, the approach presented here has the potential to estimate abundance from opportunistic animal encounters (e.g. generated via citizen science schemes) within an SCR framework.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Accurate estimation of animal abundance is fundamental to a number of ecological problems and a necessary part of assessing population status for wildlife management or conservation (Williams et al. 2002; Conroy and Carroll 2009). One challenge, however, is that detection probability is usually <1 and must also be estimated to produce unbiased estimates of population parameters (Lebreton et al. 1992; MacKenzie et al. 2002; MacKenzie et al. 2003). A number of population estimation methods can accommodate imperfect detection; for example, N-mixture modelling (Royle 2004), distance sampling (Buckland et al. 2001), time-to-detection (Garrard et al. 2008), and capture–recapture (CR) (Otis et al. 1978; Seber 1982). In the case of CR, detection probability can be estimated via captures and recaptures of recognised individuals in traps or other encounter detectors (e.g. camera traps, hair snares), where the location of encounter occurs at fixed sites where the detectors have been positioned by researchers (i.e. spatial CR) (Efford 2004; Borchers and Efford 2008).
Increasingly, however, capture–recapture data have been collected via structured searches of a study area (i.e. search–encounter) (Royle et al. 2011; Royle et al. 2014) where researchers travel a search path and record encounters with individuals (e.g. photographically) and use unique pelage patterns or body markings to identify individuals and construct capture histories (e.g. Auger-Méthé et al. 2010; Morrison and Bolger 2014; Grange et al. 2015; Marshal 2017). In a sense, the animal remains relatively stationary while the detector (e.g. a human with a hand-held camera) moves past, instead of the detector remaining stationary while the animal moves past.
Search–encounter sampling has been described as following an area search of uniform intensity (i.e. uniform search) or a fixed search path (Royle et al. 2014). With uniform search, the study area is divided into polygons with well-defined boundaries. Within each polygon, the area is searched at a uniform intensity such that, given an individual is present inside the polygon, it has a constant probability of detection. Sampling produces a location within each polygon for each detected individual on each survey occasion. This approach and spatial CR (SCR) estimation methods were used by Royle and Young (2008) to estimate abundance of flat-tailed horned lizards (Phrynosoma mcallii). An alternative uses a fixed search path identified ahead of a survey and independently of the locations of the animals in the population. For example, search paths might follow roads or trails within a nature reserve or protected area. Data collected from such sampling might consist of (1) locations of encountered individuals, (2) locations of individuals and of observers while on the search path, or (3) the closest distance from the search path to the individual location, similar to distance sampling. Royle et al. (2011) demonstrated use of approach (1) for the willow tit (Parus montanus), and Gowan et al. (2021) applied approach (3) to estimate abundance, recruitment and persistence of North Atlantic right whales (Eublaena glacialis).
For data coming from approaches (1) or (2), modelling encounters with individuals along a search path is typically via estimation of the total hazard to encounter (Royle et al. 2011; Royle et al. 2014). Instead of using encounter relative to the fixed points of a detector array, encounter is modelled relative to the search path, which is represented by a series of line segments, each delineated by two point locations. The sum of the hazard for each point location along the path yields the total hazard to encounter for the search path. Whether using fixed detector locations or points along a search path, distance from the animal to observer or detector is treated as a covariate affecting detection. For detectors, distance is that between the detector and an unobserved activity centre for the individual, whereas the distance covariate for search path sampling is between the observer and the present animal location (Royle et al. 2011). Because of this difference, analysis of search path data explicitly incorporate the movement of animals around their activity centres (Royle et al. 2014).
Search–encounter sampling along nature reserve roads and trails, combined with photographic data collection, have the potential to generate useful encounter data for abundance or demographic parameter estimation in SCR analysis. In this study, our goal was to apply this approach to estimate abundance of the plains zebra (Equus quagga), a species well-suited to non-invasive CR because of its individually unique pelage patterns (e.g. Grange et al. 2015). We apply a hazard-function analysis to model encounters with individuals by fitting Bayesian hierarchical models, and we compare a number of models to represent the hazard function. We demonstrate that data from search–encounter sampling produce abundance estimates similar to counts from independent methods. Their application to analysing photographically collected encounter data, particularly opportunistically collected data, likely depends on the availability of auxiliary information to assess spatial extent and sampling intensity within a study area.
