Abstract
Objective
To determine the variability, and preferred values, for normal liver longitudinal water proton relaxation rate R1 in the published literature.
Methods
Values of mean R1 and between-subject variance were obtained from literature searching. Weighted means were fitted to a heuristic and to a model.
Results
After exclusions, 116 publications (143 studies) remained, representing apparently normal liver in 3392 humans, 99 mice and 249 rats. Seventeen field strengths were included between 0.04 T and 9.4 T. Older studies tended to report higher between-subject coefficients of variation (CoV), but for studies published since 1992, the median between-subject CoV was 7.4%, and in half of those studies, measured R1 deviated from model by 8.0% or less.
Discussion
The within-study between-subject CoV incorporates repeatability error and true between-subject variation. Between-study variation also incorporates between-population variation, together with bias from interactions between methodology and physiology. While quantitative relaxometry ultimately requires validation with phantoms and analysis of propagation of errors, this survey allows investigators to compare their own R1 and variability values with the range of existing literature.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The liver longitudinal water proton relaxation rate R1 is important for several reasons. Native R1 is a biomarker of liver pathology [1, 2]. Also, other liver biomarkers are secondarily derived from R1 measurements: for example, increase in R1 post-gadoxetate is a biomarker of hepatocyte function [3, 4]; extracellular volume is derived by comparing R1 pre and post contrast [5]; and baseline R1 is required for rate constants in dynamic contrast-enhanced MR [6], for tissue oxygen tension in oxygen-enhanced MR [7], and for relaxivity measurements in contrast agent research[8].
Measurements of R1 in individual livers or liver regions suffer from both systematic errors and random errors [9]. Systematic errors (bias) arise because measurements are imperfectly performed. Other systematic deviations occur because different methods, even when perfectly performed, yield R1 values with different dependences on liver composition and physiology. Random (repeatability) errors arise from physiologic and instrument noise, and can be high particularly when regions-of-interest are small. In addition, even in the absence of bias and noise, there are, in each study, genuine between-subject differences in R1 due to between-subject variation in physiology or subclinical pathology.
To mitigate the effects of random error in establishing a “normal” or “baseline” liver R1, investigators sometimes employ a "compromise" R1, averaged from all subjects in their study. This likely reduces the "noise" variance, but introduces other errors by ignoring true between-subject variation. Other investigators may obtain R1 from literature reports, although this will introduce additional bias if different measurement methods had been used, or different populations had been studied.
The aim of this study was to survey values, and variabilities, of normal liver R1 from the published literature. This would give investigators an indication of whether the liver R1 or T1 values and variabilities they measure are broadly consistent with, or discordant from, the prior literature.
Methods
Literature searching
Literature was searched manually using "Ovid Medline" (www.ovid.com) for “magnetic resonance imaging” AND “liver” AND “relaxation”. Additional literature reports were retrieved from citations, supplemented by a more intensive search for data with B0 = 4.7 T, 7 T, 9.4 T, 11.7 T, 14.1 T or 21.1 T (see supplementary material 1 for further details). Liberal inclusion criteria were employed: any report, in any language, which claimed to measure liver R1 or T1 was included, irrespective of methodology or study design. Studies where B0 was unclear, or where liver R1 or T1 was measured but not reported, were necessarily excluded. Studies using Look-Locker methods were included if they reported T1 or R1, but excluded if they reported an apparent T1* only. Human and rodent subjects were included if they were normal controls of any age, if the study reported normal parts of livers with focal disease, or if they were patients in whom no liver abnormality had been found. Studies of definitely pathological liver, suspected duplicates, and ex vivo studies were excluded.
Analysis
The mean and variance of R1 across all subjects in each study was estimated from the publications, with the coefficient of variation given by \({\text{CoV}}=\sqrt{\text{variance}}/{\text{mean}}\). Where measurements were made on the same subjects using the same method (repeatability), the weighted mean ± SD was used, however where measurements were made on the same subjects using different method (e.g., different field strengths) the measurements were treated as if from two different studies. Any R1 measurement method was allowed, as long as T1 (s) or R1 (s−1) was reported. Where T1 ± SD was reported, a point estimate of R1 was estimated as T1−1 and the between-subject variance in R1 was estimated (see supplementary material 2) as:
In a few cases, the between-subject variance in R1 was estimated from a bar or scatterplot depicted in the publication, or from the range rule [10]. To aggregate the data, individual studies were weighted by the inverse of their between-subject variance in R1. Studies with N = 1, or where a variance could not be extracted, were included in Figs. 1 and 2, but their R1 was assigned zero weight in the fits. In addition, a method to account for the well-known B0-dependence of liver R1 [11,12,13,14,15] was needed. Two methods of representing this B0 dependence were used: a heuristic log–log relationship, and a biophysical power-law model developed by Diakova et al. [12]. R1 was fitted to B0 using the weighted non-linear least squares function nls() in R[16] (see supplementary material 3). The fitted parameters in the heuristic were M and C:
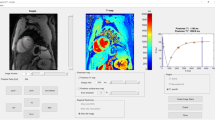
Log–log dependence of longitudinal relaxation rate on field strength. Blue: human; Red: rat; Green: mouse. Each symbol represents one study. Size of circle reflects number of subjects (some smaller symbols are occluded by larger symbols). Dashed black line: fit to Eq. 2. Solid black line: fit to Eq. 3 with \({R}_{1,\infty }\)= 0.213 s−1. The dotted line illustrates, for the benefit of investigators working at > 10 T, fits to Eq. 3 where \({R}_{1,\infty }\) was fixed at higher values of 0.4 s−1, 0.6 s−1, and 0.8 s−1, intermediate between 0.213 s−1 and the 0.9–1.0 s−1 value observed at 9.4 T in Table 1
The fitted parameters in the model were A and B:
where \({R}_{1,\infty }\) is the high-frequency asymptote, i.e., the extreme narrowing condition, set here to 0.213 s−1 at 310 K[17]; \({\tau }_{D}\) is the translational correlation time from Diakova et al. [12] adjusted for temperature to 1.43 × 10–11 s; \(k=-0.6\) also from Diakova et al. [12]; and \(\omega =2\pi \times 42.58\times {10}^{6}\times {\text{B}}_{0}\) s−1. In the summaries, lower (LQ) and upper (UQ) quartiles, and medians, are reported. For exploratory fits using other weightings, see Supplementary Material 4.
