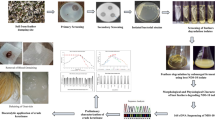
Abstract
The huge amounts of keratin-rich waste generated daily by various industries, slaughterhouses, and processing plants need to be properly managed. Most keratin degradation-related research focuses on keratin from bird feathers, but a vast minority focuses on keratin from sheep wool, which also presents a serious environmental problem. In this article, we describe the isolation, identification, and characterization of new keratinolytic microorganisms capable of sheep wool degradation from sheep wool and soil enriched with wool keratin. We isolated four bacterial species from the genus Bacillus (B. subtilis, B. altitudinis, B. mycoides, and B. wiedmannii), one streptomycete species Streptomyces coelicoflavus identified by whole genome sequencing, and a fungal species Aphanoascus reticulisporus. In some of the isolated microorganisms, we detected keratinolytic activity for the first time, and for most of them, the ability to degrade sheep wool has not been previously demonstrated. The keratinases of the new isolates are active in a wide range of temperatures (25–85 °C) and pH (6.0–10.0), so all isolates show great potential for further biotechnological use in industry and in various environmental and agricultural applications to reduce and recycle keratin-rich wastes such as sheep wool and waste woollen textiles.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Keratin-rich organic wastes are produced daily in the processing of livestock for food production and also in the wool textile industry. Waste wool, feathers, horns, animal hair, bristles, etc. are largely composed of various structural proteins such as elastin, collagen, and especially keratin [1]. Animals are more resistant to abiotic and biotic stresses because of keratin. Since microbial degradation of keratin is not widespread in nature, keratin can also serve as an effective protection against microbes. There are many known examples of preserved hair and skin on archeological material. There are also numerous examples of keratin-rich materials that have passed through the digestive tracts of animals undecomposted or only partially decomposted without providing nutritional value to the animal [2]. Keratin-rich materials are unique because of their high content of certain amino acids, particularly the sulphur-containing amino acid cysteine, but also other amino acids such as glycine, proline, arginine, and the essential amino acids valine, leucine and threonine [3]. The keratins found in wool fiber are divided into three groups. The first group, representing 50–60% of wool fiber, are alpha-keratins, which are compactly packed and stabilized by a high degree of inter- and intramolecular disulphide bonds, non-covalent hydrogen bonds, and hydrophobic interactions, have a low sulphur content and a molecular weight of 60–80 kDa, and are found in the cortex of wool fiber. The second group are beta-keratins, which have primarily a protective function and are present in small amounts in the cuticle, and third group representing 20–30% of the wool fiber, are gamma-keratins with a molecular weight of 11–28 kDa and a high sulphur content due to the high content of cysteine which act as disulphide cross-linker and help hold the keratin superstructure together, making it water-insoluble, enzymatically resistant, and mechanically stable [4]. Regardless of its compact structure, keratin-rich waste material does not accumulate in nature, which proves the presence of keratinolytic microorganisms and their enzymes [1]. These enzymes are called keratinases and are predominantly extracellular enzymes. Keratinases are the only group of proteases that can degrade keratin and belong mainly to the serine or metallo-proteases with a wide pH and temperature range [5].
Sheep in the European Union (EU) are raised primarily for meat and milk production. Because of the low quality of wool, most of the wool used in EU is Merino wool imported from Australia and New Zealand. Nevertheless, annual wool production in the EU amounts to more than 200 thousand tons, which is mainly presented as by-product and mostly landfilled or incinerated [6, 7]. Since keratin waste is mainly converted into various hydrolysates by environmentally harmful physicochemical treatments, there is a great need for biotechnological approaches such as microbial or enzymatic pretreatment, composting, or biogas production. Keratinolytic microorganisms or their enzymes can play an important role in the degradation of keratin-rich wastes by environmentally friendly processes [8].
Some experiments with the microbial or enzymatic treatment of feather or ship wool waste have been described, but there is a lack of experiments with microbial starter cultures or crude enzyme mixtures for better and faster composting or biogas production. Gousterova et al. [9] described the degradation of wool by some thermoactinomycete strains, Queiroga et al. [10] tested five wool-degrading Bacillus strains for degradation of chicken feathers, Fakhfakh et al. [11] described the production of wool protein hydrolysate by Bacillus pumilus A1, Fang et al. [12] described the degradation of wool by three keratinolytic enzymes isolated from Stenotrophomonas maltophilia BBE11-1, and Zhang et al. [13] tested the enzymatic degradation of wool using esperase in combination with L-cysteine and urea. Keratinolytic microorganisms have been isolated from various environments, from soils of sheep pens [14], soils enriched with keratin-containing material or keratinous hydrolysate [9], from poultry farms or processing plants [12, 15], from raw wool [16, 17] or directly from healthy sheep [10, 18]. The vast majority of research articles related to keratin-rich waste are focused on waste bird feathers, which represent an even greater environmental problem than waste sheep wool. Some detailed review articles have also been published describing the potential uses of keratinases in various biotechnological applications [19,20,21] but even here sheep wool is only included in a few cases. As we reported in a previous article [8], our review was the first really focused on the use of keratinolytic microorganisms or their enzymes exclusively on sheep wool. Due to the different composition of keratins in the structure of bird feathers, we cannot directly apply these results to sheep wool, so there is a lack of highly active keratinolytic microorganisms capable of degrading the hard structure of wool keratin. Therefore, in this article, we report the isolation of keratinolytic microorganisms from the sheep wool compost pile and the soil under the pile, the identification of the isolates by 16S rRNA genes and whole genome sequencing, and the characterization (determination of the pH and temperature optimum, the molecular weight of keratinases and the effect of various substances on keratinolytic activity) of several new highly keratinolytic bacterial and fungal isolates with high potential for biotechnological approaches of processing waste sheep wool.
Materials and methods
Isolation of keratinolytic microorganisms
Microorganisms were isolated twice from the sheep wool compost pile and the soil under the pile, at room temperature (21 °C), 30 °C and 55 °C. Samples were first inoculated onto skim milk agar (12.5 g/L skim milk; 3 g/L yeast extract; 5 g/L NaCl; 15 g/L agar technical) to detect proteolytic activity. The proteolytic microorganisms were then inoculated onto keratinolytic activity agar with wool keratin as the sole carbon and nitrogen source (KAA-WK) (1.5 g/L K2HPO4 × 3H2O; 0.025 g/L MgSO4 × 7H2O; 0.025 g/L CaCl2; 0.015 g/L FeSO4 × 7H2O; 0.005 g/L ZnSO4 × 7H2O; 2 g/L wool keratin; 15 g/L agar technical) [17]. The KAA-WK plates were incubated under the same conditions as during isolation. The clearing zones produced by the keratinolytic microorganisms were recorded daily for a period of 7 days. The isolates that showed visible clearing zones for three consecutive inoculations at KAA-WK were selected for further studies.
