Abstract
The phytopathogenic fungus Ustilago maydis causes corn smut by suppressing host plant defenses, including the oxidative burst response. While many studies have investigated how U. maydis responds to oxidative stress during infection, the consequences of heightened resistance to oxidative stress on virulence remain understudied. This study aimed to identify the effects on virulence in U. maydis strains exhibiting enhanced resistance to hydrogen peroxide (H2O2).
To achieve this, we exposed U. maydis SG200 to 20 escalating H2O2 shocks, resulting in an adapted strain resistant to concentrations as high as 60 mM of H2O2, a lethal dose for the initial strain. Genetic analysis of the adapted strain revealed five nucleotide substitutions, two minor copy number variants, and a large amplification event on chromosome nine (1–149 kb) encompassing the sole catalase gene. Overexpressing catalase increased resistance to H2O2; however, this resistance was lower than that observed in the adapted strain. Additionally, virulence was reduced in both strains with enhanced H2O2 resistance.
In summary, enhanced H2O2 resistance, achieved through either continuous exposure to the oxidative agent or through catalase overexpression, decreased virulence. This suggests that the response to the oxidative stress burst in U. maydis is optimal and that increasing the resistance to H2O2 does not translate into increased virulence. These findings illuminate the intricate relationship between oxidative stress resistance and virulence in U. maydis, offering insights into its infection mechanisms.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Fungal plant pathogens are widespread and exert substantial impacts on crop yield and quality (Avery et al. 2019). Among these pathogens is Ustilago maydis, a biotrophic fungus widespread in the Americas, Asia, and Europe, responsible for corn smut disease in Zea mays. U. maydis has served as an invaluable model organism for investigating the cellular physiology and molecular biology of host–pathogen interactions (Holliday 1961; Schuster et al. 2016; Olicón-Hernández et al. 2019; Dutheil et al. 2020; Lin et al. 2021; López-Martínez et al. 2022; Villagrán et al. 2023), and more recently, the evolutionary history of crop infecting fungi through population genomics (Munkacsi et al. 2006, 2008; Depotter et al. 2021; Schweizer et al. 2021; Momeni and Nazari 2022). The current reference genome of U. maydis spans 20.5 Mb and includes 23 nuclear chromosomes, one mitochondrial chromosome, and three unmapped contigs (Kahmann and Kämper 2004; Kämper et al. 2006). Within the annotation of this reference, there are 6,902 protein-encoding genes and two putative functional mobile elements: the retrotransposons TigR and HobS. This last mobile element consistently appears in either complete or incomplete form, with one copy per chromosome. Mobile elements are commonly found in fungal pathogen genomes and play critical roles in their adaptive processes (Möller and Stukenbrock 2017; Tralamazza et al. 2023). These elements have been associated with various chromosomal rearrangements including amplifications, deletions, translocations, and inversions, in yeasts and other fungi (Daboussi 1996; Wang et al. 2020a, b).
Fungal pathogens have a remarkable ability to adapt to their hosts and to changing environments influenced by drugs or stressors (Stukenbrock and Croll 2014; Grandaubert et al. 2019; Habig et al. 2021; Möller et al. 2021). These adaptations frequently encompass significant changes at the chromosomal level, including gains or losses of entire chromosomes or a fraction of them. They may also entail epigenetic modifications affecting regulation of gene expression and alterations in mutation rate (Selmecki et al. 2009; Gilchrist and Stelkens 2019; Kramer et al. 2023; Vande Zande et al. 2023).
While there has been extensive research on U. maydis and its infection process in maize, the consequences of increasing oxidative stress resistance on virulence remain elusive. Notably, specific gene deletions have been shown to decrease both virulence and the ability to withstand oxidative stress. For instance, disruptions in tps2, a gene involved in trehalose biosynthesis (Cervantes-Chávez et al. 2016), and deletions of the yap1 transcription factor, which plays a role in oxidative stress response (Molina and Kahmann 2007), have been identified. While numerous studies have explored the impact of specific genes on pathogenicity, it remains unclear whether increased resistance to oxidative stress directly translates into enhanced virulence.
This study investigates the adaptive response of U. maydis to oxidative stress and its implications in virulence. We used as the initial strain the haploid and solopathogenic U. maydis SG200 (Kämper et al. 2006), which is capable of infecting without mating, enabling subsequent experiments in maize plants. We exposed this initial strain to periodic H2O2 shocks over 200 generations, resulting in significant H2O2 adaptation. In this study, we compared the virulence between SG200, a H2O2-adapted strain (UmH2O2-R) and a strain with an extra catalase copy (oexUMAG_11067).
Results
Hydrogen peroxide adaptation and inheritance of H2O2-resistance in Ustilago maydis
Adaptation is a biological process through which organisms gradually acquire physiological and genetic changes over successive generations, enhancing their fitness. This enhanced fitness is reflected in their capacity to thrive and propagate in specific environmental conditions. In this study, we took advantage of the adaptive potential of U. maydis SG200 to obtain the strain UmH2O2-R, showing increased resistance to H2O2. This strain was obtained by selecting surviving cells after periodic exposures to increasing concentrations of H2O2, starting at 5 mM and reaching 60 mM after twenty treatments.
Adaptation to H2O2 increased by the end of the 20 treatments. As depicted in Fig. 1A, we assessed the percentage of viable cells by quantifying colony-forming units (CFUs) following a three-hour exposure to 5 and 60 mM H2O2, using the number of CFUs without exposure as baseline. At a 5 mM concentration, the survival rate increased significantly from 42% in the initial strain, to 91% in the adapted strain UmH2O2-R. A 95% confidence interval (32.39033, 52.34300) was calculated for the survival percentage at 5 mM for the initial strain. Notably, the measurement obtained for the adapted strain after a shock at the same concentration lies outside of this confidence interval. Exposure to 60 mM H2O2 shock was lethal for the initial strain however, we observed a survival rate of 5% in the evolved strain UmH2O2-R. These observations indicate that periodic exposure to H2O2 significantly improved the resistance of U. maydis to this oxidative agent. The adapted strain, UmH2O2-R, even survived a shock with a concentration that was previously lethal for the initial strain.
