Abstract
Species introductions outside their native ranges, often driven by trade and other anthropogenic activities, present significant ecological challenges. Reptiles, frequently traded as pets for their attractiveness, are particularly susceptible to such introductions, leading to shifts in distribution patterns and potential ecological impacts. The common chameleon (Chamaeleo chamaeleon), which has been historically introduced in several European countries, is such an example, yet no overall assessments are available to date for this species. In this study, we used ecological niche models to assess habitat suitability for the common chameleon in the Mediterranean basin for current and future scenarios. Concurrently, circuit theory techniques were employed to evaluate habitat connectivity in two historically introduced areas. We identified areas of high habitat suitability and dispersal corridors in introduced regions. Our results reveal a latitudinal gradient in habitat suitability changes, with the species facing both expansion and decline in different parts of its range, depending on the ecozone considered. Severe declines are noted in southeastern Spain, Tunisia, and Israel, while habitat suitability increases westwards in Portugal, Morocco, and Southern Italy. These insights contribute to a better understanding of the common chameleon’s ecological dynamics, providing a foundation for targeted management and conservation efforts. Our study highlights the importance of integrating ecological niche modelling and circuit theory techniques to predict habitat suitability and identify critical dispersal corridors for effective conservation strategies. Considering the ongoing challenges posed by human-mediated dispersals for the common chameleon, our research establishes a foundation for future studies to enhance our understanding of this elusive species.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The introduction of species outside their native range as a direct or indirect consequence of human action (defined as non-native species) may cause changes in the ecosystems to which they are introduced. These effects may be observed when a non-native species becomes invasive (Blackburn et al. 2011) but still occur even if species are not firmly established (Jeschke et al. 2013; Ricciardi et al. 2013). In some cases, these changes could be dramatic, resulting in the replacement of native species or radical changes in ecosystem functioning (Dorcas et al. 2012; Jeschke et al. 2014).
Reptiles constitute a significant portion of voluntarily introduced species, often for their aesthetic appeal (Reed 2005; Luiselli et al. 2012). Global trade in living reptiles exceeds half a million individuals annually (Karesh et al. 2005). The presence of reptiles outside their natural distribution can pose threats to ecosystems, either through their interactions with native species (Kraus 2015) or by introducing alien pathogens (Burridge et al. 2000; Nowak 2010).
The common chameleon (Chamaeleo chamaeleon) is one of the species that has been at the center of the pet trade process in recent decades (Carpenter et al. 2004). Native to the Mediterranean basin and Middle Asia, this species has been historically traded as a pet for its attractiveness. Consequently, it has been repeatedly released outside of its native range, so that, currently, it occurs outside of its native range in Spain, Portugal, Malta, and Italy (Paulo et al. 2002; Sindaco et al. 2008; Andreone et al. 2016; Basso et al. 2019).
In the Iberian Peninsula, the common chameleon has been introduced historically with two main events from North Africa (Paulo et al. 2002). Of these two events, one resulted in the establishment of Mediterranean populations in Iberia, while the other led to the formation of Atlantic populations (Paulo et al. 2002). Four subspecies of Chamaeleo chameleon are recognized, the nominotypical subspecies, C. c. chamaeleon, C. c. musae, C. c. orientalis, and C. c. rectricrista; the Iberian chameleons are attributed to the nominate subspecies. The presence of the common chameleon in Spain was even reported by Linnaeus in his System Naturae of 1766, and then several subsequent authors reported the species as present in southern Spain. It seems that the Malaga region was the first where the species was introduced in historical times (Pleguezuelos 1997). This event was carried out using individuals from North Africa, probably from a population of Erfoud, south of the Atlas Mountains (Paulo et al. 2002). The second introduction was reported in the nineteenth century, first in the Cadiz region and then in the Algarve and the Huelva regions. These populations originated from individuals of the southwestern Atlantic coast of Morocco, probably from the Essaouira province (Paulo et al. 2002).
In Italy, the common chameleon is reported historically in Sicily, even if its naturalization was never confirmed (Razzetti and Sindaco 2006; Sindaco et al. 2008). However, the species is present in Southern Italy, also in Apulia (since 1940s) and Calabria regions (since 2010) (Basso and Calasso 1991; Fattizzo and Marzano 2002; Basso et al. 2019; Sperone et al. 2010; Pellegrino et al. 2016), with other occurrences reported for other Italy’s regions (e.g., Bologna et al. 2000). Indeed, preserved specimens were found in central and northern territories (Corti et al. 2011). Moreover, the species is still traded, as a single individual has been found in Sicily about 10 years ago (Di Giuseppe 2013). Previous genetic analysis suggested the Italian populations’ independent origin with the Calabrian population probably originated by individuals from Tunisia, while individuals from North Israel founded the Apulian population (Andreone et al. 2016). However, a more recent genetic study indicated that the Apulian population had a multiple origin, with samples belonging to C. c. chamaeleon, C. c. recticrista, and C. c. musae subspecies (Basso et al. 2019), supporting a multiple-release hypothesis in Apulia.
Common chameleons have also been introduced in Malta, where individuals belong to two subspecies (Dimaki et al. 2008), while the individuals currently present in Cyprus and Greek islands, originated both from natural expansion and human-derived introductions, share mitochondrial haplotypes with the Turkish population (Andreone et al. 2016).
To address this knowledge gap, a crucial initial step involves the assessment of the general ecological requirements and then a possible trend of its expansion. This evaluation should encompass regions where the common chameleon is documented as introduced, such as Italy (Blackburn et al. 2014), and areas where it is considered native, particularly in the Iberian Peninsula. It is noteworthy to highlight that the species is under strict protection, being listed in Appendix II of the Bern Convention and included in Annex IV of the Habitat Directive (92/43/EEC).