Methods
Study area
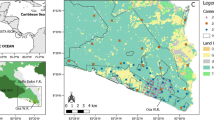
The study occurred at Telperion and Ezemvelo nature reserves, Gauteng and Mpumalanga provinces, South Africa (Fig. 1). Together, the reserves constituted c. 13,000 ha of protected predominantly grassland biome. The Wilge River separated an east section, containing Telperion, and a west section, containing Ezemvelo. Based on data from nearby Bronkhorstspruit weather station, the region experiences distinct wet and dry seasons, with c. 90% of rain falling during the months October to March (austral spring and summer). Average annual rainfall was 650 mm (range: 412 [1998]–949 [1989]), and range in average daily temperatures was \(4 \ ^\circ \text{C}\) in July to \(26 \ ^\circ \text{C}\) in January (Helm 2006).
The vegetation within Telperion and Ezemvelo is classified as Rand Highveld Grassland (Mucina et al. 2006) and Loskop Mountain Bushveld (Rutherford et al. 2006). Common grass species were Elionurus muticus, Eriagrostis curvula and Setaria sphacelata, and common woody species included Englerophytum magalismontanum, Vangueria infausta, Faurea saligna, Burkea africana, Combretum apiculatum, Cussonia paniculata, Strychnos pungens, Protea caffra, Acacia caffra and Gymnosporia spp. (Helm 2006). In addition to plains zebras, common large herbivores include blesbok (Damaliscus pygargis phillipsi), greater kudu (Tragelaphus strepsiceros), blue wildebeest (Connochaetes taurinus taurinus), black wildebeest (Connochaetes gnou), red hartebeest (Alcelaphus buselaphus caama), common eland (Taurotragus oryx), giraffe (Giraffa camelopardalis) and springbok (Antidorcas marsupialis). Leopards (Panthera pardus) occur in the reserve but are not common. Smaller carnivores include African civet (Civettictis civetta), black-backed jackal (Canis mesomelas) and caracal (Caracal caracal) (Helm 2006).
Data collection
We collected photographic data of plains zebras by driving a set route through both sections of the study area. We drove the entire route over 10 daily occasions in July–August 2017, recording our route with a global positioning system (GPS). We defined an encounter as a clear photograph of a zebra’s right flank (Fig. 2). We also recorded perpendicular distance between the vehicle and the animal with a range finder and the GPS coordinate of the observers.
Each photographed zebra was compared to a database of known individuals. If a photograph matched a recognised animal, we updated the encounter history with the new photograph; otherwise, we created an encounter history for a new individual. To ensure accuracy and consistency of identifications, all matching was conducted by one experienced observer (JPM). We also performed two additional checks after matching was complete: (1) within encounter histories for each individual to ensure that photographs were of the same individual, and (2) between encounter histories to ensure that an individual was not represented in more than one encounter history (Marshal 2017).
Model formulation
Following Royle et al. (2011), we formulated a Bayesian hierarchical model that represented the observation process and the ecological process. The observation level consisted of a hazard function to represent encounters with individual animals as a function of distance between the animal location (\(\textbf{u}_{ik}\)) and a search path (\(\textbf{X}\)) consisting of segments delineated by two-dimensional coordinates (\(\textbf{x}\)). Our choice of hazard model was to accommodate heterogeneity in detection caused by a non-linear search path, and the consequential variable exposure of animals along the path to sampling (Kéry and Royle 2021). We used the Gompertz formulation for the hazard model:
where \(h(\textbf{u}_{ik}, \textbf{x})\) is the hazard of encounter at location \(\textbf{x}\) for individual i occasion k, \(||\textbf{u}_{ik} - \textbf{x}||\) is the distance between individual i and the search path, and \(\beta _0\) and \(\beta _1\) are parameters to be estimated. We modelled probability of detection (\(p_{ik}\)) from the cumulative hazard (\(H_{ik}\)) and the data augmentation inclusion variable (\(z_i\); described below):
Observed encounters (\(y_{ik}\)) given the animal location were distributed as a Bernoulli random variable:
We defined the ecological process model assuming the individual activity centres (\(\textbf{s}_i\)) are distributed over a two-dimensional state space (S) and followed a uniform distribution:
where S was defined by the perimeter of the study area (Royle and Young 2008). Because some of the individual locations (\(\textbf{u}_{ik}\)) were unobserved, we used a bivariate normal distribution for \(\textbf{u}_{ik}\):
where \(\mathbf{\sigma }^2_{move}\) represented variability in movement around the activity centre and \(\textbf{I}\) was a \(2 \times 2\) identity matrix.
We used data augmentation to estimate population size (N) within S (Royle and Young 2008; Royle et al. 2009; Gardner et al. 2009). We defined M as the number of individuals encountered during the study (n) plus the augmented individuals with all-zero capture histories (\(M-n\)), some of which represented animals in the population available to be encountered but were not. We defined a binary latent variable (\(z_i\)) to identify which individuals in M were in the population N. The \(z_i\) were assumed to follow a Bernoulli distribution with probability \(\psi\). We estimated N as the sum of the latent variables: \(N = \sum _i^M z_i\).
Sensitivity analysis
We assessed the effects of different models for the hazard function in the detection process, with the goal of choosing a model that adequately fit the observation data. Specifically, we considered the normal kernel and Weibull functions, and a function based on the squared distance (model details in Royle et al. (2011)). We divided the search path into segments with length 1.5 km, based on preliminary analysis that \(\mathbf{\sigma }^2_{move} \approx 0.8\) km; 1.5-km spacing kept segment length \(<2\mathbf{\sigma }^2_{move}\) (Sun et al. 2014). We evaluated the sensitivity of the abundance estimate to segment length by running the analysis with 1.25- and 1-km segments.
Model implementation
We conducted analyses in JAGS (Plummer 2003), running through R (R Core Team 2023) by using the library jagsUI (Kellner 2021). We used noninformative uniform priors for parameters \(\beta _0, \beta _1, \log (\sigma )\) and \(\psi\), and augmented encounter histories to a total of \(M = 1500\). We ran three independent chains of sufficient length to allow for convergence (10,000–20,000 iterations) following a burn-in period (5000 iterations). We thinned one value for every 10 iterations to reduce the effects of autocorrelation on posterior estimates. We assessed convergence with the Brooks-Gelman-Rubin diagnostic (\(\hat{R} < 1.1\)) (Gelman and Shirley 2011).
We used Bayesian P-values to assess goodness of fit of the detection models (Gelman and Shirley 2011), based on a variation suggested by Royle et al. (2014) that assessed discrepancies between expected and observed number of encounters across individuals. An adequate-fitting model was indicated by Bayesian P-values within the range 0.1\(-\)0.9 (Royle et al. 2014). To the assess spatial randomness assumption for the ecological process model, we calculated an index-of-dispersion test based on the ratio of the variance to mean (Illian et al. 2008) in the number of activity centres across a defined number of grid cells covering the study area. We also calculated a Bayesian P-value, based on a Freeman-Tukey statistic, for the discrepancy between counts from the posterior sample of activity centres in grid cells and simulated counts under spatial randomness (Royle et al. 2014). We calculated both statistics with function SCRgof in library scrbook (Royle et al. 2014). We report estimated posterior means and 95% credible intervals for model parameters. R and JAGS scripts to fit the models are in the Supplementary Material.
Results
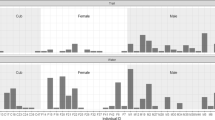
We analysed a total of 821 encounters from 383 recognised individuals. Number of occasions on which individuals were encountered ranged from 11 for 1 individual to 2 for 99 individuals. One-hundred eighty-one zebras were encountered on 1 occasion. Our assessment of survey line point coverage revealed little difference between estimates using 1-, 1.25-, and 1.5-km segment lengths (Table S1). Presented here are estimates for the 1.5-km analysis (Table 1).
Based on three detection models, the posterior mean of zebra abundance was just over 1000 individuals (c. 960, 1220), and did not vary much between the models. Interestingly, the Weibull detection model produced a substantially lower estimate than the other three models: 811 (719, 917).
For all detection models, \(\sigma _{move}\) was c. 0.8 km, and again it did not vary much among detection models (Table 1). The Weibull model produced a slightly higher estimate of 0.88. Bayesian P-values for all models indicated adequate fit to observed detection frequencies (Table 1), with values ranging between 0.21 (Weibull) and 0.49 (Gompertz). However, the ecological process model appeared to deviate from the spatial randomness assumption (index-of-dispersion and Freeman–Tukey: \(P < 0.001\) for all models), with local density appearing to be relatively high in the south-central and north-east portions of the study area (Fig. 3).
Discussion
Our application of photographic and search–encounter sampling for plains zebras at Telperion and Ezemvelo showed that three of the four detection models estimated abundance in the range of 1080–1088 animals, numbers that were similar to counts generated from aerial surveys conducted earlier the same year (1374, February 2017) and early the following year (1106, January 2018; Oppenheimer Generations Research and Conservation, unpublished data). Because the aerial counts occurred around the annual peak in zebra births, the lower SCR estimates could reflect in part higher mortality and reduction in first-year animal numbers by the food-scarce dry season. The Weibull model produced a somewhat lower SCR estimate, an outcome similar to that of Royle et al. (2011). All four detection models showed evidence of adequate fit to our encounter data; however, the assumption of spatial randomness was not supported for the ecological process model.
Ideally, search paths are located randomly with respect to animal locations (Conroy and Carroll 2009), something that often is not possible in protected areas where vehicles are required to remain on designated routes. We argue, however, that potential for bias in our study was minimal. A histogram of perpendicular distances indicated no evidence that zebras avoided roads (Fig. S1), and the open grassland landscape and relatively flat terrain of the study area suggested little benefit to using roads as travel routes to the extent that it would distort encounter rates.
Spatial randomness
Because animals are selective about where they occur on the landscape, higher densities of individuals in favourable parts of the environment should be expected. This selectivity could generate the appearance of clustering even with independent activity centres (Royle et al. 2014). Alternatively, animals expressing territoriality or avoidance of conspecifics could exhibit activity centres that are more evenly distributed on the landscape than expected from spatial randomness, in which case locations of activity centres are not likely independent between individuals (Reich and Gardner 2014).
However, the uniform statistical distributions describing the locations of activity centres are able to accommodate a variety of spatial patterns, including a range of clustering and spacing patterns (Royle et al. 2014). Moreover, the uniform priors have minimal effects on locations of activity centres if models are fitted to a large enough data set (Royle et al. 2014). Moreover, data simulation by Kéry and Royle (2021) suggests a minimal influence on estimation if spatial variation in density is ignored. Estimates based on uniform distribution of activity centres, habitat categories and density surface modelling produce estimates with considerable overlap of posterior distributions, differing mainly in the variation explained in the data. They did not do an exhaustive simulation study, but the analysis is supportive of minimal bias. Thus we expect the consequence of varying density in the study area to have a minimal influence on estimation of overall abundance, affecting the locations of the activity centres rather than their number.
Although not our objective to investigate drivers of local variation in zebra density across the study area, relationships between landscape attributes and density might be a question of ecological interest and could be accommodated in SCR modelling (Royle et al. 2014). Including such relationships in the ecological process model might produce an intensity surface for the point process model that accounts for the inhomogeneity in the underlying intensity or density function. An alternative to modelling intensity as a function of environmental covariates might be to use cluster process models having an intensity function with multiple clusters, each cluster representing a high-intensity region of state space (Illian et al. 2008).
A potential consequence of unmodelled spatial heterogeneity in activity centres is an effect of proximity to survey path on individual detection probability. If preferential space use produces heterogeneity in encounters because of differing probabilities of detection between individuals, such heterogeneity could bias estimates of abundance (Royle et al. 2013) and should be apparent though poor fit between a detection model and the observed numbers of encounters across individuals. Our detection models did not account for individual heterogeneity and yet there was no evidence of poor fit to the encounter data, suggesting a minimal influence on bias.
Zebra herding and cohesion
Plains zebras occur in herds, and so independent encounters of animals might be violated because of zebras occurring in social groups (aggregation) or having non-independent encounters (cohesion) (Bischof et al. 2020). Estimates of abundance from SCR are, however, robust to low–moderate spatial dependence: there is low bias with social groups, and low–moderate amounts of cohesion or dispersion have minimal effects on bias or precision; however, overdispersion strongly affects coverage of confidence intervals around parameter estimates (Bischof et al. 2020).
Zebras generally occur in groups of approximately a half-dozen animals, and rarely more than a dozen (Estes 1991). This degree of social grouping suggests a minimal effect on bias, but a probable effect on coverage. Modelling groups explicitly (e.g. Hickey and Sollmann 2018; Emmet et al. 2022) is probably not currently feasible for this population; additional data might assist in identifying zebra herds and assigning individuals to herds. However, we encountered many individuals once, and a single observation would not be sufficient to distinguish herd mates from animals in separate herds but encountered coincidentally. In the examples of both Hickey and Sollmann (2018) and Emmet et al. (2022), study animals had been the subject of long-term monitoring, and so social structure and group membership had been well established.
Search–encounter and opportunistic data
Citizen science schemes and online wildlife databases (e.g. iNaturalist, Macaulay Library) are a growing source of photographic data to which search–encounter SCR models might be applied, particularly with species that have individually unique identifying features. Because of photo-recorded animal encounters, geographic coordinates for each encounter via GPS-enabled devices, and machine learning to ease the effort of matching individual animals based on pelage patterns, there is the possibility of using opportunistically collected data to develop spatially explicit encounter histories of wildlife species. With sufficiently photographed species in highly visited areas, such data could yield estimates of abundance or other demographic parameters in situations where resources to conduct structured surveys are inadequate. Data generation via such schemes, however, amounts to unstructured spatial sampling, where locations, intensity and protocols of sampling occur opportunistically, rather than according to a pre-defined systematic approach (Royle et al. 2014), which could lead to spatially biased sampling.
The problem of spatial sampling bias is well-recognised in species distribution research (Kéry et al. 2010; Botts et al. 2011; Hugo and Altwegg 2017; Binley and Bennett 2023), and it has been addressed for some atlas databases by explicitly modelling separate observation and ecological processes (Kéry et al. 2010; van Strien et al. 2013; Péron and Altwegg 2015). Others integrate multiple data sets, where systematic presence-absence data (PA) are combined with opportunistic presence-only (PO) data in a model representing an ecological process for abundance or density and conditional observation processes for each of the PA and PO datasets (Sun et al. 2019). Despite the problems, however, potential data for such analyses are widely available if they can be analysed appropriately. Recognising that photographic encounters are the product of an observation process and developing models that describe that process will contribute substantially to producing more reliable estimates of abundance or demographic parameters from photographic encounter data.
Data availability
The datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request.
References
Auger-Méthé M, Marcoux M, Whitehead H (2010) Nicks and notches of the dorsal ridge: promising mark types for the photo-identification of narwhals. Mar Mamm Sci 26(3):663–678. https://doi.org/10.1111/j.1748-7692.2009.00369.x
Binley AD, Bennett JR (2023) The data double standard. Methods Ecol Evol 14(6):1389–1397. https://doi.org/10.1111/2041-210X.14110
Bischof R, Dupont P, Milleret C et al (2020) Consequences of ignoring group association in spatial capture-recapture analysis. Wildl Biol 2020(1). https://doi.org/10.2981/wlb.00649
Borchers DL, Efford MG (2008) Spatially explicit maximum likelihood methods for capture-recapture studies. Biometrics 64(2):377–385. https://doi.org/10.1111/j.1541-0420.2007.00927.x
Botts EA, Erasmus BFN, Alexander GJ (2011) Geographic sampling bias in the South African Frog Atlas Project: implications for conservation planning. Biodivers Conserv 20(1):119–139. https://doi.org/10.1007/s10531-010-9950-6