Hydrogen Peroxide Resistance in Ustilago maydis. A Percentage of cells surviving H2O2 shocks at 5 and 60 mM in the initial strain U. maydis SG200 and the adapted strain (UmH2O2-R). B Inhibition zone areas produced by H2O2 of the initial strain and of UmH2O2-R immediately after isolation, and after 7 days of culture with daily dilutions in the absence of H2O2 (UmH2O2-R + 7 days)
To assess the stability of H2O2-resistance phenotype, UmH2O2-R was cultured for seven days with daily dilutions in the absence of H2O2. We evaluated the H2O2 resistance by measuring the area of inhibition in an agar diffusion assay. Figure 1B shows that there was no significant difference in the inhibition zone between UmH2O2-R right after the twentieth treatment and following its propagation for seven days without H2O2 (as determined by paired t-test, p-value = 0.260). Furthermore, in both scenarios H2O2 inhibition was lower than in the initial strain U. maydis SG200 (with p-values of 0.011 and 0.002, respectively).
Genomic variants found after hydrogen peroxide adaptation in Ustilago maydis
Due to the hereditary nature of the resistance phenotype, we aimed to identify the genetic variants that accumulated in the short adaptation period. We conducted whole-genome sequencing on both the initial strain (SG200) and the H2O2-adapted strain (UmH2O2-R). After a rigorous filtering we identified five bona fide single nucleotide variants (SNVs) distributed throughout the nuclear and mitochondrial genomes of the adapted strain (Supplementary Table 1). Three SNVs were non-synonymous substitutions, while the other two SNVs were present in intergenic regions.
The first non-synonymous substitution was located at position Chr01:2,430,690, within the UMAG_00813 gene (Chr01:2,430,525–2,432,966 ( +)). This gene encodes a putative protein containing the FAD/NAD(P)-binding domain, as predicted by AlphaFold2 (reference entry: A0A0D1E974). This mutation resulted in an amino acid substitution, specifically changing glycine to arginine at position 56 (Gly56Arg).
The second non-synonymous substitution was found at Chr08:464,088, within the UMAG_10823 gene (Chr08:463,697 – 464,216 (-)). This gene encodes a putative long chronological lifespan protein, as predicted by AlphaFold2 (reference entry: A0A0D1C4W6). The mutation led to an amino acid substitution, replacing glutamic acid with aspartic acid at position 43 (Glu43Asp).
The last non-synonymous substitution occurred at Chr18:330,272 within the UMAG_05545 gene (Chr18:328,759 – 331,155 ( +)). This gene encodes a putative histone-lysine N-methyltransferase with a Dot1 domain responsible for histone H3 lysine 79 (Lys-79) methylation. AlphaFold2 (reference entry: Q4P2W8) predicted this function. Specifically, the nucleotide substitution involved a C to A transversion at position 330,272, resulting in the replacement of the threonine residue with an asparagine residue (Thr505Asn) within the Dot1 domain, a critical region for the catalytic activity of the protein (Lee et al. 2018; Min et al. 2003).
The intergenic substitutions were a T to A transversion located at Chr10:623,352( +), and a G to A transition in the mitochondrial genome at position 15,246( +).
In addition to single nucleotide variants, copy number variants (CNVs) can be induced in response to environmental stressors, conferring rapid adaptive advantages. We found three CNVs that were only present in the adapted strain (Supplementary Table 2). Among these, two were relatively small, with less than 5 kb, while the third CNV was substantially larger, spanning approximately 150 kb.
One of the CNVs was situated on chromosome twenty from base 50,551 to base 51,600 (1,050 bp), exhibiting a copy number of 3.74. Although this sequence lacks annotated genes, it contains a 127 bp segment that is recurrently found throughout the genome. The second CNV was identified on chromosome seventeen from position 151 to 4,650 (4,500 bp), with a copy number of 1.80. This sequence overlaps with the UMAG_04693 gene (Chr17:1396–4642 ( +)), which encodes a putative protein containing a helicase ATP-binding domain.
The third CNV, identified by CNVnator ranged from position 1 to 149,100 of chromosome nine, and displayed a copy number of 2.99 (as illustrated in Supplementary Fig. 1A). This CNV presented a log2 ratio of 1.54 in the normalized coverage, signifying a total copy number of three for the corresponding sequence, consistent with CNVnator’s results. In this 150 kb region, we identified a total of 55 protein-coding genes, which are listed in Supplementary Table 3. Among these genes, UMAG_11067 (Chr9:18,661–20,910 (-)) stood out as the sole catalase enzyme within the U. maydis reference genome. This enzyme plays a critical role in the breakdown of hydrogen peroxide into water and oxygen, making it a promising candidate for studying the impact of its duplication on resistance to H2O2 and virulence.
To explore the breakpoint of the CNV on chromosome 9 in greater detail, we conducted an in-depth analysis within the amplified region, at one bp resolution. The end of the amplification found by CNVnator (Chr9:149,100) falls within a 4.5 kb region lacking annotated protein-coding genes. Nevertheless, within the region spanning positions 147 kb to 150 kb, we observed a slight decrease in the log2 ratio of normalized coverage, as visualized in Supplementary Fig. 1B. Inside this region, we also identified two remnants of the 750 bp-HobS direct repeats, oriented in an inverted direction at positions Chr9:147,098–147,496 and Chr9:149,294–148,883. Two fragments of 400 bp and 412 bp of these remnants align with the HobS direct repeats with 91% and 92% identity, respectively; they may be responsible for the genomic instability of the region and its triplication.
Triplication of the catalase gene enhances expression in response to hydrogen peroxide in the adapted strain
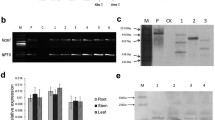
To determine whether the observed increase in the copy number of the catalase gene in the adapted strain UmH2O2-R resulted in elevated transcript levels, we conducted a gene expression analysis using qPCR. After a 10 mM H2O2 shock, we observed a 2.1-fold increase in catalase gene expression within the strain bearing the triplication compared to the initial strain U. maydis SG200. This difference in expression levels was found to be statistically significant with a p-value of 0.010 as determined by t-test (Fig. 2).
Increased catalase expression enhances hydrogen peroxide resistance but diminishes virulence in maize
To test the potential role of the catalase gene in enhancing H2O2 resistance, we engineered an overexpression strain carrying an additional copy of UMAG_11067 (oexUMAG_11067). Both the oexUMAG_11067 and UmH2O2-R strains exhibited significant differences in the survival rates following exposure to the 10 mM H2O2 shock, compared to U. maydis SG200 (t-test p-value = 0.020 for oexUMAG_11067, and p-value = 0.004 for UmH2O2-R), where this concentration proved completely inhibitory (Fig. 3A). Specifically, the oexUMAG_11067 strain displayed an 8% survival rate, whereas survival rate in UmH2O2-R was 38.6%. These results confirmed that catalase overexpression enhanced resistance to H2O2. However, it is important to note that despite this enhancement, the overexpression mutant failed to achieve the same level of resistance observed in the adapted strain UmH2O2-R.