Preliminary identification of bacterial isolates by FAMEs profiles
Selected isolates were cultured in tryptic soy broth or agar at 30 °C for 24 h. Colonies were harvested from agar plates, or pellets were collected from the broth. FAMEs were prepared and analysed strictly according to the guidelines of MIDI Inc. (Newark, USA) for their Microbial Identification System [22] using gas chromatography with flame ionisation detector (GC-FID). For identification of bacteria RTSBA6 and for streptomycete ACTIN1 database was used.
Identification of bacterial isolates by 16S rRNA genes sequencing
DNA was extracted using the DNeasy UltraClean Microbial Kit (Qiagen, Hilden, Germany). The 16S rRNA genes of the bacterial isolates were amplified by PCR using primers fD1 [23] and 1492R [24]. PCR products were sequenced with the same primers by Microsynth AG (Balgach, Switzerland). Sequences were compared with the 16S rRNA database of type and reference bacterial strains in the NCBI database. All obtained 16S rRNA sequences of isolates were deposited in the GenBank database: OQ875782 (S3021), OQ875783 (S3027), OQ875784 (RS308), OQ875785 (RS3014).
Identification of fungal isolate by several taxonomic markers sequencing
Culture morphology was observed on MEA, OA and PDA after 7 and 14 days of incubation at 25 and 37 °C. Microscopic characters of slides prepared with water and 60% lactic acid were observed with Nomarski interference contrast optics on an Olympus microscope BX-51, and images were taken with a DP72 camera (Olympus). DNA was extracted after mechanical lysis in CTAB buffer according to the protocol described by Gerrits van den Ende and de Hoog [25]. The following genes were amplified and Sanger sequenced for identification: internal transcribed spacer 1 and 2 including the 5.8S rDNA (ITS; primer pair ITS1/ITS4 [26]), partial large ribosomal subunit rDNA including its D1/D2 domains (LSU; primer pair NL1/NL4 [26]), partial small ribosomal subunit (SSU; primer pair NS1/NS24 [27]), partial sequences of genes encoding β-tubulin 2 (TUB2; primer pair Btub2Fd/Btub2Rd [28]) and the largest subunit of RNA polymerase II (RPB1; primer pair RPB1-Af/RPB1-Cr [29]). All obtained DNA sequences of isolate S3014 were deposited in the GenBank database: OP303311 (ITS rDNA), OP303343 (D1/D2 domains of LSU rDNA), OP327550 (SSU rDNA), OP321170 (RPB1), OP321169 (bTUB2). Isolate S3014 was deposited in the Ex Culture Collection of the Infrastructural Centre Mycosmo, MRIC UL, Slovenia, at the Department of Biology, Biotechnical Faculty, University of Ljubljana (http://www.ex-genebank.com) as EXF-15693.
Whole genome sequencing (WGS) of bacterial isolates
DNA was isolated using the DNeasy UltraClean Microbial Kit (Qiagen, Hilden, Germany). Whole genomes were sequenced by Novogene (Beijing, China) using Illumina MiSeq paired end technology. The quality of raw reads was checked using the FASTQC tool, version 0.11.2 [30]. Assembly was done by SPAdes Assembler, version 3.1.1 [31]. Final identification was performed by calculating average nucleotide identity (ANI) [32] for whole genome comparisons with selected type strains using pyani [33]. Genome annotations were performed using Prokka 1.7 [34]. Core genome analysis was done with Roary tool [35]. Phylogenetic trees were prepared with Molecular Evolutionary Genetics Analysis (MEGA 11) software. The genome sequences were deposited in the GenBank database under BioProject: PRJNA970193 under accession numbers JASDBW000000000 (S3018), JASDBX000000000 (S3021), JASDBY000000000 (S3027), JASDBO000000000 (RS308), and JASDBR000000000 (RS3014). Accession numbers for other sequenced isolates are listed in Online resource 1.
Keratinolytic activity determination
All isolates were tested for keratinolytic activity by various qualitative methods, by growing on media containing wool keratin powder (KAA-WK), raw wool, or keratin azure. Selected isolates were further tested by a quantitative method by detecting the increase in soluble proteins using the modified method of Bohacz and Korniłłowicz-Kowalska [36]. Isolates were grown in keratinolytic activity broth with raw wool as the sole source of carbon and nitrogen (KAB-RW) (1.5 g/L K2HPO4 × 3H2O; 0.025 g/L MgSO4 × 7H2O; 0.025 g/L CaCl2; 0.015 g/L FeSO4 × 7H2O; 0.005 g/L ZnSO4 × 7H2O; 1.5 g/L raw wool) for 5 days at 30 °C and 150 rpm. Although different temperatures were used in the isolation, the most active isolates included in further studies were isolated at 30 °C, therefore, all the following studies were performed at 30 °C and 150 rpm. The culture samples were centrifuged (14,000 g, 5 min) and the supernatants were analysed for soluble protein concentration and used as an enzyme source for keratinolytic activity assays. Soluble protein concentration was determined by measuring absorbance at 280 nm [37]. Absorbance was measured with the Spark 10 M microplate reader (Tecan, Austria) using a reference blank of sterile medium. Soluble protein concentration was calculated using bovine serum albumin (BSA) standard curve. To test keratinolytic activity, 500 µL of the culture supernatant was mixed with 500 µL of 1% (w/v) wool keratin (TCI, Japan) in 50 mM TRIS–HCl buffer (pH 8.0). The mixtures were incubated at 55 °C and 50 rpm for 1 h. After incubation, the reaction was terminated with 1 mL of 10% (w/v) trichloroacetic acid (TCA), then the mixtures were incubated at 4 °C for 30 min. Controls were treated the same way, except that the TCA was added before incubation. Samples were centrifuged (14,000 g, 3 min) and the supernatants were used to detect the concentration of soluble proteins by measuring absorbance at 280 nm [37]. One unit of keratinolytic activity (KAU/mg) was defined as the amount of enzyme per mg of soluble protein in one mL of culture supernatant that increases the absorbance at 280 nm for 0.01 in one hour under the assay conditions.
