Abstract
Overexpression of the GA20-OXIDASE gene under control of the constitutive cauliflower mosaic virus 35S promoter in poplar leads to increased shoot growth and biomass production, however, the trees suffer from unstable shoots and poor root growth. Transgenic hybrid poplar (Populus tremula L. × P. alba L.) plants overexpressing the GA20-OXIDASE gene from pine under control of a poplar-xylem-specific DX15-promoter also revealed a superior effect on growth and biomass production but without changing the overall phenotype. We tested seven DX15::GA20-OXIDASE-transgenic lines for growth and biomass production in the glasshouse in 2017, and repeated the experiment in 2018 with the “best-four” lines. Plants from one DX15::GA20-OXIDASE-transgenic line, N457‑4, turned out to be the tallest, with highest biomass, in both years under investigation. In contrast to the other lines tested in 2017 and 2018 carrying two or more copies of the transgene, N457‑4 carries only one copy. We suggest that transcriptional or post-transcriptional transgene silencing in the two- or more copies carrying lines might be responsible for lower GA20-OXIDASE transgene expression and that the single-copy-line N457‑4 has stable GA20-OXIDASE-gene expression.
Zusammenfassung
Die Überexpression des GA20-OXIDASE-Gens unter der Kontrolle des konstitutiven 35S-Promotors des Blumenkohlmosaikvirus in Pappeln führt zu einem erhöhten Sprosswachstum und einer erhöhten Biomasseproduktion. Jedoch leiden die Bäume unter instabilen Sprossen und schlechtem Wurzelwachstum. Transgene Hybridpappelpflanzen (Populus tremula L. × P. alba L.), die das GA20-OXIDASE-Gen aus der Kiefer unter der Kontrolle eines Pappel-Xylem-spezifischen DX15-Promotors überexprimieren, zeigten ebenfalls eine erhöhtes Sprosswachstum und Biomasseproduktion, jedoch ohne den Gesamtphänotyp zu verändern. Wir haben 2017 sieben DX15::GA20-OXIDASE-transgene Linien in Hinblick auf Wachstum und Biomasseproduktion im Gewächshaus getestet und das Experiment 2018 mit den „besten-vier“-Linien wiederholt. Pflanzen der DX15::GA20-OXIDASE-transgenen Linie N457‑4 erwiesen sich in beiden Untersuchungsjahren als die mit größten Sprosswachstum und mit der höchsten Biomasse. Im Gegensatz zu den anderen 2017 und 2018 getesteten Linien, die zwei oder mehr Kopien des Transgens trugen, beinhaltet N457‑4 nur eine Kopie des Gens. Wir vermuten, dass transkriptionales oder posttranskriptionales Transgen-Silencing in den zwei oder mehr Kopien-tragenden Linien für eine geringere GA20-OXIDASE-Transgenexpression verantwortlich ist und dass die Linie N457‑4 mit nur einer Kopie eine stabile GA20-OXIDASE-Genexpression aufweist.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Plant growth and development is one of the most fascinating aspects of developmental biology. The diversity of the plant habitus ranges from small and ground-creeping to upright and tall growth between different plant species (Linné 1764). Within a species, plant habitus can also be very flexible depending on environmental conditions (Darwin 1897), e.g., a plant is tall when growing at low elevation, and dwarf and creepy in the mountains. Various environmental stimuli like gravity, light, water, and touch, also called plant tropisms, help plants to adapt to environmental changes (Chelakkot and Mahadevan 2017). Plant tropisms influence plant shoot and root growth in the way that they grow towards or away from such a stimulus (Gilroy 2008).
Shoot growth is strongly controlled by the status, distribution and relative ratios of specific plant hormones, e.g., cytokinins promote cell division, and auxins and gibberellins promote shoot elongation (Santner et al. 2009; Davies 2012). Factors like plant age, dormancy, drought, and temperature, but also externally applied hormones or hormone inhibitors, influence endogenous hormone levels.
One of the most famous examples shoot alteration is the effect of chlorocholine chloride (CCC) on the growth and yield of spring wheat in the field (Humphries et al. 1965). The effect of CCC can partly be reverted by the application of gibberellic acid (GA) (Dyson 1965). Besides other effects on seed germination, leaf development and transition from vegetative to flowering, bioactive GA has well-known effects on shoot growth and elongation, in particular on increased internode extension (Brian 1959; Ram and Mehta 1978; Gupta and Chakrabarty 2013). The phenotype of many dwarf mutants can be reverted by external application of GA leading to plants not distinguishable from that of the tall phenotype (Brian 1959).
Shoot growth is, in particular, of high interest for forest trees because of woody biomass production (Hansen 1991; Scarascia-Mugnozza et al. 2000) and cellulose fibre characteristics for the pulp and paper industry (Doblin et al. 2002; Somerville 2006). A classical approach to modify shoot growth and cellulose fibre length is to change hormone content, mainly by application of GA (Wang et al. 2017). Classical breeding has a limited application value to genetically improve forest trees in the mentioned characteristics because of the long generation cycles of many forest tree species, ranging from one to several decades (Flachowsky et al. 2009). Genetic engineering and genome editing are attractive tools for the modification of wood and other tree components (Strauss et al. 2001; Campbell et al. 2003; Bruegmann et al. 2019a), but are, however, not always feasible due to the lack of genetic transformation protocols for most forest trees. Therefore, members of the genus Populus have become “model tree” species, e.g., because of availability of a complete genome sequences, genetic maps, and the ease of genetic engineering (Taylor 2002).