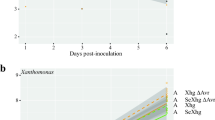
Hydrogen peroxide resistance and virulence profiles of oexUMAG_11067 and UmH2O2-R. A Percentage of viable cells determined through CFU counts after exposure to 10 mM H2O2 shock. B Virulence assessment on maize seedlings analyzed at 12 days post infection. Color coding represents the severity of infection
To evaluate the virulence of U. maydis SG200, UmH2O2-R, and oexUMAG_11067, we monitored symptoms in maize seedlings at 12 days post-infection. Figure 3B illustrates that the two strains exhibiting increased H2O2 resistance, specifically oexUMAG_11067 and UmH2O2-R, elicited milder symptoms when compared to the U. maydis SG200 strain (chi-square, p-value = 2.64 × 10–4 for oexUMAG_11067 and chi-square, p-value = 4.8 × 10–3 for UmH2O2-R). The most significant difference was the reduction of deceased maize plants. Furthermore, we observed an increase in the proportion of symptom-free plants and a decrease in the number of individuals displaying severe tumors (Supplementary Fig. 2). Moreover, the disparities in symptoms between oexUMAG_11067 and UmH2O2-R were not statistically significant (chi-square, p-value = 0.299). These collective findings conclusively indicate that enhanced resistance to hydrogen peroxide, whether achieved through catalase overexpression (oexUMAG_11067) or adaptive exposure (UmH2O2-R), did not translate into increased virulence on maize.
Discussion
The main goal of this study was to enhance H2O2 resistance in Ustilago maydis SG200 strain, identify the genetic changes occurring during adaptation, and subsequently evaluate the impact of the enhanced H2O2 resistance on virulence in maize.
Adaptation occurs when individuals of a species increase their fitness within a specific environment. In this work we induced oxidative stress using H2O2, a highly reactive molecule that at high concentrations, can cause damage to various cellular components, ultimately leading to cell death. Despite being lethal at high concentrations, exposure to milder concentrations of this reagent can lead to adaptation in different fungal species (Linder et al. 2017; Huang and Kao 2018; Qi et al. 2019; Zhang et al. 2019). In our study, we observed that the adapted UmH2O2-R strain exhibited increased survival rate when exposed to H2O2 concentrations that were lethal to the U. maydis SG200. These findings align with research conducted on the pathogenic yeast Candida glabrata (Huang and Kao 2018). In C. glabrata, an increase in H2O2 resistance was observed when the organism was subjected to 22 periodic H2O2 shocks, starting from an initial concentration of 80 mM and reaching 350 mM by the end of the experiment. Both our observations in U. maydis and the results in C. glabrata (Huang and Kao 2018) evidence the capacity of fungal species to rapidly adapt to oxidative stress induced by H2O2. In both cases, within just 180 generations for C. glabrata, and 200 generations for U. maydis, both species were able to survive to a previously lethal concentration of H2O2.
Then, we analyzed the genomes of the initial strain and the adapted strain, UmH2O2-R. Variants already present in the initial strain were considered as the genetic background and were removed from the set of identified variants in UmH2O2-R. Regarding single nucleotide variants, we identified two intergenic mutations and three non-synonymous mutations. One intergenic SNV was located within the promoter region of the gene UMAG_03851, a putative adenylosuccinate synthase. The other intergenic SNV was found in the mitochondrial genome (Mt:15,246), with no annotated genes in the 2 kb periphery.
The non-synonymous nucleotide substitutions were detected in three distinct genes: UMAG_00813, which encodes a putative protein containing the FAD/NAD(P)-binding domain; UMAG_10823, coding for a putative long chronological lifespan protein; and UMAG_05545, encoding a histone-lysine N-methyltransferase (HKMTs) with a Dot1 domain. These functional annotations were all predicted by AlphaFold2 (Jumper et al. 2021), nevertheless the prediction confidence is low for UMAG_00813 and UMAG_10823, but more reliable for UMAG_05545.
The UMAG_00813 protein with a FAD/NAD(P)-binding domain, falls under the gene ontology category of oxidoreductase activity, suggesting a connection to oxidative stress. In the adapted strain, UmH2O2-R, the identified mutation occurred in the protein’s amino-terminal region.
In the case of UMAG_10823, the amino acid substitution occurred within the signal peptide. Although AlphaFold2 prediction for UMAG_10823 as a putative long chronological lifespan protein has a low confidence, previous research has observed an association between antioxidant defense and extended long chronological lifespan in the yeast S. cerevisiae (Mirisola and Longo 2022).
The last non-synonymous substitution occurred within the catalytic domain of the protein encoded by UMAG_05545, which is a putative histone-lysine N-methyltransferase with a Dot1 domain. This protein is also known as Dot1 and catalyzes the methylation of lysine 79 on histone H3 (Ng et al. 2002). Histone methylation plays a critical role in the regulation of transcription and the structure of chromatin (Brosch et al. 2008; Li et al. 2019). Disrupting the catalytic activity of Dot1 leads to the repression of telomeric silencing in both S. cerevisiae and Penicillium oxalicum (Ng et al. 2002; Li et al. 2019). In Aspergillus flavus deletion of dot1 reduces the virulence on maize seeds and increases resistance to oxidative stress induced by tert-butyl hydroperoxide (Liang et al. 2017). Furthermore, previous studies have reported that the inactivation of genes associated with chromatin remodeling reduces the virulence of fungal plant pathogens (Gu et al. 2017; Wang et al. 2020a, b). The mutation within the Dot1 domain of the protein encoded by UMAG_05545 could potentially impact the activity of this protein and consequently have an impact in the chromatin structure of U. maydis.
In addition to single nucleotide mutations, we identified copy number variants upon H2O2 exposure in the UmH2O2-R strain. In mammalian cells, abnormal chromosome numbers or large chromosomal duplications often have deleterious effects (Tosh et al. 2022; Williams et al. 2008). However, in the case of filamentous fungi and yeast, chromosomal duplications play a key role in facilitating rapid adaptation to stressful environments by increasing the copy number of genes associated with stress response (Selmecki et al. 2009; Chen et al. 2012; Linder et al. 2017; Gilchrist and Stelkens 2019; Khateb et al. 2023).
