Abstract
When observational studies are used to establish the causal effects of treatments, the estimated effect is affected by treatment selection bias. The inverse propensity score weight (IPSW) is often used to deal with such bias. However, IPSW requires strong assumptions whose misspecifications and strategies to correct the misspecifications were rarely studied. We present a bootstrap bias correction of IPSW (BC-IPSW) to improve the performance of propensity score in dealing with treatment selection bias in the presence of failure to the ignorability and overlap assumptions. The approach was motivated by a real observational study to explore the potential of anticoagulant treatment for reducing mortality in patients with end-stage renal disease. The benefit of the treatment to enhance survival was demonstrated; the suggested BC-IPSW method indicated a statistically significant reduction in mortality for patients receiving the treatment. Using extensive simulations, we show that BC-IPSW substantially reduced the bias due to the misspecification of the ignorability and overlap assumptions. Further, we showed that IPSW is still useful to account for the lack of treatment randomization, but its advantages are stringently linked to the satisfaction of ignorability, indicating that the existence of relevant though unmeasured or unused covariates can worsen the selection bias.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
When the goal is to establish the causal effects of treatments, randomized controlled trials (RCTs) are the gold standard (Kovesdy et al. 2012). Due to various reasons, including costs, ethicality, and the growing easiness of access to registers and large follow-up data, observational studies are increasingly used for the evaluation of treatment effect differences between groups of individuals (Austin 2019). The present work is inspired by an observational study, where the objective was to explore the potential of oral anticoagulant treatment (OAT) in reducing the risk of mortality due to atrial fibrillation in patients with end-stage renal disease (ESRD) (Genovesi et al. 2014). The observational nature of the study introduces a selection bias into the estimated average treatment effect as the lack of randomization can cause the treated and control groups to be different in terms of baseline characteristics (Lunceford and Davidian 2004). This treatment selection bias is commonly known as confounding bias or non-exchangeability problem (Hernán and Robins 2020).
Several methods have been used to account for baseline differences between treated and untreated subjects. These include from the simple model adjustment to g-computation (Hernán and Robins 2020), propensity score-based matching and stratification (Austin and Small 2014; Imbens and Rubin 2015), and doubly robust estimators (Saarela et al. 2016). In particular, the inverse propensity score weighting [(IPSW, Rosenbaum and Rubin (1983)], under the potential outcomes approach, has been widely used to address the selection bias of the estimated average treatment effect. IPSW uses weights based on the propensity score to balance baseline covariates between the treated and control groups so that the two groups are similar in terms of pre-treatment covariates (Joffe et al. 2004; Hernán and Robins 2020). The use of IPSW is theoretically appealing as it intends to make the groups comparable (Kovesdy et al. 2012). In practice, however, the approach requires strong assumptions in order to successfully balance baseline covariate differences and allow estimation of the treatment effect with reduced selection bias (Rosenbaum and Rubin 1983; Frölich 2007). Specifically, the performance of IPSW critically depends on the ‘strongly ignorable treatment assignment’ condition, which requires the validity of two main assumptions. The first one is the ignorability restriction, which implies that there is no unobserved covariate that affects both the outcome and treatment simultaneously (Rosenbaum and Rubin 1983; Joffe et al. 2004). The second is known as the overlap assumption (Hernán and Robins 2020), which indicates that after IPSW rebalance, the distributions of the baseline covariates are comparable between the treated and control groups (McDonald et al. 2013). It has been suspected that the IPSW lacks robustness against the misspecification of these assumptions (Rubin 2004). Indeed, Morgan and Todd (2008) warn that applying the propensity score methods without due attention to the underlying assumptions, under the justification that these models are more advanced than the unweighted analysis (dubbed naïve henceforth), may provide a worse biased estimate. Mao et al. (2019) and Zhou et al. (2020) suggested the use of stabilized propensity score weights to limit the impact of the misspecification of the overlap assumption.
One statistical tool that can be used to improve the performance of the propensity score methods in estimating the average treatment effect for observational data is the bootstrap technique. The bootstrap is a well-known resampling method to assess the precision, whether bias or variability, of estimators with a complex analytic structure and unknown probability distribution (Efron and Tibshirani 1994). Some researchers have shown a promising impact of the bootstrap to enhance the propensity score-based treatment effect estimate. Peng and Jing (2011) adopted the bootstrap to estimate standard error for the average treatment effect, while Austin and Small (2014) used it to study the sampling variability of treatment effect focusing on propensity score matching. For the case of IPSW, a simulation by Gubhinder and Voia (2018) indicated that the bootstrap reduces the bias of the treatment effect on a Gaussian outcome. Apart from these limited efforts, the potential of bootstrap has not been sufficiently explored in conjunction with IPSW to correct bias due to the misspecification of key assumptions. Further, a broader simulation is lacking that illustrates the impact of failure to hold key assumptions that can be useful for applied researchers to understand the drawbacks and benefits of the propensity score.
