Abstract
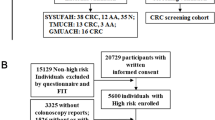
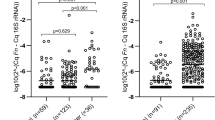
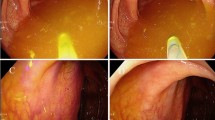
Norway has one of the world’s highest incidences of colorectal cancer (CRC). Accumulating research suggests that the intestinal microbiota may have an important role in initiation and progression of colorectal cancer. In order to evaluate microbiome-based biomarkers for non-invasive detection of CRC, the levels of Fusobacterium nucleatum and selected Escherichia coli toxin genes in stool and mucosa from a small cohort of Norwegian patients were investigated. The study cohort included 72 patients scheduled for colonoscopy. The patients were divided into three groups upon their examinations: cancer, polyp, and control groups. Levels of F. nucleatum in stool samples were significantly higher in the cancer group compared with the control group and the polyp group. High levels of F. nucleatum in stool reflected detection of F. nucleatum in the tumor tissues of colorectal cancer patients. However, no difference in the levels of E. coli toxin genes in neither stool nor biopsy samples between the patient groups was observed. This study suggests that a quantitative PCR assay targeting F. nucleatum in stool samples has the potential to be included in a larger panel of biomarkers for non-invasive testing for colorectal cancer.





Similar content being viewed by others
Change history
28 November 2019
Two affiliations of author John Christopher Noone were not included in the original article and have been added here. Also, Acknowledgments of the originally published article is not complete. Please see the corrected section below.
28 November 2019
Two affiliations of author John Christopher Noone were not included in the original article and have been added here. Also, Acknowledgments of the originally published article is not complete. Please see the corrected section below.
References
Ahlquist DA (2015) Multi-target stool DNA test: a new high bar for noninvasive screening. Dig Dis Sci 60(3):623–633
Niederreiter L, Adolph TE, Tilg H (2018) Food, microbiome and colorectal cancer. Dig Liver Dis 50(7):647–652
Tilg H, Adolph TE, Gerner RR, Moschen AR (2018) The intestinal microbiota in colorectal cancer. Cancer Cell 33(6):954–964
Louis P, Hold GL, Flint HJ (2014) The gut microbiota, bacterial metabolites and colorectal cancer. Nat Rev Microbiol 12(10):661–672
Mima K, Ogino S, Nakagawa S, Sawayama H, Kinoshita K, Krashima R et al (2017) The role of intestinal bacteria in the development and progression of gastrointestinal tract neoplasms. Surg Oncol 26(4):368–376
Leung A, Tsoi H, Yu J (2015) Fusobacterium and Escherichia: models of colorectal cancer driven by microbiota and the utility of microbiota in colorectal cancer screening. Expert Rev Gastroenterol Hepatol 9(5):651–657
Bashir A, Miskeen AY, Bhat A, Fazili KM, Ganai BA (2015) Fusobacterium nucleatum: an emerging bug in colorectal tumorigenesis. Eur J Cancer Prev 24(5):373–385
Bashir A, Miskeen AY, Hazari YM, Asrafuzzaman S, Fazili KM (2016) Fusobacterium nucleatum, inflammation, and immunity: the fire within human gut. Tumour Biol 37(3):2805–2810
Ma CT, Luo HS, Gao F, Tang QC, Chen W (2018) Fusobacterium nucleatum promotes the progression of colorectal cancer by interacting with E-cadherin. Oncol Lett 16(2):2606–2612
Nosho K, Sukawa Y, Adachi Y, Ito M, Mitsuhashi K, Kurihara H et al (2016) Association of Fusobacterium nucleatum with immunity and molecular alterations in colorectal cancer. World J Gastroenterol 22(2):557–566
Rubinstein MR, Wang X, Liu W, Hao Y, Cai G, Han YW (2013) Fusobacterium nucleatum promotes colorectal carcinogenesis by modulating E-cadherin/beta-catenin signaling via its FadA adhesin. Cell Host Microbe 14(2):195–206
Kostic AD, Chun E, Robertson L, Glickman JN, Gallini CA, Michaud M et al (2013) Fusobacterium nucleatum potentiates intestinal tumorigenesis and modulates the tumor-immune microenvironment. Cell Host Microbe 14(2):207–215
Flanagan L, Schmid J, Ebert M, Soucek P, Kunicka T, Liska V et al (2014) Fusobacterium nucleatum associates with stages of colorectal neoplasia development, colorectal cancer and disease outcome. Eur J Clin Microbiol Infect Dis 33(8):1381–1390
Bullman S, Pedamallu CS, Sicinska E, Clancy TE, Zhang X, Cai D et al (2017) Analysis of Fusobacterium persistence and antibiotic response in colorectal cancer. Science 358(6369):1443–1448
Buc E, Dubois D, Sauvanet P, Raisch J, Delmas J, Darfeuille-Michaud A et al (2013) High prevalence of mucosa-associated E. coli producing cyclomodulin and genotoxin in colon cancer. PLoS One 8(2):56964
Raisch J, Buc E, Bonnet M, Sauvanet P, Vazeille E, de Vallee A et al (2014) Colon cancer-associated B2 Escherichia coli colonize gut mucosa and promote cell proliferation. World J Gastroenterol 20(21):6560–6572
Arthur JC, Gharaibeh RZ, Muhlbauer M, Perez-Chanona E, Uronis JM, McCafferty J et al (2014) Microbial genomic analysis reveals the essential role of inflammation in bacteria-induced colorectal cancer. Nat Commun 5:4724
Gomez-Moreno R, Robledo IE, Baerga-Ortiz A (2014) Direct detection and quantification of bacterial genes associated with inflammation in DNA isolated from stool. Adv Microbiol 4(15):1065–1075
Kostic AD, Gevers D, Pedamallu CS, Michaud M, Duke F, Earl AM et al (2012) Genomic analysis identifies association of Fusobacterium with colorectal carcinoma. Genome Res 22(2):292–298
Castellarin M, Warren RL, Freeman JD, Dreolini L, Krzywinski M, Strauss J et al (2012) Fusobacterium nucleatum infection is prevalent in human colorectal carcinoma. Genome Res 22(2):299–306
McCoy AN, Araujo-Perez F, Azcarate-Peril A, Yeh JJ, Sandler RS, Keku TO (2013) Fusobacterium is associated with colorectal adenomas. PLoS One 8(1):53653
Wu N, Yang X, Zhang R, Li J, Xiao X, Hu Y et al (2013) Dysbiosis signature of fecal microbiota in colorectal cancer patients. Microb Ecol 66(2):462–470
Baxter NT, MTt R, Rogers MA, Schloss PD (2016) Microbiota-based model improves the sensitivity of fecal immunochemical test for detecting colonic lesions. Genome Med 8(1):37
Suehiro Y, Sakai K, Nishioka M, Hashimoto S, Takami T, Higaki S et al (2017) Highly sensitive stool DNA testing of Fusobacterium nucleatum as a marker for detection of colorectal tumours in a Japanese population. Ann Clin Biochem 54(1):86–91
Noone JC, Bemanian V, Smith Tunsjø H (2017) DNA-ekstraksjon: Mengde er ikke synonymt med mangfold. