Abstract
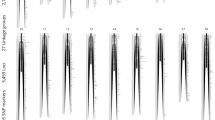
Small abalone, Haliotis diversicolor, is naturally distributed along the coastal waters of East Asia from Japan to the Philippines. It is an economically important maricultured species in southern China and Taiwan. Genetic linkage maps for small abalone were constructed using a total of 308 simple sequence repeat markers including 297 novel markers. Segregation data on 96 progeny were genotyped using a pseudo-testcross strategy. Sixteen linkage groups were identified in both female and male maps, consistent with the haploid chromosome number. The female linkage map covered 758.3 cM, with an average interval of 5.2 cM. The male linkage map spanned a total genetic distance of 676.2 cM, with an average interval of 4.5 cM. An integrated map was constructed by incorporating the homologous parental linkage groups, resulting in 16 linkage groups with a total of 762.1 cM. Genome coverage of the integrated linkage map was approximately 80.7%. The genetic linkage maps of small abalone may facilitate marker-assisted selection and quantitative trait loci mapping.



Similar content being viewed by others
References
Baranski M, Loughnan S, Austin CM, Robinson N (2006) A microsatellite linkage map of the blacklip abalone, Haliotis rubra. Anim Genet 37:563–570
Beckmann JS, Soller M (1990) Toward a unified approach to genetic mapping of eukaryotes based on sequence tagged microsatellite sites. Biotechnology 8:930–932
Bert PF, Charmet G, Sourdille P, Hayward MD (1999) A high-density molecular map for ryegrass (Lolium perenne). Theor Appl Genet 99:445–452
Cai J, Zhou J, Yang H (2006) Isolation and characterization of Vibrio alginolyticus as a pathogen to the massive mortality of abalone (Haliotis diversicolor) postlarvae. Trans Oceanology Limnol 3:54–59, in Chinese with English abstract
Chakravarti A, Lasher LK, Reefer JE (1991) A maximum likelihood method for estimating genome length using genetic linkage data. Genetics 128:175–182
Chen CS, Yan ZL, Liu GZ, Liu H, Ji DH (2003) Karyotypes of diploid and triploid Haliotis diversicolor aquatilis. J Jimei Univ (Natural Science) 8(4):291–294
Darrin PR, Smith CA, Rommens M, Blanchard B, Robichaud DM, Reith M (2007) A genetic linkage map of Atlantic halibut (Hippoglossus hippoglossus L.). Genetics 177:1193–1205
Faris JD, Laddomada B, Gill BS (1998) Molecular mapping of segregation distortion loci in Aegilops taushii. Genetics 149:319–327
Hubert S, Hedgecock D (2004) Linkage maps of microsatellite DNA markers for the pacific oyster Crassostrea gigas. Genetics 168:351–362
Kosambi DD (1994) The estimation of map distances from recombination values. Ann Eugen 12:172–175
Koshimizu E, Strüssmann CA, Okamoto N, Fukuda H, Sakamoto T (2010) Construction of a genetic map and development of DNA markers linked to the sex-determining locus in the Patagonian pejerrey (Odontesthes hatcheri). Mar Biotechnol 12:8–13
Lallias D, Gomez-Raya L, Haley CS, Arzul I, Heurtebise S, Beaumont AR, Boudry P, Lapègue S (2009) Combining two-stage testing and interval mapping strategies to detect QTL for resistance to bonamiosis in the European flat oyster Ostrea edulis. Mar Biotechnol 11:570–584
Li J, Boroevich KA, Koop BF, Davidson WS (2010) Comparative genomics identifies candidate genes for infectious salmon anemia (ISA) resistance in Atlantic salmon (Salmo salar). Mar Biotechnol. doi:10.1007/s10126-010-9284-0
Liao X, Ma HY, Xu GB, Shao CW, Tian YS, Ji XS, Yang JF, Chen SL (2009) Construction of a genetic linkage map and mapping of a female-specific DNA marker in half-smooth tongue sole (Cynoglossus semilaevis). Mar Biotechnol 11:699–709
Lindahal KF (1991) His and hers recombinational hotspots. Trends Genet 7:273–276