This paper presents a bootstrap-corrected IPSW (BC-IPSW) to address selection bias when the propensity score weights are considered for the estimation of treatment effect under the misspecification of the ignorability and overlap assumptions. The BC-IPSW will be applied on the real ESRD dataset to pursue an unbiased estimate of the OAT effect on time-to-event mortality endpoint. The goal is twofold: (a) to evaluate whether OAT could lead to a reduction in mortality. The effect of OAT estimated by BC-IPSW will be compared with the estimates obtained using IPSW and the naïve (basic unweighted) methods. (b) To examine the impact of failure to hold the ignorability and overlap assumptions by a simulation study and to assess to what extent the BC-IPSW can eliminate or reduce the bias of these misspecifications under various scenarios. Both goals will be fostered based on real ESRD data.
In Sect. 2, we describe the ESRD application dataset. Section 3 presents the methodologies: the naïve approach, the IPSW and BC-IPSW methods. The results of the ESRD application dataset are presented in Sect. 4. The simulation study is discussed in Sect. 5, covering the simulation design and associated results. Section 6 provides a concluding discussion.
2 Motivating application data
The motivating application is based on determining the effectiveness of oral anticoagulant treatment (OAT) in reducing mortality due to atrial fibrillation (AF) in patients with end-stage renal disease (ESRD). OAT has been the treatment of choice to prevent thromboembolic events in ESRD patients with AF, but its beneficial effects are uncertain, also due to the high risk of bleeding in ESRD (Genovesi et al. 2014). The data are coming from a prospective cohort study of 290 patients aged 44–93 years with atrial fibrillation and ESRD in ten Italian hemodialysis centers. The patients were followed up for 4 years from October 31, 2010, to October 31, 2014. At recruitment, 134 patients (\(46.2\%\)) received OAT prescription and 72 \((53\%)\) died during the 4-year follow-up. Among the 156 control patients, 98 \((63\%)\) died. The median survival time was 2.62 years, with a \(95\%\) confidence interval of \(2.20-3.86\). We considered the overall mortality as the endpoint, and several baseline covariates were measured at patient recruitment. After preliminary analyses, nine covariates were considered which are also known confounders by the medical studies on the topic (Camm et al. 2012). Table 1 reports these patient characteristics of the ESRD dataset. The research interest is to quantify the estimate of the OAT effect in reducing mortality considering the observational nature of the study. The data have been previously analyzed by Genovesi et al. (2014, 2017).
3 Methods
In this section, we outline the methods we considered to address the two aims discussed in the Introduction section. We start with a recollection of the potential outcomes framework to causal inference in Subsect. 3.1. We then discuss in Subsect. 3.2 the model we focus on for causal inference on OAT treatment and what we call the naïve estimation from observational data, which ignores the risk of treatment selection bias. Then, Sect. 3.3 describes the IPSW to account for the selection bias, and Sect. 3.4 presents BC-IPSW to further improve treatment effect estimation under the risk of misspecification of crucial requirements to IPSW. Let \(T^*_i\) be the failure time of a subject i \((i=1,\dots ,n)\) in the cohort of size \(n=290\). Since the survival times are affected by the right censoring, the observed survival time is \(T_i=\text{ min }(T^*_i, C_i)\), where \(C_i\) is the right censoring time. \(T^*_i\) and \(C_i\) are assumed to be independent, as commonly done in survival analysis (Marubini and Valsecchi 1996; Arisido et al. 2019). The outcome is denoted by \(D_{it}\), which is the mortality event for subject i taking values 1 if \(T_i\le t\) and 0 if \(T_i>t\). Let \(z_i\) be the indicator of observed treatment status taking value 1 if subject i received treatment and 0 if assigned to control, and \(\mathbf{x }_i\) be the individual vector of baseline covariates. The interest is to estimate the unbiased effect of OAT on mortality, denoted by \(\beta\), using the hazard ratio scale.
3.1 Treatment effect estimation and assumptions
In the potential outcomes framework (Imbens and Rubin 2015), for each subject i there exist two potential outcomes: \(D^1_{it}\) and \(D^0_{it}\), the mortality outcome if \(z_i=1\) and \(z_i=0\), respectively. In the context of survival analysis (e.g., Hernán and Robins 2020), the causal effect for subject i is the hazard ratio (HR) between \(D^1_{it}\) and \(D^0_{it}\), which is never observed since either \(D^1_{it}\) or \(D^0_{it}\) can be observed, but not both (Lunceford and Davidian 2004). The actual observed outcome \(D_{it}\) is the one that would have been seen under the actual treatment assignment:
The parameter of interest is the average treatment effect in the population (ATE), i.e., the average HR if all individuals were to receive the treatment against if all individuals were control. Another measure of treatment effect is the average treatment effect for the treated (ATT) as given by averaging HR over the treated (\(z_i=1\)) group. Since ATE and ATT do not coincide in an observational study, the choice between them depends on the specific application (e.g., Pirracchio et al. 2016). In this paper, we focus on identifying the ATE of OAT in the ERSD dataset. For the unbiased estimation of ATE, two key assumptions are required. First let \(e(\mathbf{x }_i)=Pr(z _i= 1|\mathbf{x }_i)\) define the propensity score, namely the probability of treatment assignment conditional on \(\mathbf{x }_i\) (Hernán and Robins 2020). The first assumption is the ignorability or conditional independence assumption:
where \(\bot \!\!\!\bot\) denotes the statistical independence. The assumption states that provided that \(\mathbf{x }_i\) captures all confounders and that there are no omitted relevant covariates, assignment to treatment and the potential responses are unrelated. Under this assumption, the treatment assignment is ignorable (Rosenbaum and Rubin 1983). The second is the overlap or positivity assumption, which implies that conditional on \(e(\mathbf{x }_i)\), the distribution of \(\mathbf{x }_i\) does not depend on \(z_i\). In other words, any subject in the treatment group has the potential match in the control group (McDonald et al. 2013), indicating that \(e(\mathbf{x }_i)\) of the two groups overlaps. When the combination of ignorability and overlap assumptions holds, the treatment assignment is said to be ‘strongly ignorable’(Rosenbaum and Rubin 1983).
