Abstract
Taxus baccata L. is a highly valuable species with wide distribution but scattered and locally rare occurrence. Human intervention, including forest management practices and fragmentation, can significantly impact the species’ genetic diversity, structure, and dynamics. In this study, we investigated these factors within T. baccata populations in the Bavarian Forest National Park (NP) in Germany and their implications for conservation. We used 13 EST-SSRs to assess the genetic diversity and structure of the population. Our analysis revealed a scarcity of small-diameter trees, indicating limited natural regeneration over time. However, conservation efforts, like selectively removing competitor species and using protective fencing, have improved growth conditions and promoted seedling emergence. The NP’s natural zone has no active management, which is confined to the development and management zones. Genetic diversity assessments revealed high genetic diversity (He: 0.612 and 0.614 for seedlings and adults, respectively) compared to other studies in Taxus baccata, dispelling concerns of significant inbreeding and showcasing a stable genetic structure. However, significant spatial clustering of related individuals (family structures) in both cohorts and low effective population size in the progeny hints at restricted gene flow, necessitating conservation efforts prioritizing safeguarding and promoting natural regeneration in development and management zones. Limited natural regeneration and the recent decrease in effective population size in the NP populations indicate habitat fragmentation and human interventions. Effective population size estimates emphasize the need for diverse conservation strategies. Conservation efforts should prioritize protecting natural regeneration and enhancing gene flow by actively promoting European yew, e.g., by shelterwood cutting, to ensure the long-term viability of T. baccata in the region outside the NP.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Loss of genetic variation increases the risk of extinction (Frankham 1995), and currently, the biological diversity is being depleted because of direct or indirect consequences of human activities (Crist et al. 2017; Semper-Pascual et al. 2021). Genetic diversity of trees has been recognized as an essential conservation issue (Ledig 1992; Wang and Szmidt 2001) as the loss of forest genetic diversity threatens the health and resilience of forest species (Porth and El-Kassaby 2014).
Therefore, the maintenance of genetic variation is crucial to ensure the long-term existence of species. The amount and pattern of genetic variation determine the adaptability of tree populations to the environment (Müller-Starck et al. 1992). To conserve diversity, monitor population dynamics, and characterize gene flow, genetic markers are critical for identifying genetic diversity hotspots (Porth and El-Kassaby 2014). These markers can provide insights into the level and distribution of genetic variation within and among populations, the level of inbreeding, effective population size, and bottleneck events (Gailing et al. 2009; Mariette et al. 2010; Eliades et al. 2011; Porth and El-Kassaby 2014).
Stand-forming and common tree species have received more attention than rare and scattered species like Taxus baccata L. (English yew), which is widely distributed but typically rare (Ruprecht et al. 2010). One of the most notable characteristics of the species is its longevity and very slow growth rate (Hulme 1996). In the middle age due to the wide range of artifacts made from yew wood, the species was on the verge of extinction in Europe (Hartzell 1991; Benham et al. 2016). Today, it remains an important tree species that is widely used, e.g. as a source of anti-cancer taxane alkaloid (Suffness 1995; Hageneder 2013; Gardner et al. 2019) alongside its role in hedging and gardening.
English yew is native to European temperate and Mediterranean forests; from Scandinavia and North Africa to the Iberian Peninsula and the Caspian Sea, this species is present in a wide range of geographical regions. Therefore, it occurs under various environmental conditions, including oceanic, continental, and Mediterranean environments (Deforce and Bastiaens 2007; Thomas and Garcia-Marti 2015; Chybicki and Oleksa 2018). Although the species is found throughout a wide range of habitats, it is in decline, and may face local extinction, as some parts of Europe have witnessed extinction and the isolation of populations (Svenning and Magård 1999; Dubreuil et al. 2010; Schirone et al. 2010; Alavi et al. 2019; Mayol et al. 2019; Ahmadi et al. 2020).
The species is wind-pollinated, and seeds are dispersed mainly by birds and also by gravity (Hulme 1996; Thomas and Polwart 2003; Chybicki et al. 2011). Despite the considerable potential for seed and pollen dispersal, yew populations demonstrate pronounced kinship structure (Chybicki et al. 2011). In addition, the species has limited gene flow, in contrast to other often wind dispersed gymnosperms, and the genetic diversity is highly structured at the local and regional scale (Dubreuil et al. 2010; González-Martínez et al. 2010; Chybicki et al. 2012).
In some areas, conservation management targets the protection of yew populations in Germany. To this end, a fence surrounding mature trees and seedlings is used to prevent deer from eating or trampling them; while the fences will not have an effect on bird movement and seed dispersal, they reduce the browsing pressure on seedlings and saplings (Dhar et al. 2008). This approach fosters natural regeneration only in close vicinity of seed trees by protecting seedlings partially from animal damage. However, they are vulnerable to snails, mice, fungi, and even dry periods after germination. In an isolated population in Poland, birds and other animals dispersed 95% of seeds up to 109 m from the mother trees, while the pollen traveled up to 704 m (Chybicki and Oleksa 2018).
Yew expanded its range during several Pleistocene interglacial periods and played a prominent role in the vegetation of Europe (de Beaulieu et al. 2001; Deforce and Bastiaens 2007; Delahunty 2007). However, it has experienced effective population size declines since the last interglacial (Mayol et al. 2019). In addition to environmental change, human interventions have aggravated this population reduction (Linares 2013; Thomas and Garcia-Marti 2015). Nevertheless, there has been an expansion in population size in certain parts of southern Europe, attributed to favorable environmental conditions (Kassioumis et al. 2004). Similarly, targeted management and conservation initiatives, such as selective cutting of beech trees in northern Europe (Svenning and Magård 1999), have contributed to positive developments in specific regions.
Climate change also threatens the yew populations, i.e. the species has limited capacity to migrate north of its distribution range in Europe because of its slow migration rate, specific soil requirements, and limited potential to adapt to the anticipated climate change (Thomas and Garcia-Marti 2015). The scattered distribution, along with less favorable climate, slow growth, weak reproductive capacity, illegal trade, and over-harvesting of Taxol from the bark and leaves, have caused the species to be threatened in most countries (Deforce and Bastiaens 2007; Jia et al. 2022).
T. baccata is principally a dioecious species (strictly outcrossing); thus, monoecious individuals are extremely rare (Thomas and Polwart 2003; Zatloukal and Vančura 2004; Deforce and Bastiaens 2007; Dubreuil et al. 2008). Dioecious species face a greater threat from habitat fragmentation than monoecious species. In large populations, the representation of both sexes is roughly equal (Zatloukal and Vančura 2004); small stands, however, may deviate from a 1:1 distribution (Leinemann and Hattemer 2006). Usually, the sex of trees can be distinguished at sexual maturity; T. baccata reaches maturity at around 30–35 years when it receives sufficient light. However, if it grows in heavily shaded areas, it takes much longer to mature, sometimes not until 70 or even 120 years (Zarek 2016). Its scattered distribution, unequal numbers of male and female trees and low population density in small fragments may promote biparental inbreeding even in an outcrossing species (Dubreuil et al. 2008).
A 2013 Federal Office for Agriculture and Food (BLE 2013) survey found roughly 60,000 naturally growing adult yew trees across 342 populations in Germany. Only 50 populations have more than 100 trees, with 13 having over 1,000. The German federal state of Thuringia harbors most yew trees (about 33,000), while most of the stands (128 mapped occurrences), and about a quarter of all trees (14,761) are located in Bavaria. Lack of natural regeneration is observed in many yew populations in Germany, and less than 12% fall into the age group of 1–60 years (BMELV 2012).
The Bavarian Forest National Park (NP) was the first national park in Germany (established in 1970) and together with the Šumava National Park in the Czech Republic, it forms the largest continuous protected area for woodland in Central Europe (Heurich et al. 2015). Forests cover 97% of the NP. The major part of the forest habitat is a mix of montane beech, silver fir and spruce trees (Krenova and Kiener 2012; Van der Knaap et al. 2020). There is no management in the natural zone of the NP, and conservation, bark beetle control measures, and wildlife management are restricted to the development and management zones. Moreover, management practices for T. baccata conservation have been implemented in the development and management zones since 2016, when the first Taxus inventory was conducted, and competing trees of other species were cut down and fences were constructed to promote natural regeneration.
Research on T. baccata includes the study of population genetic diversity and structure at a limited set of nuclear SSRs (Dubreuil et al. 2008), e.g. González-Martínez et al. (2010) in the western Mediterranean Basin, Gargiulo et al. (2019) in Britain, Maroso et al. (2021) in Cantabrian-Atlantic Region, Komárková et al. (2022) in Czech Republic, and Mayol et al. (2015) used them in 238 sampling sites covering the entire distribution range. Mayol et al. (2019) also analyzed single nucleotide polymorphims (SNPs) to infer the demographic history and to test for molecular signatures of selection at different geographical scales. In Germany, studies have been conducted on the population genetic structure and diversity of English yew at allozyme and cpSSR markers (Hertel and Kohlstock 1996; Cao et al. 2004; Leinemann and Hattemer 2006). A broader geographical sampling in Europe, including German populations revealed that most of the populations from Germany harbor high levels of heterozygosity and comprise trees with admixed ancestry from the western and eastern European gene pools of the species (Romšáková and Paule 2009; Mayol et al. 2015).
T. baccata, as a woody plant with a long lifespan, wide geographical range, outcrossing breeding system, wind-pollination and animal dispersed seeds, is expected to maintain a high level of genetic diversity even in moderately fragmented populations (Hamrick et al. 1992). In order to assess the conservation status of the population in the NP, to evaluate and develop suitable management strategies and to deal with the potential threats, we investigated the genetic diversity and structure in different zones, i.e. in the natural zone (unmanaged), the management zone, the development zone and outside of the NP. To characterize the genetic variation of T. baccata in the NP, we used 13 EST-SSRs, developed specifically for Taxus spp. (T. baccata, T. chinensis var. mairei (Lemee & Levl.) Cheng & L.K.Fu and T. cuspidata Siebold & Zucc (Ueno et al. 2015)). The trees sampled in this study represent all yew trees and seedlings exhaustively sampled in the NP and a random subsample from one adjacent population. In addition, close-by private forests may contain yew trees that were not accessible. Since we collected samples from three management zones, a stand outside of the NP and grouped them into two age cohorts (adults and seedlings), we can compare the population genetic diversity between two generations and investigate the pattern of genetic diversity in populations with different management intensity. We also assessed the sex distribution and estimated the genetic variation in male and female trees. Assessing the two cohorts’ samples provides a better understanding of the effect of forest management and conservation programs. During environmental changes, dispersal capacity plays a critical role in species’ establishment, persistence, and range dynamics. Hence, we characterized the fine-scale spatial genetic structure (SGS) to determine the patterns of family relationships in both cohorts. Effective population size estimates always reflect the effective population size of the previous generation. While the adult trees established a long time ago from a potentially diverse parental generation, we hypothesize a lower effective population size estimate based on the seedling generation due to periods with insufficient natural regeneration and non-random mating.
Materials and methods
Study population and plant material
Situated in South-East Bavaria (48.9597° N, 13.3949° E), the Bavarian Forest NP extends along the border to Czech Republic. The total area of the NP is 245 km2 and the elevation ranges from 600 to 1453 m a.s.l. An exhaustive sampling was conducted in the area to include all T. baccata individuals (Fig. 1 and Supplementary material, Fig. S1). According to the DBH histogram (Supplementary material, Table S3 and Figs. S2 and S3), in adult trees, the DBH ranged from 4 to 74.4 cm, and the mean DBH was 36.31 cm. The adult trees’ height was also recorded, it ranged from 2 to 22.1 m.
Needle samples (183 seedlings and 216 adult trees, 399 samples in total) were collected in three management zones of the NP and in a private forest in the vicinity of the NP from October to November. Despite collecting all seedling and adult samples from five forestry districts (1, 5, 7, 8 and 9), no natural regeneration was found in the central area of the NP in forestry district 7 (Supplementary material, Fig. S1A). In addition, natural regeneration is not uniform throughout the NP area; instead, it is confined to a limited number of patches surrounding mature trees.
The management zones in the NP encompass the natural zone, the management zone, and the development zones. The natural zone is an area where no forest management or human intervention occurs. The management zone, on the other hand, acts as a protective barrier for the forests adjacent to the NP. Its purpose is to perform conservation management (e.g., to protect T. baccata) and to safeguard adjacent forests from damage caused by bark beetles (König et al. 2023b), which can naturally develop within the NP. Effective forest protection measures, including bark beetle control (e.g., debarking of trees), are implemented within the management zone, which extends 500 m wide. Lastly, the NP’s development zones consist of forests designated to undergo a gradual transition towards natural development. These areas are intended to be incorporated into the natural zone over time (Table 1). As of the annual report in 2022, it was determined that 75% of the NP is now classified as natural zone (König et al. 2023a).
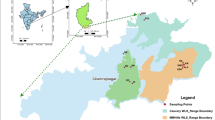
Taxus baccata sampling locations. Colors represent the different management zones (development zone, management zone, natural zone and outside of the Bavarian Forest National Park (NP), Germany, see Table 1) in the NP. Trees protected by fences are shown with black circles. The dark grey color depicts the area of the NP. For a map showing the distribution of seedlings see Supplementary material, Fig. S1A. Copyright: © 2023 Nationalparkverwaltung Bayerischer Wald, © Geobasisdaten: Bayerische Vermessungsverwaltung 2023, © Europäische Union, 1995–2023
Sex determination was based on the observation of male cones and seeds between October and November 2020. Trees with no reproductive structures observed were labeled with NA.
In total, the sex of 142 adult trees (93 females and 49 males) was identified and for the rest of the trees, the sex could not be determined (NA) (Fig. 2). Twenty adult trees in the development, management zones and outside of NP with their seedlings were protected by 10 × 10 m fences since 2016 (Supplementary material, Table S1), although, no seedlings were found around two female and four fenced adult trees with unknown sex.
The maximum distance between sampled trees from the North (forestry districts 1, 5 and 7) to the South (forestry districts 8 and 9) of the sampling area was 22.7 km. All the samples were collected from within the protected natural forests with the exception of samples (21 adult trees and 44 seedlings) collected from a privately owned forest (von Poschinger; https://www.poschinger.de/land-und-forst/), outside the protected area. Development zones belong to the northern part of the NP and individuals from outside the NP are located South of the sampling area (Supplementary material, Fig. S1B). In addition to diameter at breast height (DBH at 1.40 m), the height of the adult trees and GPS (GPSMAP® 65s, Garmin, USA) coordinates for each individual (seedling and adult) were recorded. In T. baccata the DBH is significantly correlated with tree age (Iszkuło and Boratyński 2005), hence, trees with a greater DBH indicate advanced age.
DNA extraction, marker testing, and genotyping
DNeasy 96 Plant Kit (Qiagen, Germany) was used to extract genomic DNA from dried needles according to the manufacturer’s protocol (Qiagen, Germany). We assessed DNA quality and quantity in 1% agarose gels stained with Roti®Gelstain (Carl ROTH, Germany), then visualized under UV light and compared them with a Lambda DNA size marker (Roche, Germany). Isolated DNA was diluted 1:10 for PCR amplification. Forty-one SSR markers (Supplementary material, Table S2), including 25 EST-SSRs originally developed for T. baccata, T. cuspidata, a close relative of T. baccata, T. chinensis var. mairei and T. x media (Ueno et al. 2015), eight chloroplast SSRs developed for Pinus thunbergii (cpSSRs) (Vendramin et al. 1996; Liu et al. 2019) and eight nuclear SSRs developed for T. baccata (Dubreuil et al. 2008; Huang et al. 2008), were tested in eight samples to amplify and detect polymorphisms. For all markers, amplification success was evaluated by visualizing PCR products on agarose gels (Supplementary material, Table S2). 13 EST-SSR markers originally developed for Taxus spp. (Supplementary material, Table S2) showed clear amplification and polymorphism when amplification products were separated on an ABI 3130xl Genetic Analyzer (Applied Biosystems, USA). Ultimately, these 13 polymorphic markers were genotyped in all samples. Polymerase chain reactions (PCRs) were performed with M13 tails (5’-CACGACGTTGTAAACGAC-3’) and dye-labeled adaptors complementary to the forward primers (Schuelke 2000; Kubisiak et al. 2013) and PIG-tail sequences (5’-GTTTCT-3’) (Brownstein et al. 1996) added to the 5’ end of reverse primers. A touchdown program was used for amplification with the following protocol in a Biometra thermocycler (Analytik Jena, Germany): first denaturation at 95 °C for 15 min, followed by ten cycles including a denaturation step of 1 min at 94 °C, an annealing step at 60 °C for 1 min (-1 °C per cycle), an extension step at 72 °C for 1 min, then 25 cycles with the same denaturation and extension time and temperature, but 50 °C annealing for 1 min, and a final extension at 72 °C for 20 min.
The PCR mix consisted of 1.0 µL genomic DNA (ca. 10 ng/µL), 1.5 µL 10x reaction buffer B (Solis BioDyne, Estonia), 1.5 µL MgCl2 (25 mM), 1.0 µL dNTPs (2.5 mM each dNTP), 0.2 µL (5 U/ µL) HOT FIREPol® Taq DNA polymerase (Solis BioDyne, Estonia), 0.2 µL tailed (Schuelke 2000; Kubisiak et al. 2013) forward primer (5 pM/µL), 0.5 µL PIG-tailed (Brownstein et al. 1996) reverse primer (5 pM/µL), 1 µL (5 pM/µL) dye-labeled (6-FAM or HEX) M13 primer, and HPLC grade H2O (filled up to a volume of 13 µL). GS 500 ROX (Applied Biosystems, USA) was used as an internal size standard in the genetic analyzer ABI 3130xl. We visualized and called fragment sizes of PCR products using GeneMapper v.4.0 software (Applied Biosystems, USA).
Genetic variation and differentiation
Among the 25 EST-SSRs (Ueno et al. 2015) 13 showed polymorphism. Five of the polymorphic loci were derived from T. baccata (B-16372-214B, B-26,615–282 A, B-23775-301D, B-20,918–231 C and B-17,952–496 C), five from T. cuspidata (C-52987-385B, C-29073-907B, C-703-364D, C-56,747–365 A and C-58157-573B) and three from T. mairei (Ma-14186-166D, Ma-10038-2563B and Ma-10,506–40 C). These were the 13 markers used in subsequent analyses. Other markers did not yield reliable genotypes. One of the EST-SSRs (B-12953-780B) did not amplify and three were monomorphic (C-52641-193B, Ma-10302-425D and Ma-18,795–254 C), while the remaining markers produced weak or non-specific amplification products. Out of eight nuclear SSRs, six produced non-specific bands and two did not amplify, even though all of them were originally developed for T. baccata (Dubreuil et al. 2008; Huang et al. 2008). One of the cpSSRs (cpSSR-33), originally developed for Pinus thunbergii was polymorphic, five were monomorphic (cpSSR‐00, cpSSR‐34 from P. thunbergii) and the remaining did not amplify. The cpSSR‐33 was not highly polymorphic and only two haplotypes were found without clear geographic pattern in the adult cohort (Supplementary material, Fig. S4).
MICROCHECKER v.2.2.3 (Van Oosterhout et al. 2004) was used to check for the presence of null alleles. Null alleles were detected with low frequencies in four markers in seedlings (C-52987-385B, B-23775-301D, C-58157-573B and B-17,952–496 C) and in three markers (C-56,747–365 A, C-58157-573B and B-17,952–496 C) in adult cohorts. We used SPAGeDi 1.5d (Hardy and Vekemans 2002) to assess the genetic diversity parameters, number of alleles per locus and averaged over loci (Na), effective number of alleles (Nae), allelic richness on a standardized sample of gene copies (AR), and observed and expected heterozygosity (Ho, He) over all loci. The inbreeding coefficient (Fis) was calculated at the eight markers with no indication of null alleles, for age cohorts, forest management zones and for the two sexes and also for each EST-SSR marker separately. Furthermore, due to the limited number of samples available from the management zone (27 samples), we randomly selected 27 samples from each forest zone and assessed genetic diversity characteristics accordingly. Also, analysis of molecular variance (AMOVA) was performed using GenAlEx v.6.5 (Peakall and Smouse 2006). FST was used to determine genetic differences between age cohorts, different management zones and the two sexes.
The significance of differences in genetic diversity parameters, observed and expected heterozygosity were tested, among the age cohorts and the two sexes (females and males and non-recognized sex) using a Kruskal-Wallis test with multiple comparisons implemented in the R statistical software (v4.2.2; R Core Team 2022) package pgirmess v.2.0.0 (Giraudoux 2022). Also, associations between individual heterozygosity (proportion of heterozygous loci/sample) and DBH (4–40 cm and 40–74.4 cm, small and large DBH groups, respectively) were evaluated by Pearson’s correlation coefficient. To visualize the results of the correlation between heterozygosity and DBH, a scatter smooth plot was generated in the R package stats v.4.2.2 using the function ggscatter from the ggpubr package v.0.4.0 (Kassambara 2020).
To understand the relationships and genetic structure between adults and seedlings and also between forest zones, a Principal Coordinate Analysis (PCoA) and a Bayesian clustering algorithm implemented in STRUCTURE v2.3.4 (Pritchard et al. 2000) were used. PCoA was performed using GenAlEx v.6.5 (Peakall and Smouse 2006) based on Nei’s unbiased genetic distances between individual samples (Nei 1978). In STRUCTURE, we ran the admixture model with correlated allele frequencies and tested K 1 to 10 with 10 replicates per K. Each run consisted of a length of burn-in period of 10,000 followed by 100,000 Markov Chain Monte Carlo (MCMC) replicates. For the choice of the most likely K, the ΔK method described by Evanno et al. (2005), implemented in STRUCTURE HARVESTER v.0.6.94 (Earl and VonHoldt 2012), was used. Bar plots for the optimum K were visualized using CLUMPAK pipeline (Kopelman et al. 2015). Pie charts representing admixture results for K = 2 were generated by the R package scatterpie (Yu 2021) and ggplot2 (Wickham 2016), showing the spatial distribution of seedling and adult individuals in forest zone 5 in the NP.
The fine-scale spatial genetic structure (SGS) was estimated using SPAGeDi 1.5d (Hardy and Vekemans 2002), with the pair-wise Loiselle kinship coefficient F between individual samples (Loiselle et al. 1995) regressed on the logarithm of spatial distances. This was done separately for the two age cohorts (adult and seedlings). In order to quantify the fine-scale SGS, the Sp statistic was estimated. The Sp statistic was obtained from the formula Sp = -bf/(F1-1), where F1 is the mean kinship coefficient of individual pairs belonging to the first distance class and bf is the regression slope (Vekemans and Hardy 2004). 10,000 permutations of individuals among spatial positions were used to assess the significance of the regression slope (bf). This analysis was performed on each cohort separately to avoid parent-offspring pairs. In SPAGeDi, the distance classes were chosen ensuring that each distance class contained at least thirty individual pairs. Therefore, the first distance class for seedlings was 10 m and for adult trees it was 25 m.
Two approaches were used to detect signals of past demographic events. First, the T2 statistic implemented in INEST v.2.2 (Chybicki and Burczyk 2009), initially described by Cornuet and Luikart (1996), was used for the detection of a recent bottleneck. When using SSR markers, and the number of SSRs is less than 20, the recommended test and model to investigate the bottleneck event are Wilcoxon signed-rank test and the Two-Phase Model (TPM, Piry et al. 1999). However, some microsatellites are known to follow an Infinite Allele Model (IAM). Therefore, we report the bottleneck test statistics of both models, TPM and IAM. INEST v.2.2 provides improved p value estimates for the Wilcoxon signed-rank test based on 106 permutations (Chybicki 2017). We used the default settings for the TPM (proportion of multi-step mutations = 0.22 and average multi-step mutation size = 3.1) to assess the bottleneck effect.
Additionally, we tested for ancient bottleneck events by calculating the M ratio according to Garza and Williamson (2001) under TPM and IAM. The M ratio (M = k/r) is defined as the number of observed alleles (k) over the number of expected alleles in the allele size range (r), which was also assessed using INEST. Both tests were run on the eight loci that showed no signs of null alleles.
Given the inherent complexities and challenges associated with estimating effective population size (Ne) in large continuous tree populations or small genetically connected units within such populations, it is imperative that Ne estimates are interpreted with a high degree of caution (Santos-del-Blanco et al. 2022). Liesebach et al. (2024) stated that the sibship assignment (SA) method provides more robust results than the other Ne estimators. The unique characteristics of tree populations, including their extensive spatial distribution and often vast numbers, pose specific methodological and statistical hurdles that demand careful consideration.
It is worth emphasizing that in the case of large populations, the estimation of contemporary Ne based on genetic data may introduce a downward bias that persists even as more genetic loci are used (Santos-del-Blanco et al. 2022). Considering these challenges, NeEstimator v.2.1 1 (Do et al. 2014) and COLONY 2.0.6.9 (Jones and Wang 2010) software were used to estimate the effective population size (Ne). However, it is important to mention that even with the use of these software and statistical methods, interpreting Ne estimates in such complex populations should be approached with a cautious and nuanced perspective.
The linkage disequilibrium (LD) method (Hill 1981; Waples 2006; Waples and Do 2010) for seedlings and adult trees was used as implemented in NeEstimator (Ne(LD)). Assuming random mating/absence of inbreeding, the most suitable method for estimating Ne is the LD method. We reported parametric confidence intervals based on a chi-square approximation (Waples 2006). The threshold of 0.02 was used as the lowest allele frequency to estimate the Ne values.
Additionally, the adult cohort was divided into two groups based on DBH size, smaller and larger than 36 cm. COLONY uses the sibship assignment method to estimate Ne (Ne(SA)) which is described by Wang (2009) as proportion of full-, half-sib and unrelated pairs among a cohort of genotyped individuals. This method can provide additional insights into Ne when combined with the LD estimates. It offers a different perspective by considering the familial relationships among individuals and can contribute to a more comprehensive understanding of the population dynamics. Based on sibship assignments, the number of full-siblings and half-siblings in the sample, Ne was estimated. We used the following settings for COLONY: female and male polygamy, with inbreeding system, dioecious species, medium length of run, the full-likelihood analysis method, medium likelihood precision, only one run and 1000 random number seed. We assumed a random mating population so deviations from Hardy Weinberg equilibrium are negligible (Alpha = 0). Among the adult trees, 93 were identified as females, 49 as males and for 74 trees the sex could not be determined (NA) (Supplementary material, Table S3). To estimate Ne, the sample includes males, females, and seedlings. Individuals with NA sex are included in both the male and female groups as potential mothers and fathers.
Results
Genetic diversity
Fis estimates for EST-SSRs ranged from − 0.028 to 0.235 for individual markers and in combination did not indicate inbreeding (Supplementary material, Table S4). The Fis values for four markers were negative, but not significantly different from zero (B-20,918–231 C, B-26,615–282 A, Ma-10038-2563B, and C-29073-907B).
The level of genetic diversity was not significantly different between the adults and seedlings (Table 2). In the seedlings, He was 0.612 and Ho was 0.587; in the adult cohort, He and Ho were 0.614 and 0.587, respectively. Mean He and Ho, in all samples, were 0.615 and 0.588, respectively. Neither Ho nor He significantly differed between development, management, natural zones and outside of the NP. Fis values based on the 8 loci without evidence of null alleles in seedlings and adult trees were not significantly different from zero (Table 2).
A total of 72 alleles were observed across the 13 markers in 399 individuals. Genetic diversity characteristics based on all samples and also based on 27 samples, between different zones were similar and did not differ significantly (Table 2). He and Ho and allelic richness were similar in female and male groups, and no significant difference was observed (Supplementary material, Table S5). Based on the Kruskal-Wallis test, differences in observed and expected heterozygosity between the two age cohorts and female and male trees were insignificant. The genetic variation over all 13 loci for seedling and adult cohorts is presented in the Supplementary material, Tables S6 and S7.
Allelic richness values (AR, k = 364) in seedlings and adults, were 5.31 and 5.13, respectively, showing slightly higher AR in seedlings (Table 2). Comparing the genetic diversity of adults and seedlings in different zones showed slightly higher allelic richness for adults in the Natural and the Development zone (Supplementary material, Table S8). Among the different zones, the natural zone has a higher AR (4.42) than the other zones, with the lowest AR being 4.31 for the management zone, however, none of the differences was significant (Table 2).
Seedling and adult cohorts and also different management zones showed low but significant differentiation (between age cohorts: FST = 0.007, p < 0.001 and among management zones: FST = 0.008, p < 0.001). In addition, a low and non-significant FST value was found between sexes (female, male, FST < 0.0001, p = 0.394).
Correlation analyses revealed a significant positive but weak association between individual heterozygosity and DBH for larger adult trees (DBH: 40.0–74.4 cm), but not for smaller adult trees (4.0–40 cm). However, based on all samples (4.0 to 74.4 cm) a significant correlation between DBH and individual heterozygosity was observed (Supplementary material, Figs. S5, S6 and S7).
