Abstract
The cold periods of the Pleistocene have had a striking impact on the diversification of most organisms in temperate regions. Phylogeographic patterns and postglacial expansion are poorly understood in the Black Grouse (Tetrao tetrix). This species is widely distributed across Eurasia, and has been divided into a number of subspecies on the basis of morphological differences and geographic isolation. To investigate the evolutionary history of the species, 143 samples from different regions were examined for a mtDNA control region fragment. Overall, analyses of mtDNA gave support for the divergence between Tetrao tetrix tetrix, T. t. ussuriensis and T. t. mongolicus. The analyses reveal the effects of colonisation out of glacial refugia on the genetic diversity and genetic structure of Black Grouse. The phylogeographical results are consistent with a demographic population expansion following a bell-shaped mismatch distribution, a star-shaped phylogeny and low nucleotide diversity. Patterns of postglacial dispersal imply that Black Grouse from southern Europe have been restricted to this area, and did not contribute to the genetic diversity of northern Europe. Instead, Black Grouse spread out to northern Europe from a refugium in the east and a possible one in western Europe, following the retreat of glacial ice sheets, although both refugia remain unidentified. We suggest that T. t. britannicus and T. t. viridanus correspond to northern T. t. tetrix, and that this lineage has diverged from the other subspecies. This division is tentative due to limited sampling, but it will facilitate the management of different evolutionary significant units of the species.
Zusammenfassung
Phylogeographie und Status der Birkhühner-Unterarten
Die kalten Perioden im Pleistozän hatten einen starken Einfluss auf die Ausbreitung der meisten Organismen in den gemäßigten Breiten. Für die Birkhühner sind die phylogeographischen Muster und die Ausbreitung nach der Eiszeit noch überwiegend unbekannt. Die Art ist in Eurasien weit verbreitet und wurde aufgrund morphologischer Unterschiede und geographischer Isolationen in mehrere Unterarten eingeteilt. Zur Untersuchung der Entwicklungsgeschichte der Spezies wurden für 143 Proben aus unterschiedlichen Gegenden die Fragmente der mtDNA Kontrollregionen untersucht. Generell unterstützten die mtDNA-Analysen die Unterscheidung zwischen Tetrao tetrix tetrix, T. t. ussuriensis und T. t. mongolicus. Die Analysen verdeutlichten die Auswirkungen der Ausbreitung aus den eiszeitlichen Rückzugsgebieten heraus auf die genetische Vielfalt und die genetische Struktur der Birkhühner. Die phylogeographischen Ergebnisse passen zu einer demographischen Populations-Ausbreitung entsprechend einer glockenförmig ungleich verteilten Verbreitung, einer sternförmigen Stammesentwicklung und einer geringen Nukleotid-Vielfalt. Die Muster der postglazialen Ausbreitung legen nahe, dass die südeuropäischen Birkhühner auf diese Gebiete beschränkt blieben und nicht zur genetischen Vielfalt der nordeuropäischen beitrugen. Stattdessen breiteten sich die Birkhühner aus einem Zufluchtsgebiet in Ost- und möglicherweise auch einem in Westeuropa nach Nordeuropa aus und folgten so dem Rückzug der Eismassen, wobei allerdings die genaue Lage beider Zufluchtsgebiete noch unklar ist. Wir schlagen vor, daß T. t. britannicus und T. t. viridanus der nördlichen T. t. tetrix entsprechen und sich diese Linie von der anderen Unterart abgespalten hat. Diese Einteilung muss wegen der begrenzten Stichprobengröße noch als vorläufig gelten, wird aber die Arbeit mit den unterschiedlichen entwicklungsbiologisch signifikanten Einheiten dieser Spezies erleichtern.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The cold periods of the Pleistocene have had a striking impact on most organisms in temperate regions (Hewitt 2000). The Arctic ice sheet started to grow about 2.5 Mya, with more harsh climate fluctuations during the last 700,000 years, producing major changes in species distributions (Hewitt 1996). The places where species persist during glaciations that denote the species’ maximum contraction in geographical range have been described as refugia (Stewart et al. 2010). With the amelioration of climate, species expanded northwards from one or several refugia and colonised uninhabited areas (Hewitt 1996). The major glacial refugia for many species have been identified in the southern peninsulas of Iberia, Italy and the Balkans, where climatic conditions were less extreme during the Last Glacial Maximum (LGM, 23–18 kya) (Hewitt 1996, 1999, 2000; Taberlet et al. 1998). However, it has been suggested that glacial refugia in Asia and northeastern Europe may have contributed to the colonisation of northern Europe (Bilton et al. 1998; Naydenov et al. 2007). Examples of more northern glacial refugia near Moscow have been found for Norway spruce (Picea abies) (Lagercrantz and Ryman, 1990) and in Scandinavia for the Norwegian lemming (Lemmus lemmus) (Fedorov et al. 1996). In Asia, refugia have been suggested in the Middle East and the Caspian Sea (Durand et al. 1999; Liukkonen-Anttila et al. 2002; Hansson et al. 2008). However, few data are available on the contribution of potential easternmost refugia (Taberlet et al. 1998).
There have been several studies assessing patterns of genetic variation to infer population expansions from refugia (Taberlet et al. 1998), allowing inferences about the processes that led to current levels of genetic diversity and population structure (Johnson et al. 2007). This information is important for conservation management purposes, and for revising the traditional species and subspecies designations (Avise and Ball 1990; Moritz 1994).
In the present study, we focus on Black Grouse (Tetrao tetrix) which is a forest bird species distributed across Europe, from Britain to Siberia. It occurs as far north as Norway (at 70°N) and as far south as Kyrgyzstan and Manchuria (at 40°N). It inhabits a continuous range in large parts of the boreal forest, but is patchily distributed in Central Europe (Höglund et al. 2007b). The phylogeographic history, refugial isolation and post glacial expansion have not been studied so far in this species. Following its wide continental distribution, seven subspecies of Black Grouse have been described and defined by differences in body size and plumage coloration (del Hoyo et al. 1994). In Europe and north Siberia, the dominant subspecies is T. t. tetrix, with an isolated subspecies described in Great Britain, T. t. britannicus. The other five subspecies are located in Asia: T. t. viridanus in northwest Kazakhstan and southeast Russia (from the Don river to the Irtysh river), T. t. mongolicus in southeast Kazakhstan and west Mongolia, T. t. tschusii in south-central Russia (from the Irtysh river to Irkutsk and the Baikal lake), T. t. baikalensis in the eastern part from the Baikal lake and northern Manchuria and T. t. ussuriensis in southern Manchuria and North Korea (Peters 1934). Since morphological distinctions have been based on few specimens and geographically restricted samples, the exact boundaries of the Asian subspecies are not clear and, therefore, the adequacy of these subspecies designations is tentative. The reliability of these subspecies designations and their relevance for conservation purposes are thus not certain.