3.2 The Naïve estimation
To evaluate the impact of OAT treatment, the survival endpoint is specified as a function of treatment status and baseline characteristics, typically using the Cox proportional hazards model (Cox 1972)
where \(h_0(t)\) denotes the unspecified baseline hazard function and \(\beta _{nv}\) is the log hazard measuring the association between the treatment \(z_i\) and the hazard \(h_i(t)\). Further, \(\mathbf{x }_i\) denotes the \(n\times 9\) matrix of the nine baseline covariates as listed in Table 1 and \(\varvec{\theta }\) denotes a column vector of the corresponding regression coefficients. With the naïve approach without accounting for the selection bias, parameters are estimated based on maximizing the partial likelihood (Cox 1972), and the hazard ratio \(\text{ HR }=\exp (\beta _{nv})\) is interpreted as the relative change in the hazard at any time t for a subject who received OAT relative to a subject in the control group in the presence of confounding covariates.
3.3 Propensity score weighted estimation
The inverse propensity score weighting (IPSW) is a balancing strategy based on the propensity score \(e(\mathbf{x }_i)\) to match baseline characteristics between treated and control patients in observational studies (Imbens and Rubin 2015; Hernán and Robins 2020). In practice, the probability distribution of \(z_i|\mathbf{x }_i\) is unknown and the propensity score \(e(\mathbf{x }_i)\) has to be estimated from the observed data, usually by modeling the conditional probability \(Pr(z_i=1|\mathbf{x }_i)\) via a parametric multivariable logistic regression model (Joffe et al. 2004; Hernán and Robins 2020)
where \(\varvec{\alpha }\) is the column vector of regression coefficients measuring the effect of each covariate on the probability of treatment assignment. We then used the estimated \(\hat{\varvec{\alpha }}\) to evaluate the propensity score \({\hat{e}}(\mathbf{x }_i)\)
Figure 1a shows the estimated distribution of \({\hat{e}}(\mathbf{x })\) in the ERSD dataset. The treated group appears to have higher propensity scores than the control group, except for a small number of control patients who act as outliers. The inverse propensity score weight (IPSW) \({\hat{\omega }}(\mathbf{x }_i)=\frac{1}{{\hat{e}}(\mathbf{x }_i)}\) was used to estimate the weighted treatment effect. For the ERSD dataset, this is achieved by fitting \({\hat{\omega }}(\mathbf{x }_i)\)-adjusted Cox model, i.e., maximizing the weighted partial likelihood of the form
where R(t) is the risk set at time t and \(\delta _i\) indicates the censoring status of a subject. Notice that by setting \({\hat{\omega }}_j(t)=1\) for all \(j\in R(t)\), Eq. (5) is the usual unweighted partial likelihood. \(\exp ({\hat{\beta }}_{ps})\) is the marginal hazard ratio for a subject who received OAT relative to a subject in the control group accounting for the selection bias. The standard error of \({\hat{\beta }}_{ps}\) was computed by the robust variance estimation to account for the weighting (Joffe et al. 2004; Buchanan et al. 2014). The IPSW adjusts for the treatment selection bias provided that the ignorability and overlap assumptions, as discussed in Sect. 3.1, hold. On the other hand, misspecification of either one or both has the potential to lead to a biased \({\hat{\beta }}_{ps}\). For instance, the ignorability assumption in our ERSD data anticipates that the nine-dimensional covariate vector in Table 1 used to estimate \({\hat{e}}(\mathbf{x }_i)\) is complete and represents all relevant covariates that affect the treatment status. This is quite a strong constraint, and its possible deviation from this assumption poses the risk of not eliminating or even worsening the selection bias.
3.4 Bootstrap-corrected IPSW estimation
When IPSW fails to eliminate the selection bias due to the misspecification of one or more key assumptions, we consider a bootstrap bias correction of the IPSW (BC-IPSW). The bootstrap, and particularly its nonparametric original version, is a well-known computer-intensive methodology to assess estimation accuracy. Its main advantage relies on its ability to provide a resampling simulation of the unknown probability distribution of an estimator, known as bootstrap distribution, regardless of its analytical complexity. Consequently, bootstrap estimates of the estimator’s expectation and variance are offered upon the bootstrap distribution, as well as percentiles and confidence intervals. The underlying inferential idea is based on a combination of plug-in frequentist estimation and Monte Carlo (MC) simulation (Efron 1979; Efron and Hastie 2016). For the purposes of this paper, we focus on the bias of the IPSW estimator, i.e., the difference \(\text {bias}\left( {{\hat{\beta }}}_{ps}\right) = E\left( {{\hat{\beta }}}_{ps}\right) - \beta\) where the expectation is under the probability distribution of the IPSW estimator. We now illustrate how to compute the bootstrap estimate of such bias in order to improve upon the IPSW estimate by constructing a BC-IPSW estimate.