As early as at the beginning of this century, Eriksson et al. (2000) described the transfer of the GIBBERELLIC ACID 20 OXIDASE (GA20-OXIDASE) into poplar via genetic engineering. The authors showed that transgenic poplar plants constitutively overexpressing the GA20-OXIDASE gene revealed increased growth and biomass production, and longer xylem fibre lengths. These results could be confirmed in Arabidopsis by Biemelt et al. (2004) and poplar by Dünisch et al. (2006), Fladung (2006), and Jeon et al. (2016), however, also negative effects on shoot stability and on root formation were reported. To overcome the problems of unstable shoots, Jeon et al. (2016) placed the GA20-OXIDASE gene from Pinus densiflora (PdGA20ox1) under the control of the xylem-specific DX15 promoter from poplar and transferred this gene construct into the hybrid poplar clone BH (P. alba × P. tremula var. glandulosa). Transgenic plants still revealed high increases in biomass and altered cell wall composition but with a quite normal phenotype (Jeon et al. 2016). Alternatively, Fladung (2018) showed that mixoploid GA20-OXIDASE overexpressing transgenic poplar still revealed increased growth but relatively thicker shoots compared to diploid GA20-OXIDASE transgenic poplar.
In this paper, we used the DX15::PdGA20-OXIDASE gene construct (Jeon et al. 2016) and transformed the hybrid poplar clone 717-1B4 (Populus tremula L. × P. alba L.). Various independent transgenic lines were obtained and transferred to the glasshouse and plant height and shoot diameter were recorded monthly from April to September (2017), and April to November (2018). At the end of the experiments, fresh-and dry weight of shoots were determined and compared to non-transgenic controls. The aim of the study was to highlight that only the DX15::PdGA20-OXIDASE transgenic line revealing just one copy of the transgene show increased shoot height and biomass production in the two years, 2017 and 2018, of morphological investigations in the glasshouse.
Material and Methods
Plant Material and Poplar Transformation
A leaf disc co-cultivation method was used for Agrobacterium-mediated transformation of Populus × canescens clone INRA 717-1B4 (P. tremula L. × P. alba L.; Leplé et al. 1992) (in the following called “P1”) as described in Fladung et al. (1996, 1997). The Agrobacterium tumefaciens strain C58 carrying the PdGA20-OXIDASE gene from Pinus densiflora under control of the developing xylem (DX)-specific DX15-promoter from hybrid poplar (Ko et al. 2012) was used for genetic leaf-disc transformation. The bacterial strain was kindly provided by Hyung-Woo Jeon (Prof. J. H. Ko Lab., Department of Plant & Environmental New Resources, Kyung Hee University, Yongin, Korea) and described in detail in Jeon et al. (2016). The plasmid also carries the HYGROMYCIN B PHOSPHOTRANSFERASE (HPT) gene leading to hygromycin (hyg) resistance (Jeon et al. 2016). The regeneration media (Woody Plant Medium [WPM, Duchefa, Haarlem, The Netherlands; M0220] supplemented with 0.01% Pluronics F‑68 (Sigma P‑7061), thidiazuron [0.01 μM]) contained hygromycin (20 mg/l) for transgenic plant selection, and cefotaxime (500 mg/l) for bacterial removal. Plates were cultivated under in vitro conditions at 23–24 °C, 24 h constant light, and 8–10 µE m−2 s−1 irradiation.
Two transformation experiments were performed with P1 by applying the A. tumefaciens strain carrying the DX15::PdGA20-OXIDASE gene construct (N456, N457). Independent hyg-resistant transgenic lines putatively containing the PdGA20ox gene were termed as N456‑X and N457‑X. In this study, growth behavior of the transgenic lines N456‑1, N457‑4, -5, -6, -12, -13, and -14 (2017) and of N457‑4, -6, -12, and -14 (2018), in comparison to the non-transgenic control P1, were investigated in detail in the glasshouse under natural daylight and temperature conditions (with open windows).
