Abstract
The overarching goal of this study is to predict the risk of developing oral squamous cell carcinoma (OSCC) in Fanconi anemia (FA) patients. We have compared the microRNA (miRNA, miR) expression levels in saliva samples from FA patients (n = 50) who are at a low-moderate and/or high risk of developing OSCC to saliva samples from healthy controls (n = 16). The miRNA expression levels in saliva samples were quantified using qPCR. We observed that miR-744, miR-150-5P, and miR-146B-5P had the best discriminatory capacity between FA patients and controls, with an area under the curve (AUC) of 94.0%, 92.9% and 85.3%, respectively. Our data suggest that miR-1, miR-146B-5P, miR-150-5P, miR-155-5P, and miR-744 could be used as panel to predict the risk of developing OSCC in FA patients, with a 89.3% sensitivity and a 68.2% specificity (AUC = 81.5%). Our preliminary data support the notion that the expression levels of salivary miRNAs have the potential to predict the risk of developing OSCC in FA patients and in the future may reduce deaths associated with OSCC.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Fanconi anemia (FA) is a rare inherited genetic condition characterized by physical abnormalities, bone marrow failure syndrome and a very high risk of developing cancer [1]. It is usually inherited as an autosomal recessive genetic disorder, but X-linked inheritance has also been reported. It is caused by the loss of function of at least one of 23 genes involved with the FA/BRCA pathway [1]. This pathway is involved in the repair of double-stranded DNA breaks and homologous recombination, as well as repair of interstrand DNA cross-linking (ICL) caused by exogenous agents [2, 3]. If the ICLs are left unrepaired, they will promote genotoxic stress, genomic instability and tumorigenesis [4]. Approximately, 20% of FA patients have some types of malignancy, mainly leukemias [5] or oral squamous cell carcinoma (OSCC) [6, 7] as these are the two most common types of solid tumor in FA patients.
MicroRNAs (miRNAs) are non-coding, small RNAs that regulate the gene expression by modulating many cellular processes, such as controlling and modulating immunity, cell proliferation, promoting survival and growth of malignant cells, and cancer metastasis [8, 9]. A single miRNA can bind to many mRNA sequences and each mRNA sequence is a target of many miRNAs. Compelling evidence has demonstrated that the expression of miRNA is commonly dysregulated in multiple disease states, including FA [10,11,12]. However, there are few studies analyzing miRNAs expression levels in tumor tissues, plasma or serum of FA patients [13, 14]. For instance, miR-1, miR-146B-5P and miR-150-5P have been indicated as novel plasma biomarkers for aplastic anemia, a condition often found in FA patients [15]. Furthermore, low expression levels of miR-155-5P and miR-181C have been reported in FA patients [13, 14].
Salivary analysis has been proposed as an alternative method of choice to blood based analysis for unravelling disease specific biomarkers [16,17,18,19]. Saliva is an ideal diagnostic medium for biomarker detection studies in FA patients because of its non-invasiveness and ease of collection so multiple samples can be collected from an individual cost effectively [20,21,22]. A further advantage of this biofluid is its proximity to the oral cavity, as this region is one of the most common sites for the development of OSCC [23,24,25]. To the best of our knowledge, there is thus far no study has attempted to investigate the relationship between salivary FA-associated miRNAs and OSCC risk factors in FA patients. The overarching objective of this study was to investigate whether FA-associated salivary miRNAs signature could be used as a biomarker in FA patients to predict the risk of developing OSCC.
Materials and methods
Participants
This study was approved by the Ethics Review Board of the Complexo Hospital de Clínicas da Universidade Federal do Paraná (approval number 2.426.777) as well as by the University of Queensland Medical Ethical Institutional Board [HREC No: 2014000679 and 2014000862]; Queensland University of Technology [HREC No: 1400000617 and 1400000641] and by the Princess Alexandra Hospital Ethics Review Board [HREC Number: HREC/12/QPAH/381]. All participants provided written informed consent before saliva sample collection, following the Declaration of Helsinki. FA patients who attended the Bone Marrow Transplantation Unit, at the Hospital de Clínicas from November 2017 to June 2018, participated in the study. The participants were classified into two groups: low-moderate and high risk of developing OSCC, based on the criteria proposed by Furquim et al. [17]. Briefly, high risk group consisted of FA patients who underwent hematopoietic stem-cell transplantation (HSCT) > 5 years before the most recent examination or ≥ 18 years old and have developed oral potentially malignant disorders (OPMD). The low-moderate risk group consisted of patients over 18 years of age without OPMD, or under 18 years of age with OPMD, or without OPMD but HSCT within the last 5 years. We also recruited a control group, and the inclusion criteria included no self-reported illness or history of any oral or systemic disease.