The presence of catalase UMAG_11067 within the amplified segment of chromosome 9, along with its elevated gene expression, suggests a potential role in the observed H2O2 resistance in UmH2O2-R. Our findings regarding the increased copy number of catalase in U. maydis is analogous to observations in S. cerevisiae. In this yeast, partial chromosomal amplifications harboring H2O2-degrading proteins, such as cytoplasmic thioredoxin peroxidase (tsa2) and cytosolic catalase (ctt1), have been reported in response to oxidative stress induced by H2O2 (Linder et al. 2017; Zhang et al. 2019). The contribution of catalase to H2O2 resistance was evaluated by overexpressing this gene in the oexUMAG_11067 strain. While oexUMAG_11067 showed increased resistance, it was not as pronounced as the one achieved through adaptive evolution in the UmH2O2-R strain, suggesting that the other identified genetic variants, or any other epigenetic mechanisms that arose during this process may also play a role.
In the final stage of our research, we evaluated the consequences of enhanced H2O2 resistance on the pathogenicity of U. maydis in maize seedlings. Our findings indicate that both the UmH2O2-R and oexUMAG_11067 strains of U. maydis show a decrease in virulence. The common genetic feature in these strains, not present in the initial strain SG200, is an increased number of catalase gene copies. This suggests a correlation between the overexpression of catalase and the observed reduction in virulence, a phenomenon also present in Candida albicans where the demand for iron by catalase—a cofactor scarce in host tissues—has been linked to decreased virulence (Pradhan et al. 2017). Both C. albicans and U. maydis rely on iron as a catalase cofactor. Additionally, the research by Eichhorn et al. (2006) supports the need of iron for virulence, showing that U. maydis strains with compromised iron uptake are less virulent.
If iron levels are low, the metabolic expense of catalase production becomes detrimental, as the enzyme will be non-functional without its cofactor. Moreover, catalase intense iron requirement would reduce its availability for other essential metabolic processes such as DNA synthesis and respiration. While we lack specific data on the iron content in maize plants and whether it meets the requirements of strains with high catalase expression, increasing iron content in the plant might potentially reverse the reduced virulence. Nonetheless, it is still an open question whether U. maydis can efficiently use iron from supplemented maize. It is important to consider that supplying external iron can suppress the expression of the entire iron-related gene cluster (Eichhorn et al. 2006).
Our research evidenced U. maydis capacity for rapid adaption to H2O2 and the genetic variants linked to this process. To uncover the potential roles of the identified genetic variants, and of any potential epigenetic modifications that may have emerged during the adaptation process further investigation would be required.
In conclusion, our study presents a strategy for selecting fungi adapted to harsh environmental conditions. This research also highlights the remarkable adaptive potential of U. maydis in response to H2O2 stress through large chromosomal duplications. Additionally, it illustrates the trade-offs associated with this adaptation, notably the reduction in virulence. These findings enrich our understanding of how fungi rapidly adapt and paves the way for forthcoming research aiming to study the molecular mechanisms involved in these processes.
Materials and methods
Cultures and growth conditions
The solopathogenic strain Ustilago maydis SG200 (Kämper et al. 2006) was employed for both, H2O2 adaptation (UmH2O2-R) and the generation of a strain with catalase overexpression (oexUMAG_11067). Liquid cultures of U. maydis were cultivated in CM broth (Holliday 1961) and incubated at 28 °C with continuous agitation at 180 rpm using an orbital shaker. For agar-based assays, CM broth was supplemented with 2% microbiological agar, and the plates were incubated at 28 °C for 48 h.
Assessment of H2O2 resistance through area of inhibition in agar diffusion assay
To evaluate resistance to H2O2, we inoculated standard round petri dishes (CM plates) with a 100 μL solution containing 107 cells/mL, using a sterile L-shaped spreader. Subsequently, we placed a 5 μL droplet of 30% (w/w) H2O2 (H1009, Sigma Aldrich) on a sterile filter paper disk at the center of the plate. The area of the inhibition halo produced by the H2O2 droplet was measured. To determine the statistical significance of differences of the inhibition area, a t-test was conducted using the rstatix package (Kassambara 2020) within R software, version 4.3.0 (R Development Core Team 2023).
Generation of the adapted strain UmH2O2-R
An isogenic culture of U. maydis SG200 served as the starting point for generating the adapted strain UmH2O2-R. This adaptation process encompassed a series of 20 treatments, during which H2O2 concentrations were incrementally raised, starting at 5 mM and reaching a peak of 60 mM (refer to Supplementary Table 4). Each treatment was divided into two phases: a shock phase and a recovery phase (see Supplementary Fig. 3). In the shock phase, 106 cells of the previous treatment were exposed to H2O2 for 3 h in 1 mL of CM broth and incubated at 28 °C with continuous agitation at 500 rpm, in a 1.5 mL tube inside a thermoblock (ThermoMixer C, Eppendorf). At the end of the shock phase, we used 0.1 mL to estimate the survival rate of the exposed cells through quantification of CFUs. For this, the cells were diluted 1:1000 and 0.1 mL of that dilution was evenly distributed on CM agar plates. Subsequently, the recovery phase started by transferring the remaining 0.9 mL of the shock-surviving cell population into 19.1 mL of CM broth, which was then incubated at 28 °C with agitation at 180 rpm for 45 h. This entire process was iterated 20 times, with H2O2 concentrations being raised in incremental steps every two treatments. The H2O2-adapted strain, UmH2O2-R was isolated from the shocked cells after the twentieth treatment with H2O2 at 60 mM. The colony was randomly selected from the plate on which CFUs were quantified. Then it was streaked twice on CM agar to obtain single isolated colonies. One colony from the last plate was randomly chosen and named UmH2O2-R. Then, UmH2O2-R was cultured in CM broth for 24 h. A portion of this last culture was cryo-preserved, while the remaining cells were used for DNA extraction and sequencing.
Inheritance of H2O2 resistance in the adapted strain UmH2O2-R
To investigate whether the observed H2O2 resistance in the adapted strain UmH2O2-R was a stable inheritable trait, it was propagated for seven days in CM broth without exposure to H2O2, with daily dilutions. Subsequently, we assessed its resistance using agar diffusion assays, measuring the area of inhibition both immediately after isolation and at the end of the seven-day propagation period without any exposure to H2O2.