Buckland ST, Anderson DR, Burnham KP et al (2001) Introduction to distance sampling: estimating abundance of biological populations. Oxford University Press, Oxford, United Kingdom
Conroy MJ, Carroll JP (2009) Quantitative conservation of vertebrates. Wiley-Blackwell, Chichester, United Kingdom
Efford MG (2004) Density estimation in live-trapping studies. Oikos 106(3):598–610. https://doi.org/10.1111/j.0030-1299.2004.13043.x
Emmet RL, Augustine BC, Abrahms B et al (2022) A spatial capture-recapture model for group-living species. Ecology 103(10):e3576. https://doi.org/10.1002/ecy.3576
Estes RD (1991) The behavior guide to African mammals. University of California Press, Berkeley, USA
Gardner B, Royle JA, Wegan MT (2009) Hierarchical models for estimating density from DNA mark-recapture studies. Ecology 90(4):1106–1115. https://doi.org/10.1890/07-2112.1
Garrard GE, Bekessy SA, McCarthy MA et al (2008) When have we looked hard enough? A novel method for setting minimum survey effort protocols for flora surveys. Austral Ecol 33(8):986–998. https://doi.org/10.1111/j.1442-9993.2008.01869.x
Gelman A, Shirley K (2011) Inference from simulations and monitoring convergence. In: Brooks S, Gelman A, Jones GL et al (eds) Handbook of Markov chain Monte Carlo. Chapman & Hall/CRC, Boca Raton, Florida, USA, pp 163–174
Gowan TA, Crum NJ, Roberts JJ (2021) An open spatial capture-recapture model for estimating density, movement, and population dynamics from line-transect surveys. Ecol Evol 11(12):7354–7365. https://doi.org/10.1002/ece3.7566
Grange S, Barnier F, Duncan P et al (2015) Demography of plains zebras (Equus quagga) under heavy predation. Popul Ecol 57(1):201–214. https://doi.org/10.1007/s10144-014-0469-7
Helm CV (2006) Ecological separation of the black and blue wildebeest on Ezemvelo Nature Reserve in the highveld grasslands of South Africa. Master’s thesis, University of Pretoria, Pretoria, South Africa
Hickey JR, Sollmann R (2018) A new mark-recapture approach for abundance estimation of social species. PLoS One 13(12):e0208726. https://doi.org/10.1371/journal.pone.0208726
Hugo S, Altwegg R (2017) The second Southern African Bird Atlas Project: causes and consequences of geographical sampling bias. Ecol Evol 7(17):6839–6849. https://doi.org/10.1002/ece3.3228
Illian J, Penttinen A, Stoyan H et al (2008) Statistical analysis and modelling of spatial point patterns. John Wiley & Sons, Chichester, United Kingdom
Kellner K (2021) jagsUI: a wrapper around ’rjags’ to streamline ’JAGS’ analyses. https://CRAN.R-project.org/package=jagsUI
Kéry M, Royle JA (2021) Applied hierarchical modeling in ecology: analysis of distribution, abundance and species richness in R and BUGS, vol 2: dynamic and advanced models. Academic Press, London, United Kingdom
Kéry M, Royle JA, Schmid H et al (2010) Site-occupancy distribution modeling to correct population-trend estimates derived from opportunistic observations. Conserv Biol 24(5):1388–1397. https://doi.org/10.1111/j.1523-1739.2010.01479.x
Lebreton J, Burnham K, Clobert J et al (1992) Modeling survival and testing biological hypotheses using marked animals: a unified approach with case studies. Ecol Monogr 62(1):67–118. http://www.jstor.org/stable/2937171
MacKenzie DI, Nichols JD, Lachman GB et al (2002) Estimating site occupancy rates when detection probabilities are less than one. Ecology 83(8):2248–2255. https://doi.org/10.1890/0012-9658(2002)083[2248:ESORWD]2.0.CO;2
MacKenzie DI, Nichols JD, Hines JE et al (2003) Estimating site occupancy, colonization, and local extinction when a species is detected imperfectly. Ecology 84(8):2200–2207. https://doi.org/10.1890/02-3090
Marshal JP (2017) Survival estimation of a cryptic antelope via photographic capture-recapture. Afr J Ecol 55(1):21–29. https://doi.org/10.1111/aje.12304
Morrison TA, Bolger DT (2014) Connectivity and bottlenecks in a migratory wildebeest Connochaetes taurinus population. Oryx 48(04):613–621. https://doi.org/10.1017/S0030605313000537
Mucina L et al (2006) Grassland biome. No. 19 in Strelitzia, South African National Biodiversity Institute, p 349–437
Otis D, Burnham K, White G et al (1978) Statistical inference from capture data on closed animal populations. Wildl Monogr (62):3–135. http://www.jstor.org/stable/3830650