The rationale behind bootstrapping the bias can be simplified as rooted on the frequentist interpretation of \(E\left( {{\hat{\beta }}}_{ps}\right)\) which, ideally, is what we could observe by repeatedly sampling new version of treated and control groups of patients to observe the estimator’s distribution. With the bootstrap, we simulate this by repeatedly re-sampling from the available data stratified by the treatment status that gives a MC-induced estimator’s distribution (Tu and Shao 1995). The first step of the bootstrap algorithm consists of repeatedly drawing with replacement from the single original sample a sufficiently large number B of bootstrap samples (Efron and Tibshirani 1994; Conti et al. 2020). As applied to our original ESRD data, we set a number \(B=2000\) of bootstrap samples each with the same size \(n=290\). For each of the B bootstrap samples, we computed a replication of the IPSW-adjusted treatment effect \({\hat{\beta }}_{ps_1}^*, {\hat{\beta }}_{ps_2}^*, \dots , {\hat{\beta }}_{ps_B}^*\), which provides the bootstrap distribution of \({\hat{\beta }}_{ps}\). Let \({\hat{\beta }}_{ps}^*\) denote the average of the bootstrap distribution \(B^{-1} \sum _{b=1}^B {\hat{\beta }}_{ps_b}^*\), which is in fact the bootstrap version \(E_B\left( {{\hat{\beta }}}_{ps}\right)\) of the expectation \(E\left( {{\hat{\beta }}}_{ps}\right)\). The bootstrap estimate for the bias of the IPSW estimator is given by
The bootstrap bias estimate is then used to update IPSW estimator \({\hat{\beta }}_{ps}\) with the aim to correct the residual bias. Thus, the bootstrap-corrected BC-IPSW average treatment effect estimator \({\tilde{\beta }}_c\) is given in the form
In other words, \({\tilde{\beta }}_c\) is obtained after a double bias correction in a single framework. First, IPSW was adopted to produce comparable treated and control groups in terms of pre-treatment differences. Second, the bootstrap is applied in an attempt to improve the IPSW estimate against violation of crucial assumptions. As a remark, a BC-IPSW-estimated effect confidence interval can be computed as bootstrap interval based on bias-corrected percentiles distribution \(({\hat{\beta }}^*_{ps_b}, b=1,\dots , B )\) (Efron 1979). More specifically, the bias-corrected 95% confidence interval, according to Eq. 6, is given by \([{\hat{\beta }}^*_{ps,0.025} - \text{ bias}_B({\hat{\beta }}_{ps}), {\hat{\beta }}^*_{ps,0.975} - \text{ bias}_B({\hat{\beta }}_{ps})].\) The BC-IPSW approach can be computationally intensive for choices of very large B. However, the approach does not require analytic computation, which should appeal to applied researchers for its broader applicability (Kim and Sun 2016).
4 Results of the application data
The results of the analysis of the ESRD dataset, which was described in Sect. 2, are shown in Table 2. The results are reported in terms of parameter estimate, the hazard ratio (HR) along with the \(95\%\) confidence interval (\(95\%\) HR CI). In general, the estimated HR from the three methods indicated that OAT treatment has the benefit of improving survival. However, the impact of the estimated treatment effect on mortality varies across the methods, ranging from a HR of 0.65 for the BC-IPSW to 0.90 for the naïve analysis. The naïve estimate gives indication for little difference in the mortality risk between treated and untreated groups, possibly minimizing the effect of the treatment due to selection bias. We noticed a great variability among the \(95\%\) confidence intervals, which indicated that both the naïve and IPSW methods estimated a statistically nonsignificant OAT effect on mortality, although the IPSW interval is narrower than the corresponding naïve confidence interval. The BC-IPSW estimated a HR\(=\)0.65(95% CI 0.421–0.978), which implies a statistically significant effect of OAT on mortality. This indicates that the rate of mortality for patients receiving the OAT decreased by 35% as compared to the same in the control group.
IPSW is intended to adjust for the selection bias in treatment assignment provided that the main assumptions as discussed in Sect. 3.1 hold. One way to examine whether IPSW successfully balanced the distribution of baseline covariates between the treated and control is to compare the standardized mean difference (Austin 2019; Ridgeway et al. 2017). When the propensity score achieves a perfect balance, the standardized mean difference will be zero. Figure 1b displays the standardized mean differences for the nine baseline covariates in ERSD data. The absolute standardized differences ranged from 0.01 for age to a maximum of 0.1 for HF. For most covariates such as age and bleeding, IPSW achieved a high degree of balance as the mean differences are closer to zero, but the balance in some covariates such as HF and DM is less satisfactory as the mean differences of these covariates are relatively large. Thus, the lack of balance between treated and control for some covariates may indicate a failure of an assumption to hold. The distributional difference between treated and control of the estimated propensity score \(e(\mathbf{x }_i)\) in Fig. 1b suggests that the overlap assumption was not satisfied. A simulation study may provide detailed insights on the extent of the impact of failure to hold for propensity score assumptions. Further, the three methods provided different estimates, which results in different conclusions regarding the causal effect of OAT on mortality. Thus, a simulation study is necessary to validate the approaches and compare the performance of each method with and without misspecification of key assumptions.