DNA extraction and sequencing
Genomic DNA extraction was carried out using a mechanical lysis-based protocol using 0.3 mm glass beads (SigmaAldrich, G1277) and phenol:chloroform:isoamyl alcohol (SigmaAldrich, 77,617). DNA concentration was quantified using a fluorometric assay with the Qubit 2.0 Fluorometer (Life Technologies) and the Qubit dsDNA BR Assay kit (ThermoFisher, Q32853). DNA purity was determined by assessing the absorbance ratio at 260/280 nm using a Nanodrop, while DNA integrity was verified through electrophoresis on a 0.8% agarose gel stained with SYBR Safe (ThermoFisher, S33102). Whole genome sequencing, for the initial strain U. maydis SG200 and for the adapted strain UmH2O2-R, was conducted using short-read 150 bp paired-end sequencing on the DNBseq platform at BGI.
Genome sequence analysis
Read preprocessing and alignment
The raw short reads, for initial strain U. maydis SG200 and for the adapted strain UmH2O2-R, were preprocessed using fastp v.0.20.0 (Chen et al. 2018) with default parameters. Subsequently, they were aligned to the U. maydis 521 v.2 reference genome (available at https://fungi.ensembl.org/Ustilago_maydis/Info/Index, download date: 19–10-2019; last accessed on 22–08-2022) using BWA-MEM v.0.7.8 (Li and Durbin 2010). Duplicated reads were identified and flagged using the picard toolkit v.2.6.0 (Broad Institute 2016). The final alignment outputs were first exported as a bam files and subsequently compressed into cram files.
SNV identification
To identify Single Nucleotide Variants (SNVs), we used the cram-formatted alignment files as input for bcftools mpileup v.1.9 (Li et al. 2009). Base quality (-Q) and mapping quality (-q) thresholds were set at 20 and 30, respectively. SNVs were called from the mpileup file using bcftools call with the multiallelic caller option (-m) and specifying ploidy as 1 (–ploidy 1). SNVs coming from the genetic background of SG200, 858 out of 899, were identified and subsequently removed using vcfR v.1.14.0 (Knaus and Grünwald 2017), dplyr v.1.1.2 (Wickham et al. 2023), and data.table v.1.14.8 (Dowle and Srinivasan 2023) packages with a customized R script. Subsequently, the remaining 41 SNVs were further filtered by setting a quality score threshold greater than 200 (Q > 200). We then determined whether the SNVs were located within a gene or were intergenic using the gene annotation file of U. maydis v.2, accessible at: https://ftp.ensemblgenomes.ebi.ac.uk/pub/fungi/release-57/gff3/ustilago_maydis, last accessed on 2022–08-22). This annotation was processed with pandas v.2.0.3 (McKinney 2010) in a customized Python script (Python Software Foundation 2016).
CNV detection
To detect the CNVs (Copy Number Variants) arising during the generation of the adapted strain UmH2O2-R, we employed CNVnator, following the methodology outlined by Abyzov et al. (2011), with a bin width of 150 bp. We subsequently removed the CNVs that were already present in the initial SG200 strain, representing the departing genetic background. Additionally, variants with an e-value greater 0.05 were excluded. We also computed the log2 ratio of normalized coverage at base-pair resolution to visualize the breakpoints of the large chromosomal amplifications.
For a functional assessment of the genes involved in SNVs or CNVs, we consulted the U. maydis UniProt database obtained with AlphaFold2 (https://www.uniprot.org/taxonomy/237631, last accessed 2023/01/20).
Quantification of catalase gene (UMAG_11067) expression
To assess the expression of UMAG_11067 (catalase) in U. maydis strains SG200 and UmH2O2-R, quantitative polymerase chain reaction (qPCR) was performed on the StepOnePlus™ system (Applied Biosystems™) using Maxima SYBR Green/ROX qPCR Master Mix (2X). The UMAG_04869 gene that encodes the elongation factor subunit 2b (elF2b) served as the reference for normalization. Total RNA was extracted from 108 cells after a three-hour exposure to 0 and 10 mM of H2O2, following the manufacturer’s instructions for the TRIzol™ reagent. Subsequently, first-strand cDNA synthesis was carried out using SuperScript® III Reverse Transcriptase with 300 ng of total RNA as initial template. The efficiencies of the qPCR reactions for both the target and reference genes were determined through standard curves generated with 1 µg of cDNA. Changes in gene expression were calculated by comparing UMAG_11067 gene expression levels without H2O2 exposure (0 mM), following the method described by Pfaffl, (2001). Primer sequences are provided in Supplementary Table 5.
Generation of catalase overexpression strain (oexUMAG_11067)
To create the catalase overexpression strain (oexUMAG_11067), we introduced the open reading frame (ORF) of UMAG_11067 into the ip locus, under the regulatory control of the OMA promoter and the NOS terminator. Initially, the UMAG_11067 ORF was amplified via PCR and linked with the NOS terminator, as well as a segment located one kilobase upstream of the ip locus. This resulting construct was then inserted into the plasmid pUMa2625 at the NcoI – EcoRI sites. The recombinant vector obtained was subsequently transformed into Escherichia coli by heat shock and selected on LB agar containing 100 μg/mL of ampicillin. E. coli clones were subcloned twice on plates with antibiotics and cultured in LB broth supplemented with 100 μg/ml of ampicillin. Plasmid identity was confirmed through restriction analysis using NcoI, NcoI + EcoRI, and NcoI + SdaI, and validated by Sanger sequencing. The transforming molecule was excised from the resultant plasmid through AgeI + EcoRI digestion and purified from a 0.8% agarose gel.
Protoplasts of U. maydis strains were prepared during the mid-exponential growth phase using Lytic enzymes from Trichoderma harzianum (SigmaAldrich, L1412). The protoplasts were combined with 3 μg of DNA and selected on media containing 3 μg/mL of carboxin. Transformant clones were streaked three times on antibiotic-containing plates. Subsequently, overexpressing strains were verified through end-point PCR. Oligonucleotide sequences used in generating this strain are provided in Supplementary Table 5.