Saliva sample collection
Participants were asked to refrain from eating, drinking or brushing their teeth for at least one hour prior to saliva sample collection. All samples were collected in the morning between 7:30 am to 11:30 am to minimize diurinal variations [24, 26,27,28]. In brief, participants were asked to refrain from eating 1-h prior to saliva collection. Participants were asked to tilt their heads down and pool saliva in the mouth for 2–3 min and expectorate (2–5 mL of saliva) into a 50 mL Falcon tube kept on ice. Samples were transported on dry-ice to the laboratory for further processing. Saliva collection, preprocessing and storage was conducted using previously published protocols [24, 26,27,28]. After collection, all saliva samples were immediately stored on dry ice and transferred to the laboratory for down stream processing.
miRNA extraction and cDNA synthesis
In order to avoid batch to batch variations, we have isolated miRNA from saliva samples at one time point. miRNA was isolated and enriched by QIAzol lysis reagent (Qiagen) in combination with the NucleoSpin miRNA kit (Macherey–Nagel, Düren, Germany) as previously described [29]. Briefly, 800μL QIAzol lysis reagent was added to 200μL saliva. Then 200μL chloroform was added and the mixture was then centrifuged at 10,000 × g for 10 min at 4 °C. The upper aqueous layer was transferred into a new microcentrifuge tube containing 200 μL ethanol and subsequently transferred into a Nucleospin column for miRNA enrichment as per manufacturer protocol. The quantity of the isolated miRNA was determined using both Qubit® 3.0 Fluorometer (Thermo Fisher Scientific, Waltham, MA, USA) and Nanodrop ND-1,000 spectrophotometer (Thermo Fisher Scientific). OD 260:280 ratios ≥ 1.8 were accepted as pure.cDNA synthesis was carried out via a miScript II RT Kit (Qiagen) according to the manufacturers’ instructions. Briefly, miRNAs were polyadenylated with polymerase A and transcribed into cDNA using oligo-dT primers (in parallel in the same tube). The oligo-dT had a 3′degenerate anchor and a universal tag sequence at the 5′end allowing amplification of mature miRNA through a real-time PCR step (60 min at 37ºC followed by heat inactivation for 5 min at 95ºC). The polyadenylation and universal tag primer ensure that genomic DNA is not amplified. Therefore, DNAase treatment was omitted from the miRNA isolation protocol.
miRNA qPCR analysis
The qRT-PCR for miRNA quantitation was performed using target-specific miScript Primer Assays (miR-1, Cat. No. MS00008358; miR-146B-5P, Cat. No. MS00003542; miR-150-5P, Cat. No. MS00003577; miR-155-5P, Cat. No. MS00031486; miR-181C-5P, Cat. No. MS00008841; miR-744 Cat. No. MS00010549) and the miScript SYBR Green PCR Kit (Qiagen), according to the manufacturer’s protocol. For the identification of optimal miRNA normalizer in saliva samples a panel of six miScript™ primer PCR assays (SNORD 61, Cat. No. MS00033705; SNORD68, Cat. No. MS00033712; SNORD72, Cat. No. MS00033719; SNORD95, Cat. No. MS00033726; SNORD96A, Cat. No. MS00033733 and miR-489 Cat. No. MS00007700) were used. Briefly, 3 ng cDNA was used as a template across control and FA patients groups for PCR amplification and the thermocycling conditions were as follows: 95 °C for 15 min to activate hot start Taq DNA polymerase and this was followed by 40 cycles of 94 °C for 15 s, 55 °C for 30 s and 70 °C for 30 s and a final melting curve analysis with the following conditions: 95 °C for 15 s, 60 °C for 60 s, and 95 °C for 15 s using QuantStudio™ 7 Flex Real-Time PCR System (Applied Biosystems). A value of 40 was assigned when the threshold cycle (Ct) value was underdetermined and the average Ct value of the reactions in duplicate were determined. The comparative Delta Ct method was adopted and miR-489 was used as a normalizer as published by us previously. In this method, we have subtracted the ct value of target gene form the normalizer (miR 489). If we get a negative delta Ct value, this would mean that the target gene Ct value is higher (lower expression) than the normalizer.
Statistical analysis
Statistical analyses were performed using JMP Pro version 16 (SAS Institute, Cary, NC, USA) and GraphPad Prism version 8.0 (GraphPad Software Inc., La Jolla, CA, USA). miRNA expression levels of miR-1, miR-146B-5P, miR-150-5P, miR-155-5P, miR-181C-5P and miR-744 in saliva samples were compared between two cohorts (controls and FA patients) using t-tests; and comparison of the three cohorts (controls, low-moderate and high risk FA patients) were compared using one way ANOVA, followed by the Tukey–Kramer HSD multiple comparison procedure [30, 31]. P-values less than 0.05 were considered significant. Correlation between the predictors was calculated using Pearson correlations. Lasso penalized multiple logistic regression [32] was used to identify a panel of miRNA markers for distinguishing between FA patients classified as low-moderate risk of OSCC from those classified as high risk. The Lasso penalty parameter was chosen to minimize the Akaike's Information Criterion [33]. Sensitivity and specificity were calculated for the predictive model, with the cutoff chosen to maximize the Youden’s Index [34]. Confidence intervals were calculated using the score method [35]. Receiver operating characteristic (ROC) curves were generated for all logistic regressions, and the area under the curve (AUC) with 95% confidence intervals presented to indicate the predictive ability of the models. The AUC of the ROC is also known as the concordance or c-statistic [36].