Genetic structure of the population
Based on Nei’s unbiased genetic distance, PCoA (Principal Coordinates Analysis) did not reveal any genetic structure between cohorts and between forest zones (Supplementary material, Fig. S8). Furthermore, no clear genetic clusters were found in the STRUCTURE analysis (Fig. 3 and Supplementary material, Fig. S9). However, as opposed to the rest of the population, some seedlings and mature trees particularly in forestry district 5 belong to a different cluster (Supplementary material, Fig. S9). The ΔK approach by Evanno (Evanno et al. 2005) indicated K = 2 as best explaining our data (Supplementary material, Figs. S11 and S12). Visual inspection showed some individuals may belong to different clusters, a pattern reflected in both age cohorts, but more common in seedlings (Fig. 3 and Supplementary material, Fig. S9). In general, seedlings were found in the vicinity of a limited number of adults (183 seedlings around only 39 adults). The distribution of these seedlings is very dense. This composition is better seen in forestry district 5, where half of the seedlings are located. Here, a large fraction of seedlings are assigned to the same genetic cluster as a small fraction of adjacent adults (Fig. 3). Using COLONY software, the analysis of seedlings and adults in this district confirmed their parent-offspring relationship.
Structure barplot of ancestry proportions of two genetic clusters (K = 2) for 399 Taxus baccata samples sorted by cohort and forestry zones in the Bavarian Forest National Park (NP), Germany. K: 2; A.1, adult trees, forest zone 1; S.1, seedlings, forest zone 1; A.5, adult trees, forest zone 5; S.5, seedlings, forest zone 5; A.7, adult trees, forest zone 7; A.8 adult trees, forest zone 8; S.8, seedlings, forest zone 8; A.9, adult trees, forest zone 9; S.9, seedlings, forest zone 9
Fine-scale spatial genetic structure
The fine-scale SGS indicated a significant family structure in both cohorts. The results showed a strong and significant Sp value in seedling and adult cohorts of 0.0120 and 0.0063, respectively (Table 3; Fig. 4), revealing a stronger fine-scale SGS in the seedling cohort. The minimum number of pairwise comparisons per distance class in the seedling cohort was 83, whereas it was 34 in the adult cohort. A notable fine-scale spatial genetic structure, extending roughly 1000 m in seedlings and 400 m in adults was observed.
Fine-scale spatial genetic structure in seedling and adult Taxus baccata cohorts in the Bavarian Forest National Park (NP), Germany using EST-SSRs. The mean Loiselle kinship coefficient per distance class, FD, was plotted against the logarithm of geographical distances between individuals. Dashed gray lines represent the 95% confidence interval
Bottleneck and effective population size
It has been suggested that a significant heterozygosity excess in populations can be attributed to a bottleneck that occurred recently. Under TPM and IAM, significant p values and positive T2 values showed signs of recent population size reduction in the NP; although in adult trees, under TPM, it was statistically not significant (T2 p value = 0.054), there is a clear tendency. Interestingly, Garza-Williamson’s M ratio, most suitable for detecting more ancient bottleneck signals, showed no significant M ratio under TPM but significance under IAM (Table 4).
Effective population size based on linkage disequilibrium (Ne(LD)) when allowing for 0.02 as the lowest allele frequency was estimated as 50 (95% CI = 42.1–59.7) and 234 (95% CI = 167.6–358.4) for seedling and adult cohorts, respectively (Table 5). Also, Ne for the small DBH size class of the adults of 42.2 (95% CI = 15.5–174.5), is less than for the large DBH size class of adults equaling 279.3 (95% CI = 33.9 – ∞). Additionally, effective population size, according to the sibship assignment method (Ne(SA)), was estimated as 61 (95% CI = 43–86) for the whole population.
Discussion
The genetic diversity of species is a crucial component of biodiversity, as it enables adaptation to different environments (Hoban et al. 2020). This is especially important during an era of rapid global change that challenges local adaptation of natural populations. To conserve and manage forest genetic resources, it is critical to understand the genetic impacts of forest management practices on genetic diversity and counteract these negative effects (Ratnam et al. 2014).
Dynamic natural forests should exhibit a greater abundance of trees with smaller diameters, indicating abundant natural regeneration. Throughout the NP, T. baccata exhibited a scattered natural regeneration and a low representation of small-diameter classes, indicating that the level of natural regeneration is low. A total of only 29 young trees are present (less than 25 cm DBH), which is considerably fewer than the 150 trees in the intermediate size class (25–50 cm DBH). In all categories, by sex and forest zone, low DBH individuals are less abundant. In the forest zone outside the NP, average DBH is 6 cm larger (ranges from 31.5 to 42.5 cm, equally distributed) than that of other inner areas of the NP (Supplementary material, Table S3 and Figs. S2 and S3).
The results indicated that there were low levels of natural regeneration in the NP for a considerable period of time. Nevertheless, the conservation management in the NP and outside has led to the emergence of seedlings in recent years. The fencing protected seedlings that are vulnerable to damage, such as browsing by deer.
Genetic diversity
The population of T. baccata in the NP, characterized using EST-SSRs, revealed comparatively high levels of heterozygosity. The heterozygosity in this study (He = 0.615) was higher than in other yew populations studied using 18 EST-SSRs in Poland (He = 0.564) (Stefanowska et al. 2021) with partly the same markers. Furthermore, the application of 13 isoenzymes by Rau et al. (2021) on 19 Bavarian yew populations revealed considerable diversity in the NP.
The different cohorts studied here (seedling and adult) did not show statistically significant differences in genetic diversity. Consequently, the genetic makeup of individuals within a population is relatively uniform, regardless of their age or life stage. However, a positive correlation exists between individual heterozygosity and DBH at later developmental stages (see below). Therefore, we conclude that the population will not undergo genetic erosion if natural regeneration is protected and promoted in development, management, and natural zones and outside of the NP.
Cao et al. (2004), reported nearly equal numbers of female and male trees in six German T. baccata populations. There were 116 female and 108 male individuals, with only one population showing an exception by deviating from the 1:1 sex ratio. Furthermore, based on isozyme and SSR loci, significant disparities in genetic structure between the female and male trees within the stands were detected, a phenomenon that may be attributed to genetic drift and the small size of the populations. Populations with greater numbers of individuals displayed higher levels of heterozygosity, highlighting the influence of population size on the genetic diversity of T. baccata in Germany. In the NP, the genetic diversity of females (He = 0.616) and males (He = 0.614) and trees with unknown sex (He = 0.611) of T. baccata did not show any significant difference.
Although both male and female T. baccata individuals are found in the NP, they may respond differently to environmental stress and adversity (Juvany and Munné-Bosch 2015). The sex ratio in the study, at least for recognizable individuals, was found to deviate from the expected 1:1 ratio, with a higher proportion of female individuals present (~ 2:1). Since male trees are difficult to recognize, the number of female trees could have even been underestimated (immature female trees). Factors contributing to similar levels of expected heterozygosity in T. baccata despite different male and female sample sizes include its outcrossing nature and efficient means of gene dispersal, possibly resulting in similar levels of genetic variation in males and females and low genetic differentiation between sexes. Factors such as efficient pollen dispersal by wind, the species’ longevity, and slow growth may help maintain genetic diversity over time.
Fis estimates in our study (Table 2) did not deviate significantly from zero and were similar to the reported Fis (0.007) in Poland (Stefanowska et al. 2021) with partly the same EST-SSRs. The Fis in the present study and in Stefanowska et al. (2021) are the lowest inbreeding coefficients estimated for T. baccata in the distribution range of the species, while other studies using nSSRs have estimated higher values (Dubreuil et al. 2010; Chybicki et al. 2012; Gargiulo et al. 2019; Maroso et al. 2021; Hematzadeh et al. 2023) possibly as result of a higher frequency of null alleles at non-genic nuclear SSRs. Besides, other Taxus species have been observed to exhibit different levels of inbreeding. For example in southwestern China species T. yunnanensis, Miao et al. (2016) observed high Fis values, with extreme mean Fis values reaching 0.702. Similar observations were reported in T. contorta from Pakistan (Fis = 0.219–0.418; Poudel et al. 2014) and in T. mairei from eastern and southern China (Fis = 0.24; Luo et al. 2021).
The absence of signatures of inbreeding and the comparatively high level of genetic variation suggest efficient gene flow within the NP and possibly with populations from Šumava National Park on the other side of the border with hundreds of yew individuals (Komárková et al. 2022). The nearest natural population to the NP in Germany is approximately 100 km away (Regensburg), and other species occurrences are mostly plantations or overgrown garden forms.
Correlation between gene diversity and DBH
The correlation observed between a trees’ DBH and its individual multilocus heterozygosity suggests a discernible pattern of selection favoring higher levels of individual heterozygosity over time (Supplementary Figs. S5, S6 and S7). Although, the correlation is not significant for trees with DBH from 4 to 40 cm, it shows a significant and positive relationship for all samples (DBH from 4 cm to 74.4 cm) and especially for DBH from 40 to 74.4 cm. These results can be interpreted as a selective advantage of heterozygous individuals, especially at later developmental stages.
Similar positive correlations between ontogenic stages and diversity have been observed in other species. For example, in Pitch pine (Pinus rigida Mill), trees with higher heterozygosity exhibit greater mean annual growth, with the relationship between growth rate and heterozygosity increasing with trees age (Ledig et al. 1983). Similarly, the tropical tree species Hymenaea courbaril and Jacaranda copaia show an increase in heterozygosity with the age of individuals (Jones and Hubbell 2006; de Lacerda et al. 2008). Furthermore, it has been observed that seedlings of Ardisia escallonioides exhibit an excess of homozygosity compared to the adult trees in most populations (Pascarella 1997). Similarly, in offspring populations of Thuja occidentalis, a reduced level of heterozygosity has been reported compared to the parent stands (Perry and Knowles 1990).
Genetic structure of the population
In forestry district 5 (Supplementary material, Fig. S1) most of the seedlings were assigned to only one genetic cluster (62 out of 91 seedlings) that is underrepresented in the NP and shared by only a small group of adult trees (11 out of 76 trees) in this district (Supplementary material, Fig. S10). These seedlings are clustered around the few adult trees of the same genetic cluster and apparently originate from matings between a limited number of adults. The close genetic relatedness of these adult trees and seedlings assessed by COLONY confirms this conclusion.