Detection of optimum temperature and pH range of keratinolytic activity
Optimal pH for keratinolytic activity of selected isolates was determined using the method described above, with culture supernatants as source of enzymes, at 55 °C, except that 1% (w/v) wool keratin solutions were prepared with different buffers in the pH range 3.0–10.0 (i) 0.1 M glycine–HCl buffer for pH 3.0, (ii) 0.1 M acetate buffer for pH 4.0, 5.0, (iii) 0.1 M phosphate buffer for pH 6.0, 7.0, (iv) 0.1 M Tris–HCl buffer for pH 8.0, 9.0 and (v) 0.1 M glycine–NaOH buffer for pH 10.0. The optimum temperature was also determined using the same method, in 50 mM TRIS–HCl buffer (pH 8.0), except that the culture supernatant and keratin mixtures were incubated at different temperatures ranging from 30–90 °C.
Effect of inhibitors, detergents, oxidizing agents and metal ions on keratinolytic activity
The effect of various inhibitors (EDTA, β-mercaptoethanol; 1 mM and 10 mM), detergents (Tween 20, Triton X-100, SDS; 0.1% (w/v) and 1% (w/v)), organic solvents (DMSO; 0.1% (w/v) and 1% (w/v)), monovalent cations (Na+, K+; 1 mM and 10 mM) and divalent cations (Fe2+, Mg2+, Zn2+, Cu2+, Co2+, Mn2+; 1 mM and 10 mM) on keratinolytic activity was studied by pre-incubating the culture supernatant with these chemicals for 30 min at room temperature. After incubation, 500 µL of 1% (w/v) wool keratin (TCI, Japan) in 50 mM TRIS–HCl buffer (pH 8.0) was added to the reactions and keratinolytic activity was detected by method described above. Keratinolytic activity obtained without additives was considered as 100%.
Purification of keratinases by ion exchange high-performance liquid chromatography (HPLC)
After growing, microbial cultures were centrifuged and microfiltered (filter (Sartorius, Germany) with 0.22 µm pore size). The cell-free supernatants (CFS) were stored at − 25 °C until use. Prior to use the CFS were thawed and re-microfiltered as needed (0.22 µm CA syringe filters). Approximately 100 mL of each CFS (pH approximately 8.5) was loaded onto a 1 mL CIMmultus™ QA-1 anion exchange column (BIA Separations, Slovenia, now part of Sartorius) using the 10 mL mobile phase pump of a Knauer Smartline HPLC system (Knauer, Germany) with a 2550 UV/VIS detector with deuterium and halogen lamp. The column was equilibrated in 25 mM TRIS pH 8.5 (mobile phase A1). We collected the flow-through fraction and refrigerated or frozen it until further use. After loading, the column was washed with mobile phase A1 and the bound material was eluted with a gradient of 0–50% mobile phase B (25 mM TRIS, 2 M NaCl, pH 8.5) in 6 min, followed by a steeper gradient from 50–100% of mobile phase B in 1 min. We manually collected the eluted fractions and then washed the column with 100% mobile phase B for 2 min before re-equilibrating the column in mobile phase A1 for the optional next run. We monitored absorbance at 226 nm and 280 nm and maintained the flow rate at 5 mL per minute throughout the experiment. The total collected flow-through from the anion exchange chromatography was further fractionated in a second chromatography step, using the CIMmultus™ SO3-1 cation-exchange column from the same manufacturer. The pH of the feed was adjusted to 6.5 with 1 M NaOH and loaded onto the column, equilibrated in 25 mM TRIS pH 6.5 (mobile phase A2). We collected the flow-through and then eluted the bound material using mobile phase A2 and the same mobile phase B and gradient elution program as above. All collected fractions were tested for keratinolytic activity using the method described before.
HPLC fractions separation by SDS-PAGE
SDS-PAGE was performed with 4% stacking gel and 12.5% separating gel according to the method described by Laemmli [38]. All HPLC fractions were mixed with loading buffer (0.5 mM Tris–HCl, pH 6.8; 20% (v/v) glycerol; 2% (w/v) SDS; 5% (v/v) β-mercaptoethanol; 0.006% (w/v) bromophenol blue) and heated at 100 °C for 5 min before electrophoresis. The gel was stained with 0.1% (w/v) Coomassie Brilliant Blue R-250 in 60:30:10 ratio of water/methanol/acetic acid and destained in the same solution without dye.
Results and discussion
Isolation and identification of bacterial isolates
A total of 116 keratinolytic microbial strains were obtained from both isolations. 60 isolates were isolated at 30 °C, 12 isolates at 55 °C and 44 isolates at room temperature (21 °C). After preliminary testing for keratinolytic activity (detection of visible clearing zones (Fig. 1) on media containing wool keratin), 38 isolates (all of them were isolated at 30 °C) with the largest clearing zones were selected for identification and further studies. Sequencing of 16S rRNA genes revealed 33 isolates from the genus Bacillus, 3 isolates from the genus Stenotrophomonas, and 1 isolate each from the genera Pseudomonas and Streptomyces. Isolation of new bacterial and fungal strains from sheep wool or wool keratin-enriched soil has been reported before. Queiroga et al. [10] reported the isolation of 156 strains from raw Portuguese Merino wool, and further research of Queiroga et al. [39] yielded 115 caseinolytic isolates, including 5 with high keratinolytic activity. Analysis of 16S rRNA genes revealed that they all belonged to the genus Bacillus and showed a high degree of homology with B. subtilis/licheniformis. Two additional Bacillus strains capable of growing on basal salt medium with wool as the sole carbon and nitrogen source were isolated from raw wool by Infante et al. [16]. They were identified as B. megaterium and B. thuringiensis by sequencing the 16S rRNA gene. Iglesias et al. [17] described the isolation of 66 proteolytic isolates out of 135 bacterial isolates from Patagonian Merino wool. They belonged to the genera Bacillus, Exiguobacterium, Deinococcus and Micrococcus. The highest wool-keratinolytic activity was observed in Bacillus sp. G51.
Preliminary identification of keratinolytic isolates by FAMEs profiles at the genus level gave exactly the same results as sequencing of 16S rRNA genes. More precise identification was not possible, due to the high similarity of profiles within the same genus and the lack of data for different Bacillus species until then. Notwithstanding the results of Slabbinck et al. [40], who showed that machine learning proved to be very useful for bacterial species identification based on FAMEs profiles, Santos et al. [41] successfully identified Bacillus strains using gas chromatography only after replacing the originally used FID detector with a vacuum ultraviolet detector (GC-VUV). Our 16 bacterial isolates (15 Bacillus and 1 Streptomyces) that showed better keratinolytic activity than the positive control (Streptomyces fradiae, DSM 40758) were selected for WGS. According to the WGS results (Online resource 1), the sequenced isolates belonged to four different Bacillus species (B. altitudinis, B. mycoides, B. subtilis, B. wiedmannii) and one Streptomyces species (S. coelicoflavus). The most active strains of each microbial species (Table 1) were isolate RS3014 with an identity of 0.979 to B. altitudinis strain GR-8, isolate S3021 with an identity of 0.972 to B. mycoides strain BPN36-3, isolate RS308 with an identity of 0.986 to B. subtilis subsp. subtilis strain 168, isolate S3027 with an identity of 0.986 to B. wiedmannii strain SR 52, and isolate S3018 with an identity of 0.966 to S. coelicoflavus strain NBRC 15399 (Online resource 2).
Core genome analysis revealed greater similarity between B. mycoides and B. wiedmannii (Online resource 3), which is to be expected given that both belong to the larger B. cereus group, while other isolates belong to the larger Bacillus subtilis group. As mentioned earlier, many species of the genus Bacillus with keratinolytic activity have already been described, but most of them are related to the decomposition of bird feathers and only a few to the decomposition of sheep wool. B. altitudinis, isolated from a poultry farm [42], was the source of the thermostable keratinase KBALT, expressed in recombinant E. coli. The wool degrading B. subtilis MA21 was isolated from soil and well characterized by Hassan et al. [43]. Kim et al. [44] reported the isolation of three Bacillus species (B. pumilis, B. cereus and B. subtilis) with great potential for degrading feathers. B. subtilis has also been used as a recombinant strain for keratinase expression [45, 46]. The keratinolytic activity of B. mycoides and B. wiedmannii has never been reported before. Jadhav et al. [47] reported the isolation of a keratinolytic isolate identified as Streptomyces coelicoflavus.
Isolation and identification of the fungal isolate
A single fungal strain S3014 was isolated from sheep wool on Skim milk agar and KAA-WK medium incubated at 30 °C. Its white to off-white colonies reached 26/25/25 and 10/7/5 mm in diameter on PDA/MEA/OA in 7 days at 25 °C and 37 °C, respectively. The thin-walled hyphae disintegrated into arthroconidia on OA and PDA, whereas no sporulation was observed on MEA. The rhexolytically dehiscent arthroconidia were smooth, thin-walled, pear-shaped, oblong or cylindrical, 4 (3‒5) × 8 (5‒14) µm (Fig. 2). The sexual stage was not observed.
Isolate S3014 was identified as Aphanoascus reticulisporus (Onygenaceae, Onygenales, Eurotiomycetidae, Eurotiomycetes, Pezizomycotina, Ascomycota), based on above described morphological characteristics and ITS rDNA sequence comparisons (Fig. 3, Table 1). The species is considered heterothallic, what explains the absence of ascomata in our isolate. However, Cano and Guarro [48] reported homothallic strains as well. Although the main morphological characters of the species, the size of the ascospores and their regularly reticulate surface, could not be studied, morphology of colonies and abundant production of arthroconidia corresponds to the species description. Identification of Aphanoascus species is most often based on ribosomal gene sequences such as ITS or LSU [49,50,51], while partial protein-encoding genes are sporadically available only for certain isolates and not for the ex-type strain of Aphanoascus reticulisporus, CBS 392.67 [52, 53], thus not allowing systematic comparisons. Phylogenetic placement of strain S3014 is therefore presented based on ITS sequences (Fig. 3).
The SSU sequence of the strain S3014 was identical to sequences of the strains of A. fulvescens (strain NBRC 30411; JN941601) and A. reticulosporus (strain NRBC 32373; JN941598), and differed in 1 of 1542 nt from the type strain of A. reticulisporus (strain NBRC 32372; JN941599) and A. verrucosus (strain NBRC 32382; JN941595). The LSU sequence was 100% identical to the strain A. reticulisporus (strain NBRC 32372 T, CBS 392.67; JN941549, JN941549), 99.4% similar to A. keratinophilum (strain CBS 104.62 T, NG 064030), and 98.7% to A. verrucosus (strain NBRC 32381; JN941553). The ITS sequence was 100% identical to A. reticulisporus (strain NBRC 32372 T; JN943434), 99.3–99.7% to the other strains idnentified as A. reticulisporum and Chrysosporium articulatum, but also had a high similarity to the sequences of type strains of A. keratinophilum (99.2% to CBS 104.62; MH838113), A. jingzhouense (98.9% to GZAC EB1303M; NR 158334), and was also closely related to A. fulvescens, and A. verrucosus (Fig. 4). The RPB1 sequence of the strain was 99.4% identical to the type strain of A. reticulisporus (NBRC 32372 T, JN992357), and differed by 6.5% to A. keratinophilum type strain sequence (CBS 104.62 T), and by 2.8–7.9% nts to A. fulvescens (NBRC 31723, JN992358; NRBC 8390, JN992361). The closest bTUB2 sequence available from GenBank was of A. keratinophilus (strain CBS 219.62 T, KT155513) and had 9.4% differing substitutions in comparison to the strain S3014, while the bTUB2 sequences of A. reticulisporus were not available for comparison.
The species A. reticulisporus was described as Anixiopsis reticulispora from arable soil in New Zealand; it was combined into Aphanoascus by Hubalek [54]. Cano and Guarro [48] examined numerous isolates from agricultural soils in Spain and accepted A. reticulisporus as a species. Vidal et al. [49] proposed the synonymy of this species with C. articulatum based on ITS sequences. Identity at the species level is also evident from Fig. 3. Although numerous isolates have been studied from soil [48], only a limited number of strains are available in culture collections. Nevertheless, the data suggest that it is a cosmopolitan species. Aphanoascus and other fungi previously assigned to the genus Chrysosporium [48, 49, 52] are otherwise known soil fungi that decompose keratin. The prevalence of these genera on sheep wool is also evident from the study of Abdel-Gawad [18], who reported Aphanoascus keratinophilus (before C. keratinophilum), Aphanoascus terreus (before C. indicum), and Chrysosporium (C. tropicum) as the most abundant fungi.
Keratinolytic activity
The qualitative methods used for preliminary detection of keratinolytic activity revealed clear evidence of keratin degradation in the form of clearing zones around colonies (Fig. 1) on agar plates with wool keratin as the sole C and N source, as well as visible damage on keratin azure fibers (Fig. 4) viewed under the microscope.
B. wiedmannii (S3027) was the most active among the isolates with a keratinolytic activity of 600.1 KAU/mg, followed by B. mycoides, B. subtilis, and B. altitudinis with keratinolytic activities of 529.3, 489.4, and 415.5 KAU/mg, respectively. Fungal and streptomycete isolates with keratinolytic activities of 237.5 and 259.4 KAU/mg were the least active isolates (Fig. 5).