5 Simulation study
In this section, we illustrate a simulation study with the aim of evaluating the impact of the misspecification of key assumptions and of assessing to what extent the BC-IPSW can reduce or eliminate the bias. Specifically, we explore the performance of the three methods presented in Sect. 3 under the misspecification of the ignorability and the overlap assumptions.
5.1 Simulation protocol
The simulation was designed according to the motivational application in Section 2. We considered a sample size of \(n = 290\) subjects with follow-up time of 4 years (\(t=0,\dots , 4\)). Data were generated by the following five steps:
-
1.
A vector of nine binary baseline covariates \(\mathbf{x }_i\) was generated from a vector of independent Bernoulli distribution as \(\mathbf{x }_i \sim \text{ Bernoulli }(\mathbf{p }_x)\) with \(\mathbf{p }_x=(0.21, 0.4, 0.35, 0.19, 0.48, 0.81, 0.31, 0.15, 0.39)\). These values are estimated from the ESRD data. For simplicity, interactions were not considered for the simulation study.
-
2.
Treatment status \(z_i\) was generated as \(z_i\sim \text{ Bernoulli }({\hat{p}}_i)\), where \({\hat{p}}_i\) is estimated from model (3) using covariates generated in step 1 with coefficients \(\varvec{\alpha }=(1.4,0.22,0.08,0.14,0.32,0.39,0.15,0.07,0.39)\). To assess the impact of ignorability misspecification, we considered two levels of misspecifications: 1) mild, where two covariates weakly related to survival outcome with coefficients 0.08 and 0.14 were omitted in the calculation of \({\hat{e}}(\mathbf{x })\); and 2) gross, where a strongly related covariate with the highest coefficient 1.4 was omitted.
-
3.
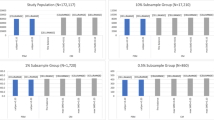
To assess the impact of failure to hold the overlap assumption, \(\mathbf{p }_x\) in step 1 was altered to reflect a certain covariate imbalance between treatment and control groups. We consider two scenarios in generating \(\mathbf{x }_i\): 1) We set \(\mathbf{p }_x=0.2\) for treated and \(\mathbf{p }_x=0.3\) for control. We call this scenario large overlap as the disparity of \(\mathbf{p }_x\) between the two groups is small. 2) \(\mathbf{p }_x=0.2\) for treated and \(\mathbf{p }_x=0.5\) for control. We call this situation partial overlap as higher \(\mathbf{p }_x\) disparity between the two groups creates small overlap. Figure 2b and c depicts the distribution of \({\hat{e}}(\mathbf{x })\) using \(\mathbf{x }\) generated under the two overlap misspecification scenarios, while Fig. 2a shows the distribution of \({\hat{e}}(\mathbf{x })\) for a perfect overlap scenario.
-
4.
Survival times \(T^*_i\) were generated by a Weibull proportional hazard model \(h_i(t)=\lambda \rho t^{\rho -1}\exp (\beta z_i+ \mathbf{x }_i^t\varvec{\alpha })\), by evaluating the inverse of the cumulative hazard (Bender et al 2005; Arisido et al. 2019). The rate and shape parameters are set as \(\lambda =0.1\) and \(\rho =1.4\), respectively. The true log hazard was fixed at \(\beta \in (-1.2, -0.3, 0.4)\), corresponding to a situation of strongly negative, weakly negative and a positive association between treatment and mortality.
-
5.
Censoring times \(C_i\) were generated according to a uniform distribution in (0, 4), leading to about 20% of censoring proportion before year 4. The censored observed survival times were obtained as \(T_i = \text{ min }(T^*_i, C_i)\).
We run \(M = 1000\) Monte Carlo simulations. For each simulation, we fitted a naïve model that ignores the selection bias and the IPSW adjusted model. To fit the BC-IPSW model, \(B=2000\) bootstrap runs have been nested into the M simulations. To assess the performance of each method, simulation results are summarized using bias, absolute percentage bias (%Bias), mean-squared error (MSE), and the Monte Carlo coverage of 95% confidence intervals (CP). The description and computational details of these metrics are described in Burton et al. (2006); Arisido (2016).