Results
Population characteristics
The demographic and clinical characteristics of controls (n = 16) and FA patients (n = 50) are summarized in Table 1. Controls and FA patients had a mean age of 23 years (range 18–31 years) and 20 years (range 7–37 years), respectively. Gender distribution was approximately even in each group. All FA patients were considered as non-smokers and non-drinkers while; the majority of controls were regarded as non/former smokers and non-drinkers. Nearly all FA patients were single (95%) while marital status was not known for controls. Nearly all controls were identified as Caucasian (88%) compared to only 50% of FA patients. All high-risk FA patients underwent HSCT, compared to only 50% of the low-moderate risk FA patients. Graft vs host disease (GVHD) and OPMD were diagnosed in 6 (12%) and 9 (18%) FA patients, respectively.
The stability of the miRNA housekeeping gene in saliva samples collected from FA patients and controls
The distributions of the Ct values of the five reference genes from the SNORD family and miR-489 in saliva samples of controls and FA patients are shown in Supplementary Fig. 1. All reference genes from the SNORD family showed statistical differences between controls and FA patients, but the salivary expression levels of miR-489 were quite similar between controls and FA patients. Furthermore, miR-489 showed the highest level of relative expression (mean Ct = 22.56) with the lowest standard deviation (SD) values of 1.11 compared to the other reference genes from the SNORD family (Supplementary Table 1). Previous findings are consistent with our study showing that miR-489 as the best normalizer for miRNA research when using the body fluids [37].
The expression levels of FA-associated miRNAs in FA patients
Dysregulation of miRNAs has been associated with a number of diseases including FA [13], however their expression levels in saliva samples collected from FA patients still remains to be elucidated. To address this question, we first investigated the expression levels of six literature documented FA-associated miRNAs (miR-1, miR-146B-5P, miR-150-5P, miR-155-5P, miR-181C-5P and miR-744) in saliva samples collected from controls and FA patients. All six FA-associated miRNAs were signficantly downregulated (P < 0.05) in saliva samples collected from FA patients compared to controls as shown in Fig. 1A. The diagnostic values of miR‑1, miR-146B-5P, miR-150-5P, miR-155-5P, miR-181C-5P and miR-744 were also investigated using logistic regressions analysis between groups (control vs FA) against each of the miRNAs. Each miRNA was statistically significant, however, the best predictors were miR-744, miR-150-5P, and miR-146B-5P (ROC AUC (95% CI) = 94.0% (85.1%, 97.7%), 92.9% (80.7%, 97.6%), and 85.3% (66.8%, 94.3%), respectively, P < 0.0001 in each case, Supplementary Fig. 2).
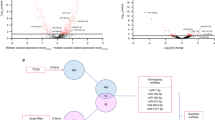
Dysregulation of miRNAs in saliva samples collected from FA patients. A The relative expression (ΔCT) of miR-1, miR-146B-5P, miR-150-5P, miR-155-5P, miR-181C-5P and miR-744 in saliva collected from controls and FA patients. B The six miRNA statistically significantly overlap with genesets from the MsigDB catalog. Data were normalized with miR-489 and are presented as delta Ct values. Statistically significant differences (P < 0.05) between two groups were determined using Mann–Whitney U-test. (P values: * < 0.05, ** < 0.005, *** < 0.001, **** < 0.0001)
We computed the overlap of these six FA-associated miRNAs with the molecular signature database (MSigDB) which contains 31,117 gene sets [38]. We present in Fig. 1B the 19 statistically significant overlapping gene sets, (FDR q-value ≤ 0.05). In addition to the expected gene sets related to gene silencing and post-transcriptional regulation, we also observed that miR-1 and miR-150-5P are involved in mesoderm development and gastrulation, as well as the regulation of endothelial cell differentiation. Importantly, miR-146B-5P and miR-181C-5P are involved in negative regulation of IL-17.
Salivary miRNA expression differences in FA patients at low-moderate vs high risk of developing OSCC
As shown in Fig. 2, the expression levels of miR-146B-5P, miR-155-5P, miR-181C-5P and miR-744 were significantly downregulated (P < 0.05) in saliva samples collected from high risk patients compared to low-moderate risk FA patients. Logistic regressions of OSCC risk levels (low-moderate/high risk) versus individual miRNAs found that miR-146B-5P, miR-155-5P, miR-181C-5P and miR-744 to be significant predictors (P < 0.01, in each case), but with relatively low ROC AUCs (< 75%, Supplementary Fig. 3). Furthermore, as the expression levels of the miRNAs are quite strongly correlated (Supplemetary Fig. 4), we used a lasso penalized logistic regression to identify a set of markers that could serve as a potential biomarker panel to discriminate among controls, low-moderate and high risk groups. The model selected miR-1, miR-146B-5P, miR-150-5P, miR-155-5P, and miR-744 as predictors, yielding an ROC AUC = 81.5% (95% CI 66.1%, 90.9%), sensitivity of 89.3% and specificity of 68.2%, based on using Youden’s J statistic to select the cutoff (Fig. 3).
The relative expression of miR-1, miR-146B-5P, miR-150-5P, miR-155-5P, miR181C-5P and miR-744 in saliva collected from controls, low-moderate and high risk of developing OSCC groups. Data were normalized with miR-489 and are presented as delta Ct values. Statistically significant differences (P < 0.05) among these three groups were determined using ordinary one way ANOVA. (P values: * < 0.05, ** < 0.005, *** < 0.001, **** < 0.0001)
Discussion
FA is a rare and a complex tumor-prone disease defined by an interlaced genotype and phenotype. Accumulating evidence supports the notions that salivary miRNAs expression changes can be used as potential biomarkers for the detection of oral and systemic diseases [8, 10, 39,40,41]. However, there is a dearth of available studies investigating the utility of salivary miRNAs as useful biomarkers to identify FA patients who are likely to develop OSCC. Studies have shown that early detection of OSCC improves five-year survival rates from 20 to 80% [42]. As such, it then becomes important to early identify FA patients who are at low-moderate vs high risk of developing OSCC. For the first time, we demonstrate that by combining miR-1, miR-146B-5p, miR-150-5p, miR-155-5P, and miR-744 as a biomarker panel can predict FA patients who are at a higher risk of developing OSCC (AUC = 81.5%, Sensitivity 89.3% and Specificity 68.2%).
In our study, miR-1 and miR-146B-5P were under expressed in the saliva samples from high-risk FA patient group. This is further supported by previous studies demonstrating that the downregulation of both miRNAs is associated with aggressive OSCC [43, 44]. Previous studies have demonstrated that the dysregulation of miR-1 and miR-146B-5P to be associated with the development of aplastic anemia, a common hematological problem in FA patients [15]. Furthermore, the alterations of miR-1 and miR-146B-5P levels in serum have been linked to chronic inflammation [45, 46]. While miR-1 targets B-cell lymphoma 2 (BCL2), miR-146B-5P targets and represses TNF-receptor associated factor 6 (TRAF6) and Interleukin 1 receptor-associated kinase 1 (IRAK1) and has shown to be involved in feedback regulation of the innate immune response.miR-150-5P is mainly expressed in the megakaryocytic lineage, driving megakaryocyte-erythrocyte progenitor differentiation towards megakaryocytes [47]. Previous research has reported that miR-150-5P was expressed at a higher level in plasma of patients with aplastic anemia compared to controls [15]. Conversely, we observed a lower expression level of miR-150-5P in the saliva samples of high-risk FA patients compared to controls. In fact, miR-150-5P was reported to be silenced in OSCC tissues, further supporting the potential of miR-150-5P as a biomarker to predict the FA patients who are at a greater risk of developing OSCC [48].
We found significantly lower levels of miR-155-5P in saliva samples from high-risk FA patients. Similarly, a study by Degan et al. reported lower expression of miR-155-5P in blood samples of FA patients compared to healthy controls [13]. Furthermore, the under expression of miR-155-5P was associated with an increased risk of OSCC metastasis development [49]. Earlier studies have identified miR-155-5P as a critical regulator of inflammation as well as regulating the innate and adaptive immune responses [50]. Several studies have demonstrated that miR-155-5P is required for normal functions of B and T lymphocytes and is up-regulated upon B and T cell activation [51, 52]. Since FA is characterized by chromosomal breakage, miR-155-5P downregulation is obvious.
We found in our study that miR-181C-5P levels to be significantly lower in high-risk FA group compared to both control and low-moderate risk FA groups. Similarly, a study has shown significantly lower levels of miR-181C in saliva samples collected from patients with oral low-grade dysplasia (LGD) who are at a higher risk of developing OSCC compared to those patients with non-progressive oral LGD [53]. This then suggests that miR-181C levels in saliva samples may be used as a non-invasive biomarker to predict OSCC in OPMD patients. It is also reported that miR-181C is downregulated in lymphoblastoid cell lines and fresh peripheral blood cells from FA patients. Also, miR-181C has shown to play an important part in regulating the FA hematopoietic progenitors by targeting the TNFα [14].