Results
Approximately 500 publication abstracts were read, from which around 270 publications were selected and reviewed. After exclusions, 116 publications remained, with publication dates between 1981 and 2020. Some publications reported multiple studies, or multiple groups within a single study, so that 143 studies were available to contribute to this analysis. These represented 3392 humans [1,2,3,4, 7, 11, 14, 15, 18,19,20,21,22,23,24,25,26,27,28,29,30,31,32,33,34,35,36,37,38,39,40,41,42,43,44,45,46,47,48,49,50,51,52,53,54,55,56,57,58,59,60,61,62,63,64,65,66,67,68,69,70,71,72,73,74,75,76,77,78,79,80,81,82,83,84,85,86,87,88,89,90,91,92,93,94], 99 mice [95,96,97,98,99,100,101,102,103,104,105] and 249 rats [5, 33, 105,106,107,108,109,110,111,112,113,114,115,116,117,118,119,120,121,122,123,124,125,126]. The number of subjects per study varied between 1 and 1037 (median 12). A very wide variety of T1 measuring methods was used. Frequently used approaches (see supplementary material 5) were inversion-recovery (18% of studies), saturation-recovery (21%) or variable-flip-angle (10%), which compare signal arising respectively when inversion time, repetition time, or flip angle are incremented. The median number of increments was 3 (range 2–20). Various read-outs were employed including spin-echo, gradient-echo, echo-planar or localized spectroscopy. Other studies employed variants of Look-Locker (24%) or MR fingerprinting (1%). Some studies reported that they suppressed fat, and/or corrected for iron-induced T1-shortening; some reported motion suppression, registration, triggering, gating or breath-hold; some reported B1 correction or phantom-based validation. Some studies analysed quite small regions of interest often avoiding blood vessels and bile ducts; others included most or all of the liver. Seventeen field strengths were included between 0.04 T and 9.4 T. No values were found in reports using B0 > 9.4 T: one report of \({T}_{1}^{*}=1.0\pm 0.1\mathrm{ s}\) at 14.1 T was excluded[127]. Figures 1 and 2 show plots of R1 against B0, in which R1 shows the expected decrease with increasing field: Table 1 gives values for the most important field strengths. The fit to Eq. 2 gave \(M=-0.3611\pm 0.0115\) and \(C=0.2956\pm 0.0073\). The fit to Eq. 3 gave \(A=(8.663\pm 0.681)\times {10}^{4}\) and \(B=(1.294\pm 0.082)\times {10}^{9}\). An exploratory attempt at a three-parameter fit to Eq. 3 (i.e., to A, B, and \({R}_{1,\infty }\)) failed to provide evidence for \({R}_{1,\infty }>0\) (supplementary material 4). When data were subgouped by species or by method, no evidence was found that the subgoup R1 values deviated systematically from Eq. 3 (supplementary material 6). Across all studies, the median between-subject CoV was 9.1% (LQ 5.9%, UQ 16.5%, rms 17.0%). There was, however, a tendency for early studies to report high between-subject CoV (Fig. 3 and supplementary material 7): no study published after 1992 had CoV ≥ 20%, and for post-1992 studies the median between-subject CoV was 7.4% (LQ 5.6%, UQ 11.0%, rms 9.6%). In half those studies, the measured R1 deviated from Eq. 3 by 8.0% or less (LQ 2.8%, UQ 16.6%).
At each field strength, there was considerable variation in R1 between studies: the between-study CoV was 16% for post-1992 studies. Six publications[2, 37, 98, 119, 128, 129] also reported liver R1 repeatability (same subject, different scan, same measurement conditions): the rms CoV was 1.9%. These CoVs allowed a crude estimate (supplementary material 8) of the relative size of the three main variance components: repeatability variance contributed ~ 1%; within-study-between-subject variance contributed ~ 25%; and between-study variance contributed ~ 74%.
Discussion
In liver, as in pure water, both intramolecular and intermolecular water 1H-1H dipolar relaxation contribute to R1. Specific additional contributors to water 1H R1 in liver arise from 1H-1H dipolar relaxation between water and other molecules, and 1H-electron dipolar relaxation between water and various iron- or copper-containing substances or dioxygen. These 1H-containing and unpaired-electron-containing substances differ in concentration between subjects. The liver 1H resonance arises mostly from tissue water in hepatocytes. Other contributions come from water in other intracellular compartments (e.g., Kupffer cells, erythrocytes), and in extracellular compartments (e.g., bile, plasma, space of Disse). Signal from triglyceride and inflowing blood may contribute, depending on the sequence used. Macromolecules contribute to the signal, notably collagen and glycogen which have different concentrations in different subjects. These factors likely account for some of the variation between subjects and between studies.
Fits from the heuristic and from the model were very similar. The main difference is that the heuristic forces R1 to zero at infinite field, while the model forces R1 to asymptote in the extreme narrowing condition. This difference might become important at fields above 7 T (Fig. 1). In this study, following Diakova et al.[12], the asymptote \({R}_{1,\infty }\) was fixed at 1/4.7 s−1, equal to the R1 of pure deoxygenated water at 310 K at high field [17]: a slightly higher value would be more appropriate if R1 values from liver water and pure water do not converge as illustrated in Fig. 1.
The relative magnitude of the major variance components was estimated. This is very crude, and given the heterogeneity and variable quality of the raw data, should be considered a rough guide only. The within-study between-subject CoV reflects not only repeatability error (~ 1% of the variance), but also the expected between-subject variation (~ 25% of the variance). Between-study variation (~ 74% of the variance) also includes between-population variation, together with bias from interactions between each study’s measurement method and its livers’ variation in flow, motion, fat, oedema, collagen, glycogen and iron. R1 may also change after a meal [89], during the menstrual cycle [25] or with drug treatment [25].
The literature survey was not fully PRISMA-compliant [130] and is unlikely to be complete. Studies explicitly of liver R1 or T1 as a biomarker are readily retrieved, because appropriate keywords are generally used in the title and abstract. However, for studies where liver R1 or T1 measurement is incidental to another objective, for example extracellular volume, relaxivity, or dynamic contrast-enhanced studies, suitable keywords may not have been included.