Here, we report a phylogeographical study of Black Grouse, including seven described subspecies: T. t. tetrix, T. t. britannicus, T. t. viridanus, T. t. mongolicus, T. t. tschusii, T. t. baikalensis and T. t. ussuriensis. Due to limited sampling, mongolicus was combined with tschusii, and baikalensis with ussuriensis for all the genetic analyses. Although the definition of a subspecies is problematic, we hold to the subspecies concept defined by Avise and Ball (1990) and O’Brien and Mayr (1991), to include populations below the species level that share a distinct geographic distribution and a unique natural history relative to other subdivisions of the species. In addition, subspecies at the molecular level are revealed when all sequences from a subspecies share a common ancestral sequence not found in individuals from any other subspecies (Avise 2000). This reciprocal monophyly would imply that a subspecies has been evolving independently for more than 2N ef generations, where N ef is the effective size of the female population, giving support to the subspecies concept (Avise 2000; Zink 2004).
We estimated the genetic relationships and population structure among Black Grouse subspecies using a fragment of the mtDNA control region. Mitochondrial DNA analyses will reveal whether the subspecies are evolving independently or are in an intermediate stage of isolation due to an ongoing exchange of breeding individuals (Zink 2004). In addition, we address the phylogeography, and in particular we ask whether southern and/or eastern refugia acted as sources of colonisation after deglaciation. The results of this study will provide a better understanding of the history and the systematic classification of Black Grouse, which will be useful for conservation and management purposes in the near future.
Methods
Tissue collection and DNA extractions
Samples were obtained from the collections of the Natural History Museum in Stockholm, Helsinki, Århus, Hamburg, Kharkiv and Almaty. All samples were footpad scrapes (5 mm2) of dried Black Grouse specimens that have been stored for periods ranging from few years to decades (Appendix 1, supplementary material). In total, 43 museum samples were obtained from Finland, Denmark, Germany, Estonia, Russia, Kazakhstan and Korea. In addition, 97 samples (blood, muscle and feathers) from Sweden, Norway, Netherlands, France, Italy, Switzerland, Ukraine, Wales, Scotland and England were collected in the field (Fig. 1). We also included in our analyses three Black Grouse sequences from Russia and Mongolia (Drovetski 2002; GenBank accession numbers AF532457, AF532458, AF532459). These samples represent the subspecies T. t. tetrix, T. t. britannicus, T. t viridanus, T. tschusii, T. t. mongolicus, T. t. baikalensis and T. t. ussuriensis (Appendix 1, supplementary material).
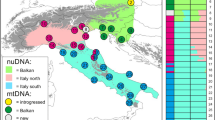
Sampling area and Black Grouse (Tetrao tetrix) clades distribution throughout Eurasia (after Storch 2007). Each subspecies is represented by a number: 1 T. t. tetrix, 2 T. t. britannicus, 3 T. t. viridanus, 4 T. t. tschusii, 5 T. t. mongolicus, 6 T. t. baikalensis, 7 T. t. ussuriensis. Limits between subspecies are not shown due to ambiguity. Dots indicate clades belonging to different lineages: eastern (yellow), western (green), southern (red), mongolicus (turquoise), ussuriensis (blue) (color figure online)
Skin, muscle and feather samples were cut into small pieces using a sterile scalpel followed by a proteinase K-DTT 1 M digestion with an extended incubation interval at 56 °C for up to 48 h until little solid tissue remained. DNA was extracted using the Qiagen DNeasy® blood and tissue kit following the different manufacturer protocols for tissue, feathers and blood (Qiagen, Hilden, Germany).
Sequencing
We obtained a fragment of 304 nucleotides of the control region (CR) with the primers 186L/521H (Quinn and Wilson 1993). Sequences started at position 170 and ended at position 471 when comparing with the complete CR sequence for Black Grouse (GenBank accession no. AJ297153). Amplification was performed in 25-μl reactions following Johnson et al. (2003) and 35 cycles. PCR products were run on 2 % agarose gel containing ethidium bromide. The ethanol/salt and isopropanol precipitation protocol was used for pre-sequencing cleanup. Samples were sequenced in a MegaBace (Amersham Biosciences) using the DYEnamic ET Dye Terminator Kit (Amersham Biosciences) following the manufacturer’s indications. All amplified fragments were sequenced twice with both light- and heavy-stranded primers and compared with the complete Black Grouse CR sequence from GenBank for accuracy (No. AJ297153). We sequenced the same CR fragment for the closely related species Caucasian Grouse (T. mlokosiewiczi), and Capercaillie (T. urogallus) to be used as outgroups. We also used one sequence of Causasian Grouse (AF532460) to compare with our results. Unique sequences have been deposited at GenBank under the accession numbers HQ889288, JF440304, JF440311–JF440313, JF440315–JF440333, JQ966110–JQ966120, JX312695–JX312698 and KC514352–KC514353.
To avoid contamination, the equipment was exposed to ultraviolet radiation and DNA extractions, pre-PCR and post-PCR, were done in different rooms, and aerosol-resistant filter pipette tips were used throughout.
Statistical analyses
Each complete sequence was generated by aligning and editing in MEGA 3 (Kumar et al. 2004). Nucleotide diversity (π) and haplotype diversity (h) were estimated with their respective standard deviations for each population using ARLEQUIN 3.0 (Excoffier et al. 2005) and DnaSP version 4.0 (Rozas et al. 2003). π is the average pairwise nucleotide difference between individuals within samples and h is the probability that two randomly chosen individuals have different haplotypes (Nei 1987). θ ST values were calculated using a distance matrix between haplotypes as implemented in ARLEQUIN 3.0 (Excoffier et al. 2005). Differences between populations were tested using 10,000 permutations with Fisher’s exact test. We tested for significant variance in the distribution of control region sequences between groups of populations, populations and individuals using AMOVA, carried out in ARLEQUIN 3.0 (Excoffier et al. 2005). The T. t. tschussi sample was combined with T. t. mongolicus and, T. t. baikalensis with T. t. ussuriensis due to geographical closeness, in order to obtain better resolution. We created a minimum spanning network with all unique haplotypes. For this analysis, we used the Median-Joining (MJ) network algorithm implemented in the program NETWORK v.4.1.1.0 (Bandelt et al. 1999).