The distribution of the propensity score \({\hat{e}}(\mathbf{x })\) estimated from the simulated data showing various levels of overlap between treated and control. a Nearly perfect overlap as covariate imbalance was eliminated, b large overlap with slight covariate imbalance, c partial overlap with high covariate imbalance. The x-axis shows the range of \({\hat{e}}(\mathbf{x })\) from 0 to 1, and the y-axis shows its probability density distribution
5.2 Simulation results
Table 3 shows basic simulation results without model misspecification with the true \(\beta \in (-1.2, -0.3, 0.4)\). In general, the estimated treatment effect obtained by the naïve analysis has a higher bias compared with the biases estimated by the other two approaches across the three \(\beta\) values. For a strong beneficial impact of treatment (\(\beta =-1.2\)), the naïve model that fails to account for the treatment selection bias overestimates the true effect of the treatment by \(4.8\%\), while the IPSW and BC-IPSW methods underestimated the true effect by \(3.5\%\) and \(1.4\%\), respectively. Further, IPSW and BC-IPSW showed slightly better accuracy as measured by MSE and good coverage properties. For \(\beta =-0.3\), the naïve method estimated a percentage bias of 5% and this bias was lowered to 3.33% when IPSW was implemented. The BC-IPSW resulted in a further lowest percentage bias of 0.67%. The three methods showed a minor difference in terms of accuracy and 95% coverage probability for \(\beta \in (-0.3, 0.4)\). This may indicate that the treatment selection bias severely affects the estimated treatment effect, with the impact less pronounced in precision and coverage properties.
Table 4 reports the simulation results for the misspecification of the ignorability assumption. The data simulation under this misspecification was stated in step 2 of Sect. 5.1. In addition to the mild and gross misspecification scenarios, increasing sample size \((n=100, 290, 600)\) was considered and \(\beta\) was fixed at \(\beta =0.4\). For \(n=100\) (relatively small), the IPSW estimator was the most affected under mild misspecification, as its percentage bias of \(22\%\) was far higher than the biases estimated by the naïve and BC-IPSW methods. The bias decreased as the sample size n increased, reaching a minimum bias of \(1.7\%\) by the BC-IPSW estimator with \(n = 600\), and the coverage probabilities were closer to the nominal 95% for the three methods. Under the gross misspecification, the amount of bias for the IPSW and BC-IPSW estimator was increased further, while the increase was small for the naïve estimator. To better understand the impact of the ignorability misspecification, one can compare this misspecification result for \(n=290\) with the basic simulation results obtained under no misspecification in Table 3 at \(\beta =0.4\) scenario. The percentage bias of \(11.3\%\) for the IPSW under mild misspecification corresponds to an extra \(8\%\) bias as compared with the same scenario without misspecification in Table 3. The extra bias could be as large as \(13\%\) under gross misspecification. BC-IPSW resulted in reduced biases of \(5\%\) and \(7.3\%\) for the mild and gross misspecifications, respectively.
The performance of the three approaches improves with n, but the sensitive nature of the ignorability assumption is reflected by the fact that the estimates by IPSW are more biased than the naïve estimates for all the scenarios. BC-IPSW substantially reduced the bias, although it does not lead to a negligible bias even for the largest n scenario. Figure 3 shows the distribution of the estimated treatment effect using the BC-IPSW method under the mild (a) and gross misspecification (b) across various n with a maximum size 1200. It is evident that the estimate under the mild scenario approaches the true \(\beta =0.4\) as n increases and close to coincide at \(n=1200\). However, the estimate appears to deviate from the true value under the gross misspecification.
Table 5 shows the simulation results of the overlap misspecification for \(\beta \in (-0.3, 0.4)\) and \(n=290\). The data simulation under this misspecification was stated in step 3 of Sect. 5.1. Figure 2 shows distribution of the propensity score \({\hat{e}}(\mathbf{x })\) for the treated and control groups under various overlap scenarios. The performance of the naïve method was comparable to the previous results (see Tables 3 and 4). The performance of IPSW strictly depends on the overlap assumption. For instance, for \(\beta =-0.3\) and when most of the subjects in treated and control overlap (large overlap scenario), the estimated effect was \(9\%\) biased with coverage probability lower than \(90\%\). The BC-IPSW method reduced this bias by \(8\%\) with a good empirical coverage probability close to the nominal \(95\%\). For the corresponding partial overlap scenario, the percentage bias of the IPSW rose to \(16\%\) with a poor coverage probability of just \(75\%\). Again, BC-IPSW was able to shrink the IPSW bias to \(12\%\) for the partial overlap scenario.
6 Discussion
Investigating the effects of a treatment based on observational studies requires statistical tools that take into account selection bias due to the lack of randomization. The propensity score-based IPSW has been widely used to address such selection bias (Imbens and Rubin 2015). While the propensity score is a viable way to rely on observational studies when RCT is not feasible to conduct, it necessitates strong assumptions whose misspecification might severely bias the treatment effect. In this context, there have been some efforts to jointly use the bootstrap and the propensity score to describe either the bias (Gubhinder and Voia 2018) or variability (Peng and Jing 2011; Austin and Small 2014) of the average treatment effect. The former paper showed that bootstrap reduces the bias of the treatment effect on a Gaussian outcome under treatment response misspecifications such as ignorability and endogeneity.
In this paper, we present a more complex bootstrap-corrected IPSW (BC-IPSW) approach for a time-to-event endpoint to improve the performance of IPSW in dealing with selection bias in observational studies. The method first adopts IPSW to balance baseline characteristics between treated and control groups, and then it applies the bootstrap to improve the estimate of the treatment effect when the IPSW fails to achieve adequate balance. The work was motivated by an observational real cohort study, where the objective was to investigate the potential of OAT in reducing mortality in patients with ESRD. We found that OAT treatment has the benefit of improving the survival rate, although the estimated effect was different across the methods used to analyze the data. The naïve method, which does not account for the observational nature of the study, estimated a very weak effect of the treatment. The combined use of the IPSW and the bootstrap resulted in an estimate that showed a statistically significant reduction in mortality for patients receiving the treatment.
