Abstract
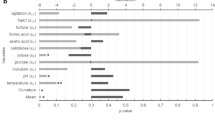
Flavins in the form of flavin mononucleotide (FMN) and flavin adenine dinucleotide (FAD) play an important role in metabolism as cofactors for oxidoreductases and other enzymes. Flavin nucleotides have applications in the food industry and medicine; FAD supplements have been efficiently used for treatment of some inheritable diseases. FAD is produced biotechnologically; however, this compound is much more expensive than riboflavin. Flavinogenic yeast Candida famata synthesizes FAD from FMN and ATP in the reaction catalyzed by FAD synthetase, a product of the FAD1 gene. Expression of FAD1 from the strong constitutive promoter TEF1 resulted in 7- to 15-fold increase in FAD synthetase activity, FAD overproduction, and secretion to the culture medium. The effectiveness of FAD production under different growth conditions by one of these recombinant strains, C. famata T-FD-FM 27, was evaluated. First, the two-level Plackett–Burman design was performed to screen medium components that significantly influence FAD production. Second, central composite design was adopted to investigate the optimum value of the selected factors for achieving maximum FAD yield. FAD production varied most significantly in response to concentrations of adenine, KH2PO4, glycine, and (NH4)2SO4. Implementation of these optimization strategies resulted in 65-fold increase in FAD production when compared to the non-optimized control conditions. Recombinant strain that has been cultivated for 40 h under optimized conditions achieved a FAD accumulation of 451 mg/l. So, for the first time yeast strains overproducing FAD were obtained, and the growth media composition for maximum production of this nucleotide was designed.






Similar content being viewed by others
References
Abbas C, Sibirny A (2011) Genetic control of biosynthesis and transport of riboflavin and flavin nucleotides and construction of robust biotechnological producers. Microbiol Mol Biol Rev 75:321–360. doi:10.1128/MMBR.00030-10
Brooke AG, Dijkhuizen L, Harder W (1986) Regulation of flavin biosynthesis in the methylotrophic yeast Hansenula polymorpha. Arch Microbiol 145(2):162–170
Cao G, Ren N, Wang A, Guo W, Yao J, Feng Y, Zhao Q (2010) Statistical optimization of culture condition for enhanced hydrogen production by Thermoanaerobacterium thermo-saccharolyticum W16. Bioresour Technol 101(6):2053–2058. doi:10.1016/j.biortech.2009.11.031
De Colibus L, Mattevi A (2006) New frontiers in structural flavoenzymology. Curr Opin Struct Biol 16(6):722–728. doi:10.1016/j.sbi.2006.10.003
Dmytruk KV, Voronovsky AY, Sibirny AA (2006) Insertion mutagenesis of the yeast Candida famata (Debaryomyces hansenii) by random integration of linear DNA fragments. Curr Genet 50(3):183–191. doi:10.1007/s00294-006-0083-0
Dmytruk KV, Yatsyshyn VY, Sybirna NO, Fedorovych DV, Sibirny AA (2011) Metabolic engineering and classic selection of the yeast Candida famata (Candida flareri) for construction of strains with enhanced riboflavin production. Metab Eng 13(1):82–88. doi:10.1016/j.ymben.2010.10.005
Exinger F, Lacroute F (1992) 6-Azauracil inhibition of GTP biosynthesis in Saccharomyces cerevisiae. Curr Genet 22(1):9–11. doi:10.1007/BF00351735
Faeder EJ, Siegel LM (1973) A rapid micromethod for determination of FMN and FAD in mixtures. Anal Biochem 53(1):332–336. doi:10.1016/0003-2697(73)90442-9
Frago S, Velázquez-Campoy A, Medina M (2009) The puzzle of ligand binding to Corynebacterium ammoniagenes FAD synthetase. J Biol Chem 284(11):6610–6619. doi:10.1074/jbc.M808142200
Gonzalez-Cabo P, Ros S, Palau F (2010) Flavin adenine dinucleotide rescues the phenotype of frataxin deficiency. PLoS One 5(1):e8872. doi:10.1371/journal.pone.0008872
Haaland PD (1990) Experimental design in biotechnology. Elsevier, New York
Hagihara T, Fujio T, Aisaka K (1995) Cloning of FAD synthetase gene from Corynebacterium ammoniagenes and its application to FAD and FMN production. Appl Microbiol Biotechnol 42(5):724–729
Huang YF, Liu SY, Yen CL, Yang PW, Shieh CC (2009) Thapsigargin and flavin adenine dinucleotide ex vivo treatment rescues trafficking-defective gp91phox in chronic granulomatous disease leukocytes. Free Radic Biol Med 47(7):932–940. doi:10.1016/j.freeradbiomed.2009.06.037
Ishchuk OP, Dmytruk KV, Rohulya OV, Voronovsky AY, Abbas CA, Sibirny AA (2008) Development of a promoter assay system for the flavinogenic yeast Candida famata based on the Kluyveromyces lactis β-galactosidase LAC4 reporter gene. Enzym Microb Technol 42(3):208–215. doi:10.1016/j.enzmictec.2007.09.008
Ishchuk OP, IatsyshynViu, Dmytruk KV, Voronovs’kyĭ Aia, Fedorovych DV, Sybirnyĭ AA (2006) Construction of the flavinogenic yeast Candida famata strains with high riboflavin kinase activity using gene engineering. Ukr Biokhim Zh 78(5):63–69 (Article in Ukrainian)
Kalingan AE, Liao C-M (2002) Influence of type and concentration of flavinogenic factors on production of riboflavin by Eremothecium ashbyii NRRL 1363. Bioresour Technol 82(3):219–224. doi:10.1016/S0960-8524(01)00194-8