Assessment of H2O2 resistance through CFU counting
To evaluate resistance to H2O2 in the strains SG200, UmH2O2-R and oexUMAG_11067, we employed a quantification method based on colony-forming units (CFU) following exposure to a three-hour shock with 10 mM H2O2. To establish control conditions for CFU comparisons, an equivalent number of cells (106) was exposed to 0 mM H2O2. After the exposure period, both the shocked and untreated cell populations underwent serial dilution until 10–3. Subsequently, 100 µL of each dilution were uniformly spread onto CM agar plates using sterile 0.5 mm-diameter glass beads. These plates were then incubated at 28 °C for 48 h. The percentage of surviving cells after the 10 mM H2O2 shock treatment was estimated by referencing the CFU counts obtained under 0 mM H2O2 conditions as a baseline reference. To discern statistical variations in the percentage of surviving cells, a t-test was performed using R software v.4.3.0 (R Development Core Team 2023).
Pathogenicity test
We assessed the virulence symptoms produced by U. maydis SG200, U. maydis UmH2O2-R, and U. maydis oexCAT on nine-day-old cacahuazintle maize seedlings, that were infected with 300 μL of a cellular suspension at 108 cells/mL of each strain injected at the stem. The pots with maize seedlings were maintained in a green-house. Disease symptoms were recorded at 12 days post-infection. Statistical significance was tested with a chi-squared test in R software v.4.3.0 (R Development Core Team 2023). All plots were generated using the package ggplot2 (Wickham 2016).
Data availability
The code and input files required to replicate the analyses conducted in this study are accessible at: https://github.com/JLuisCuamatzi/USMA_H2O2_Adaptation. The raw reads for U. maydis strains SG200 and UmH2O2-R genomic data can be obtained from the National Center for Biotechnology Information (NCBI) under accession numbers SRR25650020 and SRR25650011, respectively.
References
Abyzov A, Urban AE, Snyder M, Gerstein M (2011) CNVnator: an approach to discover, genotype, and characterize typical and atypical CNVs from family and population genome sequencing. Genome Res 21(6):974–984. https://doi.org/10.1101/gr.114876.110
Avery SV, Singleton I, Magan N, Goldman GH (2019) The fungal threat to global food security. Fungal Biol 123(8):555–557. https://doi.org/10.1016/j.funbio.2019.03.006
Brosch G, Loidl P, Graessle S (2008) Histone modifications and chromatin dynamics: a focus on filamentous fungi. FEMS Microbiol Rev 32(3):409–439. https://doi.org/10.1111/j.1574-6976.2007.00100.x
Cervantes-Chávez JA, Valdés-Santiago L, Bakkeren G, Hurtado-Santiago E, León-Ramírez CG, Esquivel-Naranjo EU, Landeros-Jaime F, Rodríguez-Aza Y, Ruiz-Herrera J (2016) Trehalose is required for stress resistance and virulence of the basidiomycota plant pathogen Ustilago maydis. Microbiology 162(6):1009–1022. https://doi.org/10.1099/mic.0.000287
Chen G, Rubinstein B, Li R (2012) Whole chromosome aneuploidy: big mutations drive adaptation by phenotypic leap. BioEssays 34(10):893–900. https://doi.org/10.1002/bies.201200069
Chen S, Zhou Y, Chen Y, Gu J (2018) fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34(17):i884–i890. https://doi.org/10.1093/bioinformatics/bty560
Daboussi MJ (1996) Fungal transposable elements: generators of diversity and genetic tools. J Genet 75:325–339. https://doi.org/10.1007/BF02966312
Depotter JRL, Zuo W, Hansen M, Zhang B, Xu M, Doehlemann G (2021) Effectors with different gears: divergence of Ustilago maydis effector genes is associated with their temporal expression pattern during plant infection. Journal of Fungi 7(1):1–14. https://doi.org/10.3390/jof7010016
Dowle M, Srinivasan A (2023) data.table: Extension of ‘data.frame’. https://CRAN.R-project.org/package=data.table
Dutheil JY, Münch K, Schotanus K, Stukenbrock EH, Kahmann R (2020) The insertion of a mitochondrial selfish element into the nuclear genome and its consequences. Ecol Evol 10(20):11117–11132. https://doi.org/10.1002/ece3.6749
Eichhorn H, Lessing F, Winterberg B, Schirawski J, Kamper J, Muller P, Kahmann R (2006) A ferroxidation/permeation iron uptake system is required for virulence in Ustilago maydis. Plant Cell 18(11):3332–3345. https://doi.org/10.1105/tpc.106.043588
Gilchrist C, Stelkens R (2019) Aneuploidy in yeast: segregation error or adaptation mechanism? Yeast 36(9):525–539. https://doi.org/10.1002/yea.3427
Grandaubert J, Dutheil JY, Stukenbrock EH (2019) The genomic determinants of adaptive evolution in a fungal pathogen. Evol Lett 3(3):299–312. https://doi.org/10.1002/evl3.117
Gu Q, Ji T, Sun X, Huang H, Zhang H, Lu X, ... & Gao X (2017) Histone H3 lysine 9 methyltransferase FvDim5 regulates fungal development, pathogenicity and osmotic stress responses in Fusarium verticillioides. FEMS Microbiol Lett 364(19):fnx184. https://doi.org/10.1093/femsle/fnx184
Habig M, Lorrain C, Feurtey A, Komluski J, Stukenbrock EH (2021) Epigenetic modifications affect the rate of spontaneous mutations in a pathogenic fungus. Nat Commun 12(1):5869. https://doi.org/10.1038/s41467-021-26108-y
Holliday R (1961) The genetics of Ustilago maydis. Genet Res 2(2):204–230. https://doi.org/10.1017/S0016672300000719
Huang M, Kao KC (2018) Identifying novel genetic determinants for oxidative stress tolerance in Candida glabrata via adaptive laboratory evolution. Yeast 35(11):605–618. https://doi.org/10.1002/yea.3352
Jumper J, Evans R, Pritzel A, Green T, Figurnov M, Ronneberger O, ... & Hassabis D (2021) Highly accurate protein structure prediction with AlphaFold. Nature 596(7873):583–589. https://doi.org/10.1038/s41586-021-03819-2
Kahmann R, Kämper J (2004) Ustilago maydis: how its biology relates to pathogenic development. New Phytol 164(1):31–42. https://doi.org/10.1111/j.1469-8137.2004.01156.x
Kämper J, Kahmann R, Bölker M, Ma LJ, Brefort T, Saville BJ, Banuett F, Kronstad JW, Gold SE, Müller O, Perlin MH, Wösten HAB, De Vries R, Ruiz-Herrera J, Reynaga-Peña CG, Snetselaar K, McCann M, Pérez-Martín J, Feldbrügge M, … Birren BW (2006) Insights from the genome of the biotrophic fungal plant pathogen Ustilago maydis. Nature 444(7115):97–101. https://doi.org/10.1038/nature05248
Kassambara A (2020) rstatix: Pipe-friendly framework for basic statistical tests (R package version 0.6. 0.)