Plant Cultivation and Growth Conditions
Putative transgenic plants carrying the DX15::PdGA20-OXIDASE gene construct obtained after four to eight weeks culture on hyg-containing regeneration medium were propagated in vitro on WPM medium without hormones at 25 °C and with continuous light. Regenerated plants from the poplar hybrid clone P1 derived from tissue culture were used as non-transgenic controls. Following in vitro culture, plants of about 5 to 10 cm in height were potted into soil (Container substrate with clay (No. 2008), Heinrich Harden, Hamburg, Germany), transferred to growth chambers (light period: 16/8 [day/night], light intensity: 300 µE m−2 sec−1, lamps: Phillips TLM 140W/33RS, relative humidity: 70%, temperature: 22/19 °C, recorded with thermal hydrograph), fully covered by plastic lids and cultivated in boxes at 25 °C/16 °C and 16/8 h day/night cycle. For acclimatization to ambient air conditions (relative humidity of air: 70%), lids were opened daily for increasing time periods over one week. Following acclimatization, plants were watered daily and cultivated for 2 to 4 weeks in the growth chamber. At age of about 4 to 6 weeks, plants were transferred into the glasshouse (min.–mean–max. air temperature, April to October: 15–19–24 °C, November to March: 8–17–21 °C, recorded with thermal hydrograph). During the growing season, the plants were cultivated in pots with sizes of 13 cm, watered daily, and supplemented one to two times per week with mineral fertilizer (Universol Blue, Everris, Nordhorn, Germany).
Morphological Investigations of Glasshouse Plants
Total height and diameter of the main shoot (5 cm above soil) were measured from all plants cultivated in glasshouse (2017: 14 to 16 plants per line, 2018: 13 to 15 plants per line) of the seven independent transgenic lines N456‑1, N457‑4, -5, -6, -12, -13, and -14 (2017) and of N457‑4, -6, -12, and -14 (2018) once per month from April to September (2017) and April/May to November (2018). After the last measurements, stems were cut short just above ground and defoliated. Leafless stems were segmented into 20 cm pieces and stem fresh weight was determined. The stem segments were then placed into a drying cabinet at 105 °C for 16 h. To assess dry weight, dried stem segments were adapted to ambient air conditions and subsequently weighted again (2017: September 20–21; 2018: November 27–28).
Molecular Analyses of Transgenic Plants
Genomic DNA was extracted from leaves of one to two plants of the 16 different putative DX15::PdGA20-OXIDASE transgenic lines and the non-transgenic control P1 line grown in vitro for PCR and Southern-blot analyses according to Fladung et al. (1996) and Dünisch et al. (2006). PCR was performed as described in detail in Fladung et al. (1997), but with annealing temperatures of 52 °C for amplification of a partial PdGA20-OXIDASE-fragment and 54 °C for HPT. The sequences of the primer pairs used in PCR reactions to amplify fragments of the DX15::PdGA20-OXIDASE gene construct are the following: forward primer PdGA20ox (f-PdGA20ox): 5′-ATG GGT ACT TCG ACT GTG AGT-3′; reverse primer PdGA20ox (r-PdGA20ox): 5′-TTA TGG CTG GTT TCT TGA GG-3′, with an expected amplicon size of ~1100 bp. Amplification of the HPT gene was performed using the primer pair: forward (f-hpt) 5′-AAA GCC TGA ACT CAC CGC GA-3′ and reverse (r-hpt) 5′-TCG GTT TCC ACT ATC GGC GA-3′ (expected amplicon size: ~950 bp).
For Southern blot analyses, twenty or 40 µg genomic DNA were digested from PCR-confirmed DX15::PdGA20-OXIDASE transgenic lines and the non-transgenic control line P1 with the restriction enzymes EcoRV (New England Biolabs, Frankfurt M, Germany) or XhoI (Thermo Fisher Scientific, Braunschweig, Germany), according to the respective supplier’s instructions. DNA electrophoresis and transfer of DNA to Biodyne A membranes (Pall Europe Limited, Portsmouth, UK) were performed as described elsewhere (Fladung et al. 1996, 1997). Prehybridisation and hybridisation of Southern-blots were performed with the non-radioactive DIG (digoxigenine) system using a DIG-dUTP PCR partial-labeled PdGA20-OXIDASE-probe as described in other papers (Fladung and Ahuja 1995; Fladung et al. 1997). Primer pair f‑PdGA20ox/r-PdGA20ox was used for synthesis of DIG-labeled PdGA20ox-probes. The gels were stained with ethidium bromide shortly before blotting (to confirm similar DNA amounts loaded and uniform restriction patterns) and after blotting (to confirm complete transfer of DNA to the membrane).
Data Analysis and Statistics
Data are presented in Figures as mean ± SD. Differences of the last measurements were tested for significance by ANOVA at P < 0.05 (Fisher’s F‑test). Values in the Figures labelled by different letters differ significantly at P < 0.05 (Fisher’s F‑test).
Results
Molecular Analysis of DX15::PdGA20-OXIDASE-transgenic Plants
In total, 16 independent putative transgenic lines were obtained in the two transformation experiments with the Agrobacterium strain containing the DX15::PdGA20-OXIDASE gene construct (N456, N457). All transgenic lines were regenerated and clonal propagated on hygromycin-containing media. Transgenic plants grew well and formed roots on the antibiotic-containing media. Transgenic plants of all lines revealed no apparent phenotypic alterations compared to not-transgenic control plants. PCR analyses amplifying part of the HPT gene (expected size about 950 bp) of one to two plants of all 16 putative transgenic lines confirmed the presence of the antibiotic selection gene (Fig. 1a). Using the primer pair f‑PdGA20ox/r-PdGA20ox in PCR experiments of one to two plants of the 16 lines, the amplification products reveal the expected size of about 1100 bp (Fig. 1b), indicating the presence of the DX15::PdGA20-OXIDASE gene.