Keratinolytic activity of the most active isolates of each species, after 5 days of cultivation in media with wool as the sole source of carbon and nitrogen at 30 °C and 150 rpm. Activity was determined by measuring absorbance at 280 nm after 1 h of incubation at 55 °C (S3014 – A. reticulisporus, S3018 – S. coelicoflavus, S3021 – B. mycoides, S3027 – B. wiedmannii, RS308 – B. subtilis, RS3014 – B. altitudinis). Results are the average of 3 replicates with error bars to indicate the standard deviation (n = 3)
Characterization of the keratinases
Since the method of Bohacz and Korniłłowicz-Kowalska [36] proved to be a successful quantitative method for determining keratinolytic activity, we also used it for enzymatic characterization (determination of optimal temperature and pH, and verification of the effect of various chemicals on keratinolytic activity). For B. mycoides and B. wiedmannii, a temperature 55 °C was optimal, whereas for the other isolates the optimal temperature was 65 °C (Fig. 6a). Temperature did not have a strong effect on activity, as activity was detected at all tested temperatures. For most isolates, relative activity greater than 50% was detected over a range between 35 °C and 85 °C, while relative activity greater than 80% was detected between 55 °C and 75 °C. A greater effect of temperature was noted for fungal and streptomycete isolates, particularly at temperatures of 75 °C and above. A greater effect of pH on activity was noted. None of the isolates showed activity at pH below 5. The optimal pH was 8.0 for most isolates and 10.0 for A. reticulisporus and B. altitudinis (Fig. 6b). Our results are similar to those of Iglesias et al. [17], who reported the best keratinolytic activity of Bacillus sp.G51 at 60 °C and pH 9, and to those of Hassan et al. [43], who reported the isolation of a B. subtilis strain with the best activity at 60 °C and pH 9.0, and partially to those of Gegeckas et al. [55], who described the keratinase BtKER from B. thuringiensis with the best activity at 30 °C and pH 7.0. Kim et al. [44] described the optimal conditions for enzyme production by B. subtilis at 40 °C and pH 5–9, but they did not describe the optimal conditions for enzyme activity. Pawar et al. [42] reported the highest activity of B. altitudinis keratinase KBALT (tested at 55 °C) at pH 8.0, while it was still very high at pH 9.0. The keratinase was highly active at temperatures ranging from 65 to 95 °C, with an optimum at 85 °C (tested at pH 8.0), which is very similar to the keratinase from our B. altitudinis isolate with optimum at 65 °C and pH 8.0. S. coelicoflavus isolated by Jadhav et al. [47] was most active at 40 °C and pH from 6.0 to 9.0 with an optimum at 8.0. The keratinolytic enzymes of A. reticulisporus have not yet been characterized, so no data are available on the optimal pH and temperature for keratinolytic activity.
Effect of temperature (a) and pH (b) on keratinase activity of isolated microorganisms (♦ A. reticulisporus – S3014, ◊ S. coelicoflavus – S3018, ● B. mycoides – S3021, ○ B. wiedmannii – S3027, ■ B. subtilis – RS308, □ B. altitudinis – RS3014). Optimal pH was determined at 55 °C in the pH range 3.0–10.0 and the optimal temperature was determined at pH 8.0 at temperatures ranging from 30–90 °C. The maximum KAU value for an individual strain was set as 100%. Results are the average of 3 replicates with error bars to indicate the standard deviation (n = 3)
Table 2 shows the results of the influence of different substances that keratinases can potentially come into contact with in different applications and that can affect their activity both in a positive and a negative way. As expected, the addition of EDTA had an inhibitory effect on the keratinolytic activity of all 6 isolates, whereas the addition of β-mercaptoethanol (β-ME) increased the activity in most isolates, even by almost 30% in S. coelicoflavus and B. subtilis. The effect of detergents on keratinolytic activity was mostly very similar among the isolates. While Triton X-100 and Tween 20 showed no significant effect on activity, the addition of SDS significantly decreased activity, especially at a concentration of 1% (w/v), where it was zero in some cases. The effect of DMSO increased the keratinolytic activity with increasing concentration. Monovalent cations increased the activity of most isolates, especially at a concentration of 1 mM. The addition of Zn2+ and Cu2+ cations decreased the keratinolytic activity in all cases. Co2+ and Mn2+ had no effect on the activity, or it was only minimally increased. No differences were observed for cations tested in combination with different anions (Cl− or SO42−), so it can be assumed that the differences in activity are due to the presence of cations. The influence of Fe2+ cations on activity was difficult to detect. We assume that the presence of Fe2+ cations interferes with the measurements of proteins in the UV spectrum at 280 nm, so the results are not reliable. S. coelicoflavus, described by Jadhav et al. [47] had very similar results when they tested the effect of DMSO on keratinolytic activity, showing a minimal increase in activity (11%) at a DMSO concentration of 5 mM, while we found an increase in activity of 13.6% at 10 mM. In their research, the addition of SDS and EDTA also caused a minimal increase in activity of 5% and 9%, respectively, while in our case, the addition of both substances significantly reduced the activity to 0–50% of initial activity. Similar differences also exist with the addition of different metal cations. In our case, the addition of all metal cations, except Mg2+ and Co2+, which were not tested, resulted in a significant decrease in keratinolytic activity, while in their case, the addition of various metal cations had no significant effect on the activity or even increased it. Our results in B. altitudinis are broadly similar to those of Pawar et al. [42] describing KBALT keratinase. In both experiments, the addition of Mg2+ cations significantly increased keratinolytic activity (up to 20%), while the effect of DMSO and β-ME on increasing keratinolytic activity was greater in our case (where the DMSO concentration was the same as in our case, but β-ME concentration was higher in our case). Mn2+ cations had no significant effect in any case. Greater differences occurred with the addition of Zn2+, which decreased activity in our case and increased it in theirs, and with EDTA and SDS, where the effect was reversed. Hassan et al. [43], in characterizing the keratinolytic protease of B. subtilis, found a negative effect at a concentration of 5 mM when EDTA was added, while 1 mM had no effect on the activity and even 1 mM greatly reduced the activity in our case. The addition of SDS and Cu2+ cations caused the same effect as in our case, both at a lower and at a higher concentration. Zn2+ cations in combination with Cl− had no effect on the activity, while in our case they had a negative effect on the activity. The opposite effect was caused by the addition of Mg2+, Mn2+, Tween 20 and Triton X-100, which showed no effect on the activity in our case, but Hassan et al. [43]reported a decreased activity. In their case, the addition of β-ME and DMSO had a negative effect on the activity, while in our case it was positive, especially at higher concentrations. Since the keratinolytic enzymes of A. reticulisporus have not been characterized so far, there are also no data on the influence of different substances on keratinolytic activity.