To examine general evolutionary relationships among subspecies, we estimated genetic distances between individuals. Neighbor-joining (NJ) trees were constructed in MEGA 3 (Kumar et al. 2004) with 10,000 bootstrap replicates to assess the statistical significance of internal nodes. We performed Bayesian phylogenetic inference with MrBAYES v.3.2 (Ronquist and Huelsenbeck 2003) to confirm levels of NJ bootstrap support. The TPM1uf+G model (=K81 with unequal base frequencies and gamma distribution rates) (Kimura 1981) was the best-fit model of nucleotide substitution for the CR sequences tested in MODELTEST v.3.7 (Posada and Crandall 1998) with selection based on the Akaike Information Criterion (AIC; Akaike 1974). Analysis in MrBayes included two separate runs, each containing 1 cold chain and 3 heated chains using a temperature of 0.15. Markov chains were sampled every 500 generations and run for 2 million generations. A tree was drawn using FigTree v.1.2.2 (available at http://tree.bio.ed.ac.uk/software/figtree/).
To examine whether the species showed any sign of historical population expansion, we estimated Tajima’s D (Tajima 1989), Fu’s Fs (1997) and a mismatch distribution analysis (Rogers and Harpending 1992) for every subspecies using ARLEQUIN 3.0 (Excoffier et al. 2005). Negative values of Tajima’s D can be interpreted as evidence of population expansions (Fu 1997), and negative values of Fs indicate an excess of recent mutations and reject population stasis. A diagram of frequencies of pairwise genetic differences was drawn using DnaSP v.4.0 (Rozas et al. 2003). A thousand bootstrap replicates were used to generate an expected distribution using a model of sudden demographic expansion (Excoffier et al. 2005). The sum of squared deviation (SSD) and the raggedness index were also calculated. These measures quantify the smoothness of the observed mismatch distribution. Small raggedness values are typical of an expanding population whereas higher values are observed among stationary or bottlenecked populations (Harpending et al. 1993).
To estimate the time of expansion, we used the formula t = τ/2u (Rogers and Harpending 1992), where t is the number of generations, τ is the mode of the mismatch distribution, expressed in units of evolutionary time, and u = μk where μ is the mutation rate per nucleotide and k is the length of the analysed sequence. The mutation rate of the avian control region is uncertain (Hansson et al. 2008), but we used a mutation rate between 9.12 × 10−5 and 3.65 × 10−4 (3 years generation time and 304 bp) for the domain I (Randi et al. 1999), which is a hypervariable region in the CR (Quinn 1992; Holder et al. 1999; Randi et al. 1999; Baba et al. 2002; Ruokonen and Kvist 2002).
Results
Unambiguous DNA sequences could be generated for all 143 samples. The haplotypes had only 33 polymorphic sites (of which 22 were parsimony informative) out of 304 bp. There were 30 transitions, 4 transversions and 4 indels at the variable sites. We found 38 haplotypes and from these 26 belonged to T. t. tetrix, 5 to T. t. britannicus, 4 to T. t. viridanus, 6 to T. t. mongolicus, 1 to T. t. tschusii, 1 to T. t. baikalensis and 1 to T. t. ussuriensis. One haplotype (H1) occurred in all study areas except for southern Europe, Kazakhstan, Mongolia and North Korea. There were 22 unique haplotypes for T. t. tetrix (H3–H15, H22–28, H34–H35), 3 for T. t. viridanus (H30, H32, H37), 1 for T. t. tschusii (H31), 5 for T. t. mongolicus (H19–H21, H31, H36), 2 for T. t. britannicus (H17, H18), 1 for T. t baikalensis (H38) and 1 for T. t. ussuriensis (H29) (Appendix 2, supplementary material). Total haplotype diversity was 0.827 ± 0.029 and nucleotide diversity was 0.0877 ± 0.050. The highest diversity values were found in T. t. viridanus, T. t. mongolicus and southern T. t. tetrix. Sweden and Netherlands had the highest values within T. t. tetrix (Table 1).
The overall fixation index was 0.655 (P ≤ 0.001 AMOVA). Pairwise θ ST values (Table 2) ranged from 0 to 0.771. We detected that all samples from southern Europe were highly differentiated from the northern European ones (θ ST = 0.713, P < 0.05). Therefore, we divided T. t. tetrix in northern and southern samples for the subsequent analyses. Southern T. t. tetrix, T. t. viridanus, T. t. mongolicus and T. t. ussuriensis were greatly differentiated from northern T. t. tetrix, and T. t. britannicus (Table 2). When the θ ST pairwise comparison were performed between populations of northern T. t. tetrix, we found that Sweden–Netherlands (θ ST = 0.142, P = 0.018), Sweden–Estonia (θ ST = 0.129, P = 0.020), Russia–Germany (θ ST = 0.027, P = 0.017) and Estonia–Germany (θ ST = 0.457, P = 0.033) had significant values (data not shown). Analysis of molecular variance of the CR was consistent with high levels of genetic structure. We found highly significant structure among groups (45.64 % of variance), among populations within groups (19.89 % of variance), and within populations (34.47 % of variance).
We did not find many differences between NJ and Bayesian phylogenetic topologies of Black Grouse. Although the NJ tree was capable to discern between subspecies, the bootstrap values were low (<75 %; not shown) and only gave support to the division between southern and northern T. t. tetrix (bootstrap value 75 %; not shown). Consequently, only the Bayesian tree is displayed (Fig. 2). The most remarkable aspect of the topology was the well-supported separation of southern and northern T. t. tetrix. Haplotypes H26 and H28 were only found in southern T. t. tetrix, but they were placed together within the northern T. t. tetrix–T. t. britannicus clade. In addition, five samples from T. t. viridanus (H1, H30, H32) and three samples from T. t. mongolicus (H21, H36, H33) were found within the northern T. t. tetrix clade. Three more clades could also be differentiated: one clade with samples from T. t. ussuriensis and T. t. baikalensis, another clade with samples from T. t. tschusii and T. t. mongolicus, and a clade formed by three individuals from T. t. tetrix, indicating a possible origin unrelated to the northern and southern clades. The Minimum Spanning network (Fig. 3) gave complementary information and verified the existence of the clusters suggested in the Bayesian tree.
