Abstract
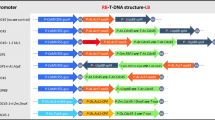
The stability of transgenes in the genome of transformed plants depends strongly on their correct physical integration into the host genome as well as on flanking target DNA sequences. For long-lived species like trees, however, no information is available so far concerning inactivation or loss of transgenes due to gene silencing or somatic genome rearrangement events. In this study, four independently transformed 35S-rolC transgenic hybrid aspen plants (Populus tremula L. × tremuloides Michx.), each harbouring one copy of the transgene, were investigated during continuous growth in the greenhouse. In one of these transgenic lines (Esch5:35S-rolC-##1) individuals frequently show phenotypic reversions, while in the remaining three lines (Esch5:35S-rolC-#3, -#5, -#16) the gene was essentially stable. Molecular analysis including PCR, Southern and Northern assays clearly showed that the transgene had been lost in the revertant tissue of the unstable line. Sequencing of T-DNA right and left borders, and flanking DNA regions, in all four transgenic aspen lines revealed no differences either in the type of flanking DNA (G-C to A-T ratio) or with respect to the presence of enhancers or MAR (matrix associated repeats)-like structures. Primers located within the left and right flanking regions in the three stable lines could be used to recover the target sites from the untransformed plants. This was not possible, however, with the unstable line, indicating that at least one flanking sequence does not derive from the plant target DNA but is of unknown origin. PCR using other primer pairs, and inverse PCR analysis, revealed an additional truncated T-DNA copy of 1050 nucleotides adjacent to the left border of the complete copy in this line. Sequencing of this truncated T-DNA revealed that it represented an inverted copy of part of the right half of the original construct. This special feature would allow the inverted repeat to pair with right border sequences of the complete copy. This would explain the frequently observed reversion resulting in transgene loss as due to intrachromosomal base-pairing leading to double-stranded loops of single-stranded DNA during mitotic cell divisions.
Similar content being viewed by others
Author information
Authors and Affiliations
Additional information
Received: 9 June 1998 / Accepted: 6 October 1998
Rights and permissions
About this article
Cite this article
Fladung, M. Gene stability in transgenic aspen (Populus). I. Flanking DNA sequences and T-DNA structure. Mol Gen Genet 260, 574–581 (1999). https://doi.org/10.1007/s004380050931
Issue Date:
DOI: https://doi.org/10.1007/s004380050931