Predictive models, such as ecological niche models (ENMs), have become a crucial tool to assess the distribution of non-native species by quantifying species-environment relationships and to predict suitable areas outside the known distribution range of the target species (Guisan and Thuiller 2005; Elith and Leathwick 2009; Barbet‐Massin et al. 2012; Mainali et al. 2015; Iannella et al. 2020; Farashi and Alizadeh-Noughani 2021; Serva et al. 2023). Moreover, the ENMs can be further refined in a GIS environment, converting them into more precise species distribution models (SDMs), which increases the predictive performance and allows the creation of more realistic models (Iannella et al. 2021). This approach, widely employed in current research, has demonstrated its effectiveness in providing accurate insights into the potential distribution of target species (Broennimann et al. 2007; Farashi and Alizadeh-Noughani 2021; Biber et al. 2023).
In this study, we used ENMs to model habitat suitability for the common chameleon within its current range, exploring both current and future scenarios, to predict possible expansion, especially in the areas where the species has been introduced. Additionally, we focused on habitat connectivity in Southern Italy and the southern part of the Iberian Peninsula, where the species is referred as native, using a robust connectivity-assessment algorithm at the landscape scale.
Material and methods
Study area and spatial data
We selected the Mediterranean basin as the main study area, considering the range of the target species as assessed within the IUCN Red List of Threatened Species. Specifically, we focused on three of the four subspecies, C. c. chamaeleon, C. c musae, and C. c recticrista to which the populations of the Iberian Peninsula and Southern Italy belong (Paulo et al. 2002; Andreone et al. 2016; Basso et al. 2019).
The nominative subspecies has a range across Algeria, Egypt, Libya, Malta, Morocco, Tunisia, the Western Sahara, and Yemen. At the same time, it has been historically introduced in Spain, Portugal, and Italy. The other subspecies show smaller ranges, with C. c musae distributed in Jordan, Israel, and Egypt and C. c recticrista spanning between Greece, Turkey, Cyprus, Israel, Lebanon, and Syria. While the species is native to North Africa and the Middle East, the populations in the Iberian Peninsula and Italy result from historical introduction. Nowadays, the species is considered native to the Iberian Peninsula (Paulo et al. 2002).
Data for Chamaeleo chamaeleon were gathered from this paper authors’ sampling campaigns, published literature, and the Global Biodiversity Information Facility (GBIF 2023). From GBIF, we selected only recent records (from 2000) and removed duplicate records, as well as those with uncertain geographic information about the occurrence locality, by using a 1-km spatial filter. From the literature occurrences, we selected only those with precise geographic information (Miraldo et al. 2005; Qninba et al. 2013), removing those reporting presence within Provinces/Regions.
The final dataset was then processed through the “Spatially Rarefy Occurrence Data for SDMs” tool (set at = 1 km) of the SDMtoolbox 0.9.1 (Brown et al. 2017) for ArcGIS Pro 2.9.3 (Esri Inc. 2023) to make the spatial resolution of both predictors and occurrences comparable, according to Sillero and Barbosa (2021).
Environmental predictors
We used climatic, topographic, and habitat predictors to investigate the ecological needs of the target species. The climatic aspect was assessed by downloading the 19-bioclimatic variables from WorldClim 2.1 (Fick and Hijmans 2017) archive (https://www.worldclim.org/data/) at 30 arc-seconds resolution (~ 1 km) for the “current” scenario (i.e., 1970–2000 average climatic conditions) as well as for three future time projections (i.e., 2030, 2050, and 2070). For each future projection, we downloaded raster data representing predicted climatic conditions under three Shared Socioeconomic Pathways (SSPs). Specifically, we selected the SSPs 2.45, 3.70, and 5.85 to involve all but one (the 1.26, the most optimistic) of the different possible trajectories (Riahi et al. 2017).
As for topography, we downloaded a Digital Elevation Model (DEM, at ~ 90-m resolution) from the European Space Agency and Sinergise (2021) (https://spacedata.copernicus.eu/collections/copernicus-digital-elevation-model). We then used the “Surface Parameters” tool in ArcGIS Pro to calculate the Aspect, starting from the DEM.
About the habitat predictors, we took advantage of the 100-m resolution Global Corine Land Cover map of the Copernicus repository (https://land.copernicus.eu/global/products/lc), which contains a discrete classification with 23 classes according to the UN-FAO Land Cover Classification System (Buchhorn et al. 2020).
Ecological niche modelling
The ecological niche modelling step was performed in R (R Core Team 2013). We built the ENMs with the “gbm” R package (Greenwell et al. 2019). This package implements the gradient boosting model (GBM) algorithm, also known as boosted regression trees (Elith and Leathwick 2009). This algorithm is one of the best-performing ENM algorithms with presence-pseudo-absence data, once properly tuned (Elith and Leathwick 2009; Hao et al. 2020), and the tuning parameters are easy to set in the R environment.
The 19-bioclimatic variables from WorldClim were checked for multicollinearity through the Variance Inflation Factor (using the “vifstep” algorithm of the “usdm” R package (Naimi 2015), with a threshold ≥ 10, as it is deemed a suitable threshold to deal with multicollinearity in ENM (Guisan et al. 2017). To reduce the variability caused by using individual General Circulation Models (GCMs) in future projections (Stralberg et al. 2015), we selected and managed three different GCMs, namely the BCC-CSM2-MR (Wu et al. 2019), the IPSL-CM6A-LR (Boucher et al. 2020), and the MIROC6 (Tatebe et al. 2019).
Then, we generated 10,000 pseudo-absences through the “disk” strategy of the “BIOMOD_FormatingData” function of the “biomod2” R package (Thuiller et al. 2016), setting 1 and 50 km as the minimum and maximum radius, respectively. Then, we weighed presences and pseudo-absences so that the sum of the weights of the previous equals the one of the latter. In fact, it has been demonstrated that assigning the same overall weight to presences and pseudo-absences usually increases ENMs’ predictive performance when the generated pseudo-absences are far more numerous than the available presences (Cerasoli et al. 2017; Gouvêa et al. 2020; Thiault et al. 2020).