A comprehensive simulation study was presented that examined the impact of failure to hold key propensity score assumptions on the estimation of treatment effect and evaluated the extent to which the BC-IPSW is able to reduce the bias of IPSW estimator due to violation of these assumptions. The performance of the three methods was compared by considering various scenarios under the misspecification of the ignorability and overlap assumptions. The strong reliance of the IPSW on the ignorability assumption was disclosed by the simulation results, in which even omitting covariates that are weakly associated with the treatment status resulted in a biased estimate of treatment effect. Under such mild misspecification, the BC-IPSW method substantially reduced the bias to a negligible magnitude. In the event of a gross misspecification, when a covariate that is strongly associated with the treatment status was missing, BC-IPSW performed well in shrinking the bias, but a non-negligible bias was still observed. The implication of this is that the existence of unmeasured or unused, though relevant, covariates could affect the treatment effect, which is frequently the case for observational data (Morgan and Todd 2008). The simulation results additionally indicated the risk of failure to hold the overlap assumption. We noted that when a greater imbalance of baseline covariate exists between the treated and control groups, the IPSW is unable to eliminate the disparity to achieve an overlap between the groups, which in turn resulted in a severely biased treatment effect. This was particularly evident when we attempted to correct a highly divergent distribution of covariates between the two groups. The BC-IPSW reduced the misspecification bias even though the estimated effect was still relatively biased.
In conclusion, we investigated that the benefit of the propensity score adjustment to account for the selection bias associated with observational studies is linked to a careful consideration of its main assumptions. Both the application and the simulation results suggest that the BC-IPSW approach markedly improved the performance of the IPSW. However, the approach did not lead to an unbiased estimate of treatment effect under serious misspecifications. Thus, efforts to improve the BC-IPSW performance should be a future research focus. Further, the bootstrap can be used in conjunction with other estimating methods. To illustrate this, we compared the stabilized propensity score weights (Hernán and Robins 2020) with the unstabilized IPSW which we have adopted. The stabilized IPSW was suggested to address the lack of overlap between the treated and control groups (Mao et al. 2019; Zhou et al. 2020). The performance of the bootstrap with the stabilized IPSW was mostly similar to bootstrap with the unstabilized IPSW, though the former estimated slightly better coverage probability. The results are shown in Table 1 of the Supplemental Appendix. We also note that we adopted a uniform censoring with administratively known in advance as in ERSD data. In other real settings, censoring can be a random non-administrative with no pre-specified end point (Worms and Worms 2018; Stupfler 2019). Table 2 of the Supplemental Appendix shows simulation results obtained with and without administrative censoring. For a lower censoring rate, the results obtained with both censoring types are relatively comparable. A higher censoring rate resulted in a more biased estimate of the treatment effect for both censorings, but the bias is stronger for the non-administrative censoring. It should be noted that non-random censoring (endogenous) was not addressed here, but could be of interest for further work on the topic.
Code availability
The code and complete simulation results of the current study are available from the corresponding author on reasonable request.
References
Arisido, M., Antolini, L., Bernasconi, D., Valsecchi, M., Rebora, P.: Joint model robustness compared with the time-varying covariate Cox model to evaluate the association between a longitudinal marker and a time-to-event endpoint. BMC Med. Res. Methodol. 19, 222–235 (2019)
Arisido, M.W.: Functional measure of ozone exposure to model short-term health effects. Environmetrics 27, 306–17 (2016)
Austin, P.C., Small, D.S.: The use of bootstrapping when using propensity-score matching without replacement: a simulation study. Statist. Med. 33, 4306–4319 (2014)
Austin, P.C.: Assessing covariate balance when using the generalized propensity score with quantitative or continuous exposures. Stat. Methods Med. Res. 28, 1365–1377 (2019)
Bender, R., Augustin, T., Blettner, M.: Generating survival times to simulate Cox proportional hazards models. Stat. Med. 24, 1713–1723 (2005)
Buchanan, A.L., Hudgens, M.G., Cole, S.R., Lau, B., Adimora, A.A.: Women’s Interagency HIV Study.: Worth the weight: using inverse probability weighted Cox models in AIDS research. AIDS Res. Human Retroviruses. 30, 1170–1177 (2014)
Burton, A., Altman, D.G., Royston, P., Holder, R.L.: The design of simulation studies in medical statistics. Stat. Med. 25, 4279–4292 (2006)