Lindberg DR (1992) Evolution, distribution and systematics of Haliotidae. In: Abalone of the World: Biology, Fisheries and Culture
Liu F, Shao Z, Zhang H, Liu J, Wang X, Duan D (2010) QTL mapping for frond length and width in Laminaria japonica aresch (Laminarales, Phaeophyta) using AFLP and SSR markers. Mar Biotechnol 12:386–394
Lyttle TW (1991) Segregation distorters. Annu Rev Genet 25:511–557
Morgante M, Olivieri AM (1993) PCR-amplified microsatellites as markers in plant genetics. Plant J 3:175–182
Morishima K, Nakayama J, Arai K (2008) Genetic linkage map of the loach Misgurnus anguillicaudatus (Teleostei: Cobitidae). Genetica 132:227–241
Reece, KS, Morrison, CL, Ribeiro WL, Gaffney PM, Allen SK (2002) Microsatellite markers for the eastern oyster Crassostrea viginica: linkage mapping and genetic monitoring for restoration projects. Presented at the International Conference on the Status of Plant, Animal and Microbe Genome X, January 12–16
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning a laboratory manual, 2nd edn. Cold Spring Harbor Laboratory Press
Sekino M, Hara M (2007) Linkage maps for the Pacific abalone (genus Haliotis) based on microsatellite DNA markers. Genetics 175:945–958
Shi YH, Guo XM, Gu ZF, Wang AM, Wang Y (2010) Preliminary genetic linkage map of the abalone Haliotis diversicolor Reeve. Chin J Oceanol Limnol 28(3):549–557
Van Ooijen JW, Voorrips RE (2001) JoinMap 3.0, software for the calculation of genetic linkage maps. Plant Research International, Wageningen
Voorrips RE (2002) MapChart: software for the graphical presentation of linkage maps and QTLs. J Hered 93:77–78
Wang CM, Lo LC, Zhu ZY, Pang HY, Liu HM, Tan J, Lim HS, Chou R, Orban L, Yue GH (2011) Mapping QTL for an adaptive trait: the length of caudal fin in Lates calcarifer. Mar Biotechnol 13:74–82
Woram RA, McGowan C, Stout JA, Gharbi K, Ferguson MM, Hoyheim B, Davidson EA, Davidson WS, Rexroad C, Danzmann RG (2004) A genetic linkage map for Arctic char (Salvelinus alpinus): evidence for higher recombination rates and segregation distortion in hybrid versus pure strain mapping parents. Genome 47:304–315
You WW, Ke CH, Luo X, Wang DX (2009) Growth and survival of three small abalone Haliotis diversicolor populations and their reciprocal crosses. Aquac Res 40:1474–1480
Zhan A, Hu J, Hu X, Hui M, Wang M, Peng W, Huang X, Wang S, Lu W, Sun C, Bao Z (2009a) Construction of microsatellite-based linkage maps and identification of size-related quantitative trait loci for Zhikong scallop (Chlamys farreri). Anim Genet 40:821–831
Zhan X, Hu HY, Ke CH, Hu SN, Wang DX, Chen F (2009b) Isolation and characterization of 11 microsatellite loci in small abalone, Haliotis diversicolor Reeve. Conserv Genet 10:1185–1187
Zhang Y, Xu P, Lu C, Kuang Y, Zhang X, Cao D, Li C, Chang Y, Hou N, Li H, Wang S, Sun X (2010) Genetic linkage mapping and analysis of muscle fiber-related QTLs in common carp (Cyprinus carpio L.). Mar Biotechnol. doi:10.1007/s10126-010-9307-x
Acknowledgments
The project was supported by the National High Technology Research and Development Program of China (2006AA10A407), the Innovative Platform for Science and Technology, Fujian Province (2008N2004), and the earmarked fund for Modern Agro-Industry Technology Research System (nycytx-47).
Author information
Authors and Affiliations
Corresponding author
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
Supplemental Table 1
Characterization of 297 novel microsatellite markers genotyped in the mapping family (DOC 724 kb)
Rights and permissions
About this article
Cite this article
Zhan, X., Fan, F., You, W. et al. Construction of an Integrated Map of Haliotis diversicolor Using Microsatellite Markers. Mar Biotechnol 14, 79–86 (2012). https://doi.org/10.1007/s10126-011-9390-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10126-011-9390-7