The best GBM algorithm parametrization was obtained by creating three different matrices containing several combinations of “gbm” parameters and the respective set of values (for brevity; here, we show the ones for the first matrix only: shrinkage = 0.01, 0.1, 0.3; interaction.depth = 1, 3, 5; n.minobsinnode = 5, 10, 15; bag.fraction = 0.65, 0.8, 1). Then, we ran as many GBM models as the combinations, increasing the n.trees value from 1000 to 15,000 but keeping the train.fraction = 0.8 and the cv.folds = 10 as fixed. Finally, we chose the set of parameters resulting in the lowest root mean square error (RMSE) (Friedman 2001; Greenwell et al. 2019; Cervellini et al. 2021).
Successively, we checked the discrimination power of the optimized GBM model through the Boyce index (Boyce et al. 2002), which is particularly suited for ENMs built on presence and pseudo-absence data (Hirzel et al. 2006; Leroy et al. 2018). Moreover, we measured the relative contribution of the selected variables through the randomization algorithm implemented in the “summary.gbm” function of the “gbm” R package.
Then, we projected the optimized GBM model across the entire study area for both current climatic conditions and various future scenarios represented by the combinations of year (2030, 2050, and 2070) and SSP (SSP3.70 and SSP5.85) by using the GCMs listed above. To assess the uncertainties deriving from future projections calibrated on each GCM and concurrently merge them into 1 year/SSP single model, we first checked for model extrapolation (i.e., the dissimilarity from the calibration conditions) by assessing the Multivariate Environmental Surface Similarity (MESS) (Elith et al. 2010), computed through the function “mess” of the “dismo” package (Hijmans et al. 2023). Then, we used the resulting MESS maps to implement the Multivariate Environmental Dissimilarity Index (MEDI). This index weighs ENMs’ projections under different GCMs based on the corresponding MESS, finally returning a combined weighted projection (Iannella et al. 2017). We repeated this process of ENMs’ fine-tuning for each year × SSP combination. We averaged each future projection between year’s scenarios to obtain a consensus map for 2030, 2050, and 2070. ENMs’ predicted suitability, ranging from 0 to 1 (low-to-high suitability), was then reclassified on a 1-to-10 scale for post-modelling purposes (see below) using the “Reclassify” tool in ArcGIS Pro.
Post-modelling analysis
Predictions from the climate-based ENMs were then refined in a post-modelling phase by including topographic and habitat-related predictors. We thus applied the “couple-and-weigh” framework following Iannella et al. (2021). This process permits to refine models based on a single predictors’ family (in this case, climatic-related variables) by incorporating others. Thus, we selected the topographic and habitat-related variables mentioned above, which are known to influence the common chameleon, as reported by Hódar et al. (2000).
Specifically, we extracted elevation values at occurrence localities from the DEM to obtain an elevation preference curve, converting the “raw” occurrence frequencies of the elevation gradient (from 0 to 500 m, bin size = 50 m) to a 1-to-10 scale. We repeated the same process for the Copernicus habitat predictor. Similarly, we assigned to each habitat category a value from 1 (low suitability) to 10 (high suitability), matching information extracted by occurrence data to the ones of Hódar et al. (2000). Taking advantage of these authors’ findings, we similarly reclassified the Aspect data, to match the common chameleon ecological preferences.
Finally, the “Weighted overlay” tool in ArcGIS Pro was used. This tool merges a given set of rasters, sharing a common evaluation scale, through a weighted averaging process in which each input raster is also assigned a specific percentage set by the operator. Thus, we entered the 1-to-10 information obtained as reported above to let the tool reclassify the supplied rasters for current and future conditions. The predictive performances of the so-obtained weighted models were further reassessed through the Boyce index, considering the presence-only nature of the dataset (Hirzel et al. 2006; Leroy et al. 2018).
To highlight the range shifts in future projections, we used the “BIOMOD_RangeSize” function of the R package “biomod2” (Thuiller et al. 2016). This function uses binarized suitability map of current and future projections, returning maps with areas predicted to be lost, remaining stable, and gained in each future projection. We used the 10th percentile value as a threshold to binaries the current weighted suitability. To detect the direction of the climate-triggered shifts, we used the “Centroid changes” function of the SDMtoolbox 0.9.1 (Brown et al. 2017). Starting from binarized suitability maps for each scenario, this tool provides both the direction and the intensity of the changes in suitability, using the centroids of the study areas. To better understand the possible different population dynamics, we divided the Mediterranean basin in three sections (Western, Central, and Eastern) to focus on the Iberian and North African populations (Western), Italian populations (Central), and Greek and Middle Eastern populations (Eastern).
Landscape connectivity assessment
We assessed landscape connectivity in the areas where the common chameleon was introduced historically, i.e., in the Southern side of the Iberian Peninsula and Southern Italy. We obtained the corresponding resistance surfaces from the current and future weighted suitability maps. First, we downloaded the road and railway layers from Open Street Map (https://www.openstreetmap.org/), selecting the major roads (i.e., motorway, trunk, primary, and secondary) for both study areas. Then, we used the “Mosaic to new raster” function in ArcGIS to merge these layers into the weighted suitability maps. We converted these weighted suitability layers into resistance surfaces using a negative exponential function following Keeley et al. (2016):
In this function, R represents the final resistance value of a pixel, h is the habitat suitability value for the same pixel, and c represents a constant factor determining the curvature of the negative exponential function. Previous studies have demonstrated that moderate values of the constant factor, c, provide the best performance (Keeley et al. 2016). Thus, we set the c factor to 4. We thus obtained high resistance values to the lowest habitat suitability ones, since it is a more accurate representation of landscape resistance, considering that habitat suitability may not be correlated with movement probability and landscape permeability (Keeley et al. 2016; Zeller et al. 2018).
We used Omniscape v.0.5.8 in Julia to compute landscape connectivity (Landau et al. 2021). This algorithm calculates omnidirectional connectivity using circuit theory (McRae et al. 2008, 2016). Omniscape is implemented with a moving window framework, where the center pixel is used as the destination and is set to “ground,” while pixels within the moving window are the sources, and the user could decide whether to use all the pixels as sources or only those with certain resistance values (Landau et al. 2021). As with other circuit theory software, the obtained current map is comparable to the probability of movement of the target species. In particular, Omniscape generates three different outputs: cumulative current flow, potential current flow, and normalized current flow. Considering the poor knowledge of the common chameleon movements, we set the search radius of the moving window to a conservative distance of 1 km.