Bayesian tree showing phylogenetic relationships among Black Grouse individuals based on 304 bp of the Control Region. Values at the nodes represent percentage of posterior probabilities. Each clade is represented by one colour: northern T. t. tetrix or eastern clade (yellow), southern T. t. tetrix (red), western T. t. tetrix (green), T. t. mongolicus (turquoise), and T. t. ussuriensis (blue) (color figure online)
Network for 38 Black Grouse CR haplotypes. The sizes of the circles are proportional to the haplotype frequencies. Each subspecies is represented by one colour: northern T. t. tetrix (yellow), southern T. t. tetrix (red), T. t. britannicus (orange), T. t. viridanus (purple), T. t. mongolicus (turquoise), and T. t. ussuriensis (blue). Black circles (MV) are inferred intermediate haplotypes calculated by NETWORK (color figure online)
The star-like phylogeny of haplotypes (Fig. 3), the reduced levels of haplotypic and nucleotide diversity demonstrate that some subspecies have undergone an expansion from a population of low genetic diversity. This historical population expansion was also confirmed by Tajima’s D and Fs only for northern T. t. tetrix (D = −1.706, P < 0.01; Fs = −14.082, P < 0.001) and T. t. britannicus (D = −1.728, P < 0.02; Fs = −1.690, P < 0.05). The mismatch distribution fitted a bell-like curve (Fig. 4) typical of a population undergoing exponential growth rather than an L-shaped one indicative of equilibrium (Slatkin and Hudson 1991). The sum of squared deviations (SSD) of mismatch distribution and the raggedness index were non-significant, indicating that the curve fits the sudden expansion model (Table 3). In addition, the multimodal mismatch distribution indicates that this expansion took place from at least three different refugia (Fig. 4a–f). We found one common refugium, when we considered samples from northern T. t. tetrix, T. t. britannicus and T. t. viridanus (Fig. 4d). However, the mismatch distribution of Great Britain, Netherlands and Switzerland revealed the presence of two different lineages, confirming the western European clade (Fig. 4c). Northern T. t. tetrix together with T. t. britannicus started to expand approximately 21129—5282 YR BP, indicating that the expansion occurred after the LGM (23–18 kya).
Observed and expected distributions of pairwise nucleotide differences between 38 Black Grouse haplotypes. The stippled line indicates the observed distribution, consistent with a growing population model. The solid line indicates the expected distribution under a constant population model. a Mismatch distribution for all samples, b mismatch distribution between northern T. t. tetrix and southern T. t. tetrix, c mismatch distribution between northern T. t. tetrix and western T. t. tetrix, d mismatch distribution between northern T. t. tetrix and T. t. viridanus, e mismatch distribution between northern T. t. tetrix, T. t. mongolicus and T. t. ussuriensis, f mismatch distribution between T. t. viridanus, T. t. mongolicus and T. t. ussuriensis
Discussion
Species status
To our knowledge, this is the first attempt to describe the phylogeography of Black Grouse in Eurasia, as previous studies in Black Grouse subspecies were based on morphological characteristics. For example, Schönwetter (1960–1962) suggested different values for egg weight depending on the subspecies (see Schreiber et al. 1998 for more information). Niewold and Nijland (1987) suggested that Dutch Black Grouse should be classified as distinct subspecies because of their heavier and larger size, longer legs and sparser plumage than the German and Swedish grouse. This was refuted by Schreiber et al. (1998) in an analysis of 38 allozyme loci in 95 individuals from southern Germany, Netherlands and Sweden.
Subspecies were often described on the basis of a few skins kept in museums, in which colours can change with the age of the bird and the time since collection, and the skeleton was not necessarily available (Duriez et al. 2007). These taxonomic divisions need to be revised in order to assess, design and implement conservation plans for a species or subspecies (Mace 2004). These early subspecies descriptions suffer various fates: in Capercaillie, there was not a clear genetic differentiation between three local subspecies in Finland (Liukkonen-Anttila et al. 2004), nor among ten Eurasian subspecies, except for the Pyrenean birds that formed a genetically distinct unit (Segelbacher and Piertney 2007; Duriez et al. 2007). In Sage-grouse, Benedict et al. (2003) did not find a genetic distinction between the described eastern and western subspecies. In contrast, other studies have found evidence for different subspecies (Black-tailed Godwits, Höglund et al. 2007a; Tigers, Luo et al. 2004; Redshank, Ottvall et al. 2005); and sometimes subspecies have been raised to species status (Gunnison Sage-grouse, Young et al. 2000; Blue Grouse, Barrowclough et al. 2004; Clouded Leopard, Wilting et al. 2007; but see Isaac et al. 2004). In the case we study here, our results agree with similar studies that found evidence to support some subspecies delineation: mtDNA tree showed a divergence between T. t. Tetrix, T. t. mongolicus and T. t. ussuriensis and a similarity between T. t. Tetrix, T. t. viridanus and T. t. britannicus. In addition, we found that T. t. tetrix may be divided in three clades: (1) the southern clade corresponding to Italy, France and Switzerland; (2) the northern clade corresponding to Netherlands, Germany, Denmark, Estonia, Russia, Ukraine, Finland, Sweden and Norway; and (3) the western clade formed by individuals from Netherlands, Switzerland and UK. The dominant northern European group included two southern birds (Italy, Switzerland). This may suggest secondary contact between both groups after the ice age, and an admixture of different populations. Gene flow does occur among populations in the Alps but it is limited due to habitat fragmentation (Caizergues et al. 2003).
On the basis of the genetic results, we recommend the recognition of three subspecies: T. t. tetrix, T. t mongolicus, and T. t. ussuriensis considering previous criteria for the designation of subspecies (Avise and Ball 1990; O’Brien and Mayr 1991). Whether T. t. tschusii and T. t. baikalensis are proper subspecies still remains unclear. Additional sampling in East Asia will be critical to determine the taxonomic status of Black Grouse subspecies poorly included in this study.
Reciprocal monophyly is a requisite to assess subspecies at the molecular level (Zink 2004). However, we consider that monophyly does not compile the validity of a subspecies for several reasons: (1) subspecies are not reproductively isolated; (2) usually continental subspecies in the Palearctic have geographical ranges that overlap and therefore, divergence will be low; and (3) there might not have been enough time for divergence and reciprocal monophyly to evolve, since Black Grouse subspecies may be young, probably appearing during the last glacial maximum (Phillimore and Owens 2006).