Several studies have reported the role of miR-744 in regulating the cellular processes which may eventually contribute to the development of human diseases [54, 55]. Moreover, miR-744 could function as a tumor suppressor or oncogene and the role vary based on the cancer types. In laryngeal squamous cell carcinoma, miR-744 functions as an oncogene and has been demonstrated to be associated with cancer metastasis [56], while miR-744 expression is downregulated in non-small lung cancer [57]. However, little attention has been paid to investigate the role of miR-744 in oral carcinogenesis. We found significantly lower expression levels of miR-744 in the saliva samples from high vs low-moderate risk FA patients and in saliva from controls. Strikingly, one of the potential targets of miR-744 is FANCG [58], which is one of the most likely affected genes in FA. This is a pilot study with some limitations: the modest sample size and a single site recruitment for controls and FA patients, respectively. It would have been appropriate to also include an OSCC group and investigate the miRNA expression changes of established biomarkers of OSCC such as miRNA-21, miR-9 [59]. However, because of the age at with OSCC occurs (55 years), it was not possible to recruit age matched controls and FA patients. Future studies using a large group of FA patients from diverse geographic regions are warranted to investigate the potential clinical utility of salivary miRNAs biomarkers panel in FA patients.
Conclusions
In conclusion, all of the miRNAs analyzed in this study are downregulated in saliva samples of FA patients when compared with controls. Additionally, miR-1, miR-146B-5P, miR-150-5P, miR-155-5P, and miR-744 have the potential to distinguish high and low-moderate risk of developing OSCC. Our data is promising to predict the risk of developing OSCC in FA patients, using non-invasive miRNA based biomarkers. However, future trial should focus on addressing the current short-comings.
Availability of data and materials
All datasets generated for this study are included in the article.
References
Amenábar JM, Torres-Pereira CC, Tang KD, Punyadeera C. Two enemies, one fight: An update of oral cancer in patients with Fanconi anemia. ACS J. 2019;125(22):3936–46. https://doi.org/10.1002/cncr.32435.
Bhattacharjee S, Nandi S. DNA damage response and cancer therapeutics through the lens of the Fanconi Anemia DNA repair pathway. Cell Commun Signal. 2017;15(1):41. https://doi.org/10.1186/s12964-017-0195-9.
Ceccaldi R, Sarangi P, D’Andrea AD. The Fanconi anaemia pathway: new players and new functions. Nat Rev Mol Cell Biol. 2016;17(6):337–49. https://doi.org/10.1038/nrm.2016.48.
Sumpter R Jr, Levine B. Emerging functions of the Fanconi anemia pathway at a glance. J Cell Sci. 2017;130(16):2657–62. https://doi.org/10.1242/jcs.204909.
Alter BP. Fanconi anemia and the development of leukemia. Best Pract Res Clin Haematol. 2014;27(3–4):214–21. https://doi.org/10.1016/j.beha.2014.10.002.
Alter BP. Inherited bone marrow failure syndromes: considerations pre- and posttransplant. Blood. 2017;130(21):2257–64. https://doi.org/10.1182/blood-2017-05-781799.
Kutler DI, Auerbach AD, Satagopan J, Giampietro PF, Batish SD, Huvos AG, Singh B. High incidence of head and neck squamous cell carcinoma in patients with Fanconi anemia. Arch Otolaryngol Head Neck Surg. 2003;129(1):106–12.
Satapathy S, Batra J, Jeet V, Thompson EW, Punyadeera C. MicroRNAs in HPV associated cancers: small players with big consequences. Expert Rev Mol Diagn. 2017;17(7):711–22. https://doi.org/10.1080/14737159.2017.1339603.
Tomasetti M, Santarelli L, Neuzil J, Dong L. MicroRNA regulation of cancer metabolism: role in tumour suppression. Mitochondrion. 2014;19:29–38. https://doi.org/10.1016/j.mito.2014.06.004.
Nagadia R, Pandit P, Coman WB, Cooper-White J, Punyadeera C. miRNAs in head and neck cancer revisited. Cell Oncol (Dordr). 2013;36(1):1–7. https://doi.org/10.1007/s13402-012-0122-4.
Salazar C, Calvopiña D, Punyadeera C. miRNAs in human papilloma virus associated oral and oropharyngeal squamous cell carcinomas. Expert Rev Mol Diagn. 2014;14(8):1033–40. https://doi.org/10.1586/14737159.2014.960519.
Wan Y, Vagenas D, Salazar C, Kenny L, Perry C, Calvopiña D, Punyadeera C. Salivary miRNA panel to detect HPV-positive and HPV-negative head and neck cancer patients. Oncotarget. 2017;8(59):99990–100001. https://doi.org/10.18632/oncotarget.21725.
Degan P, Cappelli E, Longobardi M, Pulliero A, Cuccarolo P, Dufour C, Izzotti A. A global MicroRNA profile in fanconi anemia: a pilot study. Metab Syndr Relat Disord. 2019;17(1):53–9. https://doi.org/10.1089/met.2018.0085.
Rio P, Agirre X, Garate L, Banos R, Alvarez L, San Jose-Eneriz E, Bueren JA. Down-regulated expression of hsa-miR-181c in Fanconi anemia patients: implications in TNFalpha regulation and proliferation of hematopoietic progenitor cells. Blood. 2012;119(13):3042–9. https://doi.org/10.1182/blood-2011-01-331017.