There is no single “correct” value for any liver’s 1H R1. R1 may vary spatially across the liver [60, 119]. Water 1H R1 is multiexponential, particularly with sequences where macromolecule-associated fast-relaxing water contributes to the measurement. Other substances in the liver may also contribute to the 1H signal, such as glycogen [87] or triglyceride [76, 131]. Inflowing blood [110, 132], physiologic motion [71], magnetization transfer, and iron affect the measured R1 in ways which depend both on the sequence and on the analysis employed. There may be systematic differences in R1 between fat-suppressed vs. non-fat-suppressed acquisitions; 2D acquisitions more vulnerable to inflow effects than 3D; breathhold or gated vs. free-breathing; and so on. Some investigators advocate the use of a “corrected” T1 to avoid bias caused by the relaxivity of iron-containing substances [65]. Because of these biases in the literature, studies which deviate from these survey data should not immediately be considered “incorrect”, but if large deviations are observed, then an explanation on methodological or physiological grounds should be sought.
There are some other limitations. While some publications reported carefully designed and conducted biomarker validation studies, in other publications, the precise value of T1 was only of incidental interest and possibly acquired with less care. However, in this survey, the study design and objectives were not incorporated into the weightings. Most studies did not report validation of their liver R1 by means of a phantom, so accuracy is unknown. It was difficult to explore the effect of methodology on R1, because some studies used methodology which was poorly described or did not appear robust, and because of correlation between field strength and methodology (old studies used old methodology and lower fields). Likewise, there was correlation between field strength and species (humans at low-medium fields, rats at medium–high fields and mice at high fields), so it was difficult to compare between species.
Conclusion
Quantitative relaxometry requires validation with phantoms and analysis of propagation of errors. However, it is also good scientific practice to compare one’s own findings with prior literature. An investigator who finds their average liver R1 in normal liver to be within 8% of the fit to Eq. 3, with between-subject CoV < 8%, can conclude that their measurements are in agreement with the majority of the literature: for measurements far outside these limits, a physiological or methodological explanation should be sought.
References
Smith FW, Mallard JR, Reid A, Hutchison JMS (1981) Nuclear magnetic resonance tomographic imaging in liver disease. Lancet 317:963–966
Banerjee R, Pavlides M, Tunnicliffe EM, Piechnik SK, Sarania N, Philips R, Collier JD, Booth JC, Schneider JE, Wang LM, Delaney DW, Fleming KA, Robson MD, Barnes E, Neubauer S (2014) Multiparametric magnetic resonance for the non-invasive diagnosis of liver disease. J Hepatol 60:69–77
Haimerl M, Utpatel K, Verloh N, Zeman F, Fellner C, Nickel D, Teufel A, Fichtner-Feigl S, Evert M, Stroszczynski C, Wiggermann P (2017) Gd-EOB-DTPA-enhanced MR relaxometry for the detection and staging of liver fibrosis. Sci Rep 7:41429
Haimerl M, Verloh N, Fellner C, Zeman F, Teufel A, Fichtner-Feigl S, Schreyer AG, Stroszczynski C, Wiggermann P (2014) MRI-based estimation of liver function: Gd-EOB-DTPA-enhanced T1 relaxometry of 3T vs The MELD score. Sci Rep 4:5621
Luetkens JA, Klein S, Träber F, Schmeel FC, Sprinkart AM, Kuetting DLR, Block W, Uschner FE, Schierwagen R, Hittatiya K, Kristiansen G, Gieseke J, Schild HH, Trebicka J, Kukuk GM (2018) Quantification of liver fibrosis at T1 and T2 mapping with extracellular volume fraction MRI: Preclinical results. Radiology 288:748–754
Li Z, Sun J, Chen L, Huang N, Hu P, Hu X, Han G, Zhou Y, Bai W, Niu T, Yang X (2016) Assessment of liver fibrosis using pharmacokinetic parameters of dynamic contrast-enhanced magnetic resonance imaging. J Magn Reson Imaging 44:98–104
O’Connor JPB, Naish JH, Jackson A, Waterton JC, Watson Y, Cheung S, Buckley DL, McGrath DM, Buonaccorsi GA, Mills SJ, Roberts C, Jayson GC, Parker GJM (2009) Comparison of normal tissue R1 and R2* modulation by oxygen and carbogen. Magn Reson Med 61:75–83
Ziemian S, Green C, Sourbron S, Jost G, Schütz G, Hines CDG (2021) Ex vivo gadoxetate relaxivities in rat liver tissue and blood at five magnetic field strengths from 1.41 to 7T. NMR Biomed. 34:e4401
Raunig DL, McShane LM, Pennello G, Gatsonis C, Carson PL, Voyvodic JT, Wahl RL, Kurland BF, Schwarz AJ, Gönen M, Zahlmann G, Kondratovich MV, O’Donnell K, Petrick N, Cole PE, Garra B, Sullivan DC (2015) Quantitative imaging biomarkers: A review of statistical methods for technical performance assessment. Stat Methods Med Res 24:27–67
Hozo SP, Djulbegovic B, Hozo I (2005) Estimating the mean and variance from the median, range, and the size of a sample. BMC Med Res Methodol 5:13
Araya YT, Martínez-Santiesteban F, Handler WB, Harris CT, Chronik BA, Scholl TJ (2017) Nuclear magnetic relaxation dispersion of murine tissue for development of T1 (R1) dispersion contrast imaging. NMR Biomed 30:e3789
Diakova G, Korb JP, Bryant RG (2012) The magnetic field dependence of water T1 in tissues. Magn Reson Med 68:272–277
Thomsen C (1996) Quantitative magnetic resonance methods for in vivo investigation of the human liver and spleen. Technical aspects and preliminary clinical results. Acta Radiol Suppl 401:1–34
Keevil SF, Dolke G, Brooks AP, Armstrong P, Farthing MJG, Alstead EM, Smith MA (1992) Proton NMR relaxation times in the normal human liver at 0.08 T. Clin Radiol 45:302–306
Henriksen O, de Certaines JD, Spisni A, Cortsen M, Muller RN, Ring PB (1993) V. In vivo field dependence of proton relaxation times in human brain, liver and skeletal muscle: A multicenter study. Magn Reson Imaging 11:851–856
R Development Core Team (2018) R 3.5.1., A language and environment for statistical computing. R Found Stat Comput 2. https://www.R-project.org.