Results
Occurrence records and ecological niche models
After the filtering procedure, we retained 552 occurrence localities for the common chameleon (Fig. 1).
Occurrence localities for the common chameleon in the study area and IUCN range maps. Data for Greece has been updated based on the latest information from the official page of the Atlas of Reptiles and Amphibians of Greece (Societas Hellenica Herpetologica 2024). All data are referred to the subspecies of interest (C. c. chamaeleon, C. c. recticrista, and C. c. musae)
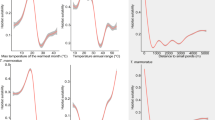
Also, considering the VIF results (Figure S1 Online Resource 1), we selected the following eight of the 19 bioclimatic variables: Bio_6 (minimum temperature of the coldest month), Bio_7 (temperature annual range), Bio_11 (average temperature of the coldest quarter), Bio_12 (annual precipitation), Bio_14 (precipitation of the driest month), Bio_16 (precipitation of the wettest quarter), Bio_18 (precipitation of the warmest quarter), and Bio_19 (precipitation of the coldest quarter).
The lowest RMSE was recorded for the GBM model fitted with: “n.trees” = 4868, “int.depth” = 7, “shrinkage” = 0.001, “bag.fraction” = 0.65, and “minobsinode” = 15. This GBM model obtained a Boyce Index of 0.91. The two most important variables for this model were Bio_12 (Annual Precipitation) (23.6%), peaking between 200 and 500 mm, and Bio_16 (precipitation of wettest quarter) (23.4%), showing a major peak from 250 to 400 mm (Figure S2a and S2b Online Resource 1).
Weighted models and future projections
Habitat preferences show that most occurrences fall within three land use categories: urban areas (22%), shrubland (21%), and cropland (18%), with other categories less represented (Figure S2c Online Resource 1). The elevation preference reports the highest number of occurrences in class 0–50 m and 50–100 m a.s.l. (Figure S2d Online Resource 1).
The weighted model for the current condition, obtained from the “couple-and-weigh” approach, scores a Boyce Index of 0.993 (Figure S3 Online Resource 1), much higher than the one obtained from the ENMs alone (B = 0.796). The current weighted suitability map (Fig. 2a) shows a generally high suitability for areas where the species is currently reported, and even for some territories where the species is not reported (e.g., Sicily). High habitat suitability values are observed in Morocco, Tunisia, and Libya in northern Africa, and between Israel and Lebanon in the Middle East (Fig. 2a). Moreover, Cyprus and the coastal zones and the islands in the Aegean Sea are reported as generally suitable for the common chameleon (Fig. 2a). Considering the countries where the species was introduced, the Iberian Peninsula shows high, continuous habitat suitability values from Murcia in Spain to the southern limit of the Algarve region in Portugal (Fig. 2a). On the other hand, in Italy, Sicily appears to be the most suitable region in the current environmental conditions (Fig. 2a).
In the future scenarios, a general and progressive increase in habitat suitability is expected in all the areas currently inhabited by the species, except for some areas in Tunisia and Turkey (Fig. 3a). Interestingly, in the Iberian Peninsula, the suitable areas would progressively shift westwards, resulting in a scenario where the Mediterranean side of the Iberian Peninsula would lose most of the suitability, in turn shifting towards the north-western Portugal (Fig. 3a). In Italy, habitat suitability is predicted to increase in Sicily, Calabria, Apulia, and Sardinia (Fig. 3a). Some losses are observed in Aegean coast of Turkey and Israel.
In detail, the range shifts calculated upon the different future inferred scenarios differ among the three Mediterranean basin sectors considered (Fig. 3b). In fact, the western and eastern parts of the study area are predicted to change mainly westwards, while the central sector, involving Tunisia and Italy, shows northeastern changes (Fig. 3b). The reduction in habitat suitability in the Mediterranean side of the Iberian Peninsula and the suitable areas “gained” in Portugal are evident (Fig. 3a).
Landscape connectivity assessment
The connectivity assessment as computed from Omniscape for the common chameleon in the current Iberian range shows high connectivity values from the Algarve region to the city of Cadiz (Fig. 4a). Furthermore, fair connectivity values occur from Gibraltar to Malaga (Fig. 4a). However, the eastern Iberian occurrences from Almeria to Murcia are less connected to the other populations, with few landscape corridors between them (Fig. 4a).
When considering the future scenarios, the standardized connectivity change index maps show an interesting pattern, with connectivity values increasing in the western area, northward across Portugal, and in the Algeciras area, but with a severe progressive decline in the Mediterranean Iberian Peninsula from Almeria to Murcia (Fig. 4b). Moreover, some reductions of connectivity occur between Algarve and Gibraltar, thus disrupting the continuous connection between these areas found for current conditions (Fig. 4b).
In southern Italy, a more complex pattern of connectivity emerges, with local high connectivity in the areas where the species currently occurs, such as the “Costa Viola” area in Calabria and in the southern side of the Apulia region (Fig. 5a). Moreover, high connectivity values are observed in the southern end and the eastern area of Calabria overlooking the Ionian Sea (Fig. 5a).
a Landscape connectivity in southern Italy where the common chameleon occurs with two separated populations in Calabria and Apulia regions and b standardized connectivity change index showing the change in connectivity (red-to-green color scale corresponding to loss and gain) between each future scenario and the current conditions
Analyzing future changes in connectivity through the SCCI, losses are clustered in the northern part of the Apulia region and in the eastern area of Calabria, where the connectivity predicted in the current scenario is lost (Fig. 5b). However, connectivity is forecasted to increase in the areas where the common chameleon is currently distributed (i.e., in the southern part of Apulia and of Calabria), where connectivity increases towards the hinterland (Fig. 5b).
Discussion
Several factors may positively influence the establishment of a new species when introduced to a novel environment, such as a long time since introduction, a high frequency of introduction events, minimal latitudinal differences between native and introduced ranges, and specific species characteristics, like phenotypic attractiveness, larger native range size, and high fecundity (Mahoney et al. 2015).
Some of these factors are pertinent to the case of the common chameleon Chamaeleo chamaeleon. The attractiveness of this species, attributed to its compact size and skilled camouflage abilities, has played a significant role in its introduction outside its natural habitat. Over time, it has been introduced to regions such as the Iberian Peninsula, Italy, Malta, and potentially to some Greek islands, where they have successfully established (Paulo et al. 2002; Dimaki et al. 2008; Sindaco et al. 2008; Andreone et al. 2016; Basso et al. 2019). However, it has also been reported in other areas, such as several northern and central regions of Italy, where no viable populations established (Bologna et al. 2000; Corti et al. 2011). This underscores the significance of smaller latitudinal differences and similar climatic conditions, which could facilitate the establishment of a species outside its native range (Mahoney et al. 2015).
Despite being classified as a species of interest in Annex IV of the EU Habitat and Species Directive (92/43/CE), strictly protected in Annex II of the Bern Convention, and assigned the highest level (C1) in the CITES Convention (3626/82/CE), the common chameleon remains poorly studied and comprehensive information guiding management actions is scarce. Existing literature offers limited insights into habitat preferences, only focusing on specific sub-regions within its range. Moreover, precise information on the species’ current distribution is lacking, with studies confined to Morocco and parts of the Iberian Peninsula (Miraldo et al. 2005; Qninba et al. 2013).
In this context, our results, which were obtained using ecological niche modelling to study the habitat suitability of the common chameleon in the Mediterranean basin, have allowed us to evaluate its possible future range expansions. Also, we assessed habitat connectivity in the areas where the species was introduced to explore how the expansion dynamics could proceed.
The climate-based ENMs identified two pivotal variables: annual mean precipitation (Bio_12) and precipitation of the wettest quarter (Bio_16). The first one reflects the total water input, offering insights about tree abundance or net primary production, concurrently indicating the wetness (or aridity) of an area. Bio_16 gives crucial information about the species’ seasonal distribution. Notably, the importance of the precipitation of the wettest quarter has been consistent across ecological niche modelling studies on reptiles, as observed by Gadsden et al. (2012) and Farashi and Alizadeh-Noughani (2021). The prominence of precipitation as a key variable aligns with the “hypothesis of water-energy dynamics,” which posits that precipitation plays a critical role, particularly for species inhabiting low altitudes (Qian 2010).
The weighted suitability maps reveal areas with high suitability values within the species’ range (see Figs. 1a and 2). Remarkably, some areas with high suitability values lack corresponding records in our ecological niche modelling dataset (and highlights that low overfit occurred during model calibration). This mismatch holds particular significance for a species, like the common chameleon, where an ascertained knowledge of its historical and current distribution is limited, especially in certain countries. The insights derived from our species distribution models could serve as a tool to guide research efforts towards areas with a higher likelihood of species’ presence, mirroring what occurred in other cases (De Siqueira et al. 2009; Fois et al. 2018). Notably, the regions of high suitability are mainly clumped in coastal areas and their surroundings.
Our results indicate an impending expansion in certain parts of the common chameleon’s range, coupled with a decline in others, predominantly following a northward latitudinal gradient. Specifically, habitat suitability will likely follow a latitudinal shift, with a northwestern shift for the western and eastern Mediterranean populations, and a northeastern shift for central ones. This directional and long-range shift aligns with patterns reported in other ecological modelling studies, where a prevailing northwestern displacement results for the Saharan-Arabic region, and a northeastern shift occurs in the Palearctic (Araújo et al. 2006; Iannella et al. 2020; Biber et al. 2023). These projected changes are consistent with the ectothermic nature of reptiles, a physiological feature which makes them particularly susceptible to the impacts of climate change (Diele-Viegas and Rocha 2018).
Careful consideration is necessary for the eastern Iberian populations because, standing to our projections, a potential reduction in habitat connectivity could progressively occur. This trend could contribute to the isolation of these eastern populations from their south-central counterparts. Conversely, populations in southern Portugal should be monitored to evaluate the possibility of expansion.
In the Italian peninsula, the habitat connectivity maps reveal an increase in the Calabria and Apulia regions but a decrease in other areas. Notably, the coastal strip towards Campania exhibits high connectivity values, suggesting unhindered connectivity. However, considering the scarce dispersal capacity of the common chameleon and the small population size, this scenario may be unlikely to occur naturally. Indeed, the human-mediated illegal trade could act rather randomly, posing even more uncertainties to the already-tangled Italian scenario.
Coastal distributions exhibit a positive relationship with common chameleon predicted shifts, reinforcing the outcomes indicated by Weil et al. (2022). In fact, its body size, life-history traits, and preferred habitat type (coastal ones) make this species particularly suited for dispersal in these environments, even though human-mediated shifts may change the natural processes. Notably, natural dispersal movements identified in chameleons tend to be continental (Weil et al. 2022).
Our study’s findings on habitat preferences align with the established habitat selection patterns of the common chameleon, as documented by Hódar et al. (2000), indicating a higher abundance in anthropized and cultivated areas. This unique habitat selection behavior implies that the common chameleon may be particularly susceptible to specific threats, such as exposure to pesticides, road kills, and illegal collections (Hódar et al. 2000; Albaba 2017). The identification of dispersal corridors holds potential significance for species conservation efforts. For instance, it could aid in preventing road kills in areas where high connectivity is detected and contribute to understanding expansion dynamics, especially for management purposes in regions where the species has recently been reported, as seen for the Calabria region. By concurrently considering both habitat suitability and connectivity, conservationists can focus on areas predicted to be both suitable and connected. However, it is crucial to note that, for certain management actions, the identified dispersal corridors should be validated with empirical data. Previous studies utilizing circuit-theory techniques and validated with empirical data have demonstrated the efficacy of theoretical predictions in identifying dispersal corridors (McClure et al. 2016) and enhancing management measures. Indeed, specific investigations are necessary given that, even today, individuals are still being taken from the wild, possibly leading the species to expand into areas beyond its native range.