Hosokawa K, Kajigaya S, Feng X, Desierto MJ, Fernandez Ibanez MDP, Rios O, Young NS. A plasma microRNA signature as a biomarker for acquired aplastic anemia. Haematologica. 2017;102(1):69. https://doi.org/10.3324/haematol.2016.151076.
Amenabar JM, Torres-Pereira CC, Tang KD, Punyadeera C. Two enemies, one fight: an update of oral cancer in patients with Fanconi anemia. Cancer. 2019. https://doi.org/10.1002/cncr.32435.
Furquim CP, Soares GM, Ribeiro LL, Azcarate-Peril MA, Butz N, Roach J, Teles FR. The salivary microbiome and oral cancer risk: a pilot study in fanconi anemia. J Dent Res. 2017;96(3):292–9. https://doi.org/10.1177/0022034516678169.
Lyko K, Bonfim C, Benelli EM, Torres-Pereira CC, Amenabar JM. Salivary detection of periodontopathic bacteria in Fanconi’s anemia patients. Anaerobe. 2013;24:32–5. https://doi.org/10.1016/j.anaerobe.2013.09.005.
Mafessoni TP, Mazur CE, Amenabar JM. Salivary lactate dehydrogenase (LDH) as a tool for early diagnosis of oral cancer in individuals with Fanconi anemia. Med Hypotheses. 2018;119:29–31. https://doi.org/10.1016/j.mehy.2018.07.021.
Tang KD, Menezes L, Baeten K, Walsh LJ, Whitfield BCS, Batstone MD, Punyadeera C. Oral HPV16 prevalence in oral potentially malignant disorders and oral cavity cancers. Biomolecules. 2020;10(2):223. https://doi.org/10.3390/biom10020223.
Tang KD, Vasani S, Taheri T, Walsh LJ, Hughes BGM, Kenny L, Punyadeera C. An occult HPV-Driven oropharyngeal squamous cell carcinoma discovered through a saliva test. Front Oncol. 2020;10:408. https://doi.org/10.3389/fonc.2020.00408.
Zhang X, Wan Y, Cooper-White J, Dimeski G, Atherton J, Punyadeera C. Quantification of D-dimer levels in human saliva. Bioanalysis. 2013;5(18):2249–56. https://doi.org/10.4155/bio.13.190.
Lim Y, Totsika M, Morrison M, Punyadeera C. The saliva microbiome profiles are minimally affected by collection method or DNA extraction protocols. Sci Rep. 2017;7(1):8523. https://doi.org/10.1038/s41598-017-07885-3.
Mohamed R, Campbell JL, Cooper-White J, Dimeski G, Punyadeera C. The impact of saliva collection and processing methods on CRP, IgE, and Myoglobin immunoassays. Clin Transl Med. 2012;1(1):19. https://doi.org/10.1186/2001-1326-1-19.
Topkas E, Keith P, Dimeski G, Cooper-White J, Punyadeera C. Evaluation of saliva collection devices for the analysis of proteins. Clin Chim Acta. 2012;413(13–14):1066–70. https://doi.org/10.1016/j.cca.2012.02.020.
Amadeu JK, Lemes AL, Schussel JL, Amenabar JM. Effect of storage time and temperature on salivary total antioxidant capacity, total oxidant status, and oxidant stress index. Acta Stomatol Croat. 2019;53(2):119–24. https://doi.org/10.15644/asc53/2/3.
Lim Y, Wan Y, Vagenas D, Ovchinnikov DA, Perry CF, Davis MJ, Punyadeera C. Salivary DNA methylation panel to diagnose HPV-positive and HPV-negative head and neck cancers. BMC Cancer. 2016;16(1):749. https://doi.org/10.1186/s12885-016-2785-0.
Pandit P, Cooper-White J, Punyadeera C. High-yield RNA-extraction method for saliva. Clin Chem. 2013;59(7):1118–22. https://doi.org/10.1373/clinchem.2012.197863.
Arunachalam SR, Tang KD, Punyadeera C. Isolation and quantification of MicroRNAs from Human Saliva. Methods Mol Biol. 2019;2054:105–14. https://doi.org/10.1007/978-1-4939-9769-5_6.
Hayter AJ. A proof of the conjecture that the Tukey-Kramer multiple comparisons procedure is conservative. Annal Stat. 1984;12(1):61–75.
Kramer CY. Extension of multiple range tests to group means with unequal numbers of replications. Biometrics. 1956;12(3):307–10. https://doi.org/10.2307/3001469.
Tibshirani R. Regression Shrinkage and Selection via the Lasso. J R Stat Soc Series B (Methodological). 1996;58(1):267–88.
Akaike H. A new look at the statistical model identification. IEEE Trans Autom Control. 1974;19(6):716–23. https://doi.org/10.1109/TAC.1974.1100705.
Youden WJ. Index for rating diagnostic tests. Cancer. 1950;3(1):32–5. https://doi.org/10.1002/1097-0142(1950)3:1%3c32::aid-cncr2820030106%3e3.0.co;2-3.
Agresti A, Coull BA. Approximate is better than “Exact” for interval estimation of binomial proportions. Am Stat. 1998;52(2):119–26. https://doi.org/10.1080/00031305.1998.10480550.
Moons KG, Altman DG, Reitsma JB, Ioannidis JP, Macaskill P, Steyerberg EW, Collins GS. Transparent Reporting of a multivariable prediction model for Individual Prognosis or Diagnosis (TRIPOD): explanation and elaboration. Ann Intern Med. 2015;162(1):W1-73. https://doi.org/10.7326/M14-0698.
Ramachandran K, Saikumar J, Bijol V, Koyner JL, Qian J, Betensky RA, Vaidya VS. Human miRNome profiling identifies microRNAs differentially present in the urine after kidney injury. Clin Chem. 2013;59(12):1742–52. https://doi.org/10.1373/clinchem.2013.210245.
Liberzon A, Subramanian A, Pinchback R, Thorvaldsdottir H, Tamayo P, Mesirov JP. Molecular signatures database (MSigDB) 3.0. Bioinformatics. 2011;27(12):1739–40. https://doi.org/10.1093/bioinformatics/btr260.
Foo JY, Wan Y, Schulz BL, Kostner K, Atherton J, Cooper-White J, Punyadeera C. Circulating fragments of N-terminal pro-B-type natriuretic peptides in plasma of heart failure patients. Clin Chem. 2013;59(10):1523–31. https://doi.org/10.1373/clinchem.2012.200204.
Punyadeera C, Slowey PD. Chapter 22 - Saliva as an emerging biofluid for clinical diagnosis and applications of MEMS/NEMS in salivary diagnostics. In: Subramani K, Ahmed W, editors. Nanobiomaterials in Clinical Dentistry. 2nd ed. Elsevier; 2019. p. 543–65.
Sun CX, Bennett N, Tran P, Tang KD, Lim Y, Frazer I, Punyadeera C. A pilot study into the association between oral health status and human papillomavirus-16 infection. Diagnostics (Basel). 2017;7(1):11. https://doi.org/10.3390/diagnostics7010011.
Thavarool SB, Muttath G, Nayanar S, Duraisamy K, Bhat P, Shringarpure K, B, S. Improved survival among oral cancer patients: findings from a retrospective study at a tertiary care cancer centre in rural Kerala. India World J Surg Oncol. 2019;17(1):15. https://doi.org/10.1186/s12957-018-1550-z.
Falzone L, Lupo G, La Rosa GRM, Crimi S, Anfuso CD, Salemi R, Candido S. Identification of novel MicroRNAs and their diagnostic and prognostic significance in oral cancer. Cancers (Basel). 2019;11(5):610. https://doi.org/10.3390/cancers11050610.
Peng CY, Liao YW, Lu MY, Yu CH, Yu CC, Chou MY. Downregulation of miR-1 enhances tumorigenicity and invasiveness in oral squamous cell carcinomas. J Formos Med Assoc. 2017;116(10):782–9. https://doi.org/10.1016/j.jfma.2016.12.003.
Georgantas RW, Streicher K, Greenberg SA, Greenlees LM, Zhu W, Brohawn PZ, Ranade K. Inhibition of myogenic microRNAs 1, 133, and 206 by inflammatory cytokines links inflammation and muscle degeneration in adult inflammatory myopathies. Arthritis Rheumatol. 2014;66(4):1022–33. https://doi.org/10.1002/art.38292.
Nakasa T, Miyaki S, Okubo A, Hashimoto M, Nishida K, Ochi M, Asahara H. Expression of microRNA-146 in rheumatoid arthritis synovial tissue. Arthritis Rheum. 2008;58(5):1284–92. https://doi.org/10.1002/art.23429.
Lu J, Guo S, Ebert BL, Zhang H, Peng X, Bosco J, Golub TR. MicroRNA-mediated control of cell fate in megakaryocyte-erythrocyte progenitors. Dev Cell. 2008;14(6):843–53. https://doi.org/10.1016/j.devcel.2008.03.012.
Koshizuka K, Nohata N, Hanazawa T, Kikkawa N, Arai T, Okato A, Seki N. Deep sequencing-based microRNA expression signatures in head and neck squamous cell carcinoma: dual strands of pre-miR-150 as antitumor miRNAs. Oncotarget. 2017;8(18):30288–304. https://doi.org/10.18632/oncotarget.16327.
Scapoli L, Palmieri A, Lo Muzio L, Pezzetti F, Rubini C, Girardi A, Carinci F. MicroRNA expression profiling of oral carcinoma identifies new markers of tumor progression. Int J Immunopathol Pharmacol. 2010;23(4):1229–34. https://doi.org/10.1177/039463201002300427.