Krynicki K (1966) Proton spin-lattice relaxation in pure water between 0°C and 100°C. Physica 32:167–178
Kamimura K, Fukukura Y, Yoneyama T, Takumi K, Tateyama A, Umanodan A, Shindo T, Kumagae Y, Ueno SI, Koriyama C, Nakajo M (2014) Quantitative evaluation of liver function with T1 relaxation time index on Gd-EOB-DTPA-Enhanced MRI: Comparison with signal intensity-based indices. J Magn Reson Imaging 40:884–889
Kim KA, Park MS, Kim IS, Kiefer B, Chung WS, Kim MJ, Kim KW (2012) Quantitative evaluation of liver cirrhosis using T1 relaxation time with 3 tesla MRI before and after oxygen inhalation. J Magn Reson Imaging 36:405–410
Heye T, Yang SR, Bock M, Brost S, Weigand K, Longerich T, Kauczor HU, Hosch W (2012) MR relaxometry of the liver: Significant elevation of T1 relaxation time in patients with liver cirrhosis. Eur Radiol 22:1224–1232
Block W, Reichel C, Träber F, Skodra T, Lamerichs R, Kreft B, Spengler U, Sauerbruch T, Schild H (1997) Effect of cytochrome P450 induction on phosphorus metabolites and proton relaxation times measured by in vivo 31P-magnetic resonance spectroscopy and 1H-magnetic resonance relaxometry in human liver. Hepatology 26:1587–1591
de Certaines JD, Henriksen O, Spisni A, Cortsen M, Ring PB (1993) IV. In vivo measurements of proton relaxation times in human brain, liver, and skeletal muscle: A multicenter MRI study. Magn Reson Imaging 11:841–850
Van Lom KJ, Brown JJ, Perman WH, Sandstrom JC, Lee JKT (1991) Liver imaging at 1.5 Tesla: Pulse sequence optimization based on improved measurement of tissue relaxation times. Magn Reson Imaging 9:165–171
Steudel A, Harder T, Träber F, Dewes W, Schlolaut KH, Koster O (1989) Relaxationszeitmessungen in Der Kernspintomographischen Differentialdiagnose Von Lebertumoren. RöFo Fortschritte auf dem Gebiete der Röntgenstrahlen und der Neuen Bildgeb Verfahren 151:449–455
Richards MA, Webb JAW, Jewell SE, Gregory WM, Reznek RH (1988) In-vivo measurement of spin lattice relaxation time (T1) of liver in healthy volunteers: The effects of age, sex and oral contraceptive usage. Br J Radiol 61:34–37
Thomsen C, Christoffersen P, Henriksen O, Juhl E (1990) Prolonged T1 in patients with liver cirrhosis: An in vivo MRI study. Magn Reson Imaging 8:599–604
Cassinotto C, Feldis M, Vergniol J, Mouries A, Cochet H, Lapuyade B, Hocquelet A, Juanola E, Foucher J, Laurent F, De Ledinghen V (2015) MR relaxometry in chronic liver diseases: Comparison of T1 mapping, T2 mapping, and diffusion-weighted imaging for assessing cirrhosis diagnosis and severity. Eur J Radiol 84:1459–1465
Henninger B, Kremser C, Rauch S, Eder R, Zoller H, Finkenstedt A, Michaely HJ, Schocke M (2012) Evaluation of MR imaging with T1 and T2* mapping for the determination of hepatic iron overload. Eur Radiol 22:2478–2486
Weinreb JC, Brateman L, Maravilla KR (1984) Magnetic resonance imaging of hepatic lymphoma. Am J Roentgenol 143:1211–1214
Belt TG, Cohen MD, Smith JA, Cory DA, McKenna S, Weetman R (1986) MRI of Wilms’ tumor: Promise as the primary imaging method. Am J Roentgenol 146:955–961
Ohtomo K, Itai Y, Furui S, Yoshikawa K, Yashiro N, Iio M (1985) Magnetic resonance imaging (MRI) of primary liver cancer. MRI- pathologic correlation. Radiat Med - Med Imaging Radiat Oncol 3:38–41
Nyman R, Ericsson A, Hemmingsson A, Jung B, Sperber G, Thuomas KÅ (1986) T1, T2, and relative proton density at 0.35 T for spleen, liver, adipose tissue, and vertebral body: Normal values. Magn Reson Med 3:901–910
Stark DD, Moseley ME, Bacon BR (1985) Magnetic resonance imaging and spectroscopy of hepatic iron overload. Radiology 154:137–142
Gilligan LA, Dillman JR, Tkach JA, Xanthakos SA, Gill JK, Trout AT (2019) Magnetic resonance imaging T1 relaxation times for the liver, pancreas and spleen in healthy children at 1.5 and 3 tesla. Pediatr Radiol 49:1018–1024
Kim JE, Kim HO, Bae K, Choi DS, Nickel D (2019) T1 mapping for liver function evaluation in gadoxetic acid–enhanced MR imaging: comparison of look-locker inversion recovery and B1 inhomogeneity–corrected variable flip angle method. Eur Radiol 29:3584–3594
Yang L, Ding Y, Rao S, Chen C, Zeng M (2020) T1 mapping on Gd-EOB-DTPA-enhanced MRI for the prediction of oxaliplatin-induced liver injury in a mouse model. J Magn Reson Imaging 53:896–902
Bradley CR, Cox EF, Scott RA, James MW, Kaye P, Aithal GP, Francis ST, Guha IN (2018) Multi-organ assessment of compensated cirrhosis patients using quantitative magnetic resonance imaging. J Hepatol 69:1015–1024
Zhou ZP, Long LL, Qiu WJ, Cheng G, Huang LJ, Yang TF, Huang ZK (2017) Comparison of 10- and 20-min hepatobiliary phase images on Gd-EOB-DTPA-enhanced MRI T1 mapping for liver function assessment in clinic. Abdom Radiol 42:2272–2278
Agrawal S, Hoad CL, Francis ST, Guha IN, Kaye P, Aithal GP (2017) Visual morphometry and three non-invasive markers in the evaluation of liver fibrosis in chronic liver disease. Scand J Gastroenterol 52:107–115
Tunnicliffe EM, Banerjee R, Pavlides M, Neubauer S, Robson MD (2017) A model for hepatic fibrosis: the competing effects of cell loss and iron on shortened modified Look-Locker inversion recovery T1 (shMOLLI-T1) in the liver. J Magn Reson Imaging 45:450–462
Chen Y, Jiang Y, Pahwa S, Ma D, Lu L, Twieg MD, Wright KL, Seiberlich N, Griswold MA, Gulani V (2016) MR fingerprinting for rapid quantitative abdominal imaging. Radiology 279:278–286