Péron G, Altwegg R (2015) Twenty-five years of change in southern African passerine diversity: nonclimatic factors of change. Glob Change Biol 21(9):3347–3355. https://doi.org/10.1111/gcb.12909
Plummer M (2003) JAGS: a program for analysis of Bayesian graphical models using Gibbs sampling. In: Hornik K, Leisch F, Zeileis A (eds) Proceedings of the 3rd International Workshop on Distributed Statistical Computing, Vienna, Austria
R Core Team (2023) R: a language and environment for statistical computing. http://www.R-project.org/
Reich BJ, Gardner B (2014) A spatial capture-recapture model for territorial species. Environmetrics 25(8):630–637. https://doi.org/10.1002/env.2317
Royle JA, Young KV (2008) A hierarchical model for spatial capture-recapture data. Ecology 89(8):2281–2289. http://www.jstor.org/stable/27650753
Royle JA (2004) N-mixture models for estimating population size from spatially replicated counts. Biometrics 60(1):108–115. https://doi.org/10.1111/j.0006-341X.2004.00142.x
Royle JA, Karanth KU, Gopalaswamy AM et al (2009) Bayesian inference in camera trapping studies for a class of spatial capture-recapture models. Ecology 90(11):3233–3244. https://doi.org/10.1890/08-1481.1
Royle JA, Kéry M, Guélat J (2011) Spatial capture-recapture models for search-encounter data. Methods Ecol Evol 2(6):602–611. https://doi.org/10.1111/j.2041-210X.2011.00116.x
Royle JA, Chandler RB, Sun CC et al (2013) Integrating resource selection information with spatial capture-recapture. Methods Ecol Evol 4(6):520–530. https://doi.org/10.1111/2041-210X.12039
Royle JA, Chandler RB, Sollmann R et al (2014) Spatial capture-recapture. Academic Press, Waltham, USA
Rutherford MC et al (2006) Savanna biome. In: Mucina L, Rutherford MC (eds) The vegetation of South Africa, Lesotho and Swaziland. No. 19 in Strelitzia, South African National Biodiversity Institute, Pretoria, South Africa, p 438–539
Seber GAF (1982) The estimation of animal abundance and related parameters, 2nd edn. The Blackburn Press, Caldwell, NJ, USA
Sun CC, Fuller AK, Royle JA (2014) Trap configuration and spacing influences parameter estimates in spatial capture-recapture models. PLoS One 9(2):e88025. https://doi.org/10.1371/journal.pone.0088025
Sun CC, Royle JA, Fuller AK (2019) Incorporating citizen science data in spatially explicit integrated population models. Ecology 100(9):e02777. https://doi.org/10.1002/ecy.2777
van Strien AJ, van Swaay CA, Termaat T (2013) Opportunistic citizen science data of animal species produce reliable estimates of distribution trends if analysed with occupancy models. J Appl Ecol 50(6):1450–1458. https://doi.org/10.1111/1365-2664.12158
Williams BK, Nichols JD, Conroy MJ (2002) Analysis and management of animal populations. Academic Press, San Diego, USA
Acknowledgements
We thank Duncan MacFadyen (Oppenheimer Generations Research and Conservation); Elsabe Bosch, Cassius Mmetle, Maroti Tau and Michael Tau (Telperion Nature Reserve); and John de Jager, Liezille Draper and Marnus Lombard (Ezemvelo Nature Reserve). Hannes Louw and Hannah Hoffmann assisted with data collection.
Funding
Open access funding provided by University of the Witwatersrand. This work was funded by the National Research Foundation of South Africa (grants 105729, 109097) and the University of the Witwatersrand.
Author information
Authors and Affiliations
Contributions
Both authors contributed to the study conception and design. JPM collected the data. Both authors conducted the analysis. JPM wrote the first draft of the manuscript and both authors contributed to revising previous versions. Both authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval
No approval of research ethics committees was required to accomplish the goals of this study because the work did not involve capture, handling or experimentation on any animal.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Marshal, J.P., Abadi, F. Abundance estimation of plains zebras via search–encounter sampling. Eur J Wildl Res 70, 40 (2024). https://doi.org/10.1007/s10344-024-01790-7
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10344-024-01790-7