PCR analyses of DX15::PdGA20-OXIDASE transgenic lines and non-transformed control using primer pairs f‑hpt/r-hpt to amplify part of the HPT gene (a) and f‑PdGA20ox/r-PdGA20ox to amplify part of the DX15::OXIDASE gene (b) in the transgenic lines. Bands in a and b are approximately 950 bp and 1100 bp in size, respectively. Marker: molecular weight marker (Smart ladder, Eurogentec, Cologne, Germany) in base pairs (bp); 0: water control, without DNA; At. Agrobacterium plasmid; P1: not-transgenic (negative) control P1; lanes N456‑1 to N457-15 indicate individual numbers of independent DX15::PdGA20-OXIDASE transgenic lines
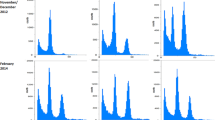
The four lines N457‑4, -6, -12, and -14 selected for the glasshouse growth experiment in 2018 were analyzed in Southern-blot analysis (Fig. 2). DNA, digested either with EcoRV or XhoI, was electrophorized, transferred to membranes and blotted against a DIG-labeled HPT-probe. The line N457‑4 revealed a single and strong hybridization signal for the EcoRV-digested DNA, but at least four weaker signals for the XhoI‑digested DNA. Lines N457‑6 and -12 both showed two signals for EcoRV, and six and two signals for XhoI, respectively. Finally, line N457-14 possessed four signals for EcoRV and six signals for XhoI.
Southern-blot analysis of four independent DX15::PdGA20-OXIDASE transgenic lines N457‑4, N457‑6, N457-12, N457-14, selected for growth and biomass experiments in the glasshouse in 2018, Agrobacterium plasmid (positive control), and clone P1 (negative control). Representative blots with EcoRV- and XhoI‑digested, and DIG-labelled OXIDASE-probed DNA isolated from Agrobacterium. (DIG‑M II DIG-labelled molecular weight marker II (Roche, Mannheim, Germany) in base pairs (bp), At Agrobacterium plasmid, P1 not-transgenic clone P1)
Morphological Investigations of Glasshouse Plants
From the 16 independent DX15::PdGA20-OXIDASE-transgenic lines obtained in total, plants from just seven lines grew well in soil when transferred to the glasshouse, namely N456‑1, N457‑4, -5, -6, -12, -13, and -14. Thus, these seven lines and the untransformed P1-control clone were selected and micro-propagated for a growth experiment in the glasshouse in 2017. Up to 16 plants per line of the seven transgenic lines and the P1 control clone were transferred into the glasshouse and recorded monthly for plant height (Fig. 3A) and shoot diameter (Fig. 3b just shows the values of the last measurement), from April 19 until September 14, 2017. Fresh and dry weights of defoliated shoots were determined at the end of the experiment (September 2017) (Fig. 3c).
Growth, shoot diameter, fresh- and dry weight of non-transgenic control P1, and seven different DX15::PdGA20-OXIDASE transgenic lines (up to 16 plants per line) in glasshouse from April until September 2017. a Height (in cm) was measured from the tallest shoot of each plant. b Shoot diameter (mm) was measured five cm above soil. c Fresh- and dry weight (g) was determined after the last measurement in September. Values (mean + SD) followed by different letters differ significantly at P < 0.05 (Fisher’s F‑test)
After five months of cultivation of the lines in the glasshouse, mean plant height revealed highest in line N457‑4 (3.57 m), followed by lines N457‑6, -12, and -14 (3.36, 3.30, 3.19 m). Lines N456‑1, N457‑5, and -13 (2.76, 2.78, 2.66 m) clustered together with the control clone P1 (2.66 m, Fig. 3a). For the highest line N457‑4, it is an increase of about 30% compared to the control P1. The line N457‑4 also revealed the highest mean shoot diameter (11.1 mm) at the end of the experiment, followed by all other lines with shoot ranging from 9.3 to 10.3 mm, however, all values were not significantly different from the control P1 (9.3 mm, Fig. 3b). Fresh and dry weight of N457‑4 was, with 158 and 60 g, respectively, also highest in the line N457‑4, and about 80% higher than in the control clone P1 (86 and 31 g, respectively). The fresh and dry weights of the other DX15::PdGA20-OXIDASE-transgenic lines ranged from 85 to 133 g, and 32 to 49 g, respectively (Fig. 3c).
In 2018, the experiment was repeated in the glasshouse with the four best-growing DX15::PdGA20-OXIDASE-transgenic lines in 2017, namely N457‑4, -6, -12, and -14, and the non-transgenic control P1. Plant height (Fig. 4A) and shoot diameter (Fig. 4b just shows the values of the last measurement) of up to 15 plants per line were measured monthly, from April/May to November 2018. Fresh and dry weights of defoliated shoots were determined at the end of the experiment (November 2018). Again, after seven months cultivation in glasshouse, line N457‑4 turned out to be highest compared to control P1. (Fig. 4a, mean plant height was increased by about 15%) with the thickest (Fig. 4b, mean shoot diameter was [not significantly] increased by about 20%), and heaviest, shoots (Fig. 4c, mean fresh and dry weights were increased by about 81 and 62%, respectively). Mean values of height, shoot diameter, and fresh and dry weight of lines N457‑6 and -12 revealed slightly lower than N457‑4 but higher than the P1 control. Surprisingly, the line N457-14 showed values for shoot height and diameter, and biomass more or less similar to the ones of the P1 control.