Ding Y, Rao SX, Chen C, Li R, Zeng MS (2015) Assessing liver function in patients with HBV-related HCC: a comparison of T1 mapping on Gd-EOB-DTPA-enhanced MR imaging with DWI. Eur Radiol 25:1392–1398
Haimerl M, Verloh N, Zeman F, Fellner C, Müller-Wille R, Schreyer AG, Stroszczynski C, Wiggermann P (2013) Assessment of clinical signs of liver cirrhosis using T1 mapping on Gd-EOB-DTPA-enhanced 3T MRI. PLoS ONE 8:e85658
Katsube T, Okada M, Kumano S, Hori M, Imaoka I, Ishii K, Kudo M, Kitagaki H, Murakami T (2011) Estimation of liver function using T1 mapping on Gd-EOB-DTPA-enhanced magnetic resonance imaging. Invest Radiol 46:277–283
Jafari F, Nayeri N, Tahsini M, Khodadoust AA (1999) Differentiation of hepatic cavernous hemangioma from metastases by rare sequence MR imaging. Magn Reson Imaging 17:669–677
Halavaara J, Lukkarinen S, Sepponen R, Markkola A, Tanttu J (2003) Contrast-to-noise ratio of multiple slice spin lock technique: Prospects for liver imaging. Br J Radiol 76:788–791
Skjold A, Vangberg TR, Kristoffersen A, Haraldseth O, Jynge P, Larsson HBW (2004) Relaxation enhancing properties of MnDPDP in human myocardium. J Magn Reson Imaging 20:948–952
De Bazelaire CMJ, Duhamel GD, Rofsky NM, Alsop DC (2004) MR Imaging Relaxation Times of Abdominal and Pelvic Tissues Measured in Vivo at 3.0 T: Preliminary Results. Radiology 230:652–659
Tadamura E, Hatabu H, Li W, Prasad PV, Edelman RR (1997) Effect of oxygen inhalation on relaxation times in various tissues. J Magn Reson Imaging 7:220–225
Morio S, Oh H, Endo N, Kawano E, Nakamura H, Asai T, Saito Y, Uchida Y, Ikehira H, Yoshida K (1997) Magnetic resonance imaging of reticulo-endothelial system in patients with idiopathic thrombocytopenic purpura. Am J Hematol 56:52–58
Tamburrini O, Andò S, Della Sala M, Maggiolini M, Sessa M (1993) Emocromatosi epatica secondaria: diagnosi e quantificazione con risonanza magnetica 0.5 T. Valore e limite Radiol Medica 86:841–846
Blüml S, Schad LR, Stepanow B, Lorenz WJ (1993) Spin-lattice relaxation time measurement by means of a TurboFLASH technique. Magn Reson Med 30:289–295
Patrizio G, Pavone P, Testa A, Marsili L, Tettamanti E, Passariello R (1990) MR characterization of hepatic lesions by T-null inversion recovery sequence. J Comput Assist Tomogr 14:96–101
Squillaci E, Cecconi L, Tipaldi L, Grandinetti ML, Orlacchio A, Squillaci S (1989) La Risonanza Magnetica Nelle Lesioni Epatiche. Esperienza Con Campo Magnetico Da 1,5 T. Radiol Medica 78:585–592
Rummeny E, Weissleder R, Stark DD, Saini S, Compton CC, Bennett W, Hahn PF, Wittenberg J, Malt RA, Ferrucci JT (1989) Primary liver tumors: Diagnosis by MR imaging. Am J Roentgenol 152:63–72
Rademaker M, Webb JAW, Lowe DG, Meyrick-thomas RH, Kirby JDT, Munro DD (1987) Magnetic resonance imaging as a screening procedure for methotrexate induced liver damage. Br J Dermatol 117:311–316
The Clinical NMR Group (1987) Magnetic resonance imaging of parenchymal liver disease: a comparison with ultrasound, radionuclide scintigraphy and X-ray computed tomography. Clin Radiol 38:495–502
Richards MA, Webb J, Reznek RH, Davies G, Jewell SE, Shand WS, Wrigley PFM, Lister TA (1986) Detection of spread of malignant lymphoma to the liver by low field strength magnetic resonance imaging. Br Med J (Clin Res Ed) 293:1126–1128
Glazer GM, Aisen AM, Francis IR, Gyves JW, Lande I, Adler DD (1985) Hepatic cavernous hemangioma: Magnetic resonance imaging. Radiology 155:417–420
Obmann VC, Mertineit N, Marx C, Berzigotti A, Ebner L, Heverhagen JT, Christe A, Huber AT (2019) Liver MR relaxometry at 3T – segmental normal T1 and T2* values in patients without focal or diffuse liver disease and in patients with increased liver fat and elevated liver stiffness. Sci Rep 9:8106
Doyle FH, Pennock JM, Banks LM, McDonnell MJ, Bydder GM, Steiner RE, Young IR, Clarke GJ, Pasmore T, Gilderdale DJ (1982) Nuclear magnetic resonance imaging of the liver: Initial experience. Am J Roentgenol 138:193–200
Ramachandran P, Serai SD, Veldtman GR, Lang SM, Mazur W, Trout AT, Dillman JR, Fleck RJ, Taylor MD, Alsaied T, Moore RA (2019) Assessment of liver T1 mapping in fontan patients and its correlation with magnetic resonance elastography-derived liver stiffness. Abdom Radiol 44:2403–2408
Huber AT, Razakamanantsoa L, Lamy J, Giron A, Cluzel P, Kachenoura N, Redheuil A (2020) Multiparametric differentiation of idiopathic dilated cardiomyopathy with and without congestive heart failure by means of cardiac and hepatic T1-weighted MRI mapping. Am J Roentgenol 215:79–86
Obmann VC, Marx C, Berzigotti A, Mertineit N, Hrycyk J, Gräni C, Ebner L, Ith M, Heverhagen JT, Christe A, Huber AT (2019) Liver MRI susceptibility-weighted imaging (SWI) compared to T2* mapping in the presence of steatosis and fibrosis. Eur J Radiol 118:66–74
Mojtahed A, Kelly CJ, Herlihy AH, Kin S, Wilman HR, McKay A, Kelly M, Milanesi M, Neubauer S, Thomas EL, Bell JD, Banerjee R, Harisinghani M (2019) Reference range of liver corrected T1 values in a population at low risk for fatty liver disease—a UK Biobank sub-study, with an appendix of interesting cases. Abdom Radiol 44:72–84
Chen Y, Lee GR, Aandal G, Badve C, WrighT KL, Griswold MA, Seiberlich N, Gulani V (2016) Rapid volumetric T1 mapping of the abdomen using three-dimensional through-time spiral GRAPPA. Magn Reson Med 75:1457–1465