Camm, A.J., Lip, G.Y., De Caterina, R.: 2012 focused update of the ESC guidelines for the management of atrial ibrillation: an update of the 2010 ESC guidelines for the management of atrial ibrillation. Developed with the special contribution of the European Heart Rhythm Association. Eur Heart J 33, 2719–2747 (2012)
Conti, P.L., Marella, D., Mecatti, F., Andreis, F.: A unified principled framework for resampling based on pseudo-populations: asymptotic theory. Bernoulli 26, 1044–1069 (2020)
Cox, D.R.: Regression models and life tables. J. R. Stat. Soc. 34, 187–220 (1972)
Efron, B.: Bootstrap methods: another look at the jackknife. Ann. Stat. 7, 1–26 (1979)
Efron, B., Tibshirani, R.J.: An Introduction to the Bootstrap. CRC Press, Boca Raton (1994)
Efron, B., Hastie, T.: Computer Age Statistical Inference. Cambridge University Press, Cambridge (2016)
Frölich, M.: On the inefficiency of propensity score matching. AStA Adv. Stat. Anal. 91, 279–290 (2007)
Genovesi, S., Rossi, E., Gallieni, M., Stella, A., Badiali, F., Conte, F., Pozzi, C.: Warfarin use, mortality, bleeding and stroke in haemodialysis patients with atrial fibrillation. Nephrol. Dial. Transp. 30, 491–498 (2014)
Genovesi, S., Rebora, P., Gallieni, M., Stella, A., Badiali, F., Conte, F., Pozzi, C.: Effect of oral anticoagulant therapy on mortality in end-stage renal disease patients with atrial fibrillation: a prospective study. J. Nephrol. 30, 573–581 (2017)
Gubhinder, K.P.R., Voia, M.C.: Bootstrap bias correction for average treatment effects with inverse propensity weights. J. Stat. Res. 52, 187–200 (2018)
Hernán, M.A., Robins, J.M.: Causal Inference: What If. Chapman and Hall/CRC, Boca Raton (2020)
Imbens, G.W., Rubin, D.B.: Causal Inference in Statistics, Social, and Biomedical Sciences. Cambridge University Press, Cambridge (2015)
Joffe, M.M., Ten Have, T.R., Feldman, H.I., Kimmel, S.E.: Model selection, confounder control, and marginal structural models: review and new applications. Am. Stat. 58, 272–279 (2004)
Kim, M.S., Sun, Y.: Bootstrap and k-step bootstrap bias corrections for the fixed effects estimator in nonlinear panel data models. Econ. Theory. 32, 1523–1568 (2016)
Kovesdy, C.P., Kalantar-Zadeh, K.: Observational studies versus randomized controlled trials: avenues to causal inference in nephrology. Adv. Chronic Kidney Dis. 19, 11–18 (2012)
Lunceford, J.K., Davidian, M.: Stratification and weighting via the propensity score in estimation of causal treatment effects: a comparative study. Stat. Med. 23, 2937–2960 (2004)
Mao, H., Li, L., Greene, T.: Propensity score weighting analysis and treatment effect discovery. Stat. Methods Med. Res. 28, 2439–2454 (2019)
Marubini, E., Valsecchi, M.G.: Analysing Survival Data from Clinical Trials and Observational Studies. Wiley, West Sussex (1996)
McDonald, R.J., McDonald, J.S., Kallmes, D.F., Carter, R.E.: Behind the numbers: propensity score analysis-a primer for the diagnostic radiologist. Radiology 269, 640–645 (2013)
Morgan, S.L., Todd, J.J.: A diagnostic routine for the detection of consequential heterogeneity of causal effects. Sociol. Methodol. 38, 231–282 (2008)
Peng, X., Jing, P.: Bootstrap confidence intervals for the estimation of average treatment effect on propensity score. J. Math. Res. 3, 52–58 (2011)
Pirracchio, R., Carone, M., Rigon, M.R., Caruana, E., Mebazaa, A., Chevret, S.: Propensity score estimators for the average treatment effect and the average treatment effect on the treated may yield very different estimates. Stat. Methods Med. Res. 25, 1938–1954 (2016)
Ridgeway, G., McCaffrey, D., Morral, A., Burgette, L., Griffin, B.A.: Toolkit for Weighting and Analysis of Nonequivalent Groups: A tutorial for the twang package. RAND Corporation, Santa Monica (2017)
Rosenbaum, P.R., Rubin, D.B.: The central role of the propensity score in observational studies for causal effects. Biometrika 70, 41–55 (1983)
Rubin, D.B.: On principles for modeling propensity scores in medical research. Pharmacoepidemiol. Drug Saf. 13, 855–857 (2004)
Saarela, O., Belzile, L.R., Stephens, D.A.: A Bayesian view of doubly robust causal inference. Biometrika 103, 667–681 (2016)
Stupfler, G.: On the study of extremes with dependent random right-censoring. Extremes 22, 97–129 (2019)
Tu, D., Shao, J.: The Jackknife and Bootstrap. Springer, New York (1995)
Worms, J., Worms, R.: Extreme value statistics for censored data with heavy tails under competing risks. Metrika 81, 849–889 (2018)
Zhou, Y., Matsouaka, R.A., Thomas, L.: Propensity score weighting under limited overlap and model misspecification. Stat. Methods Med. Res. 29, 3721–3756 (2020)
Acknowledgements
The authors wish to thank Simonetta Genovesi for making available the ESRD data for the current study. We thank the three anonymous reviewers for their useful comments.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
The authors declare that there is no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Arisido, M.W., Mecatti, F. & Rebora, P. Improving the causal treatment effect estimation with propensity scores by the bootstrap. AStA Adv Stat Anal 106, 455–471 (2022). https://doi.org/10.1007/s10182-021-00427-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10182-021-00427-3