On the other hand, our results confirm the assumption of Höglund et al. (2011) who found that British Black Grouse are significantly divergent in microsatellite data but not reciprocally monophyletic with respect to mtDNA caused by sharing of haplotypes with continental Europe. Therefore, the British population is a Management Unit (i.e. populations with significant divergence of allele frequencies at nuclear or mitochondrial loci, regardless of the distinctiveness of the alleles; Moritz 1994), but not a subspecies (Höglund et al. 2011). Regarding the division within T. t. tetrix, it would appear that southern T. t. tetrix has had a distinct evolutionary history that merits recognition. We suggest that this clade should be considered as an Evolutionary Significant Unit (i.e. populations that are reciprocally monophyletic for mtDNA alleles and show significant divergence of allele frequencies at nuclear loci; Moritz 1994). This clade has a distinct geographic distribution, and it is divergent at microsatellite loci (Caizergues et al. 2003; Höglund et al. 2007b) and mtDNA as seen in this study. However, the mtDNA relations with populations from the northern Alps still remain to be investigated.
Postglacial colonisation
Phylogeographical structure has been documented in several avian species from Europe and North America (Merilä et al. 1997; Milá et al. 2007; Pitra et al. 2000; Barrowclough et al. 2004). Our analyses distinguished five lineages in Black Grouse with high posterior probability values for the divergence between northern and southern T. t. tetrix, T. t. mongolicus, T. t. ussuriensis and a western European clade. Our results also confirm the four distinct clades (two Alpine, Norway and Kazakhstan) found by Randi et al. (1999) using CR.
On the other hand, genetic diversity values of samples from Netherlands, Switzerland and Britain were approximately the same as values of other populations. This high genetic diversity is due to the admixture of lineages from different refugia that gave rise to a hybrid zone (Provan and Bennett 2008). When haplotypes from one of the lineages were removed in the mentioned populations, the genetic diversity decreased (data not shown). On the other hand, new haplotypes could arise in situ after the colonisation process, such as in Sweden and Ukraine, due to selection shifting over time and these haplotypes do not represent those found during the LGM (Provan and Bennett 2008).
During the Pleistocene, the ice sheets advanced and receded periodically producing major changes in species distributions: some became extinct and others dispersed and survived in refugia (Hewitt 2000). At the last glacial maximum (LGM; 23–18 kya), a continuous ice sheet existed along northern Europe for over ca. 2,200 km, extending south to 52°N and permafrost south to 47°N (Hewitt 2004). Populations restricted to refugia during unfavourable climatic cycles would rapidly expand their range following the retreating glacial ice sheets, encountering a wide variety of unoccupied habitats when conditions improved (Hewitt 1999).
Analyses of mismatch distributions can be used to detect and date historical population expansions (Prairie Chicken; Johnson et al. 2007; Johnson 2008; Great Reed Warbler (Acrocephalus arundinaceus), Hansson et al. 2008). For Black Grouse, the bell-shaped mismatch distribution, the star-shaped phylogeny with a central common haplotype and low nucleotide diversity are consistent with a demographic population expansion. These are signs of an abundant species that has expanded its range rather recently from small or modest numbers of founders (Avise 2000).
Our estimated dates of population expansion (20–5 ky BP) correspond with the beginning of postglacial colonisation of available pristine habitat following the last ice retreat (14–10 ky BP). Some studies have revealed population expansions that also occurred after the last deglaciation. In wood lemmings (Myopus schisticolor), the colonisation started between 5–2.5 ky BP (Fedorov et al. 1996), in Great Reed Warblers, it started between 4 and 1.5 ky BP (Hansson et al. 2008), and in silver birch (Betula pendula) as early as 10 ky BP (see Palmé et al. 2003). These estimates should be interpreted cautiously because they are sensitive to mutation rates and generation times, and assume a constant molecular clock (Hansson et al. 2008).
Patterns of postglacial dispersal from refugia in the southern peninsulas of Iberia, Italy and the Balkans have been described for several species, such as bears, voles, oaks (see Taberlet et al. 1998) and Capercallie (Duriez et al. 2007; Segelbacher and Piertney 2007). Other taxa such as Black Grouse did not follow this pattern (Bilton et al. 1998). Our Alpine/southern clade did not expand possibly because the populations of this area had no or only limited exchange of individuals and thus evolved separately, resulting in a different genetic group (Randi et al. 1999; Schmitt 2007). Fossils of Black Grouse have been found in the Balkans (Boev 1992; Mlíkovský 1995) and in the Iberian Peninsula (Sanchez-Marco 2004), indicating that the species survived in these areas but, after climate warming, these populations became extinct.
To our knowledge, eastern European and Asian refugia that have been suggested are: (1) the Carpathians (Jaarola and Searle 2002; Deffontaine et al. 2005); (2) the Caspian region and Middle East (Durand et al. 1999, Liukkonen-Anttila et al. 2002; Hansson et al. 2008); (3) Beringia and eastern Siberia (Holder et al. 1999; Abbott et al. 2000; Holder et al. 2000; Skrede et al. 2006); (4) Moscow (Lagercrantz and Ryman 1990); (5) Manchuria and Mongolia (Baba et al. 2002); and (6) the southern Ural Mountains, where most of Taiga species might have started a postglacial expansion (Goropashnaya et al. 2004; Schmitt 2007).
Our genetic data reflect that five lineages of Black Grouse survived glaciations in separated refugia, but only two were capable to contribute to the genetic diversity of Europe. We propose the existence of two refugia based on our results, although accurate locations remains unidentified: (1) an area near the Ural Mountains with a postglacial expansion westwards to Sweden and central Europe, and (2) a possible refugium in western Europe (i.e. France) with a postglacial expansion northwards to the Netherlands and Britain.
The eastern Black Grouse expansion pattern is also similar to that seen for the Scots pine Pinus sylvestris (Naydenov et al. 2007), Norway spruce Picea abies (Lagercrantz and Ryman 1990), and mountain avens Dryas octopetala (Skrede et al. 2006), species from a boreal climate, which could not survive in the southernmost regions. Moreover, it is also known that wood lemmings colonised the north-eastern part of the Scandinavian Peninsula from Russia and Finland, and their time of expansion was estimated based on the spread of spruce forest into Scandinavia (Fedorov et al. 1996).