Fine-scale SGS
Several factors can affect the SGS in T. baccata, including limited gene flow because of restricted seed and pollen dispersal (Chybicki et al. 2016) and dioecy (Nazareno et al. 2013). Dioecious plants may face limited gene dispersal and reduced genetic variability resulting from increased genetic drift when there is an unbalanced sex ratio (Allendorf et al. 2012). Furthermore, the sex ratio influences SGS as well (Chybicki et al. 2016). In conifer populations where males are underrepresented, the intensity of SGS is higher than for populations with balanced sex ratios, especially for paternally compared to biparentally inherited DNA (Chybicki et al. 2016). In the NP, the number of male individuals is approximately equal to half of the female trees, at least among individuals whose sex has been identified. Also considering the challenges and uncertainties associated with determining the sex of male trees, it is important to note that the limited number of male trees might influence SGS. Unlike the observed distribution pattern in central Italy (Garbarino et al. 2015), where female T. baccata trees displayed a clumped distribution at a scale of 30 m and male trees exhibited random distribution, in the NP, a more random distribution pattern characterizes both the male and female trees.
In addition, species with effective means of pollen and seed dispersal (through wind or biotic vectors) show weaker SGS compared to species with gravity-dispersed seeds (Vekemans and Hardy 2004). Certain species’ populations can exhibit significant SGS despite being wind-pollinated, attributed to pollen and seed movement limitations. The two T. baccata natural populations in Poland assessed at SSR markers showed strong SGS (Sp = 0.006 and 0.009) (Chybicki et al. 2011). The Sp value (0.006) for the adult cohort is comparable to that observed in Poland. However, the Sp value exhibited by the seedlings is higher. This disparity can be attributed to the distance between seedlings and the mother tree and the high density of seedlings where natural regeneration occurs. Consequently, the family structure within the seedling population demonstrates more pronounced family structures than that observed in the adult trees. This underscores the importance of conservation efforts, particularly in the development and management zones, to prioritize conservation and promotion of natural regeneration. Population density has been frequently acknowledged as a critical factor influencing genetic structure (Vekemans and Hardy 2004).
The presence of family structures may promote biparental inbreeding and a decrease in effective population size over time. However, we found no evidence for inbreeding in the adult or seedling generation, but comparatively low values of effective population size (see below).
Bottleneck and effective population size
Various factors can cause bottlenecks, including habitat fragmentation (Luikart et al. 1998), overharvesting (Fageria and Rajora 2013), and changing environmental conditions (Canham and Murphy 2017).
The results from the bottleneck analysis suggested the occurrence of a significant heterozygosity excess in the populations, indicating a possible recent bottleneck event. However, when specifically considering adult trees under TPM, the reduction is not statistically significant, although there is a clear tendency for a population size reduction.
Interestingly, the analysis of Garza-Williamson’s M ratio, which is more suitable for detecting ancient bottleneck signals, reveals no significant M ratio under TPM but shows significance under IAM. This suggests that the population may have experienced ancient bottleneck events as detected by the IAM model.
In a recently studied T. baccata population in Northern Iran (the eastern part of the distribution range of the species), no bottleneck effect in most of the populations was observed (Hematzadeh et al. 2023), however, in Poland (the western part of the distribution range of the species) significant but variable signals of genetic bottlenecks were observed (Stefanowska et al. 2021). The findings obtained from the TPM model for the adult cohort in our study align with Stefanowska et al. (2021) findings in Poland, indicating less pronounced genetic bottlenecks and larger effective population sizes in the adult compared to the seedling generation. Thus, promoting natural regeneration is necessary to widen the population’s age distribution and maintain their genetic variation.
Human intervention can cause fragmentation of individual populations (Franklin and Gutiérrez 2002; Chybicki et al. 2011), consequently, the absence of spatial continuity among populations can result in an increase in inbreeding (Ledig 1992). This, in turn, can lead to a decline in the effective population size (Gilpin 1991). Accurate estimates of Ne in the NP face complex challenges, including the species’ metapopulation structure and overlapping generations. Overlapping generations of the species can also violate the assumption of discrete generations (Santos-del-Blanco et al. 2022). As a consequence of these challenges, accurately estimating the effective population size becomes problematic.
Ne was expected to be lower in seedlings as the genetic diversity of the seedlings is derived from only a subset of the adult population, which can result in a lower effective population size due to genetic drift.
Ne(LD) estimates for seedling and adult cohorts indicated that the Ne of adult individuals is larger than that of the seedlings. The wider confidence intervals for the adult cohort indicate greater uncertainty in the Ne estimation or larger effective population sizes. Additionally, the Ne(SA) estimate based on sibship assignment in COLONY was 61. This method takes into account the actual family structure within the population and provides a more detailed understanding of mating patterns and family sizes. It provides an estimate for the current effective population size which indeed yielded a similar estimate as the Ne(LD) estimates of the seedling cohort and of the young adult cohort. This effective population size of the natural regeneration and young adult cohort is not sufficient for the long term survival of this local population. Effective management measures such as fencing of all female trees and removing competitors must be undertaken to maintain genetic variation of the current adult tree population (Table 5, Ne = 234).
The relatively smaller Ne(LD) for seedlings and young adults compared to older adults suggested a decline and a potential vulnerability to genetic drift and reduced genetic diversity. In contrast, the larger Ne(LD) for adult and large adult stages indicated a previously more robust effective population size. These findings highlight the importance of considering different life stages when assessing population dynamics and conservation strategies.
Implications for conservation
Selective cutting of competitor species and fencing in the development zone have shown a significant positive impact on the abundance of natural regeneration around the parent trees. As a result of selectively removing trees of other species, the parent trees and their natural regeneration were provided with more resources, space, and light. Fences also protected natural regenerations from being browsed by deer. Although the development zone is comparatively small (Fig. 1), half of the sampled individuals in this study were found in this area (108 seedlings and 92 adults). In addition, the presence of shrubs is known to positively influence natural regeneration of yew (Garcı́a et al. 2000; García and Ramón Obeso 2003; Devaney et al. 2014; Calvia et al. 2023). Nevertheless, no such observations were made in the NP, where most yew locations lacked a shrub layer (Dunkel 2017). By increasing light penetration, selective cutting enhances natural regeneration.
Conservation strategies should aim to increase the genetic diversity of the population, by promoting the establishment of new seedlings from a variety of different adult trees, and by reducing human disturbances that may affect the reproductive success of the trees. In forestry district 5 (Supplementary material, Figs. S9 and S10), two genetic clusters are found in adult trees and their natural regeneration. Most seedlings spatially cluster around and originate from only a few adult trees. Seed harvest activities and in situ conservation measures should consider this genetic differentiation at a small spatial scale. Selective removal of competitors should be conducted to promote natural regeneration of underrepresented female trees.
Planting new trees using reproductive material from local sources is suitable in areas where the density of yew is low or the species has disappeared. The yew population in the NP heavily depends on the presence of fences and active management practices to ensure its survival and long-term viability as otherwise natural regeneration appeared to be heavily browsed. Effective fencing is essential for ensuring for long-term conservation, spanning a minimum of 20 years, required to support natural regeneration (Vancura et al. 2004).
In the NP, 94 seedlings from fenced adult trees and 89 seedlings from unfenced adult trees were observed. Notably, these 94 seedlings from fenced trees were derived from a relatively limited number of only 20 fenced adult trees within the sampled area (Supplementary material, Fig. S13). Without such measures, the prospects for natural regeneration would be severely hindered, ultimately jeopardizing the population’s future existence. Over time, the lack of new individuals joining the population and of gene influx from outside would lead to a decline in overall numbers, increasing the risk of local extinction and irreversible loss of this valuable species within the NP. Changing environmental conditions, particularly due to climate change, may adversely impact both the distribution and genetic diversity of this species. Yew may struggle to swiftly adopt to changing climatic conditions, potentially resulting in its extinction (Thomas and Garcia-Marti 2015; Alavi et al. 2019).
Conclusion
The genetic diversity of T. baccata is critical for its ability to adapt to changing environmental conditions, especially in the face of rapid global changes. Conservation efforts should consider the genetic impacts of forest management practices to ensure the long-term viability of this species.
The study revealed several key findings. First, the low representation of small-diameter trees in the NP indicated limited natural regeneration over an extended period. However, conservation efforts, including selective removal of competitor species and fencing, have led to the emergence of seedlings in recent years. These measures protect vulnerable seedlings from browsing, contributing to the species’ recovery.
Genetic diversity assessments using EST-SSRs revealed high heterozygosity in the NP population. Unlike prior studies in other populations, we found no significant inbreeding. Furthermore, the analysis of genetic bottlenecks indicated a relatively recent bottleneck event, possibly caused by habitat fragmentation and limited natural regeneration. The population size reduction tendency, underscored the importance of promoting natural regeneration to maintain genetic diversity.
The presence of significant family structures in both adult and seedling cohorts highlighted the potential impact of spatial aggregation and restricted gene flow on genetic structure. While inbreeding was not detected, the relatively small effective population size of seedlings and small adults suggested vulnerability to genetic drift. This effective population size (Ne ~ 50 individuals) is not sufficient for the long-term survival of this local population. Effective management measures such as fencing of all female trees, implementation and test of fenced areas of different size, and removal of competitors have to be undertaken to maintain the levels of genetic variation of the current adult tree population.
In summary, the study emphasized the need for conservation efforts that prioritize protecting and promoting natural regeneration in T. baccata populations. Selective cutting of neighboring competitor species, fencing, and habitat management have proven effective in enhancing seedling survival. These strategies should be combined with planting new trees from local sources to increase genetic diversity. The continued presence of fences and active management practices is crucial for the long-term viability of T. baccata populations, as they prevent decline, local extinction, and irreversible loss of this valuable species within the NP.
Data availability
EST-SSR data, GPS coordinates, and diameter at breast height measures for adult trees were deposited in the G?ttingen Research Online repository (Link: https://doi.org/10.25625/HQULGY).