Wiese S, Voiosu A, Hove JD, Danielsen KV, Voiosu T, Grønbæk H, Møller HJ, Genovese F, Reese-Petersen AL, Mookerjee RP, Clemmesen JO, Gøtze JP, Andersen O, Møller S, Bendtsen F (2020) Fibrogenesis and inflammation contribute to the pathogenesis of cirrhotic cardiomyopathy. Aliment Pharmacol Ther 52:340–350
Runge VM, Clanton JA, Smith FW, Hutchison J, Mallard J, Partain CL, James AE (1983) Nuclear magnetic resonance of iron and copper disease states. AJR Am J Roentgenol 141:943–948
Ebara M, Ohto M, Watanabe Y, Kimura K, Saisho H, Tsuchiya Y, Okuda K, Arimizu N, Kondo F, Ikehira H (1986) Diagnosis of small hepatocellular carcinoma: Correlation of MR imaging and tumor histologic studies. Radiology 159:371–378
Brasch RC, Wesbey GE, Gooding CA, Koerper MA (1984) Magnetic resonance imaging of transfusional hemosiderosis complicating thalassemia major. Radiology 150:767–771
Ehman RL, McNamara MT, Pallack M, Hricak H, Higgins CB (1984) Magnetic resonance imaging with respiratory gating: Techniques and advantages. Am J Roentgenol 143:1175–1182
Rödl W (1985) Differentialdiagnose von Lebererkrankungen im Kernspintomogramm. RöFo Fortschritte auf dem Gebiete der Rontgenstrahlen und der bildgeb Verfahren 142:505–510
Rupp N, Reiser M, Stetter E (1983) The diagnostic value of morphology and relaxation times in NMR-imaging of the body. Eur J Radiol 3:68–76
Buonocore E, Borkowski GP, Pavlicek W, Ngo F (1983) NMR imaging of the abdomen: Technical considerations. Am J Roentgenol 141:1171–1178
Brown DW, Henkelman RM, Poon PY, Fisher MM (1985) Nuclear magnetic resonance study of iron overload in liver tissue. Magn Reson Imaging 3:275–282
Mozes FE, Tunnicliffe EM, Moolla A, Marjot T, Levick CK, Pavlides M, Robson MD (2019) Mapping tissue water T1 in the liver using the MOLLI T1 method in the presence of fat, iron and B0 inhomogeneity. NMR Biomed 32:e4030
Ding Y, Rao SX, Zhu T, Chen CZ, Li RC, Zeng MS (2015) Liver fibrosis staging using T1 mapping on gadoxetic acid-enhanced MRI compared with DW imaging. Clin Radiol 70:1096–1103
Moss AA, Goldberg HI, Stark DB, Davis PL, Margulis AR, Kaufman L, LEC, (1984) Hepatic tumors: Magnetic resonance and CT appearance. Radiology 150:141–147
Träber F, Steudel A, Harder T (1990) In-vivo-messung von geweberelaxationszeiten mit lokalisierter 31P- Und 1H-MR-Spektroskopie. RöFo Fortschritte auf dem Gebiete der Rontgenstrahlen und der Neuen Bildgeb Verfahren 153:209–215
Wang C, Wang ZC, Ding Y, Zeng MS, Rao SX (2018) Value of gadoxetate disodium-enhanced magnetic resonance on hepatobiliary phase T1 mapping for predicting liver injury. Zhonghua Gan Zang Bing Za Zhi 26:530–534
Ehman RL, Kjos BO, Hricak H, Brasch RC, Higgins CB (1985) Relative intensity of abdominal organs in MR images. J Comput Assist Tomogr 9:315–319
Flak B, Ajzen S, Li DKB, Cooperberg PL, Clark C (1989) Hemangioma of the liver: Characteristics exhibited on a 0.15 Tesla scanner. Can Assoc Radiol J 40:135–138
Fletcher BD, Kopiwoda SY, Strandjord SE, Nelson AD, Pickering SP (1985) Abdominal neuroblastoma: Magnetic resonance imaging and tissue characterization. Radiology 155:699–703
Foley WD, Kneeland JB, Cates JD, Kellman GM, Lawson TL, Middleton WD, Hendrick RE (1987) Contrast optimization for the detection of focal hepatic lesions by MR imaging at 1.5 T. Am J Roentgenol 149:1155–1160
Schmidt HC, Tscholakoff D, Hricak H, Higgins CB (1985) Mr image contrast and relaxation times of solid tumors in the chest, abdomen, and pelvis. J Comput Assist Tomogr 9:738–748
Rödl W (1984) Differential diagnosis of liver diseases with the aid of nuclear magnetic resonance imaging. In: Demling L, Lutz H, Wenz W, Wildhirt E (eds) Diagnostic Imaging Methods in Hepatolology: proceedings of the 37th Falk Symposium, held during Basel Liver Week, Basel, September 29–October 2, 1983. MTP Press, Lancaster, MA USA, pp 153–158
Weis J, Kullberg J, Ahlström H (2018) Multiple breath-hold proton spectroscopy of human liver at 3T: Relaxation times and concentrations of glycogen, choline, and lipids. J Magn Reson Imaging 47:410–417
Hoad CL, Palaniyappan N, Kaye P, Chernova Y, James MW, Costigan C, Austin A, Marciani L, Gowland PA, Guha IN, Francis ST, Aithal GP (2015) A study of T1 relaxation time as a measure of liver fibrosis and the influence of confounding histological factors. NMR Biomed 28:706–714
O’Connor JPB, Jackson A, Buonaccorsi GA, Buckley DL, Roberts C, Watson Y, Cheung S, McGrath DM, Naish JH, Rose CJ, Dark PM, Jayson GC, Parker GJM (2007) Organ-specific effects of oxygen and carbogen gas inhalation on tissue longitudinal relaxation times. Magn Reson Med 58:490–496
Nyman R, Rhen S, Ericsson A, Glimelius B, Hagberg H, Hemmingsson A, Sundström C (1987) An attempt to characterize malignant lymphoma in spleen, liver and lymph nodes with magnetic resonance imaging. Acta Radiol 28:527–533
Hardy CJ, Edelstein WA, Vatis D, Harms R, Adams WJ (1985) Calculated T1 images derived from a partial saturation-inversion recovery pulse sequence with adiabatic fast passage. Magn Reson Imaging 3:107–116
Kinami Y, Yokota H, Takata M, Takashima S, Yamamoto I (1988) Magnetic resonance imaging in the diagnosis of tumors of the liver. Gastroenterol Jpn 23:139–146
Leung A, Bydder G, Steiner R, Bryant D, Young I (1984) Magnetic resonance imaging of the kidneys. AJR Am J Roentgenol 143:1215–1227