Growth, shoot diameter, fresh- and dry weight of non-transgenic control P1, and four different DX15::PdGA20-OXIDASE transgenic lines (up to 15 plants per line) in glasshouse from April until November 2018. a Height (in cm) was measured from the tallest shoot of each plant. Note: plants stopped growing from the end of September. b Shoot diameter (mm) was measured five cm above soil. c Fresh- and dry weight (g) was determined after the last measurement in November. Values (mean + SD) followed by different letters differ significantly at P < 0.05 (Fisher’s F‑test)
Discussion
The increase of plant biomass production is a long-cherished dream of many biotechnologists (Sticklen 2006; Demura and Ye 2010; Rojas et al. 2010; Harfouche et al. 2011; Dubouzet et al. 2013). Various pathways within the plant metabolism have been defined as starting points, in which, if the right genes are selected, plant biomass production can be increased following their overexpression or knock-out by genetic engineering. Impressive examples to increase plant biomass comprise, e.g., the introduction of the Escherichia coli glycolate catabolic pathway to reduce photorespiration in C3-plants (Kebeish et al. 2007); overexpression of the photorespiration H‑protein in tobacco (López-Calcagno et al. 2019); deregulation of endosperm ADP-glucose pyrophosphorylase in rice (Smidansky et al. 2003); knock-out of the BRASSINOSTEROID INSENSITIVE1 ortholog, OsBRI1, in rice (Morinaka et al. 2006); overexpression of SUCROSE SYNTHASE in switchgrass and cotton (Jiang et al. 2012, Poovaiah et al. 2015), or knockdown of PCBER1, a gene of neolignan biosynthesis, in poplar (Bruegmann et al. 2019b).
We have used the DX15::GA20-OXIDASE gene construct, kindly provided by Hyung-Woo Jeon (Prof. J. H. Ko Lab., Department of Plant & Environmental New Resources, Kyung Hee University, Yongin, Korea; Jeon et al. 2016), and transformed the Populus × canescens (P. tremula L. × P. alba L.) clone INRA 717-1B4 (P1). The aims of the study were (a) to prove the increased growth ability of DX15::GA20-OXIDASE gene construct in a poplar variety different from the one used by Jeon et al. (2016); (b) to monitor growth and biomass production of DX15::GA20-OXIDASE transgenic poplar plants over a complete growing season (2017), and (c) to confirm the results of this experiment with selected lines in the subsequent year 2018.
A deliberate release of GMOs into the field is, in principle, possible in Europe and, in particular, also in Germany, according to the Directive 2001/18/EC. However, to obtain approval from national authorities for field trials with transgenic trees is a very time-consuming and tedious process, as is the application procedure itself. After successful submission of the application, very detailed discussions with non-governmental organizations (NGOs) and the public follow. And, ultimately, when the field trial is finally approved, the risk of having the field trial destroyed by activists is extremely high, because information about the exact location of the field trial can be found in the public part of the location register, as can notification of date of releases or cultivation of GMOs (https://apps2.bvl.bund.de/stareg_web/showflaechen.do). Therefore, even though we were aware that field trials are not per se comparable to “in-door” experiments, we decided to test our DX15::GA20-OXIDASE-transgenic poplar in the glasshouse.
Transgenic poplar plants harbouring the GA20-OXIDASE gene under expression control of the constitutive cauliflower mosaic virus 35S-promoter reveal increased plant growth and biomass production when compared to non-transgenic control plants (Eriksson et al. 2000; Dünisch et al. 2006), but they do, however, suffer severely from unstable shoots and poor root formation (Fladung 2006; Mauriat and Moritz 2009; Mauriat et al. 2014; Jeon et al. 2016). Poor rooting may be caused by gibberellin affecting auxin transport rather than auxin signalling (Mauriat et al. 2014). Alternatively, when transgenic poplar plants specifically express the GA20-OXIDASE gene only in the xylem (under control of the poplar DX15-promoter), they also reveal increased growth and biomass production (Jeon et al. 2016). However, in contrast to the 35S::GA20-OXIDASE-transgenic poplar, we show that DX15::GA20-OXIDASE-transgenic plants reveal slightly (but not significantly) thicker shoots. In consequence, shoots of these DX15::GA20-OXIDASE-transgenic plants appear stable, so that the plants are able to grow without external stabilization. A possible explanation could be that gibberellins are involved in the wood formation processes xylogenesis (cambium) and fiber elongation (development of xylem) (Mauriat and Moritz 2009).