Code availability
Not applicable
References
Ahmadi K, Jalil Alavi S, Zahedi Amiri G, Mohsen Hosseini S, Serra-Diaz JM, Svenning JC (2020) Patterns of density and structure of natural populations of Taxus baccata in the Hyrcanian forests of Iran. Nord J Bot 38:e02598. https://doi.org/10.1111/njb.02598
Alavi SJ, Ahmadi K, Hosseini SM, Tabari M, Nouri Z (2019) The response of English yew (Taxus baccata L.) to climate change in the Caspian Hyrcanian mixed Forest Ecoregion. Reg Environ Change 19:1495–1506. https://doi.org/10.1007/s10113-019-01483-x
Allendorf FW, Luikart GH, Aitken SN (2012) Conservation and the genetics of populations. Wiley, Oxford
Benham S, Houston Durrant T, Caudullo G, de Rigo D (2016) Taxus baccata in Europe: distribution, habitat, usage and threats. https://forest.jrc.ec.europa.eu/en/european-atlas/atlas-download-page/. Accessed 25 June 2023
BLE-Bundesanstalt für Landwirtschaft und Ernährung (2013) Common Yew. https://www.genres.de/en/sector-specific-portals/trees-and-shrubs/rare-tree-species/common-yew. Accessed 19.03.2023
BMELV-Bundesministerium für Ernährung, Landwirtschaft und Verbraucherschutz (2012) Aufnahmeanweisung für die dritte Bundeswaldinventur (2011–2012). 2. geänderte Auflage
Brownstein MJ, Carpten JD, Smith JR (1996) Modulation of non-templated nucleotide addition by taq DNA polymerase: primer modifications that facilitate genotyping. BioTechniques 20:1004–1010. https://doi.org/10.2144/96206st01
Calvia G, Casula P, Farris E, Fenu G, Fantini S, Bacchetta G (2023) Shrub cover and soil moisture affect Taxus baccata L. regeneration at its southern range. Plants 12:1819. https://doi.org/10.3390/plants12091819
Canham CD, Murphy L (2017) The demography of tree species response to climate: sapling and canopy tree survival. Ecosphere 8:e01701. https://doi.org/10.1002/ecs2.1701
Cao C-P, Leinemann L, Ziehe M, Finkeldey R (2004) Untersuchungen zur genetischen variation und Differenzierung Von Eibenbeständen Mit Hilfe Von Isoenzym- und DNS-Markern. Allgemeine Forst und Jagdzeitung 175:21–28
Chybicki IJ (2017) INEST 2.2. http://www.ukw.edu.pl/pracownicy/strona/igor_chybicki/software_ukw/
Chybicki IJ, Burczyk J (2009) Simultaneous estimation of null alleles and inbreeding coefficients. J Hered 100:106–113. https://doi.org/10.1093/jhered/esn088
Chybicki IJ, Oleksa A (2018) Seed and pollen gene dispersal in Taxus baccata, a dioecious conifer in the face of strong population fragmentation. Ann Bot 122:409–421. https://doi.org/10.1093/aob/mcy081
Chybicki IJ, Oleksa A, Burczyk J (2011) Increased inbreeding and strong kinship structure in Taxus baccata estimated from both AFLP and SSR data. Heredity 107:589–600. https://doi.org/10.1038/hdy.2011.51
Chybicki IJ, Oleksa A, Kowalkowska K (2012) Variable rates of random genetic drift in protected populations of English yew: implications for gene pool conservation. Conserv Genet 13:899–911. https://doi.org/10.1007/s10592-012-0339-9
Chybicki I, Dering M, Iszkuło G, Meyza K, Suszka J (2016) Relative strength of fine-scale spatial genetic structure in paternally vs biparentally inherited DNA in a dioecious plant depends on both sex proportions and pollen-to-seed dispersal ratio. Heredity 117:449–459. https://doi.org/10.1038/hdy.2016.65
Cornuet JM, Luikart G (1996) Description and power analysis of two tests for detecting recent population bottlenecks from allele frequency data. Genetics 144:2001–2014. https://doi.org/10.1093/genetics/144.4.2001
Crist E, Mora C, Engelman R (2017) The interaction of human population, food production, and biodiversity protection. Science 356:260–264. https://www.science.org/doi/10.1126/science.aal2011
de Beaulieu J-L, Andrieu-Ponel V, Reille M, Grüger E, Tzedakis C, Svobodova H (2001) An attempt at correlation between the Velay pollen sequence and the Middle Pleistocene stratigraphy from central Europe. Quat Sci Rev 20:1593–1602. https://doi.org/10.1016/S0277-3791(01)00027-0
de Lacerda AEB, Kanashiro M, Sebbenn AM (2008) Effects of reduced impact logging on genetic diversity and spatial genetic structure of a Hymenaea courbaril population in the Brazilian Amazon Forest. For Ecol Manage 255:1034–1043. https://doi.org/10.1016/j.foreco.2007.10.009
Deforce K, Bastiaens J (2007) The Holocene history of Taxus baccata (yew) in Belgium and neighbouring regions. Belg J Bot 140:222–237
Delahunty J (2007) The ethnobotanical history and holocene extent of yew (Taxus baccata L.) on the Irish landscape. J Ethnobiol 27:204–217. https://doi.org/10.2993/0278-0771(2007)27[204:TEHAHE]2.0.CO;2
Devaney JL, Jansen MA, Whelan PM (2014) Spatial patterns of natural regeneration in stands of English yew (Taxus baccata L.); negative neighbourhood effects. For Ecol Manage 321:52–60. https://doi.org/10.1016/j.foreco.2013.06.060
Dhar A, Ruprecht H, Vacik H (2008) Population viability risk management (PVRM) for in situ management of endangered tree species-A case study on a Taxus baccata L. population. For Ecol Manage 255:2835–2845. https://doi.org/10.1016/j.foreco.2008.01.059
Do C, Waples RS, Peel D, Macbeth G, Tillett BJ, Ovenden JR (2014) NeEstimator v2: re-implementation of software for the estimation of contemporary effective population size (Ne) from genetic data. Mol Ecol Resour 14:209–214. https://doi.org/10.1111/1755-0998.12157
Dubreuil M, Sebastiani F, Mayol M, González-Martínez SC, Riba M, Vendramin GG (2008) Isolation and characterization of polymorphic nuclear microsatellite loci in Taxus baccata L. Conserv Genet 9:1665–1668. https://doi.org/10.1007/s10592-008-9515-3
Dubreuil M, Riba M, González-Martínez SC, Vendramin GG, Sebastiani F, Mayol M (2010) Genetic effects of chronic habitat fragmentation revisited: strong genetic structure in a temperate tree, Taxus baccata (Taxaceae), with great dispersal capability. Am J Bot 97:303–310. https://doi.org/10.3732/ajb.0900148
Dunkel K (2017) Das Eibenvorkommen im Nationalpark Bayerischer Wald. Dissertation, University of Freiburg
Earl DA, VonHoldt BM (2012) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Res 4:359–361. https://doi.org/10.1007/s12686-011-9548-7
Eliades N-GH, Gailing O, Leinemann L, Fady B, Finkeldey R (2011) High genetic diversity and significant population structure in Cedrus brevifolia Henry, a narrow endemic Mediterranean tree from Cyprus. Plant Syst Evol 294:185–198. https://doi.org/10.1007/s00606-011-0453-z
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620. https://doi.org/10.1007/s12686-011-9548-7
Fageria MS, Rajora OP (2013) Effects of harvesting of increasing intensities on genetic diversity and population structure of white spruce. Evol Appl 6:778–794. https://doi.org/10.1111/eva.12064
Frankham R (1995) Conservation genetics. Annu Rev Genet 29:305–327
Franklin AB, Gutiérrez R (2002) Spotted owls, forest fragmentation, and forest heterogeneity. Stud Avian Biol:203–220
Gailing O, Vornam B, Leinemann L, Finkeldey R (2009) Genetic and genomic approaches to assess adaptive genetic variation in plants: forest trees as a model. Physiol Plant 137:509–519. https://doi.org/10.1111/j.1399-3054.2009.01263.x
Garbarino M, Weisberg PJ, Bagnara L, Urbinati C (2015) Sex-related spatial segregation along environmental gradients in the dioecious conifer, Taxus baccata. For Ecol Manage 358:122–129. https://doi.org/10.1016/j.foreco.2015.09.009
García D, Ramón Obeso J (2003) Facilitation by herbivore-mediated nurse plants in a threatened tree, Taxus baccata: local effects and landscape level consistency. Ecography 26:739–750. https://doi.org/10.1111/j.0906-7590.2003.03601.x
Garcı́a D, Zamora R, Hódar JA, Gómez JM, Castro J (2000) Yew (Taxus baccata L.) regeneration is facilitated by fleshy-fruited shrubs in Mediterranean environments. Biol Conserv 95:31–38. https://doi.org/10.1016/S0006-3207(00)00016-1
Gardner M, Christian T, Hinchliffe W, Cubey R (2019) Conservation Hedges. Sibbaldia: Int J Botanic Garden Hortic 17:71–100. https://doi.org/10.24823/Sibbaldia.2019.268
Gargiulo R, Saubin M, Rizzuto G, West B, Fay MF, Kallow S, Trivedi C (2019) Genetic diversity in British populations of Taxus baccata L.: is the seedbank collection representative of the genetic variation in the wild? Biol Conserv 233:289–297. https://doi.org/10.1016/j.biocon.2019.01.014
Garza J, Williamson E (2001) Detection of reduction in population size using data from microsatellite loci. Mol Ecol 10:305–318. https://doi.org/10.1046/j.1365-294x.2001.01190.x
Gilpin M (1991) The genetic effective size of a metapopulation. Biol J Linn Soc 42:165–175. https://doi.org/10.1111/j.1095-8312.1991.tb00558.x
Giraudoux P (2022) pgirmess: spatial analysis and data mining for field ecologists. R package version 1.7.0. https://CRAN.Rproject.org/package=pgirmess
González-Martínez SC, Dubreuil M, Riba M, Vendramin GG, Sebastiani F, Mayol M (2010) Spatial genetic structure of Taxus baccata L. in the western Mediterranean Basin: past and present limits to gene movement over a broad geographic scale. Mol Phylogenet Evol 55:805–815. https://doi.org/10.1016/j.ympev.2010.03.001