We would thus expect a general correlation of expansion response of animals and suitable vegetation due to the animals’ dependence on plants (Hewitt 1996). We can conclude that the expansion of Black Grouse from eastern refugia is likely to have followed the establishment of coniferous woodland in Europe. This correlation was also suggested between the expansion patterns of oak and hedgehog (Seddon et al. 2001).
To the east of Europe and across Siberia the barriers for many species are less clear during the interglacial, but these regions would have been hostile during glacial periods (Hewitt 1996). However, the existence of refugia at mid-northern latitudes could have been possible for cold-adapted species such as silver birch (Palmé et al. 2003). Evidence from macrofossils and palynological data indicates that cold-tolerant trees survived the LGM at relatively high latitudes and that populations were probably distributed along the edge of the glaciated areas rather than restricted to a few well-defined refugia (see Palmé et al. 2003). There are also indications that the climate during the LGM was warm enough for Scots pine to survive and grow well as far north as 50°N upon permafrost (Naydenov et al. 2007). Grouse are also cold-adapted species which depend on coniferous–birch forest; therefore, it might have been possible for them to survive in this region. This condition would connect scattered “cryptic” refugia fairly close to margin of the ice sheet at its full-glacial extension (Provan and Bennett 2008).
Implications for conservation
Globally, the Black Grouse is classified as not threatened (IUCN 1996), although information in Asia is scarce. The species was discovered in the north of South Korea in 2002 (Rhim et al. 2003), and it might be still extant in the far north-east of North Korea (Duckworth and Moores, personal communication). The distribution range in Europe is fragmented, as large contractions and declines have occurred during the twentieth century (Storch 2007). Current conservation measures include habitat restoration, reintroduction projects and predator control (Storch 2007).
Our results clearly reveal that the southern European Black Grouse qualifies to be considered as an ESU (Moritz 2002), since it is geographically and genetically isolated. Caizergues et al. (2003) suggested that restricted gene flow and a large genetic drift owing to small effective population sizes were the causes of lower genetic variability in the southwestern Alps populations relative to those found in the northern Alps. Overall, this would imply that individuals from northern Eurasian populations cannot be used for exchanging or reintroduction purposes into the southern European populations or vice versa. Consequently, the use of local birds should be only considered for conservation measures in southern Europe.
Conclusion
The preliminary phylogeography of the Black Grouse demonstrates that this species has been a part of a continuous Eurasian population with Pleistocene refugia in Central-East Asia and a possible refugium in western Europe. Such refugia have been revealed in other studies but specific locations are still lacking. The recolonisation of central and north Europe started somewhere in Asia and the expansion was possibly fast because the genetic differentiation within northern Europe was low.
We found support for three subspecies/clades along the large distribution of Black Grouse, plus a possible new ESU in southern Europe. More detailed work with larger sample size throughout the Asian range is necessary to clearly understand the colonisation process and subspecies distribution of Black Grouse. A limited sampling within refugial areas might suggest the absence of haplotypes that might be found elsewhere (Provan and Bennett 2008). Moreover, the possible correlation between the distributions of Asian Black Grouse subspecies with the distribution of refugia remains to be investigated (see Holder et al. 1999).
References
Abbott RJ, Smith LC, Milne RI, Crawford RMM, Wolff K, Balfour J (2000) Molecular analysis of plant migration and refugia in the Arctic. Science 289:1343–1346
Akaike H (1974) A new look at the statistical model identification. IEEE Trans Autom Control 19:716–723
Avise JC (2000) Phylogeography: the history and formation of species. Harvard University Press, Cambridge
Avise JC, Ball RM (1990) Principles of genealogical concordance in species concepts and biological taxonomy. Oxford Surv Evol Biol 7:45–67
Baba Y, Fujimaki Y, Klaus S, Butorina O, Drovetskii S, Koike H (2002) Molecular population phylogeny of the hazel grouse Bonasa bonasia in East Asia inferred from mitochondrial control-region sequences. Wildl Biol 8:283–291
Bandelt HJ, Forster P, Röhl A (1999) Median-joining networks for inferring intraspecific phylogenies. Mol Biol Evol 16:37–48
Barrowclough GF, Groth JG, Mertz LA, Gutiérrez RJ (2004) Phylogeographic structure, gene flow and species status in blue grouse (Dendragapus obscurus). Mol Ecol 13:1911–1922
Benedict NG, Oyler-McCance SJ, Taylor SE, Braun CE, Quinn TW (2003) Evaluation of the eastern (Centrocercus urophasianus urophasianus) and western (Centrocercus urophasianus phaios) subspecies of Sage-grouse using mitochondrial control-region sequence data. Conserv Genet 4:301–310
Bilton DT, Mirol PM, Mascheretti S, Fredga K, Zima J, Searle JB (1998) Mediterranean Europe as an area of endemism for small mammals rather than a source for northwards postglacial colonisation. Proc R Soc Lond B 265:1219–1226
Boev ZN (1992) Paleornithological studies in Bulgaria. In: Campbell KE (ed) Papers in avian paleontology Honoring Pierce Brodkorb incorporating the proceedings of the II international symposium of the society of avian paleotology and evolution, pp 28–30 Sept 1988. Science Series, No 36. Natural History museum of Los Angeles County. Los Angeles, pp 459–463
Caizergues A, Rätti O, Helle P, Rotelli L, Ellison L, Rasplus JY (2003) Population genetic structure of male black grouse (Tetrao tetrix) in fragmented vs. continuous landscapes. Mol Ecol 12:2297–2305