In 2017, we tested seven independent DX15::GA20-OXIDASE-transgenic lines for growth and biomass production in the glasshouse. At the end of the growing season, the results show that plants of the line N457‑4 turned out to be highest in growth, thickest in shoot diameter and revealing most biomass among all lines. All values, with exception of shoot diameter, increased significantly compared to the P1 control. The three lines (N457‑6, -12, and -14) revealed similar values in height, shoot diameter, and fresh and dry weight, also significantly higher than the P1 control clone. Height, shoot diameter and biomass production of lines N456‑1, N457‑5, and N457-13 were similar to the ones of the P1 control.
Therefore in 2018, the growth-experiment was repeated in the glasshouse with the lines growing significant differently from the P1 control in 2017. Again, line N457‑4 revealed the highest values of plant height, shoot diameter, and fresh and dry weight, followed by lines N457‑6 and N457-12. All three lines again revealed significantly (with exception of shoot diameter) higher values than the P1 control. Surprisingly, line N457-14 grew differently from 2017, with similar values to the P1 clone. Molecular analyses of the four lines revealed that N457‑4 carried one copy of the GA20-OXIDASE gene, N457‑6 and N457-12 two copies each, and N457-14 at least four copies. Post-transcriptional and transcriptional gene silencing of transgenes is a well-known phenomenon and has been described for many transformed plant species (Vaucheret et al. 1998; Matzke and Matzke 1998; Vaucheret and Fagard 2001) carrying more than one copy of the transgene, either arranged as tandem-repeats at the same integration locus (Stam et al. 1997; Fladung 1999) or as independent integrations at different genomic loci (Butaye et al. 2005; Tang et al. 2007). Post-transcriptional gene silencing (PTGS) was mostly obtained in transgenic lines with more than three copies of T‑DNA but not in transgenic lines with one copy of T‑DNA (Tang et al. 2007). This could explain the growth-drop-down of line N457-14 (with four copies of the transgene) in 2018 (compared to 2017), and why, on the other hand, the one-transgene-copy line N457‑4 remained stable in growth behavior and biomass production in both years.
Transgene silencing caused by cytosine methylation has been reported exclusively in the promoter and not in the transgene itself (Mette et al. 2000; Mishiba et al. 2005). Methylation of the promoter may reduce transgene expression, however, doesn’t necessarily lead to 100%-drop-down of transgene expression and, thus, to transgene silencing (Fan et al. 2011). It could be that the two two-transgene-copy lines N457‑6 and N457-12 still express the GA20-OXIDASE gene but at lower levels, and, therefore, behave stably intermediate between N457‑4 and P1/N457-14 with respect to growth behavior and biomass production in the two years under investigation. In frame of the present study, unfortunately, it was not possible to confirm this hypothesis in e.g., northern blot or qRT-PCR experiments, which has to be done in a subsequent study.
Conclusions
We obtained 16 different independent DX15::GA20-OXIDASE-transgenic lines out of two transformation experiments. From these, seven lines were selected for a growth experiment in the glasshouse in 2017, and, the experiment was repeated for the “best-four” lines in 2018. Plants from one DX15::GA20-OXIDASE-transgenic line, N457‑4, turned out to be tallest, thickest, and with highest biomass in the two years under investigation. Interestingly, only this line carries only one copy of the gene construct, whereas the other lines tested in 2018 carry two or more copies. We propose that transcriptional or post-transcriptional transgene silencing probably occurred in the two- or more copies carrying lines leading to reduced GA20-OXIDASE-gene expression. In contrast, the single-copy-line N457‑4 probably stably expresses the GA20-OXIDASE gene, which has to be confirmed in subsequent e.g., northern blot or qRT-PCR experiments. In addition, due to the limited transferability of glasshouse experiments to natural environmental conditions, the experiments have also to be repeated in open fields over several years.