Preliminary molecular weights of detected keratinases
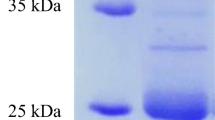
The molecular weight of potential keratinases was successfully determined by two-step ion exchange chromatography and separation of fractions on SDS-PAGE for all isolates except B. subtilis. The molecular weight of the single protein in the active fraction was determined to be approximately 55 kDa for A. reticulisporus, and approximately 30 kDa for S. coelicoflavus. The molecular weights of keratinases from Bacillus isolates were 27, 32, and 35 kDa for B. wiedmannii, B. altitudinis, and B. mycoides, respectively. An example of the molecular weight determination for isolate S3021 (B. mycoides) is shown in Fig. 7, results for other isolates are presented in Online resource 4.
a Chromatogram of the culture supernatant of B. mycoides (S3021) from the second purification step on ion exchange HPLC (absorbance at 226 nm). The obtained fractions are separated by vertical lines and the corresponding keratinolytic activities (KAU/mg) are shown for each fraction (●). Results are the average of 3 replicates with error bars to indicate the standard deviation (n = 3), b SDS-PAGE showing possible keratinase from the isolate B. mycoides after two-step ion exchange chromatography purification. Lane 1 Protein ladder with labelled molecular weights in kDa, lane 2 10 × concentrated load on the cation exchange column, lane 3 Fraction 8 (*) from cation exchange chromatography (−35 kDa)
The molecular weights of the known keratinases vary widely. As reported by Brandelli et al. [56], they range from 18 to 240 kDa. Among Bacillus species different sizes have been reported, Yoshioka et al. [57] described 26 kDa keratinase from B. licheniformis, Radha and Gunasekaran [58] described 30 kDa keratinase in B. megaterium, Kumar et al. [59] described 65 kDa keratinase in B. pumilus, and Balaji et al. [60] described 69 kDa keratinase in B. subtilis. Meanwhile, Gegeckas et al. [55] reported the isolation and characterization of the 39 kDa keratinolytic proteinase BtKER from B. thuringiensis AD-12, which, like our isolates B. mycoides and B. wiedmannii, belong to the B. cereus group. The molecular weights of the keratinases in S. coelicoflavus and A. reticulisporus have never been reported before.
Conclusions
Soils and natural materials containing keratin, such as sheep wool, harbour a variety of interesting and previously undiscovered keratin-degrading microorganisms and their beneficial and previously undescribed keratinolytic enzymes. In accordance with this prediction, we succeeded in isolating and identifying 16 microbial isolates with high keratinolytic activity from the sheep wool compost pile and the soil under the pile, belonging to six known microbial species (Bacillus altitudinis, B. mycoides, B. subtilis, B. wiedmannii, Streptomyces coelicoflavus, and Aphanoascus reticulisporus). All of them showed great potential for further use in various biotechnological applications due to their properties. The keratinases of the new isolates showed high keratinolytic activity over a wide temperature range (25–85 °C for most isolates and 25–75 °C for S. coelicoflavus and A. reticulisporus) and pH range (6.0–10.0 for most isolates and 6.0–9.0 for B. mycoides and B. wiedmannii), which is beneficial for various industrial applications.
For the species B. mycoides and B. wiedmannii keratinolytic activity on any of the keratin substrates has never been described before. And for most of the isolated species, no degradation of sheep wool keratin has been reported. For B. mycoides and B. wiedmannii, this is also the first characterization of keratinases ever described, whereas for A. reticulisporus and S. coelicoflavus, this is the first comprehensive characterization of keratinases using sheep wool keratin, since all previous studies were performed on bird feathers.
The new isolates or their purified enzymes could potentially be used as an additive in the fertilisation of cultivated land with sheep wool to accelerate the release of nutrients (especially nitrogen), as a starter culture in the composting of waste sheep wool and waste woollen textiles, or for aerobic microbial or enzymatic pretreatment of waste sheep wool and animal hair for biogas production to accelerate the process and increase methane yield. The newly described enzymes could also find applications in the cosmetic and medical industries.
References
Callegaro K, Brandelli A, Daroit DJ (2019) Beyond plucking: feathers bioprocessing into valuable protein hydrolysates. Waste Manag 95:399–415. https://doi.org/10.1016/j.wasman.2019.06.040
Lange L, Huang Y, Busk PK (2016) Microbial decomposition of keratin in nature—a new hypothesis of industrial relevance. Appl Microbiol Biotechnol 100:2083–2096. https://doi.org/10.1007/s00253-015-7262-1
Qiu J, Wilkens C, Barrett K, Meyer AS (2020) Microbial enzymes catalyzing keratin degradation: classification, structure, function. Biotechnol Adv 44:107607. https://doi.org/10.1016/j.biotechadv.2020.107607
Daroit DJ, Brandelli A (2014) A current assessment on the production of bacterial keratinases. Crit Rev Biotechnol 34:372–384. https://doi.org/10.3109/07388551.2013.794768
Vidmar B, Vodovnik M (2018) Microbial keratinases: enzymes with promising biotechnological applications. Food Technol Biotechnol 56:312–328. https://doi.org/10.17113/ftb.56.03.18.5658
Zoccola M, Montarsolo A, Mossotti R et al (2015) Green hydrolysis as an emerging technology to turn wool waste into organic nitrogen fertilizer. Waste Biomass Valorization 6:891–897. https://doi.org/10.1007/s12649-015-9393-0
Bhavsar P, Zoccola M, Patrucco A et al (2016) Superheated water hydrolysis of waste wool in a semi-industrial reactor to obtain nitrogen fertilizers. ACS Sustain Chem Eng 4:6722–6731. https://doi.org/10.1021/acssuschemeng.6b01664
Petek B, Marinšek Logar R (2021) Management of waste sheep wool as valuable organic substrate in European Union countries. J Mater Cycles Waste Manag 23:44–54. https://doi.org/10.1007/s10163-020-01121-3