Considering the heterogeneous outcomes in terms of potential distribution, landscape connectivity and expansion dynamics, it is essential to deepen our understanding of the common chameleon’s putative impact in recently colonized areas and to ascertain its updated distribution, population size, and movements.
Availability of data and materials
All the datasets and the spatial data generated during the current study are available from the corresponding author (viviana.cittadino@graduate.univaq.it) on reasonable request.
References
Albaba I (2017) Surveying wildlife roadkills in the West Bank Governorates-Palestine. J Entomol Zool Stud 5:910–913
Andreone F, Angelici F, Carlino P et al (2016) The common chameleon Chamaeleo chamaeleon in southern Italy: evidence for allochthony of populations in Apulia and Calabria (Reptilia: Squamata: Chamaeleonidae). Ital J Zool 83:372–381
Araújo MB, Thuiller W, Pearson RG (2006) Climate warming and the decline of amphibians and reptiles in Europe. J Biogeogr 33:1712–1728
Barbet-Massin M, Jiguet F, Albert CH, Thuiller W (2012) Selecting pseudo-absences for species distribution models: how, where and how many? Methods Ecol Evol 3:327–338
Basso R, Calasso C (1991) I rettili della penisola salentina. Contributi alla conoscenza dell’ambiente e della fauna salentina. Quaderni del Museo Civico di Storia Naturale del Salento 1–63
Basso R, Vannuccini ML, Nerva L et al (2019) Multiple origins of the common chameleon in southern Italy. Herpetozoa 32:11–19. https://doi.org/10.3897/herpetozoa.32.e35611
Biber MF, Voskamp A, Hof C (2023) Potential effects of future climate change on global reptile distributions and diversity. Glob Ecol Biogeogr 32:519–534
Blackburn TM, Essl F, Evans T et al (2014) A unified classification of alien species based on the magnitude of their environmental impacts. PLoS Biol 12:e1001850. https://doi.org/10.1371/journal.pbio.1001850
Blackburn TM, Pyšek P, Bacher S et al (2011) A proposed unified framework for biological invasions. Trends Ecol Evol 26:333–339
Bologna MA, Capula M, Carpaneto GM (2000) Anfibi e rettili del Lazio. Palombi Editori, p 160
Boucher O, Servonnat J, Albright AL et al (2020) Presentation and evaluation of the IPSL-CM6A-LR climate model. J Adv Model Earth Syst 12:e2019MS002010
Boyce MS, Vernier PR, Nielsen SE, Schmiegelow FKA (2002) Evaluating resource selection functions. Ecol Model 157:281–300. https://doi.org/10.1016/S0304-3800(02)00200-4
Broennimann O, Treier UA, Müller-Schärer H et al (2007) Evidence of climatic niche shift during biological invasion. Ecol Lett 10:701–709
Brown JL, Bennett JR, French CM (2017) SDMtoolbox 2.0: the next generation python-based GIS toolkit for landscape genetic, biogeographic and species distribution model analyses. PeerJ 5:e4095
Buchhorn M, Lesiv M, Tsendbazar N-E et al (2020) Copernicus global land cover layers—collection 2. Remote Sens 12:1044
Burridge MJ, Simmons L-A, Allan SA (2000) Introduction of potential heartwater vectors and other exotic ticks into Florida on imported reptiles. J Parasitol 86:700–704
Carpenter AI, Rowcliffe JM, Watkinson AR (2004) The dynamics of the global trade in chameleons. Biol Cons 120:291–301
Cerasoli F, Iannella M, D’Alessandro P, Biondi M (2017) Comparing pseudo-absences generation techniques in Boosted Regression Trees models for conservation purposes: a case study on amphibians in a protected area. PLoS ONE 12:e0187589
Cervellini M, Di Musciano M, Zannini P et al (2021) Diversity of European habitat types is correlated with geography more than climate and human pressure. Ecol Evol 11:18111–18124. https://doi.org/10.1002/ece3.8409
Corti C, Capula M, Luiselli L et al (2011) Fauna d’Italia, vol. XLV. Reptilia, Calderini, Bologna, XII, p 869
De Siqueira MF, Durigan G, de Marco JP, Peterson AT (2009) Something from nothing: using landscape similarity and ecological niche modeling to find rare plant species. J Nat Conserv 17(1):25–32
Di Giuseppe M (2013) Use of intramedullary pin for humeral fracture in a Chamaleo chamaleon. Natura Rerum 3:63–69
Diele-Viegas LM, Rocha CFD (2018) Unraveling the influences of climate change in Lepidosauria (Reptilia). J Therm Biol 78:401–414
Dimaki M, Hundsdörfer A, Fritz U (2008) Eastern Mediterranean chameleons (Chamaeleo chamaeleon, Ch. africanus) are distinct. Amphib Reptilia 29:535–540. https://doi.org/10.1163/156853808786230415
Dorcas ME, Willson JD, Reed RN et al (2012) Severe mammal declines coincide with proliferation of invasive Burmese pythons in Everglades National Park. Proc Natl Acad Sci 109:2418–2422
Elith J, Kearney M, Phillips S (2010) The art of modelling range-shifting species. Methods Ecol Evol 1:330–342
Elith J, Leathwick JR (2009) Species distribution models: ecological explanation and prediction across space and time. Annu Rev Ecol Evol Syst 40:677–697
Esri Inc. (2023) ArcGIS Pro 3.2.2 – ESRI, Redlands, California
European Space Agency, Sinergise (2021) Copernicus global digital elevation model. Distributed by OpenTopography. https://doi.org/10.5069/G9028PQB
Farashi A, Alizadeh-Noughani M (2021) Predicting the invasion risk of non-native reptiles as pets in the Middle East. Global Ecol Conserv 31:e01818. https://doi.org/10.1016/j.gecco.2021.e01818
Fattizzo T, Marzano G (2002) Dati distributivi sull’erpetofauna del Salento. Thalassia Salentina 26:113–132