References
Biemelt S, Tschiersch H, Sonnewald U (2004) Impact of altered gibberellin metabolism on biomass accumulation, lignin biosynthesis, and photosynthesis in transgenic tobacco plants. Plant Physiol 135:254–265
Brian PW (1959) Effects of gibberellins on plant growth and development. Biol Rev 34:37–77
Bruegmann T, Deecke K, Fladung M (2019a) Evaluating the efficiency of gRNAs in CRISPR/Cas9 mediated genome editing in poplars. Int J Mol Sci 20:3623
Bruegmann T, Wetzel H, Hettrich K, Smeds A, Willför S, Kersten B, Fladung M (2019b) Knockdown of PCBER1, a gene of neolignan biosynthesis, resulted in increased poplar growth. Planta 249:515–525
Butaye KM, Cammue BP, Delauré SL, De Bolle MF (2005) Approaches to minimize variation of transgene expression in plants. Mol Breed 16:79–91
Campbell MM, Brunner AM, Jones HM, Strauss SH (2003) Forestry’s fertile crescent: the application of biotechnology to forest trees. Plant Biotech J 1:141–154
Chelakkot R, Mahadevan L (2017) On the growth and form of shoots. J R Soc Interface 14:1420170001
Darwin C (1897) The power of movement in plants. Appleton
Davies PJ (2012) Plant hormones and their role in plant growth and development. Springer
Demura T, Ye ZH (2010) Regulation of plant biomass production. Curr Opin Plant Biol 13:298–303
Doblin MS, Kurek I, Jacob-Wilk D, Delmer DP (2002) Cellulose biosynthesis in plants: from genes to rosettes. Plant Cell Physiol 43:1407–1420
Dubouzet JG, Strabala TJ, Wagner A (2013) Potential transgenic routes to increase tree biomass. Plant Sci 212:72–101
Dünisch O, Funada R, Nakaba S, Fladung M (2006) Influence of overexpression of a gibberellin 20-oxidase gene on the kinetics of xylem cell development in hybrid poplar (Populus tremula L x P. tremuloides Michx). Holzforschung 60:608–617
Dyson PW (1965) Effects of gibberellic acid and (2-chloroethyl)-trimethylammonium chloride on potato growth and development. J Sci Food Agric 16:542–549
Eriksson ME, Israelsson M, Olsson O, Moritz T (2000) Increased gibberellin biosynthesis in transgenic trees promotes growth, biomass production and xylem fiber length. Nat Biotech 18:784–788
Fan J, Liu X, Xu SX, Xu Q, Guo WW (2011) T‑DNA direct repeat and 35S promoter methylation affect transgene expression but do not cause silencing in transgenic sweet orange. Plant Cell Tiss Org Cult 107:225–232
Flachowsky H, Hanke MV, Peil A, Strauss SH, Fladung M (2009) A review on transgenic approaches to accelerate breeding of woody plants. Plant Breed 128:217–226
Fladung M (1999) Gene stability in transgenic aspen-Populus. I. Flanking DNA sequences and T‑DNA structure. Mol Gen Genet 260:574–581
Fladung M (2006) Modification of cellulose. In: Fladung M, Ewald D (eds) Tree transgenesis—recent developments. Springer, Berlin, Heidelberg, New York, pp 123–136
Fladung M (2018) Growth of mixoploid GIBBERELLIC ACID 20 OXIDASE (GA20-OXIDASE) overexpressing transgenic Populus. Gesunde Pflanz 70:91–98
Fladung M, Ahuja MR (1995) “Sandwich” method for non-radioactive hybridizations. Biotechniques 18:3–5
Fladung M, Kumar S, Ahuja MR (1997) Genetic transformation of Populus genotypes with different chimeric gene constructs: transformation efficiency and molecular analysis. Transgenic Res 6:111–121
Fladung M, Muhs HJ, Ahuja MR (1996) Morphological changes observed in transgenic Populus carrying the rolC gene from Agrobacterium rhizogenes. Silvae Genet 45:349–354
Gilroy S (2008) Plant tropisms. Curr Biol 18:R275–R277
Gupta R, Chakrabarty SK (2013) Gibberellic acid in plant: still a mystery unresolved. Plant Signal Behav 8:e25504
Hansen EA (1991) Poplar woody biomass yields: a look to the future. Biomass Bioenergy 1:1–7
Harfouche A, Meilan R, Altman A (2011) Tree genetic engineering and applications to sustainable forestry and biomass production. Trends Biotechnol 29:9–17
Humphries EC, Welbank PJ, Witts KJ (1965) Effect of CCC (chlorocholine chloride) on growth and yield of spring wheat in the field. Annals Appl Biol 56:351–361
Jeon HW, Cho JS, Park EJ, Han KH, Choi YI, Ko JH (2016) Developing xylem-preferential expression of PdGA20ox1, a gibberellin 20-oxidase 1 from Pinus densiflora, improves woody biomass production in a hybrid poplar. Plant Biotechnol J 14:1161–1170
Jiang Y, Guo W, Zhu H, Ruan YL, Zhang T (2012) Overexpression of GhSusA1 increases plant biomass and improves cotton fiber yield and quality. Plant Biotechnol J 10:301–312
Kebeish R, Niessen M, Thiruveedhi K, Bari R, Hirsch HJ, Rosenkranz R, Stäbler N, Schönfeld B, Kreuzaler F, Peterhänsel C (2007) Chloroplastic photorespiratory bypass increases photosynthesis and biomass production in Arabidopsis thaliana. Nat Biotechnol 25:593–599
Ko JH, Kim HT, Hwang ID, Han KH (2012) Tissue-type-specific transcriptome analysis identifies developing xylem-specific promoters in poplar. Plant Biotechnol J 10:587–596
Leplé JC, Brasileiro ACM, Michel MF, Delmotte F, Jouanin L (1992) Transgenic poplars: expression of chimeric genes using four different constructs. Plant Cell Rep 11:137–141