Khateb A, Gago S, Bromley M, Richardson M, Bowyer P (2023) Aneuploidy is associated with azole resistance in Aspergillus fumigatus. Antimicrob Agents Chemother 67(4):e01253-e1322. https://doi.org/10.1128/aac.01253-22
Knaus BJ, Grünwald NJ (2017) VCFR: a package to manipulate and visualize variant call format data in R. Mol Ecol Resour 17(1):44–53. https://doi.org/10.1111/1755-0998.12549
Kramer HM, Cook DE, Seidl MF, Thomma BP (2023) Epigenetic regulation of nuclear processes in fungal plant pathogens. PLoS Pathog 19(8):e1011525. https://doi.org/10.1371/journal.ppat.1011525
Lee S, Oh S, Jeong K, Jo H, Choi Y, Seo HD, ... Lee D (2018) Dot1 regulates nucleosome dynamics by its inherent histone chaperone activity in yeast. Nat Commun 9(1):240. https://doi.org/10.1038/s41467-017-02759-8
Li H, Durbin R (2010) Fast and accurate long-read alignment with burrows-wheeler transform. Bioinformatics 26(5):589–595. https://doi.org/10.1093/bioinformatics/btp698
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, Marth G, Abecasis G, Durbin R (2009) The sequence alignment/map format and SAMtools. Bioinformatics 25(16):2078–2079. https://doi.org/10.1093/bioinformatics/btp352
Li Y, Hu Y, Zhao K, Pan Y, Qu Y, Zhao J, Qin Y (2019) The indispensable role of histone methyltransferase PoDot1 in extracellular glycoside hydrolase biosynthesis of Penicillium oxalicum. Front Microbiol 10:2566. https://doi.org/10.3389/fmicb.2019.02566
Liang L, Liu Y, Yang K, Lin G, Xu Z, Lan H, ... & Wang S (2017) The putative histone methyltransferase DOT1 regulates aflatoxin and pathogenicity attributes in Aspergillus flavus. Toxins 9(7):232. https://doi.org/10.3390/toxins9070232
Lin JS, Happel P, Kahmann R (2021) Nuclear status and leaf tumor formation in the Ustilago maydis–maize pathosystem. New Phytol 231(1):399–415. https://doi.org/10.1111/nph.17377
Linder RA, Greco JP, Seidl F, Matsui T, Ehrenreich IM (2017) The stress-inducible peroxidase TSA2 underlies a conditionally beneficial chromosomal duplication in Saccharomyces cerevisiae. G3: Genes, Genomes, Genetics 7(9):3177–3184. https://doi.org/10.1534/g3.117.300069
López-Martínez LX, Aguirre-Delgado A, Saenz-Hidalgo HK, Buenrostro-Figueroa JJ, García HS, Baeza-Jiménez R (2022) Bioactive ingredients of huitlacoche (Ustilago maydis), a potential food raw material. Food Chem: Mol Sci 4. https://doi.org/10.1016/j.fochms.2022.100076
McKinney W (2010) Data structures for statistical computing in python. Proceedings of the 9th Python in Science Conference, 56-61. https://doi.org/10.25080/Majora-92bf1922-00a
Mirisola MG, Longo VD (2022) Yeast chronological lifespan: longevity regulatory genes and mechanisms. Cells 11(10):1714. https://doi.org/10.3390/cells11101714
Min J, Feng Q, Li Z, Zhang Y, Xu RM (2003) Structure of the catalytic domain of human DOT1L, a non-SET domain nucleosomal histone methyltransferase. Cell 112(5):711–723. https://doi.org/10.1016/S0092-8674(03)00114-4
Molina L, Kahmann R (2007) An Ustilago maydis gene involved in H2O2 detoxification is required for virulence. Plant Cell 19(7):2293–2309. https://doi.org/10.1105/tpc.107.052332
Möller M, Stukenbrock EH (2017) Evolution and genome architecture in fungal plant pathogens. Nat Rev Microbiol 15(12):756–771. https://doi.org/10.1038/nrmicro.2017.76
Möller M, Habig M, Lorrain C, Feurtey A, Haueisen J, Fagundes WC, ... Stukenbrock EH (2021) Recent loss of the Dim2 DNA methyltransferase decreases mutation rate in repeats and changes evolutionary trajectory in a fungal pathogen. PLoS Genet 17(3):e1009448. https://doi.org/10.1371/journal.pgen.1009448
Momeni H, Nazari F (2022) Population genetic structure of Ustilago maydis in Iran. J Crop Prot 12(4):485–497
Munkacsi AB, Kawakami S, Pan JJ, Lee K, Stoxen S, Hang J, May G (2006) Genome-wide assessment of tandem repeat markers for biogeographical analyses of the corn smut fungus, Ustilago maydis. Mol Ecol Notes 6(1):221–223. https://doi.org/10.1111/j.1471-8286.2005.01199.x
Munkacsi AB, Stoxen S, May G (2008) Ustilago maydis populations tracked maize through domestication and cultivation in the Americas. Proc R Soc B: Biol Sci 275(1638):1037–1046. https://doi.org/10.1098/rspb.2007.1636
Ng HH, Feng Q, Wang H, Erdjument-Bromage H, Tempst P, Zhang Y, Struhl K (2002) Lysine methylation within the globular domain of histone H3 by Dot1 is important for telomeric silencing and Sir protein association. Genes Dev 16(12):1518–1527. https://doi.org/10.1101/gad.1001502
Olicón-Hernández DR, Araiza-Villanueva MG, Pardo JP, Aranda E, Guerra-Sánchez G (2019) New insights of Ustilago maydis as yeast model for genetic and biotechnological research: a review. Curr Microbiol 76:917–926. https://doi.org/10.1007/s00284-019-01629-4
Picard Toolkit (2019) Broad Institute, GitHub Repository. https://broadinstitute.github.io/picard/; Broad Institute
Pfaffl MW (2001) A new mathematical model for relative quantification in real-time RT–PCR. Nucleic Acids Res 29(9):e45–e45. https://doi.org/10.1093/nar/29.9.e45
Pradhan A, Herrero-de-Dios C, Belmonte R, Budge S, Lopez Garcia A, Kolmogorova A, ... Brown AJ (2017) Elevated catalase expression in a fungal pathogen is a double-edged sword of iron. PLoS Pathogens 13(5):e1006405. https://doi.org/10.1371/journal.ppat.1006405
Python Software Foundation. (2016). Python 3.6. Python Software Foundation (3.6). https://www.python.org/downloads/release/python-360/