Deffontaine V et al (2005) Beyond the Mediterranean peninsulas: evidence of central European glacial refugia for a temperate forest mammal species, the bank vole (Clethrionomys glareolus). Mol Ecol 14:1727–1739
del Hoyo J, Elliott A, Sargatal J (1994) Handbook of the birds of the world, vol 2. Lynx, Barcelona
Drovetski SV (2002) Molecular phylogeny of grouse: individual and combined performance of W-linked, autosomal, and mitochondrial loci. Syst Biol 51:930–945
Durand JD, Persat H, Bouvet Y (1999) Phylogeography and postglacial dispersion of the chub (Leuciscus cephalus) in Europe. Mol Ecol 8:989–997
Duriez O, Sachet JM, Ménoni E, Pidancier N, Miquel C, Taberlet P (2007) Phylogeography of the capercaillie in Eurasia: what is the conservation status in the Pyrenees and Cantabrian Mounts? Conserv Genet 8:513–526
Excoffier L, Laval G, Schneider S (2005) Arlequin (version3.0): an integrated software package for population genetics data analysis. Evol Bioinform 1:47–50
Fedorov V, Jaarola M, Fredga K (1996) Low mitochondrial DNA variation and recent colonisation of Scandinavia by the wood lemming. Mol Ecol 5:577–581
Fu YX (1997) Statistical tests of neutrality of mutations against population growth, hitchhiking and background selection. Genetics 147:915–925
Goropashnaya AV, Fedorov VB, Seifert B, Pamilo P (2004) Limited phylogeographical structure across Eurasia in two red wood ant species Formica pratensis and F. lugubris (Hymenoptera, Formicidae). Mol Ecol 13:1849–1858
Hansson B, Hasselquist D, Tarka M, Zehtindjiev P, Bensch S (2008) Postglacial colonisation patterns and the role of isolation and expansion in driving diversification in a passerine bird. PLoS ONE 3:e2794
Harpending HC, Sherry ST, Rogers AR, Stoneking M (1993) The genetic structure of ancient human populations. Curr Anthropol 34:483–496
Hewitt GM (1996) Some genetic consequences of ice ages, and their role in divergence and speciation. Biol J Linn Soc 58:247–276
Hewitt GM (1999) Post-glacial re-colonisation of European biota. Biol J Linn Soc 68:87–112
Hewitt GM (2000) The genetic legacy of the Quaternary ice ages. Nature 405:907–913
Hewitt GM (2004) Genetic consequences of climatic oscillations in the Quaternary. Philos Trans R Soc Lond B 359:183–195
Höglund J, Johansson T, Beintema A, Schekkerman H (2007a) Phylogeography of the black-tailed godwit Limosa limosa: substructuring revealed by mtDNA control region sequences. J Ornithol 150:45–53
Höglund J, Larsson JK, Jansman HAH, Segelbacher G (2007b) Genetic variability in European black grouse (Tetrao tetrix). Conserv Genet 8:239–243
Höglund J, Larsson JK, Corrales C, Santafé G, Baines D, Segelbacher G (2011) Genetic structure among black grouse in Britain: implications for designing conservation units. Anim Conserv. doi:10.1111/j.1469-1795.2011.00436.x
Holder K, Montgomerie R, Friesen VL (1999) A test of the glacial refugium hypothesis using patterns of mitochondrial and nuclear DNA sequence variation in rock ptarmigan (Lagopus mutus). Evolution 53:1936–1950
Holder K, Montgomerie R, Friesen VL (2000) Glacial vicariance and historical biogeography of rock ptarmigan (Lagopus lupus) in the Bering region. Mol Ecol 9:1265–1278
Isaac NJB, Mallet J, Mace GM (2004) Taxonomic inflation: its influence on macrobiology and conservation. Trends Ecol Evol 19:464–469
Jaarola M, Searle JB (2002) Phylogeography of field voles (Microtus agrestis) in Eurasia inferred from mitochondrial DNA sequences. Mol Ecol 11:2613–2621
Johnson JA (2008) Recent range expansion and divergence among North American prairie grouse. J Hered 99:165–173
Johnson JA, Toepfer JE, Dunn PO (2003) Contrasting patterns of mitochondrial and microsatellite population structure in fragmented populations of greater prairie-chicken. Mol Ecol 12:3335–3347
Johnson JA, Dunn PO, Bouzat JL (2007) Effects of recent population bottlenecks on reconstructing the demographic history of prairie-chickens. Mol Ecol 16:2203–2222
Kimura M (1981) Estimation of evolutionary distances between homologous nucleotide sequences. Proc Natl Acad Sci USA 78:454–458
Kumar S, Tamura K, Nei M (2004) MEGA3: integrated software for molecular evolutionary genetics analysis and sequence alignment. Brief Bioinform 5:150–163
Lagercrantz U, Ryman N (1990) Genetic structure of Norway spruce (Picea abies): concordance of morphological and allozymic variation. Evolution 44:38–53
Liukkonen-Anttila T, Uimaniemi L, Orell M, Lumme J (2002) Mitochondrial DNA variation and the phylogepgraphy of the grey partridge (Perdix perdix) in Europe: from Pleistocene history to present day populations. J Evol Biol 18:971–982
Liukkonen-Anttila T, Rätti O, Kvist L, Helle P (2004) Lack of genetic structuring and subspecies differentiation in the capercaillie (Tetrao urogallus) in Finland. Ann Zool Fenn 41:619–633
Luo SJ, Kim JK, Johnson WE, van der Walt J, Martenson J, Yuhki N, Miquelle DG, Uphyrkina O, Goodrich JM, Quigley HB, Tilson R, Brady G, Martelli P, Subramaniam V, McDougal C, Hean S, Huang SQ, Pan W, Karanth UK, Sunquist M, Smith JLD, O′Brien SJ (2004) Phylogrography and genetic ancestry of tigers (Panthera tigris). PLoS Biol 2:2275–2293
Mace GM (2004) The role of taxonomy in species conservation. Philos Trans R Soc Lond B 359:711–719
Merilä J, Björkland M, Baker AJ (1997) Historical demography and present day population structure of the greenfinch, Carduelis chloris—an analysis of mtDNA control-region sequences. Evolution 51:946–956
Milá B, McCormack JE, Castañeda G, Wayne RK, Smith TB (2007) Recent postglacial range expansion drives the rapid diversification of a songbird lineage in the genus Junco. Proc R Soc Lond B 274:2653–2660
Mlíkovský J (1995) Middle Pleistocene birds of Elaichoria 3, Greece. Cour Forsch Seckenberg 181:271–273
Moritz C (1994) Defining evolutionary significant units for conservation. Trends Ecol Evol 9:373–375
Moritz C (2002) Strategies to protect biological diversity and the evolutionary processes that sustain it. Syst Biol 51:238–254
Naydenov K, Senneville S, Beaulieu J, Tremblay F, Bousquet J (2007) Glacial vicariance in Eurasia: mitochondrial DNA evidence from Scots pine for a complex heritage involving genetically distinct refugia at mid-northern latitudes and in Asia Minor. BMC Evol Biol 7:233
Nei M (1987) Molecular evolutionary genetics. Columbia University Press, New York, p 512