Gousterova A, Braikova D, Goshev I et al (2005) Degradation of keratin and collagen containing wastes by newly isolated thermoactinomycetes or by alkaline hydrolysis. Lett Appl Microbiol 40:335–340. https://doi.org/10.1111/j.1472-765X.2005.01692.x
Queiroga AC, Pintado MM, Malcata FX (2007) Novel microbial-mediated modifications of wool. Enzyme Microb Technol 40:1491–1495. https://doi.org/10.1016/j.enzmictec.2006.10.037
Fakhfakh N, Ktari N, Siala R, Nasri M (2013) Wool-waste valorization: production of protein hydrolysate with high antioxidative potential by fermentation with a new keratinolytic bacterium, Bacillus pumilus A1. J Appl Microbiol 115:424–433. https://doi.org/10.1111/jam.12246
Fang Z, Zhang J, Liu B et al (2013) Biochemical characterization of three keratinolytic enzymes from Stenotrophomonas maltophilia BBE11-1 for biodegrading keratin wastes. Int Biodeterior Biodegrad 82:166–172. https://doi.org/10.1016/j.ibiod.2013.03.008
Zhang N, Wang Q, Yuan J et al (2018) Highly efficient and eco-friendly wool degradation by L-cysteine-assisted esperase. J Clean Prod 192:433–442. https://doi.org/10.1016/j.jclepro.2018.05.008
Zhang R-X, Gong J-S, Su C et al (2016) Biochemical characterization of a novel surfactant-stable serine keratinase with no collagenase activity from Brevibacillus parabrevis CGMCC 10798. Int J Biol Macromol 93:843–851. https://doi.org/10.1016/j.ijbiomac.2016.09.063
Riffel A, Brandelli A (2002) Isolation and characterization of a feather-degrading bacterium from the poultry processing industry. J Ind Microbiol Biotechnol 29:255–258. https://doi.org/10.1038/sj.jim.7000307
Infante I, Morel MA, Ubalde MC et al (2010) Wool-degrading Bacillus isolates: extracellular protease production for microbial processing of fabrics. World J Microbiol Biotechnol 26:1047–1052. https://doi.org/10.1007/s11274-009-0268-z
Iglesias MS, Sequeiros C, García S, Olivera NL (2017) Newly isolated Bacillus sp. G51 from Patagonian wool produces an enzyme combination suitable for felt-resist treatments of organic wool. Bioprocess Biosyst Eng 40:833–842. https://doi.org/10.1007/s00449-017-1748-4
Abdel-Gawad KM (1997) Mycological and some physiological studies of keratinophilic and other moulds associated with sheep wool. Microbiol Res 152:181–188. https://doi.org/10.1016/S0944-5013(97)80011-0
Gupta R, Ramnani P (2006) Microbial keratinases and their prospective applications: an overview. Appl Microbiol Biotechnol 70:21–33. https://doi.org/10.1007/s00253-005-0239-8
Anbesaw MS (2022) Bioconversion of keratin wastes using keratinolytic microorganisms to generate value-added products. Int J Biomater 2022:2048031. https://doi.org/10.1155/2022/2048031
Hassan MA, Abol-Fotouh D, Omer AM et al (2020) Comprehensive insights into microbial keratinases and their implication in various biotechnological and industrial sectors: A review. Int J Biol Macromol 154:567–583. https://doi.org/10.1016/j.ijbiomac.2020.03.116
Sasser M (1990) Identification of bacteria by gas chromatography of cellular fatty acids. MIDI Technical Note 101, Newark
Weisburg WG, Barns SM, Pelletier DA, Lane DJ (1991) 16S ribosomal DNA amplification for phylogenetic study. J Bacteriol 173:697–703. https://doi.org/10.1128/jb.173.2.697-703.1991
Lane DJ (1991) 16S/23S rRNA sequencing. In: Stackebrandt E, Goodfellow M (eds) Nucleic Acid Techniques in Bacterial Systematics. John Wiley and Sons, Chichester, England, pp 115–148
Gerrits van den Ende AHG, de Hoog GS (1999) Variability and molecular diagnostics of the neurotropic species Cladophialophora bantiana. Stud Mycol 43:151–162
White TJ, Bruns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ (eds) PCR protocols: a guide to methods and applications. Academic Press, San Diego, California, pp 315–322
Gargas A, Taylor JW (1992) Polymerase chain reaction (PCR) primers for amplifying and sequencing nuclear 18s rDNA from lichenized fungi. Mycologia 84:589–592. https://doi.org/10.2307/3760327
Woudenberg JHC, Aveskamp MM, De Gruyter J et al (2009) Multiple didymella teleomorphs are linked to the phoma clematidina morphotype. Persoonia-Mol Phylogeny Evol Fungi 22:56–62. https://doi.org/10.3767/003158509X427808
Wang QM, Theelen B, Groenewald M et al (2014) Moniliellomycetes and Malasseziomycetes, two new classes in Ustilaginomycotina. Persoonia-Mol Phylogeny Evol Fungi 33:41–47. https://doi.org/10.3767/003158514X682313
Andrews S, Lindenbaum P, Howard B, Ewels P (2011) FastQC high throughput sequence QC report Version 0.11.2
Bankevich A, Nurk S, Antipov D et al (2012) SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J Comput Biol 19:455–477. https://doi.org/10.1089/cmb.2012.0021
Richter M, Rosselló-Móra R (2009) Shifting the genomic gold standard for the prokaryotic species definition. Proc Natl Acad Sci 106:19126–19131. https://doi.org/10.1073/pnas.0906412106
Pritchard L, Glover RH, Humphris S et al (2016) Genomics and taxonomy in diagnostics for food security: soft-rotting enterobacterial plant pathogens. Anal Methods 8:12–24. https://doi.org/10.1039/C5AY02550H
Seemann T (2014) Prokka: rapid prokaryotic genome annotation. Bioinformatics 30:2068–2069. https://doi.org/10.1093/bioinformatics/btu153
Page AJ, Cummins CA, Hunt M et al (2015) Roary: rapid large-scale prokaryote pan genome analysis. Bioinformatics 31:3691–3693. https://doi.org/10.1093/bioinformatics/btv421
Bohacz J, Korniłłowicz-Kowalska T (2019) Fungal diversity and keratinolytic activity of fungi from lignocellulosic composts with chicken feathers. Process Biochem 80:119–128. https://doi.org/10.1016/j.procbio.2019.02.012
de Paiva DP, de Oliveira SS, Mazotto AM et al (2018) Keratinolytic activity of Bacillus subtilis LFB-FIOCRUZ 1266 enhanced by whole-cell mutagenesis. 3 Biotech. https://doi.org/10.1007/s13205-018-1527-1
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of Bacteriophage T4. Nature 227:680–685. https://doi.org/10.1038/227680a0