Qi L, Wu XC, Zheng DQ (2019) Hydrogen peroxide, a potent inducer of global genomic instability. Curr Genet 65:913–917. https://doi.org/10.1007/s00294-019-00969-9
R Development Core Team. (2023). R: A language and environment for statistical computing. R Foundation for Statistical Computing. https://www.R-project.org
Schuster M, Schweizer G, Reissmann S, Kahmann R (2016) Genome editing in Ustilago maydis using the CRISPR–CAS system. Fungal Genet Biol 89:3–9. https://doi.org/10.1016/j.fgb.2015.09.001
Schweizer G, Haider MB, Barroso GV, Rössel N, Münch K, Kahmann R, Dutheil JY (2021) Population genomics of the maize pathogen Ustilago maydis: Demographic history and role of virulence clusters in adaptation. Genome Biol Evol 13(5). https://doi.org/10.1093/gbe/evab073
Selmecki AM, Dulmage K, Cowen LE, Anderson JB, Berman J (2009) Acquisition of aneuploidy provides increased fitness during the evolution of antifungal drug resistance. PLoS Genet 5(10). https://doi.org/10.1371/journal.pgen.1000705
Stukenbrock EH, Croll D (2014) The evolving fungal genome. Fungal Biol Rev 28(1):1–12. https://doi.org/10.1016/j.fbr.2014.02.001
Tosh J, Tybulewicz V, Fisher EM (2022) Mouse models of aneuploidy to understand chromosome disorders. Mamm Genome 1–12. https://doi.org/10.1007/s00335-021-09930-z
Tralamazza SM, Gluck-Thaler E, Feurtey A, Croll D (2023) Copy number variation introduced by a massive mobile element underpins global thermal adaptation in a fungal wheat pathogen. bioRxiv, 2023–09. https://doi.org/10.1101/2023.09.22.559077
Vande Zande P, Zhou X, Selmecki A (2023) The dynamic fungal genome: polyploidy, aneuploidy and copy number variation in response to stress. Annu Rev Microbiol 2023(77):341–361. https://doi.org/10.1146/annurev-micro-041320
Villagrán Z, Martínez-Reyes M, Gómez-Rodríguez H, Ríos-García U, Montalvo-González E, Ortiz-Basurto RI, ... Pérez-Moreno J (2023) Huitlacoche (Ustilago maydis), an iconic Mexican fungal resource: biocultural importance, nutritional content, bioactive compounds, and potential biotechnological applications. Molecules 28(11): 4415. https://doi.org/10.3390/molecules28114415
Wang Q, Sun M, Zhang Y, Song Z, Zhang S, Zhang Q, Xu JR, Liu H (2020a) Extensive chromosomal rearrangements and rapid evolution of novel effector superfamilies contribute to host adaptation and speciation in the basal ascomycetous fungi. Mol Plant Pathol 21(3):330–348. https://doi.org/10.1111/mpp.12899
Wang S, Wu XM, Liu CH, Shang JY, Gao F, Guo HS (2020) Verticillium dahliae chromatin remodeling facilitates the DNA damage repair in response to plant ROS stress. PLoS Pathogens 16(4). https://doi.org/10.1371/journal.ppat.1008481
Wickham H (2016) Data Analysis. In: ggplot2. Use R!. Springer, Cham. https://doi.org/10.1007/978-3-319-24277-4_9
Wickham H, François R, Henry L, Müller K, Vaughan D (2023) dplyr: A Grammar of Data Manipulation. https://CRAN.R-project.org/package=dplyr
Williams BR, Prabhu VR, Hunter KE, Glazier CM, Whittaker CA, Housman DE, Amon A (2008) Aneuploidy affects proliferation and spontaneous immortalization in mammalian cells. Science 322(5902):703–709. https://doi.org/10.1126/science.1160058
Zhang K, Zheng DQ, Sui Y, Qi L, Petes TD (2019) Genome-wide analysis of genomic alterations induced by oxidative DNA damage in yeast. Nucleic Acids Res 47(7):3521–3535. https://doi.org/10.1093/nar/gkz027
Acknowledgements
We thank Alejandra Castillo Carbajal, Carina Uribe Díaz, Jair García and Luis Aguilar (LIIGH-UNAM) for their invaluable technical support. We thank LAVIS-UNAM and Max Planck Institute for Evolutionary Biology-GWDG for the use of High-Performance Computing facilities. Additionally, we would like to express our gratitude to Prof. Dr. Julien Y. Dutheil for his guidance during J.C-F.’s internship at the Max Planck Institute for Evolutionary Biology. J.C-F. acknowledges the support of the PhD Program in Biological Sciences from the Autonomous University of Queretaro.
Funding
This work was supported by CONAHCYT grants CB-2016–01 284992 and CF-103000, by UNAM-PAPIIT grants IA201019, IN212524 and IN209021, by UAQ grant FOPER FCN02469 and by CONCYTEQ grant NT/21/2022. J.C-F. has received support from CONAHCYT with the Doctoral fellowship 735110 and from DAAD with the short-term grant number 91835730, that supported his internship at the Max Planck Institute for Evolutionary Biology. M.C-G. receives a Postdoctoral Fellowship from CONAHCYT under the Program “Estancias Posdoctorales por México 2022”.
Author information
Authors and Affiliations
Contributions
All authors read, reviewed, and approved the final manuscript. J.C-F.: Design the study, performed research, analyzed data, created the repository, acquired funding, and wrote the manuscript. M.C-G.: Performed research and wrote the manuscript. S.Q-G.: Performed research. F.R-R.: Performed research. J.A.C–C.: Provided biological resources, contributed to study design, acquired funding, and wrote the manuscript. L.M.: Design the study, acquired funding, and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Cuamatzi-Flores, J., Colón-González, M., Requena-Romo, F. et al. Enhanced oxidative stress resistance in Ustilago maydis and its implications on the virulence. Int Microbiol (2024). https://doi.org/10.1007/s10123-024-00489-8
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10123-024-00489-8