Hageneder F (2013) Yew. Reaktion Books, London
Hamrick JL, Godt MJW, Sherman-Broyles SL (1992) Factors influencing levels of genetic diversity in woody plant species. Population Genetics of Forest Trees. Springer, pp 95–124
Hardy OJ, Vekemans X (2002) SPAGeDi: a versatile computer program to analyse spatial genetic structure at the individual or population levels. Mol Ecol Notes 2:618–620. https://doi.org/10.1046/j.1471-8286.2002.00305.x
Hartzell H (1991) The yew tree: a thousand whispers: Biography of a species. Hulogosi Communications, Incorporated, Eugene
Hematzadeh A, Esmailzadeh O, Jalali SG, Mirjalili MH, Walas Ł, Yousefzadeh H (2023) Genetic diversity and structure of English yew (Taxus baccata L.) as a tertiary relict and endangered tree in the Hyrcanian forests. Biodivers Conserv 32:1733–1753. https://doi.org/10.1007/s10531-023-02573-3
Hertel H, Kohlstock N (1996) Genetische Variation und geographische Struktur von Eibenvorkommen (Taxus baccata L.) in Mecklenburg-Vorpommern. Silvae Genet 45:290–293
Heurich M, Brand TT, Kaandorp MY, Šustr P, Müller J, Reineking B (2015) Country, cover or protection: what shapes the distribution of red deer and roe deer in the Bohemian Forest Ecosystem? PLoS ONE 10:e0120960. https://doi.org/10.1371/journal.pone.0120960
Hill WG (1981) Estimation of effective population size from data on linkage disequilibrium. Genet Res 38:209–216. https://doi.org/10.1017/S0016672300020553
Hoban S, Bruford M, Jackson JDU, Lopes-Fernandes M, Heuertz M, Hohenlohe PA, Paz-Vinas I, Sjögren-Gulve P, Segelbacher G, Vernesi C (2020) Genetic diversity targets and indicators in the CBD post-2020 Global Biodiversity Framework must be improved. Biol Conserv 248:108654. https://doi.org/10.1016/j.biocon.2020.108654
Huang C-C, Chiang T-Y, Hsu T-W (2008) Isolation and characterization of microsatellite loci in Taxus sumatrana (Taxaceae) using PCR-based isolation of microsatellite arrays (PIMA). Conserv Genet 9:471–473. https://doi.org/10.1007/s10592-007-9341-z
Hulme PE (1996) Natural regeneration of yew (Taxus baccata L.): microsite, seed or herbivore limitation? J Ecol 84:853–861. https://doi.org/10.2307/2960557
Iszkuło G, Boratyński A (2005) Different age and spatial structure of two spontaneous subpopulations of Taxus baccata as a result of various intensity of colonization process. Flora: Morphol Distrib Funct Ecol Plants 200:195–206. https://doi.org/10.1016/j.flora.2004.03.001
Jia X, Feng S, Zhang H, Liu X (2022) Plastome Phylogenomics Provide Insight into the Evolution of Taxus. Forests 13:1590. https://doi.org/10.3390/f13101590
Jones FA, Hubbell SP (2006) Demographic spatial genetic structure of the Neotropical tree, Jacaranda copaia. Mol Ecol 15:3205–3217. https://doi.org/10.1111/j.1365-294X.2006.03023.x
Jones OR, Wang J (2010) COLONY: a program for parentage and sibship inference from multilocus genotype data. Mol Ecol Resour 10:551–555. https://doi.org/10.1111/j.1755-0998.2009.02787.x
Juvany M, Munné-Bosch S (2015) Sex-related differences in stress tolerance in dioecious plants: a critical appraisal in a physiological context. J Exp Bot 66:6083–6092. https://doi.org/10.1093/jxb/erv343
Kassambara A (2020) ggpubr: ‘ggplot2’ Based Publication Ready Plots. R package version 0.4.0. https://CRAN.R-project.org/package=ggpubr
Kassioumis K, Papageorgiou K, Glezakos TJ, Vogiatzakis IN (2004) Distribution and stand structure of Taxus baccata populations in Greece; Results of the first national inventory. Ecol Mediterr 30:159–170. https://doi.org/10.3406/ecmed.2004.1456
Komárková M, Novotný P, Cvrčková H, Máchová P (2022) The Genetic Differences and Structure of Selected Important Populations of the Endangered Taxus baccata in the Czech Republic. Forests 13:137. https://doi.org/10.3390/f13020137
König S, Heurich M, Olszewski T (2023) Bericht zur natürlichen Walddynamik im Nationalpark Bayerischer Wald 2012 bis 2021. Nationalparkverwaltung Bayerischer Wald, Grafenau
König S, Thonfeld F, Förster M, Dubovyk O, Heurich M (2023b) Assessing combinations of Landsat, Sentinel-2 and Sentinel-1 time series for detecting bark beetle infestations. GIScience Remote Sens 60:2226515. https://doi.org/10.1080/15481603.2023.2226515
Kopelman NM, Mayzel J, Jakobsson M, Rosenberg NA, Mayrose I (2015) Clumpak: a program for identifying clustering modes and packaging population structure inferences across K. Mol Ecol Resour 15:1179–1191. https://doi.org/10.1111/1755-0998.12387
Krenova Z, Kiener H (2012) Europe’s Wild Heart-still beating? Experiences from a new transboundary wilderness area in the middle of the Old Continent. Eur J Environ Sci 2:115–124. https://doi.org/10.14712/23361964.2015.32
Kubisiak T, Nelson C, Staton M, Zhebentyayeva T, Smith C, Olukolu B, Fang G-C, Hebard F, Anagnostakis S, Wheeler N (2013) A transcriptome-based genetic map of Chinese chestnut (Castanea mollissima) and identification of regions of segmental homology with peach (Prunus persica). Tree Genet Genom 9:557–571. https://doi.org/10.1007/s11295-012-0579-3
Ledig FT (1992) Human impacts on genetic diversity in forest ecosystems. Oikos 63:87–108. https://doi.org/10.2307/3545518
Ledig FT, Guries RP, Bonefeld BA (1983) The relation of growth to heterozygosity in pitch pine. Evol 37:1227–1238. https://doi.org/10.2307/2408843
Leinemann L, Hattemer H (2006) Genetic variation and mating pattern in a stand of yew (Taxus baccata L). Allgemeine Forst und Jagdzeitung 177:217–224
Liesebach H, Eusemann P, Höltken AM, Tröber U, Kuchma O, Karopka M, Becker F, Kätzel R, Fussi B (2024) Effective population size of adult and offspring cohorts as a genetic monitoring tool in two stand-forming and wind-pollinated tree species: Fagus sylvatica L. and Picea abies (L.) Karst. Conserv Genet 1–15. https://doi.org/10.1007/s10592-024-01600-2
Linares JC (2013) Shifting limiting factors for population dynamics and conservation status of the endangered English yew (Taxus baccata L., Taxaceae). For Ecol Manage 291:119–127. https://doi.org/10.1016/j.foreco.2012.11.009
Liu L, Wang Z, Huang L, Wang T, Su Y (2019) Chloroplast population genetics reveals low levels of genetic variation and conformation to the central–marginal hypothesis in Taxus wallichiana var. mairei, an endangered conifer endemic to China. Ecol Evol 9:11944–11956. https://doi.org/10.1002/ece3.5703
Loiselle BA, Sork VL, Nason J, Graham C (1995) Spatial genetic structure of a tropical understory shrub, Psychotria officinalis (Rubiaceae). Am J Bot 82:1420–1425. https://doi.org/10.2307/2445869
Luikart G, Sherwin W, Steele B, Allendorf F (1998) Usefulness of molecular markers for detecting population bottlenecks via monitoring genetic change. Mol Ecol 7:963–974. https://doi.org/10.1046/j.1365-294x.1998.00414.x
Luo Q, Li F, Yu L, Wang L, Xu G, Zhou Z (2021) Genetic diversity of natural populations of Taxus mairei. Conserv Genet 23:63–74. https://doi.org/10.1007/s10592-021-01403-9
Mariette S, Tavaud M, Arunyawat U, Capdeville G, Millan M, Salin F (2010) Population structure and genetic bottleneck in sweet cherry estimated with SSRs and the gametophytic self-incompatibility locus. BMC Genet 11:1–13. https://doi.org/10.1186/1471-2156-11-77
Maroso F, Vera M, Ferreiro J, Mayol M, Riba M, Ramil-Rego P, Martínez P, Bouza C (2021) Genetic diversity and structure of Taxus baccata from the Cantabrian-Atlantic area in northern Spain: A guide for conservation and management actions. For Ecol Manage 482:118844. https://doi.org/10.1016/j.foreco.2020.118844
Mayol M, Riba M, González-Martínez SC, Bagnoli F, de Beaulieu JL, Berganzo E, Burgarella C, Dubreuil M, Krajmerová D, Paule L (2015) Adapting through glacial cycles: Insights from a long‐lived tree (Taxus baccata). New Phytol 208:973–986. https://doi.org/10.1111/nph.13496
Mayol M, Riba M, Cavers S, Grivet D, Vincenot L, Cattonaro F, Vendramin GG, González-Martínez SC (2019) A multiscale approach to detect selection in nonmodel tree species: Widespread adaptation despite population decline in Taxus baccata L. Evol Appl 13:143–160. https://doi.org/10.1111/eva.12838
Miao Y, Zhang Z, Su J (2016) Low genetic diversity in the endangered following a population bottleneck, a low effective population size and increased inbreeding. Silvae Genet 65:59–66. https://doi.org/10.1515/sg-2016-0008
Müller-Starck G, Baradat P, Bergmann F (1992) Genetic variation within European tree species. New For 6:23–47. https://doi.org/10.1007/BF00120638
Nazareno AG, Alzate-Marin AL, Pereira RAS (2013) Dioecy, more than monoecy, affects plant spatial genetic structure: the case study of Ficus. Ecol Evol 3:3495–3508. https://doi.org/10.1002/ece3.739
Nei M (1978) Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics 89:583–590. https://doi.org/10.1093/genetics/89.3.583