Queiroga AC, Pintado ME, Malcata FX (2012) Potential use of wool-associated Bacillus species for biodegradation of keratinous materials. Int Biodeterior Biodegrad 70:60–65. https://doi.org/10.1016/j.ibiod.2011.12.013
Slabbinck B, De Baets B, Dawyndt P, De Vos P (2009) Towards large-scale FAME-based bacterial species identification using machine learning techniques. Syst Appl Microbiol 32:163–176. https://doi.org/10.1016/j.syapm.2009.01.003
Santos IC, Smuts J, Choi W-S et al (2018) Analysis of bacterial FAMEs using gas chromatography – vacuum ultraviolet spectroscopy for the identification and discrimination of bacteria. Talanta 182:536–543. https://doi.org/10.1016/j.talanta.2018.01.074
Pawar VA, Prajapati AS, Akhani RC et al (2018) Molecular and biochemical characterization of a thermostable keratinase from Bacillus altitudinis RBDV1. 3 Biotech. https://doi.org/10.1007/s13205-018-1130-5
Hassan MA, Haroun BM, Amara AA, Serour EA (2013) Production and characterization of keratinolytic protease from new wool-degrading Bacillus species isolated from Egyptian ecosystem. BioMed Res Int 2013:175012. https://doi.org/10.1155/2013/175012
Kim JM, Lim WJ, Suh HJ (2001) Feather-degrading Bacillus species from poultry waste. Process Biochem 37:287–291. https://doi.org/10.1016/S0032-9592(01)00206-0
Liu B, Zhang J, Li B et al (2013) Expression and characterization of extreme alkaline, oxidation-resistant keratinase from Bacillus licheniformis in recombinant Bacillus subtilis WB600 expression system and its application in wool fiber processing. World J Microbiol Biotechnol 29:825–832. https://doi.org/10.1007/s11274-012-1237-5
Dong Y-Z, Chang W-S, Chen PT (2017) Characterization and overexpression of a novel keratinase from Bacillus polyfermenticus B4 in recombinant Bacillus subtilis. Bioresour Bioprocess 4:47. https://doi.org/10.1186/s40643-017-0177-1
Jadhav RS, Karad DD, Kulakrni SW (2016) Isolation, identification and characterization of keratinolytic Streptomyces coelicoflavus. Int J Curr Microbiol App Sci 5:153–163. https://doi.org/10.20546/ijcmas.2016.507.015
Cano J, Guarro J (1990) The genus Aphanoascus. Mycol Res 94:355–377. https://doi.org/10.1016/S0953-7562(09)80361-4
Vidal P, de Vinuesa los AM, Sánchez-Puelles JM, Guarro J (2000) Phylogeny of the anamorphic genus Chrysosporium and related taxa based on rDNA internal transcribed spacer sequences. Rev Iberoam Micol 17:22–29
Sutton DA, Marín Y, Thompson EH et al (2013) Isolation and characterization of a new fungal genus and species, Aphanoascella galapagosensis, from carapace keratitis of a Galapagos tortoise (Chelonoidis nigra microphyes). Med Mycol 51:113–120. https://doi.org/10.3109/13693786.2012.701767
Labuda R, Bernreiter A, Hochenauer D et al (2021) Molecular systematics of Keratinophyton: the inclusion of species formerly referred to Chrysosporium and description of four new species. IMA Fungus 12:17. https://doi.org/10.1186/s43008-021-00070-2
Kandemir H, Dukik K, de Melo TM et al (2022) Phylogenetic and ecological reevaluation of the order Onygenales. Fungal Divers 115:1–72. https://doi.org/10.1007/s13225-022-00506-z
Schoch CL, Seifert KA, Huhndorf S et al (2012) Nuclear ribosomal internal transcribed spacer (ITS) region as a universal DNA barcode marker for Fungi. Proc Natl Acad Sci U S A 109:6241–6246
Hubalek Z (1974) Fungi associated with free-living birds in Czechoslovakia and Yuogoslavia. Acta Sci Nat Brno 8:1–62
Gegeckas A, Gudiukaitė R, Citavicius D (2014) Keratinolytic proteinase from Bacillus thuringiensis AD-12. Int J Biol Macromol 69:46–51. https://doi.org/10.1016/j.ijbiomac.2014.05.024
Brandelli A, Daroit DJ, Riffel A (2010) Biochemical features of microbial keratinases and their production and applications. Appl Microbiol Biotechnol 85:1735–1750. https://doi.org/10.1007/s00253-009-2398-5
Yoshioka M, Miwa T, Horii H et al (2007) Characterization of a proteolytic enzyme derived from a Bacillus strain that effectively degrades prion protein. J Appl Microbiol 102:509–515. https://doi.org/10.1111/j.1365-2672.2006.03080.x
Radha S, Gunasekaran P (2007) Cloning and expression of keratinase gene in Bacillus megaterium and optimization of fermentation conditions for the production of keratinase by recombinant strain. J Appl Microbiol 103:1301–1310. https://doi.org/10.1111/j.1365-2672.2007.03372.x
Kumar AG, Swarnalatha S, Gayathri S et al (2008) Characterization of an alkaline active – thiol forming extracellular serine keratinase by the newly isolated Bacillus pumilus. J Appl Microbiol 104:411–419. https://doi.org/10.1111/j.1365-2672.2007.03564.x
Balaji S, Senthil Kumar M, Karthikeyan R et al (2008) Purification and characterization of an extracellular keratinase from a hornmeal-degrading Bacillus subtilis MTCC (9102). World J Microbiol Biotechnol 24:2741–2745. https://doi.org/10.1007/s11274-008-9782-7
Acknowledgements
We thank the Slovenian Research Agency (SRA/ARRS) for funding through programme P4-0097, and funding PhD study to B.P. The funder had no influence on this study and any other activities related to writing or publication.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Blaž Petek and Romana Marinšek Logar contributed to the study conception and design. Material preparation, data collection and analysis were carried out by Blaž Petek, Maja Vodušek, Tomaž Accetto, Maša Zorec, Polona Zalar and Jernej Oberčkal. The first draft of the manuscript was written by Blaž Petek and all authors commented on earlier versions of the manuscript. All authors read and approved the final manuscript. The authors declare that they have no conflict of interest in the publication.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Petek, B., Vodušek, M., Accetto, T. et al. Isolation and characterization of highly active keratinolytic microorganisms with promising potential for waste sheep wool processing. J Mater Cycles Waste Manag 26, 360–372 (2024). https://doi.org/10.1007/s10163-023-01830-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10163-023-01830-5