Okada M, Murakami T, Yada N, Numata K, Onoda M, Hyodo T, Inoue T, Ishii K, Kudo M (2015) Comparison between T1 relaxation time of Gd-EOB-DTPA-enhanced MRI and liver stiffness measurement of ultrasound elastography in the evaluation of cirrhotic liver. J Magn Reson Imaging 41:329–338
Chow AM, Gao DS, Fan SJ, Qiao Z, Lee FY, Yang J, Man K, Wu EX (2012) Measurement of liver T1 and T2 relaxation times in an experimental mouse model of liver fibrosis. J Magn Reson Imaging 36:152–158
Ding Y, Yang L, Rao SX, Zeng MS (2019) Gadoxetic disodium-enhanced MRI to characterize T1 relaxation values and expression level of organic anion transporters and multidrug resistance protein on hepatocyte surface membrane of normal C57BL/6 mice. Zhonghua Gan Zang Bing Za Zhi 27:547–551
Matsuo-Tezuka Y, Sasaki Y, Iwai T, Kurasawa M, Yorozu K, Tashiro Y, Hirata M (2019) T2* relaxation time obtained from magnetic resonance imaging of the liver is a useful parameter for use in the construction of a murine model of iron overload. Contrast Media Mol Imaging 2019:7463047
Faller TL, Trotier AJ, Miraux S, Ribot EJ (2019) Radial MP2RAGE sequence for rapid 3D T1 mapping of mouse abdomen: application to hepatic metastases. Eur Radiol 29:5844–5851
Anderson CE, Wang CY, Gu Y, Darrah R, Griswold MA, Yu X, Flask CA (2018) Regularly incremented phase encoding – MR fingerprinting (RIPE-MRF) for enhanced motion artifact suppression in preclinical cartesian MR fingerprinting. Magn Reson Med 79:2176–2182
Jackson LH, Vlachodimitropoulou E, Shangaris P, Roberts TA, Ryan TM, Campbell-Washburn AE, David AL, Porter JB, Lythgoe MF, Stuckey DJ (2017) Non-invasive MRI biomarkers for the early assessment of iron overload in a humanized mouse model of β-thalassemia. Sci Rep 7:43439
Eberhardt C, Wurnig MC, Wirsching A, Rossi C, Feldmane I, Lesurtel M, Boss A (2018) Prediction of small for size syndrome after extended hepatectomy: Tissue characterization by relaxometry, diffusion weighted magnetic resonance imaging and magnetization transfer. PLoS ONE 13:e0192847
Li H, Gray BD, Corbin I, Lebherz C, Choi H, Lund-Katz S, Wilson JM, Glickson JD, Zhou R (2004) MR and fluorescent imaging of low-density lipoprotein receptors. Acad Radiol 11:1251–1259
Oostendorp M, Douma K, Hackeng TM, Post MJ, Van Zandvoort MAMJ, Backes WH (2010) Gadolinium-labeled quantum dots for molecular magnetic resonance imaging: R1 versus R2 mapping. Magn Reson Med 64:291–298
Ramasawmy R, Campbell-Washburn AE, Wells JA, Johnson SP, Pedley RB, Walker-Samuel S, Lythgoe MF (2015) Hepatic arterial spin labelling MRI: An initial evaluation in mice. NMR Biomed 28:272–280
Polasek M, Fuchs BC, Uppal R, Schühle DT, Alford JK, Loving GS, Yamada S, Wei L, Lauwers GY, Guimaraes AR, Tanabe KK, Caravan P (2012) Molecular MR imaging of liver fibrosis: A feasibility study using rat and mouse models. J Hepatol 57:549–555
Müller A, Hochrath K, Stroeder J, Hittatiya K, Schneider G, Lammert F, Buecker A, Fries P (2017) Effects of liver fibrosis progression on tissue relaxation times in different mouse models assessed by ultrahigh field magnetic resonance imaging. Biomed Res Int 2017:8720367
Braren R, Curcic J, Remmele S, Altomonte J, Ebert O, Rummeny EJ, Steingoetter A (2011) Free-breathing quantitative dynamic contrast-enhanced magnetic resonance imaging in a rat liver tumor model using dynamic radial T1 mapping. Invest Radiol 46:624–631
Cheng HLM, Haedicke IE, Cheng W, Nofiele JT, Zhang XA (2014) Gadolinium-free T1 contrast agents for MRI: Tunable pharmacokinetics of a new class of manganese porphyrins. J Magn Reson Imaging 40:1474–1480
Nekolla S, Gneiting T, Syha J, Deichmann R, Haase A (1992) T1 maps by k-space reduced snapshot-FLASH MRI. J Comput Assist Tomogr 16:327–332
Chouhan MD, Ramasawmy R, Bainbridge A, Campbell-Washburn A, Halligan S, Davies N, Walker-Samuel S, Lythgoe MF, Mookerjee RP, Taylor SA (2020) Liver perfusion MRI in a rodent model of cirrhosis: Agreement with bulk-flow phase-contrast MRI and noninvasive evaluation of inflammation in chronic liver disease using flow-sensitive alternating inversion recovery arterial spin labelling and tissue T1. NMR Biomed 34:e4423
Marzola P, Maggioni F, Vicinanza E, Daprà M, Cavagna FM (1997) Evaluation of the hepatocyte-specific contrast agent gadobenate dimeglumine for MR imaging of acute hepatitis in a rat model. J Magn Reson Imaging 7:147–152
Hazle JD, Narayana PA, Dunsford HA (1991) In vivo NMR, biochemical, and histologic evaluation of alcohol-induced fatty liver in rat and a comparison with CCl4 hepatotoxicity. Magn Reson Med 19:124–135
Hazle JD, Narayana PA, Dunsford HA (1990) Chronic carbon tetrachloride and phospholipase D hepatotoxicity in rat: In vivo 1H magnetic resonance, total lipid analysis, and histology. Magn Reson Med 15:211–228
Ling M, Brauer M (1992) Ethanol-induced fatty liver in the rat examined by in vivo 1H chemical shift selective magnetic resonance imaging and localized spectroscopic methods. Magn Reson Imaging 10:663–677
Herfkens R, Davis P, Crooks L, Kaufman L, Price D, Miller T, Margulis AR, Watts J, Hoenninger J, Arakawa M, McRee R (1981) Nuclear magnetic resonance imaging of the abnormal live rat and correlations with tissue characteristics. Radiology 141:211–218
Davis PL, Kaufman L, Crooks LE, Miller TR (1981) Detectability of hepatomas in rat livers by nuclear magnetic resonance imaging. Invest Radiol 16:354–359