Niewold FJJ, Nijland H (1987) Die Chancen des westeuropäischen Moor- und Heidebirkhuhns. Z Jagdwiss 33:227–241
O′Brien SJ, Mayr E (1991) Bureaucratic mischief: recognizing endangered species and subsecies. Science 251:1187–1188
Ottvall R, Höglund J, Bensch S, Larsson K (2005) Population differentiation in the redshank (Tringa tetanus) as revealed by mitochondria DNA and amplified fragment length polymorphism markers. Conserv Genet 6:321–331
Palmé AE, Su Q, Rautenberg A, Manni F, Lascoux M (2003) Postglacial recolonisation and cpDNA variation of silver birch, Betula pendula. Mol Ecol 12:201–212
Peters JL (1934) Check-list of birds of the world, vol II. Harvard University Press, Cambridge
Phillimore AB, Owens IPF (2006) Are subspecies useful in evolutionary and conservation biology? Proc R Soc Lond B 273:1049–1053
Pitra C, Lieckfeldt D, Alonso JC (2000) Population subdivision in Europe’s great bustard inferred from mitochondrial and nuclear DNA sequence variation. Mol Ecol 9:1165–1170
Posada D, Crandall KA (1998) MODELTEST: testing the model of DNA substitution. Bioinformatics 14:817–818
Provan J, Bennett KD (2008) Phylogeographic insights into cryptic glacial refugia. Trends Ecol Evol 23:564–571
Quinn TW (1992) The genetic legacy of mother goose—phylogeographic patterns of lesser snow goose Chen caerulescens caurelescens maternal lineages. Mol Ecol 1:105–117
Quinn TW, Wilson AC (1993) Sequence evolution in and around the control region in birds. J Mol Evol 37:417–425
Randi E, Lucchini V, DeMarta P (1999) Evolution of the mitochondrial control-region in populations of galliforms (Alectoris, Tetrao and Lagopus). In: Adams NJ, Slotow RH (eds) Proceedings of the 22th international ornithological congress, Durban, pp 1873–1880. BirdLife South Africa, Johannesburg
Rhim SJ, Hur W-H, Lee W-S (2003) The first record of black grouse Tetrao tetrix in South Korea. Korean J Ornithol 9:69–70
Rogers AR, Harpending H (1992) Population growth makes waves in the distribution of pairwise genetic differences. Mol Biol Evol 9:552–269
Ronquist F, Huelsenbeck JP (2003) MRBAYES 3: bayesian phylogenetic inference under mixed models. Bioinformatics 19:1572–1574
Rozas J, Sanchez-del Barrio JC, Messeguer X, Rozas R (2003) DnaSP, DNA polymorphism analyses by the coalescent and other methods. Bioinformatics 19:2496–2497
Ruokonen M, Kvist L (2002) Structure and evolution of the avian mitochondrial control region. Mol Phyl Evol 23:422–432
Sanchez-Marco A (2004) Avian zoogeographical patterns during the Quaternary in the Mediterranean region and paleoclimatic interpretation. Ardeola 51:91–132
Schmitt T (2007) Molecular biogeography of Europe: Pleistocene cycles and postglacial trends. Front Zool 4:11
Schönwetter M (1960–1962) Handbuch der Oologie, vol I, IV. Akademie, Berlin
Schreiber A, Weitzel T, Strauß E (1998) Allozyme variability in Black Grouse (Tetrao tetrix), a tetraonid with lek behaviour. J Ornithol 139:55–66
Seddon JM, Santucci F, Reeve NJ, Hewitt GM (2001) DNA footprints of european hedgehogs, Erinaceus europaeus and E. concolor: Pleistoncene refugia, postglacial expansion and colonisation routes. Mol Ecol 10:2187–2198
Segelbacher G, Piertney S (2007) Phylogeography of the European capercaillie (Tetrao urogallus) and its implications for conservation. J Ornithol 148:S269–S274
Skrede I, Eidesen PB, Piñeiro-Portela R, Brochmann C (2006) Refugia, differentiation and postglacial migration in arctic-alpine Eurasia, exemplified by the mountain avens (Dryas octopetala L). Mol Ecol 13:1827–1840
Slatkin M, Hudson RR (1991) Pairwise comparisons of mitochondria DNA sequences in stable exponentially growing populations. Genetics 129:555–562
Stewart JR, Lister AM, Barnes I, Dalén L (2010) Refugia revisited: individualistic responses of species in space and time. Proc R Soc Lond B 277:661–671
Storch I (2007) Grouse: status survey and conservation action plan 2006–2010 gland. IUCN and Fordingbridge, Switzerland. World Pheasant Association, Heddon on the Wall
Taberlet P, Fumagalli L, Wust-Saucy AG, Cosson JF (1998) Comparative phylogeography and postglacial colonisation routes in Europe. Mol Ecol 7:453–464
Tajima F (1989) Statistical method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics 123:585–595
Wilting A, Buckley-Beason V, Feldhaar H, Gadau J, O′Brien SJ, Linsenmair KE (2007) Clouded leopard phylogeny revisted: support for species recognition and population division between Borneo and Sumatra. Front Zool 4:15
Young JR, Braun CE, Oyler-McCance SJ, Hupp JW, Quinn TW (2000) A new species of sage-grouse (Phasianidae: Centrocercus) from Southwestern Colorado. Wilson Bull 112:445–453
Zink RM (2004) The role of subspecies in obscuring avian biological diversity and misleading conservation policy. Proc R Soc Lond B 271:561–564
Acknowledgments
We thank Eleanor Jones and Yvonne Meyer-Lucht for their valuable comments on the manuscript. We also thank Pär Ericsson from the Natural History Museum of Stockholm, Jakob Hallermann and Cordula Bracker from the Zoological Museum at University of Hamburg, Grethe Leth Espensen from the Natural History Museum of Århus, and Martti Hilden from the Finnish Natural History Museum. We are also grateful with Gernot Segelbecher, Ron Plummer and Francois Mougeot for obtaining samples from Wales and Kazakhstan, respectively. We are especially grateful to Gunilla Engström who kindly helped us with laboratory work. This work was supported by the Swedish Research Council (grant number VR 2008–5418) to J.H.; and Stiftelsen för Zoologisk Forskning to C.C. C.C. has a PhD scholarship from the Fundación COLFUTURO and the AlBan program.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by M. Wink.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution License which permits any use, distribution, and reproduction in any medium, provided the original author(s) and the source are credited.
About this article
Cite this article
Corrales, C., Pavlovska, M. & Höglund, J. Phylogeography and subspecies status of Black Grouse. J Ornithol 155, 13–25 (2014). https://doi.org/10.1007/s10336-013-0982-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10336-013-0982-z