Kashchenko VE, Shavlovskiĭ GM (1976) Purification and properties of the riboflavin kinase of the yeast Pichia guilliermondii. Biokhimiia 41(2):376–383 Article in Russian
Kitatsuji K, Ishino S, Teshiba S, Arimoto M (1996) Method of producing flavine nucleotides. US Patent 5,514,574
Lowry OH, Rosebrough NJ, Farr AL, Randall RJ (1951) Protein measurement with the Folin phenol reagent. J Biol Chem 193(1):265–275
Massey V (2000) The chemical and biological versatility of riboflavin. Biochem Soc Trans 28:283–296. doi:10.1074/jbc.M103114200
Masuda T (1955) Application of chromatography. XXVIII. On the formation of FAD in the culture of Eremothecium ashbyii. Pharm Bull 3:434–440
Montgomery DC (1997) Design and analysis of experiments. Wiley, New York
Nagatsu T, Nagatsu-Ishibashi I, Yagi K (1963) Biosynthesis of C14-labelled flavin adenine dinucleotide by Eremothecium ashbyii. J Biochem 54:152–155 (Tokyo)
Nguyen H-V, Gaillardin C, Neuveglise C (2009) Differentiation of Debaryomyces hansenii and Candida famata by rRNA gene intergenic spacer fingerprinting and reassessment of phylogenetic relationships among D. hansenii, C. famata, D. fabryi, C. flareri (= D. subglobosus) and D. prosopidis: description of D. vietnamensis sp. nov. Closely related to D. nepalensis. FEMS Yeast Res 9(4):641–662. doi:10.1111/j.1567-1364.2009.00510.x
Plackett RL, Burman JP (1946) The design of optimum multifactorial experiments. Biometrika 33(4):305–325
Pujari V, Chandra TS (2000) Statistical optimization of medium components for enhanced riboflavin production by a UV-mutant of Eremothecium ashbyii. Process Biochem 36(1–2):31–37. doi:10.1016/S0032-9592(00)00173-4
Rai SK, Mukherjee AK (2010) Statistical optimization of production, purification and industrial application of a laundry detergent and organic solvent-stable subtilisin-like serine protease (Alzwiprase) from Bacillus subtilis DM-04. Biochem Eng J 48:173–180. doi:10.1016/j.bej.2009.09.007
Sakai T, Watanabe T, Chibata I (1973) Selection of microorganism producing flavin-adenine dinucleotide from FMN and adenine (AMP) and production of flavin-adenine dinucleotide by Sarcina lutea. Agric Biol Chem 37:849–856
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory, Cold Spring Harbor
Sandoval FJ, Zhang Y, Roje S (2008) Flavin nucleotide metabolism in plants: monofunctional enzymes synthesize FAD in plastids. J Biol Chem 283(45):30890–30900. doi:10.1074/jbc.M803416200
Shavlovskii GM, Fedorovich DV (1977) The activity of enzymes involved in synthesis and hydrolysis of flavin adenine dinucleotide in Pichia guilliermondii: studies at different levels of flavinogenesis. Mikrobiologiia 46:904–911 (In Russian)
Shimizu S (2001) Vitamins and related compounds: microbial production. In: Rehm H (ed) Biotechnology, vol 10., Wiley-VCHWeinheim, Germany, pp 322–326
Shimizu S, Yamane K, Tani Y, Yamada H (1983) Enzymatic synthesis of flavin adenine dinucleotide. Appl Biochem Biotechnol 8:237–247
Voronovsky AA, Abbas CA, Fayura LR, Kshanovska BV, Dmytruk KV, Sybirna KA, Sibirny AA (2002) Development of a transformation system for the flavinogenic yeast Candida famata. FEMS Yeast Res 2:381–388. doi:10.1016/S1567-1356(02)00112-5
Watanabe T, Uchida T, Kato J, Chibata I (1974) Production of flavine-adenine dinucleotide from riboflavine by a mutant of Sarcina lutea. Arch Microbiol 27:531–536
Wu M, Repetto B, Glerum DM, Tzagoloff A (1995) Cloning and characterization of FAD1, the structural gene for flavin adenine dinucleotide synthetase of Saccharomyces cerevisiae. Mol Cell Biol 15:264–271
Wu QL, Chen T, Gan Y, Chen X, Zhao XM (2007) Optimization of riboflavin production by recombinant Bacillus subtilis RH44 using statistical designs. Appl Microbiol Biotechnol 76:783–794. doi:10.1007/s00253-007-1049-y
Yagi K, Matsuoka Y, Kuyama S, Tada M (1956) Preparation of flavin adenine dinucleotide from Eremothecium ashbyii. J Biochem 43:93–100
Yatsyshyn VY, Fedorovych DV, Sibirny AA (2010) Medium optimization for production of flavin mononucleotide by the recombinant strain of the yeast Candida famata using statistical designs. Biochem Eng J 49(1):52–60. doi:10.1016/j.bej.2009.11.010
Yatsyshyn VY, Fedorovych DV, Sibirny AA (2009) The microbial synthesis of flavin nucleotides: a review. Appl Biochem Microbiol 45(2):115–124. doi:10.1134/S000368380902001X
Yatsyshyn VY, Ishchuk OP, Voronovsky AY, Fedorovych DV, Sibirny AA (2009) Production of flavin mononucleotide by metabolically engineered yeast Candida famata. Metab Eng 11:163–167. doi:10.1016/j.ymben.2009.01.004
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Yatsyshyn, V.Y., Fedorovych, D.V. & Sibirny, A.A. Metabolic and bioprocess engineering of the yeast Candida famata for FAD production. J Ind Microbiol Biotechnol 41, 823–835 (2014). https://doi.org/10.1007/s10295-014-1422-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10295-014-1422-7