Fick SE, Hijmans RJ (2017) WorldClim 2: new 1-km spatial resolution climate surfaces for global land areas. Int J Climatol 37:4302–4315
Fois M, Cuena-Lombraña A, Fenu G, Bacchetta G (2018) Using species distribution models at local scale to guide the search of poorly known species: review, methodological issues and future directions. Ecol Model 385:124–132
Friedman JH (2001) Greedy function approximation: a gradient boosting machine. Ann Stat 29:1189–1232. https://doi.org/10.1214/aos/1013203451
Gadsden H, Ballesteros-Barrera C, de la Garza OH et al (2012) Effects of land-cover transformation and climate change on the distribution of two endemic lizards, Crotaphytus antiquus and Sceloporus cyanostictus, of northern Mexico. J Arid Environ 83:1–9
GBIF (2023) GBIF.org (07 March 2023) GBIF Occurrence Download. https://doi.org/10.15468/dl.nsrtkk
Gouvêa LP, Assis J, Gurgel CFD et al (2020) Golden carbon of Sargassum forests revealed as an opportunity for climate change mitigation. Sci Total Environ 729:138745. https://doi.org/10.1016/j.scitotenv.2020.138745
Greenwell B, Boehmke B, Cunningham J, Developers G (2019) gbm: generalized boosted regression models. R Package Version 2:37–40
Guisan A, Thuiller W (2005) Predicting species distribution: offering more than simple habitat models. Ecol Lett 8:993–1009
Guisan A, Thuiller W, Zimmermann NE (2017) Habitat suitability and distribution models: with applications in R. Cambridge University Press
Hao T, Elith J, Lahoz-Monfort JJ, Guillera-Arroita G (2020) Testing whether ensemble modelling is advantageous for maximising predictive performance of species distribution models. Ecography 43:549–558
Hijmans RJ, Phillips S, Elith JLJ (2023) dismo: species distribution modeling. R package version 1.3-3. Retrieved from https://CRAN.R-project.org/package=dismo
Hirzel AH, Le Lay G, Helfer V et al (2006) Evaluating the ability of habitat suitability models to predict species presences. Ecol Model 199:142–152
Hódar JA, Pleguezuelos JM, Poveda JC (2000) Habitat selection of the common chameleon (Chamaeleo chamaeleon L.) in an area under development in southern Spain: implications for conservation. Biol Cons 94:63–68
Iannella M, Cerasoli F, Biondi M (2017) Unraveling climate influences on the distribution of the parapatric newts Lissotriton vulgaris meridionalis and L. italicus. Front Zool 14:1–14
Iannella M, D’Alessandro P, Biondi M (2020) Forecasting the spread associated with climate change in Eastern Europe of the invasive Asiatic flea beetle, Luperomorpha xanthodera (Coleoptera: Chrysomelidae). Eur J Entomol 117:130–138
Iannella M, Console G, Cerasoli F et al (2021) A step towards SDMs: a “couple-and-weigh” framework based on accessible data for biodiversity conservation and landscape planning. Divers Distrib 27:2412–2427
Jeschke JM, Keesing F, Ostfeld RS (2013) Novel organisms: comparing invasive species, GMOs, and emerging pathogens. Ambio 42:541–548
Jeschke JM, Bacher S, Blackburn TM et al (2014) Defining the impact of non-native species. Conserv Biol 28:1188–1194. https://doi.org/10.1111/cobi.12299
Karesh WB, Cook RA, Bennett EL, Newcomb J (2005) Wildlife trade and global disease emergence. Emerg Infect Dis 11:1000
Keeley ATH, Beier P, Gagnon JW (2016) Estimating landscape resistance from habitat suitability: effects of data source and nonlinearities. Landscape Ecol 31:2151–2162. https://doi.org/10.1007/s10980-016-0387-5
Kraus F (2015) Impacts from invasive reptiles and amphibians. Annu Rev Ecol Evol Syst 46:75–97
Landau VA, Shah VB, Anantharaman R, Hall KR (2021) Omniscape. jl: software to compute omnidirectional landscape connectivity. J Open Source Softw 6:2829
Leroy B, Delsol R, Hugueny B et al (2018) Without quality presence–absence data, discrimination metrics such as TSS can be misleading measures of model performance. J Biogeogr 45:1994–2002. https://doi.org/10.1111/jbi.13402
Luiselli L, Bonnet X, Rocco M, Amori G (2012) Conservation implications of rapid shifts in the trade of wild African and Asian pythons. Biotropica 44:569–573
Mahoney PJ, Beard KH, Durso AM et al (2015) Introduction effort, climate matching and species traits as predictors of global establishment success in non-native reptiles. Divers Distrib 21:64–74. https://doi.org/10.1111/ddi.12240
Mainali KP, Warren DL, Dhileepan K et al (2015) Projecting future expansion of invasive species: comparing and improving methodologies for species distribution modeling. Glob Change Biol 21:4464–4480
McClure ML, Hansen AJ, Inman RM (2016) Connecting models to movements: testing connectivity model predictions against empirical migration and dispersal data. Landscape Ecol 31:1419–1432
McRae BH, Dickson BG, Keitt TH, Shah VB (2008) Using circuit theory to model connectivity in ecology, evolution, and conservation. Ecology 89:2712–2724
McRae B, Popper K, Jones A et al (2016) Conserving nature’s stage: mapping omnidirectional connectivity for resilient terrestrial landscapes in the Pacific Northwest. The Nature Conservancy, Portland, Oregon. Available from http://nature.org/resilienceNW. Accessed Dec 2023
Miraldo A, Pinto I, Pinheiro J et al (2005) Distribution and conservation of the common chameleon (Chamaeleo chamaeleon) in Algarve, southern Portugal. Isr J Zool 51:157–164. https://doi.org/10.1560/EV2Y-9E2F-5DLY-P00N
Naimi B (2015) USDM: Uncertainty analysis for species distribution models. R package version 2.1-7. Retrieved from https://cran.r-project.org/web/packages/usdm/usdm.pdf