Hoy AM, McDonald N, Lennen RJ, Milanesi M, Herlihy AH, Kendall TJ, Mungall W, Gyngell M, Banerjee R, Janiczek RL, Murphy PS, Jansen MA, Fallowfield JA (2018) Non-invasive assessment of liver disease in rats using multiparametric magnetic resonance imaging: a feasibility study. Biol Open 7:bio033910
Zhou IY, Jordan VC, Rotile NJ, Akam E, Krishnan S, Arora G, Krishnan H, Slattery H, Warner N, Mercaldo N, Farrar CT, Wellen J, Martinez R, Schlerman F, Tanabe KK, Fuchs BC, Caravan P (2020) Advanced MRI of liver fibrosis and treatment response in a rat model of nonalcoholic steatohepatitis. Radiology 296:67–75
Li J, Liu H, Zhang C, Yang S, Wang Y, Chen W, Li X, Wang D (2020) Native T1 mapping compared to ultrasound elastography for staging and monitoring liver fibrosis: an animal study of repeatability, reproducibility, and accuracy. Eur Radiol 30:337–345
Gao Y, Erokwu BO, Desantis DA, Croniger CM, Schur RM, Lu L, Mariappuram J, Dell KM, Flask CA (2016) Initial evaluation of hepatic T1 relaxation time as an imaging marker of liver disease associated with autosomal recessive polycystic kidney disease (ARPKD). NMR Biomed 29:84–89
Gambarota G, Veltien A, Van Laarhoven H, Philippens M, Jonker A, Mook OR, Frederiks WM, Heerschap A (2004) Measurements of T1 and T2 relaxation times of colon cancer metastases in rat liver at 7 T. Magn Reson Mater Physics, Biol Med 17:281–287
Fan YD, Vanzieleghem B, Achten E, De Deene Y, Defreyne L, Praet M, Van Huysse J, Kunnen M, De Hemptinne B (2001) T1 relaxation times for viability evaluation of the engrafted and the native liver in a rat model of heterotopic auxiliary liver transplantation: A pilot study. NMR Biomed 14:350–359
Nakakoshi T, Kajiyama M, Fujita N, Jong-Hon K, Takeichi N, Miyasaka K (1996) Quantitative analyses of correlations of signal intensity on T1-weighted images and T1 relaxation time with copper concentration in the rat liver. Acad Radiol 3:36–39
Chamuleau RAFM, De Nie JHNCI, Moerland MA, Van der Lende OR, Smidt J (1988) Is the magnetic resonance imaging proton spin-lattice relaxation time a reliable noninvasive parameter of developing liver fibrosis? Hepatology 8:217–221
Ganesh T, Estrada M, Yeger H, Duffin J, Margaret Cheng HL (2017) A non-invasive magnetic resonance imaging approach for assessment of real-time microcirculation dynamics. Sci Rep 7:7468
Sheng RF, Wang HQ, Yang L, Jin KP, Xie YH, Fu CX, Zeng MS (2017) Assessment of liver fibrosis using T1 mapping on Gd-EOB-DTPA-enhanced magnetic resonance. Dig Liver Dis 49:789–795
Soares AF, Lei H (2018) Non-invasive diagnosis and metabolic consequences of congenital portosystemic shunts in C57BL/6 J mice. NMR Biomed 31:e3873
Steudel A, Traber F, Krahe T, Schiffmann O, Harder T (1990) Qualitatskontrolle der quantitativen mr-tomographie: in-vitro und in-vivo-uberprufung von relaxationszeitmessungen. RoFo Fortschritte auf dem Gebiete der Rontgenstrahlen und der Neuen Bildgeb Verfahren 152:673–676
Bachtiar V, Kelly MD, Wilman HR, Jacobs J, Newbould R, Kelly CJ, Gyngell ML, Groves KE, McKay A, Herlihy AH, Fernandes CC, Halberstadt M, Maguire M, Jayaratne N, Linden S, Neubauer S, Banerjee R (2019) Repeatability and reproducibility of multiparametric magnetic resonance imaging of the liver. PLoS ONE 14:e0214921
Moher D, Liberati A, Tetzlaff J, Altman DG, Altman D, Antes G, Atkins D, Barbour V, Barrowman N, Berlin JA, Clark J, Clarke M, Cook D, D’Amico R, Deeks JJ, Devereaux PJ, Dickersin K, Egger M, Ernst E, Gøtzsche PC, Grimshaw J, Guyatt G, Higgins J, Ioannidis JPA, Kleijnen J, Lang T, Magrini N, McNamee D, Moja L, Mulrow C, Napoli M, Oxman A, Pham B, Rennie D, Sampson M, Schulz KF, Shekelle PG, Tovey D, Tugwell P (2009) Preferred reporting items for systematic reviews and meta-analyses: The PRISMA statement. PLoS Med 6:e1000097
Haimerl M, Probst U, Poelsterl S, Fellner C, Nickel D, Weigand K, Brunner SM, Zeman F, Stroszczynski C, Wiggermann P (2018) Evaluation of two-point Dixon water-fat separation for liver specific contrast-enhanced assessment of liver maximum capacity. Sci Rep 8:13863
Axel L (1984) Blood flow effects in magnetic resonance imaging. Am J Roentgenol 143:1157–1166
Acknowledgements
The research leading to these results received funding from the Innovative Medicines Initiatives 2 Joint Undertaking under grant agreement No 116106 (IB4SD-TRISTAN). This Joint Undertaking receives support from the European Union’s Horizon 2020 research and innovation programme and EFPIA.
Author information
Authors and Affiliations
Contributions
Waterton, JC. Study conception and design, acquisition of data, analysis and interpretation of data, drafting of manuscript and critical revision.
Corresponding author
Ethics declarations
Conflict of interest
John Waterton holds stock in Quantitative Imaging Ltd and is a Director of, and has received compensation from, Bioxydyn Ltd, a for-profit company engaged in the discovery and development of MR biomarkers and the provision of imaging biomarker services.
Research involving human and animal participants
Not applicable, as this is a survey of previously published research.
Informed consent
Not applicable, as this is a survey of previously published research.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Waterton, J.C. Survey of water proton longitudinal relaxation in liver in vivo. Magn Reson Mater Phy 34, 779–789 (2021). https://doi.org/10.1007/s10334-021-00928-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10334-021-00928-x