Pascarella JB (1997) The mating system of the tropical understory shrub Ardisia escallonioides (Myrsinaceae). Am J Bot 84:456–460. https://doi.org/10.2307/2446021
Peakall R, Smouse PE (2006) GENALEX 6: genetic analysis in Excel. Population genetic software for teaching and research. Mol Ecol Notes 6:288–295. https://doi.org/10.1111/j.1471-8286.2005.01155.x
Perry DJ, Knowles P (1990) Evidence of high self-fertilization in natural populations of eastern white cedar (Thuja occidentalis). Canad J Bot 68:663–668. https://doi.org/10.1139/b90-086
Piry S, Luikart G, Cornuet JM (1999) Computer note. BOTTLENECK: a computer program for detecting recent reductions in the effective size using allele frequency data. J Hered 90:502–503. https://doi.org/10.1093/jhered/90.4.502
Porth I, El-Kassaby YA (2014) Assessment of the genetic diversity in forest tree populations using molecular markers. Diversity 6:283–295. https://doi.org/10.3390/d6020283
Poudel RC, Möller M, Li D-Z, Shah A, Gao L-M (2014) Genetic diversity, demographical history and conservation aspects of the endangered yew tree Taxus contorta (syn. Taxus fuana) in Pakistan. Tree Genet Genom 10:653–665. https://doi.org/10.1007/s11295-014-0711-7
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959. https://doi.org/10.1093/genetics/155.2.945
R Core Team (2022) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna. https://www.R-project.org/
Ratnam W, Rajora OP, Finkeldey R, Aravanopoulos F, Bouvet J-M, Vaillancourt RE, Kanashiro M, Fady B, Tomita M, Vinson C (2014) Genetic effects of forest management practices: global synthesis and perspectives. For Ecol Manage 333:52–65. https://doi.org/10.1016/j.foreco.2014.06.008
Rau B, Kavaliauskas D, Fussi B, Šeho M (2021) Erhaltung der Eibe in Bayern. AFZ-Der Wald. https://www.waldwissen.net/de/waldwirtschaft/waldbau/waldgenetik/erhaltung-der-eibe-in-bayern#:~:text=In%20Deutschland%20und%20Bayern%20wird,der%20oberen%20Naturschutzbeh%C3%B6rden%20eingeholt%20werden. Accessed 25 June 2023
Romšáková I, Paule L (2009) Genetische Variabilität von Populationen von Taxus baccata L. Sauteria 18:261–276
Ruprecht H, Dhar A, Aigner B, Oitzinger G, Klumpp R, Vacik H (2010) Structural diversity of English yew (Taxus baccata L.) populations. Eur J For Res 129:189–198. https://doi.org/10.1007/s10342-009-0312-4
Santos-del-Blanco L, Olsson S, Budde KB, Grivet D, González-Martínez SC, Alía R, Robledo-Arnuncio JJ (2022) On the feasibility of estimating contemporary effective population size (Ne) for genetic conservation and monitoring of forest trees. Biol Conserv 273:109704. https://doi.org/10.1016/j.biocon.2022.109704
Schirone B, Ferreira RC, Vessella F, Schirone A, Piredda R, Simeone MC (2010) Taxus baccata in the Azores: a relict form at risk of imminent extinction. Biodivers Conserv 19:1547–1565. https://doi.org/10.1007/s10531-010-9786-0
Schuelke M (2000) An economic method for the fluorescent labeling of PCR fragments. Nat Biotechnol 18:233–234. https://doi.org/10.1038/72708
Semper-Pascual A, Burton C, Baumann M, Decarre J, Gavier-Pizarro G, Gómez-Valencia B, Macchi L, Mastrangelo ME, Pötzschner F, Zelaya PV (2021) How do habitat amount and habitat fragmentation drive time-delayed responses of biodiversity to land-use change? Proceedings of the Royal Society B 288:20202466. https://doi.org/10.1098/rspb.2020.2466
Stefanowska S, Meyza K, Iszkuło G, Chybicki IJ (2021) Trunk perimeter correlates with genetic bottleneck intensity and the level of genetic diversity in populations of Taxus baccata L. Ann For Sci 78:1–14. https://doi.org/10.1007/s13595-021-01080-1
Suffness M (1995) Taxol: science and applications. CRC, Boca Raton
Svenning J-C, Magård E (1999) Population ecology and conservation status of the last natural population of English yew Taxus baccata in Denmark. Biol Conserv 88:173–182. https://doi.org/10.1016/S0006-3207(98)00106-2
Thomas P, Garcia-Marti X (2015) Response of European yews to climate change: a review. For Syst 24:eR01–eR01. https://doi.org/10.5424/fs/2015243-07465
Thomas P, Polwart A (2003) Taxus baccata L. J Ecol 91:489–524. https://doi.org/10.1046/j.1365-2745.2003.00783.x
Ueno S, Wen Y, Tsumura Y (2015) Development of EST-SSR markers for Taxus cuspidata from publicly available transcriptome sequences. Biochem Syst Ecol 63:20–26. https://doi.org/10.1016/j.bse.2015.09.016
Van der Knaap WO, van Leeuwen JF, Fahse L, Szidat S, Studer T, Baumann J, Heurich M, Tinner W (2020) Vegetation and disturbance history of the Bavarian Forest National Park, Germany. Veg Hist Archaeobot 29:277–295. https://doi.org/10.1007/s00334-019-00742-5
Van Oosterhout C, Hutchinson WF, Wills DP, Shipley P (2004) MICRO-CHECKER: software for identifying and correcting genotyping errors in microsatellite data. Mol Ecol Notes 4:535–538. https://doi.org/10.1111/j.1471-8286.2004.00684.x
Vancura K, Fady B, Koskela J, Mátyás C (2004) Conifers Network: Report of the second (20–22 September 2001, Valsaín, Spain), and third (17–19 October 2002, Kostrzyca, Poland) meetings. International Plant Genetic Resources Institute, Rome
Vekemans X, Hardy OJ (2004) New insights from fine-scale spatial genetic structure analyses in plant populations. Mol Ecol 13:921–935. https://doi.org/10.1046/j.1365-294x.2004.02076.x
Vendramin GG, Lelli L, Rossi P, Morgante M (1996) A set of primers for the amplification of 20 chloroplast microsatellites in Pinaceae. Mol Ecol 5:595–598. https://doi.org/10.1111/j.1365-294x.1996.tb00353.x
Wang J (2009) A new method for estimating effective population sizes from a single sample of multilocus genotypes. Mol Ecol 18:2148–2164. https://doi.org/10.1111/j.1365-294X.2009.04175.x
Wang X-R, Szmidt AE (2001) Molecular markers in population genetics of forest trees. Scand J For Res 16:199–220. https://doi.org/10.1080/02827580118146
Waples RS (2006) A bias correction for estimates of effective population size based on linkage disequilibrium at unlinked gene loci. Conserv Genet 7:167–184. https://doi.org/10.1007/s10592-005-9100-y
Waples RS, Do C (2010) Linkage disequilibrium estimates of contemporary Ne using highly variable genetic markers: a largely untapped resource for applied conservation and evolution. Evol Appl 3:244–262. https://doi.org/10.1111/j.1752-4571.2009.00104.x
Wickham H (2016) ggplot2: Elegant Graphics for Data Analysis. https://ggplot2.tidyverse.org
Yu G (2021) Scatterpie: Scatter Pie Plot. https://cran.r-project.org/web/packages/scatterpie/index.html
Zarek M (2016) Preliminary studies on the molecular identification of sex in Taxus baccata L. For Res Pap 77:68–75. https://doi.org/10.1515/frp-2016-0008
Zatloukal V, Vančura K (2004) Common yew (Taxus baccata L.) and its genetic diversity in the Czech Republic. In: Vančura K, Fady B, Koskela J, Mátyás C (eds) Conifers Network, Report of the second (20–22 September 2001, Valsaín, Spain) and third (17–19 October 2002, Kostrzyca, Poland) meetings. International Plant Genetic Resources Institute, Rome, pp 25–30
Acknowledgements
We would like to thank the Bavarian Forest National Park for their financial support, Christine Radler, and Alexandra Dolynska for their technical assistance in the laboratory, Olaf Schuber for his crucial help in collecting yew samples and Tabea Olszewski for her contribution to creating the maps.
Funding
Open Access funding enabled and organized by Projekt DEAL. This research was funded by the Bavarian Forest National Park (Nationalpark Bayerischer Wald).
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
Conceptualization, S.J.M., K.B.B, M.M, M.H. and O.G.; sampling, M.H; methodology, S.J.M., K.B.B, M.M and O.G.; validation, S.J.M., K.B.B, M.M, M.H. and O.G.; formal analysis, S.J.M, investigation, S.J.M., K.B.B, M.M and O.G.; resources, M.H. and O.G.; data curation, S.J.M., K.B.B, M.M, M.H. and O.G.; writing-original draft preparation, S.J.M; writing-review and editing, S.J.M., K.B.B, M.M, M.H. and O.G.; visualization, S.J.M; supervision, K.B.B, M.M and O.G.; project administration, M.H. and O.G.
Corresponding authors
Ethics declarations
Ethical approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
All authors listed in this manuscript have reviewed and approved the final version for submission. They also consent to the publication of this manuscript in the European Journal of Forest Research if accepted for publication. The authors agree to take full responsibility for the content of the manuscript and its accuracy.
Conflicts of interest
The authors declare no conflicts of interest or competing interests related to this research.
Additional information
Communicated by Marta Pardos.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Moosavi, S.J., Budde, K.B., Heurich, M. et al. Genetic variation of English yew (Taxus baccata L.) in the Bavarian Forest National Park, Germany. Eur J Forest Res 143, 1249–1265 (2024). https://doi.org/10.1007/s10342-024-01687-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10342-024-01687-9