Nowak M (2010) The international trade in reptiles (Reptilia)—the cause of the transfer of exotic ticks (Acari: Ixodida) to Poland. Vet Parasitol 169:373–381
Paulo O, Pinto I, Bruford MW et al (2002) The double origin of Iberian peninsular chameleons. Biol J Lin Soc 75:1–7. https://doi.org/10.1046/j.1095-8312.2002.00002.x
Pellegrino F, Albornoz G, Bernabò I, Iantorno A, Mazza M, Sperone E, Stepancich D, Tripepi S (2016) Prima caratterizzazione di una popolazione naturalizzata di camaleonte comune (Chamaeleo chamaeleon) in Calabria. In: Menegon M, Rodriguez-Prieto A, Deflorian MC (eds) Atti XI Congresso Nazionale della Societas Herpetologica Italica, Trento, 22–25 September 2016. 147–150
Pleguezuelos J (1997) Chamaeleo chamaeleon (Linnaeus, 1758) Camaleón común, Camaleão. Distribución y Biogeografía de los anfibios y reptiles en España y Portugal. Asociación Herpetológica Española – Universidad de Granada, Spain 190–192
Qian H (2010) Environment–richness relationships for mammals, birds, reptiles, and amphibians at global and regional scales. Ecol Res 25:629–637
Qninba A, Radi M, Amezian M et al (2013) Nouvelle limite méridionale pour le Caméléon commun Chamaeleo chamaeleon (Reptilia, Chamaeleonidae) au Maroc. Bull Soc Herp Fr 145:199–204
R Core Team R (2023) R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. https://www.R-project.org/
Razzetti E, Sindaco R (2006) Unconfirmed taxa or in need of confirmation/Taxa non confermati o meritevoli di conferma. Atlante degli Anfibi e dei Rettili d’Italia/Atlas of Italian Amphibians and Reptiles. Societas Herpetologica Italica, Edizioni Polistampa, Firenze, pp 643–653
Reed RN (2005) An ecological risk assessment of nonnative boas and pythons as potentially invasive species in the United States. Risk Analysis: an International Journal 25:753–766
Riahi K, Van Vuuren DP, Kriegler E et al (2017) The shared socioeconomic pathways and their energy, land use, and greenhouse gas emissions implications: an overview. Glob Environ Chang 42:153–168
Ricciardi A, Hoopes MF, Marchetti MP, Lockwood JL (2013) Progress toward understanding the ecological impacts of nonnative species. Ecol Monogr 83:263–282
Serva D, Iannella M, Cittadino V, Biondi M (2023) A shifting carnivore’s community: habitat modeling suggests increased overlap between the golden jackal and the Eurasian lynx in Europe. Front Ecol Evol 11:1165968
Sillero N, Barbosa AM (2021) Common mistakes in ecological niche models. Int J Geogr Inf Sci 35:213–226. https://doi.org/10.1080/13658816.2020.1798968
Sindaco R, Jeremčenko VK, Venchi A, Grieco C (2008) The reptiles of the Western Palearctic: annotated checklist and distributional atlas of the turtles, crocodiles, amphisbaenians and lizards of Europe, North Africa, Middle East and Central Asia. Vol. 1. Latina: Edizioni Belvedere, p 580
Societas Hellenica Herpetologica (2024) Chamaleo chamaleon, Herpatlas, Atlas of Reptiles and Amphibians of Greece. Available at http://herpatlas.gr/herp-finder/chamaeleo-chamaeleon/. Accessed 16 Feb 2024
Sperone E, Crescente A, Brunelli E, Paolillo G, Tripepi S (2010) Sightings and successful reproduction of allochthonous reptiles in Calabria. Acta Herpet 5(2):265–273
Stralberg D, Matsuoka S, Hamann A et al (2015) Projecting boreal bird responses to climate change: the signal exceeds the noise. Ecol Appl 25:52–69
Tatebe H, Ogura T, Nitta T et al (2019) Description and basic evaluation of simulated mean state, internal variability, and climate sensitivity in MIROC6. Geosci Model Dev 12:2727–2765
Thiault L, Weekers D, Curnock M et al (2020) Predicting poaching risk in marine protected areas for improved patrol efficiency. J Environ Manage 254:109808. https://doi.org/10.1016/j.jenvman.2019.109808
Thuiller W, Georges D, Engler R (2016) biomod2: ensemble platform for species distribution modeling. Retrieved from https://cran.r-project.org/web/packages/biomod2/biomod2.pdf
Weil S, Gallien L, Lavergne S et al (2022) Chameleon biogeographic dispersal is associated with extreme life history strategies. Ecography 2022:e06323
Wu T, Lu Y, Fang Y et al (2019) The Beijing Climate Center climate system model (BCC-CSM): the main progress from CMIP5 to CMIP6. Geosci Model Dev 12:1573–1600
Zeller KA, Jennings MK, Vickers TW et al (2018) Are all data types and connectivity models created equal? Validating common connectivity approaches with dispersal data. Divers Distrib 24:868–879
Funding
Open access funding provided by Università degli Studi dell’Aquila within the CRUI-CARE Agreement.
Author information
Authors and Affiliations
Contributions
DS and VC collected the data; DS, VC, and MI analyzed the data; IB and MB provided the scope and guidance, DS authored the first draft of the manuscript; all authors wrote the article and gave final approval for submission.
Corresponding author
Ethics declarations
Ethical approval
No approval of research ethics committees was required to accomplish the goals of this study.
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Serva, D., Cittadino, V., Bernabò, I. et al. Habitat suitability and connectivity modelling predict a latitudinal-driven expansion in the Mediterranean basin for a historically introduced reptile. Eur J Wildl Res 70, 27 (2024). https://doi.org/10.1007/s10344-024-01780-9
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10344-024-01780-9