Linné C (1764) Species plantarum: 2. Salo
López-Calcagno PE, Fisk S, Brown KL, Bull SE, South PF, Raines CA (2019) Overexpressing the H‑protein of the glycine cleavage system increases biomass yield in glasshouse and field-grown transgenic tobacco plants. Plant Biotechnol J 17:141–151
Matzke AJM, Matzke MA (1998) Position effects and epigenetic silencing of plant transgenes. Curr Opin Plant Biol 1:142–148
Mauriat M, Moritz T (2009) Analyses of GA20ox- and GID1-over-expressing aspen suggest that gibberellins play two distinct roles in wood formation. Plant J 58:989–1003
Mauriat M, Petterle A, Bellini C, Moritz T (2014) Gibberellins inhibit adventitious rooting in hybrid aspen and Arabidopsis by affecting auxin transport. Plant J 78:372–384
Mette MF, Aufsatz W, Van der Winden J, Matzke MA, Matzke AJM (2000) Transcriptional silencing and promoter methylation triggered by double-stranded RNA. Embo J 19:5194–5201
Mishiba KI, Nishihara M, Nakatsuka T, Abe Y, Hirano H, Yokoi T, Kikuchi A, Yamamura S (2005) Consistent transcriptional silencing of 35S-driven transgenes in gentian. Plant J 44:541–556
Morinaka Y, Sakamoto T, Inukai Y, Agetsuma M, Kitano H, Ashikari M, Matsuoka M (2006) Morphological alteration caused by brassinosteroid insensitivity increases the biomass and grain production of rice. Plant Physiol 141:924–931
Poovaiah CR, Mazarei M, Decker SR, Turner GB, Sykes RW, Davis MF, Stewart CN (2015) Transgenic switchgrass (Panicum virgatum L.) biomass is increased by overexpression of switchgrass sucrose synthase (PvSUS1). Biotechnol J 10:552–563
Ram HM, Mehta U (1978) Origin and development of secondary capitula in Calendula officinalis L. in response to gibberellic acid. J Exp Bot 29:653–662
Rojas CA, Hemerly AS, Ferreira PCG (2010) Genetically modified crops for biomass increase. Genes and strategies. gmcrops 1:137–142
Santner A, Calderon-Villalobos LIA, Estelle M (2009) Plant hormones are versatile chemical regulators of plant growth. Nat Chem Biol 5:301–307
Scarascia-Mugnozza G, Bauer GA, Persson H, Matteucci G, Masci A (2000) Tree biomass, growth and nutrient pools. In: Schulze ED (ed) Carbon and nitrogen cycling in European forest ecosystems. Ecological Studies (Analysis and Synthesis), vol 142. Springer, Berlin, Heidelberg, pp 49–62
Smidansky ED, Martin JM, Hannah CL, Fischer AM, Giroux MJ (2003) Seed yield and plant biomass increases in rice are conferred by deregulation of endosperm ADP-glucose pyrophosphorylase. Planta 216:656–664
Somerville C (2006) Cellulose synthesis in higher plants. Ann Rev Cell Dev Biol 22:53–78
Stam M, De Bruin R, Kenter S, Van Der Hoorn RA, Van Blokland R, Mol JN, Kooter JM (1997) Post-transcriptional silencing of chalcone synthase in Petunia by inverted transgene repeats. Plant J 12:63–82
Sticklen M (2006) Plant genetic engineering to improve biomass characteristics for biofuels. Curr Opin Biotechnol 17:315–319
Strauss SH, DiFazio SP, Meilan R (2001) Genetically modified poplars in context. For Chron 77:271–279
Tang W, Newton RJ, Weidner DA (2007) Genetic transformation and gene silencing mediated by multiple copies of a transgene in eastern white pine. J Exp Bot 58:545–554
Taylor G (2002) Populus: Arabidopsis for forestry. Do we need a model tree? Ann Bot 90:681–689
Vaucheret H, Fagard M (2001) Transcriptional gene silencing in plants: targets, inducers and regulators. Trends Genet 17:29–35
Vaucheret H, Béclin C, Elmayan T, Feuerbach F, Godon C, Morel JB, Mourrain P, Palauqui JC, Vernhettes S (1998) Transgene-induced gene silencing in plants. Plant J 16:651–659
Wang Y, Zhao J, Lu W, Deng D (2017) Gibberellin in plant height control: old player, new story. Plant Cell Rep 36:391–398
Acknowledgements
Thanks are due to Dr. Hyung-Woo Jeon (Prof. J. H. Ko Lab., Department of Plant & Environmental New Resources, Kyung Hee University, Yongin, Korea) for providing the A. tumefaciens strain carrying the DX15::PdGA20-OXIDASE construct. I thank Olaf Polak, D. Ebbinghaus, and A. Schellhorn for excellent technical assistance in the lab, and the glasshouse staff (M. Hunger, G. Wiemann, M. Spauszus; all Thuenen Institute of Forest Genetics, Grosshansdorf, Germany) for plant cultivation. I thank Dina Führmann (Thuenen Institute, Braunschweig, Germany) for thorough correction of the English language.
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
M. Fladung declares that he has no competing interests.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Fladung, M. Xylem-specific Overexpression of the GIBBERELLIN ACID 20 OXIDASE Gene (GA20-OXIDASE) from Pine in Hybrid Poplar (Populus tremula L. × P. alba L.) Revealed Reliable Increase in Growth and Biomass Production Just in a Single-copy-line. Gesunde Pflanzen 74, 239–248 (2022). https://doi.org/10.1007/s10343-022-00653-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10343-022-00653-y