Rodriguez A, Vigorito E, Clare S, Warren MV, Couttet P, Soond DR, Bradley A. Requirement of bic/microRNA-155 for normal immune function. Science. 2007;316(5824):608–11. https://doi.org/10.1126/science.1139253.
Sonkoly E, Janson P, Majuri ML, Savinko T, Fyhrquist N, Eidsmo L, Pivarcsi A. MiR-155 is overexpressed in patients with atopic dermatitis and modulates T-cell proliferative responses by targeting cytotoxic T lymphocyte-associated antigen 4. J Allergy Clin Immunol. 2010;126(3):581-589.e581-520. https://doi.org/10.1016/j.jaci.2010.05.045.
Thai TH, Calado DP, Casola S, Ansel KM, Xiao C, Xue Y, Rajewsky K. Regulation of the germinal center response by microRNA-155. Science. 2007;316(5824):604–8. https://doi.org/10.1126/science.1141229.
Yang Y, Li YX, Yang X, Jiang L, Zhou ZJ, Zhu YQ. Progress risk assessment of oral premalignant lesions with saliva miRNA analysis. BMC Cancer. 2013;13:129. https://doi.org/10.1186/1471-2407-13-129.
Lin F, Ding R, Zheng S, Xing D, Hong W, Zhou Z, Shen J. Decrease expression of microRNA-744 promotes cell proliferation by targeting c-Myc in human hepatocellular carcinoma. Cancer Cell Int. 2014;14:58. https://doi.org/10.1186/1475-2867-14-58.
Vislovukh A, Kratassiouk G, Porto E, Gralievska N, Beldiman C, Pinna G, Groisman I. Proto-oncogenic isoform A2 of eukaryotic translation elongation factor eEF1 is a target of miR-663 and miR-744. Br J Cancer. 2013;108(11):2304–11. https://doi.org/10.1038/bjc.2013.243.
Li JZ, Gao W, Lei WB, Zhao J, Chan JY, Wei WI, Wong TS. MicroRNA 744–3p promotes MMP-9-mediated metastasis by simultaneously suppressing PDCD4 and PTEN in laryngeal squamous cell carcinoma. Oncotarget. 2016;7(36):58218–33. https://doi.org/10.18632/oncotarget.11280.
Chen S, Shi F, Zhang W, Zhou Y, Huang J. miR-744-5p inhibits non-small cell lung cancer proliferation and invasion by directly targeting PAX2. Technol Cancer Res Treat. 2019;18:1533033819876913. https://doi.org/10.1177/1533033819876913.
Kleemann M, Schneider H, Unger K, Sander P, Schneider EM, Fischer-Posovszky P, Otte K. MiR-744-5p inducing cell death by directly targeting HNRNPC and NFIX in ovarian cancer cells. Sci Rep. 2018;8(1):9020. https://doi.org/10.1038/s41598-018-27438-6.
Balakittnen J, Weeramange CE, Wallace DF, Duijf PHG, Cristino AS, Kenny L, Punyadeera C. Noncoding RNAs in oral cancer. Wiley Interdiscip Rev RNA. 2022;14(3):e1754. https://doi.org/10.1002/wrna.1754.
Acknowledgements
CP is funded by the Cancer Australia (APP 1145657) and NHMRC Ideas Grants (APP 2002576 and APP 2012560), Royal Brisbane Women’s Hospital Foundation and Late Mr Jake Simpson Memorial Fund. We acknowledge the Endeavour Leadership Program through the Australian Government for the Research Fellowship to JMA.
Funding
Open Access funding enabled and organized by CAUL and its Member Institutions. No funding was obtained for this study.
Author information
Authors and Affiliations
Contributions
All authors have read and agree to the published version of the manuscript. KDT, JMA and CP: conceptualization. All authors: methodology, validation, formal analysis, data curation, investigation, and writing—review and editing. KDT, JMA, and CP: writing—original draft preparation. CP: funding acquisition.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no potential conflicts of interest.
Compliance with Ethical Standards
All procedures performed in this study involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards.
Ethics approval
This study was approved by the Ethics Review Board of the Complexo Hospital de Clínicas da Universidade Federal do Paraná (approval number 2.426.777) as well as by the University of Queensland Medical Ethical Institutional Board [HREC No: 2014000679 and 2014000862]; Queensland University of Technology [HREC No: 1400000617 and 1400000641] and by the Princess Alexandra Hospital Ethics Review Board [HREC Number: HREC/12/QPAH/381].
Informed consent
All participants provided written informed consent before saliva sample collection, following the Declaration of Helsinki.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Tang, K.D., Amenábar, J.M., Schussel, J.L. et al. Profiling salivary miRNA expression levels in Fanconi anemia patients – a pilot study. Odontology 112, 299–308 (2024). https://doi.org/10.1007/s10266-023-00834-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10266-023-00834-9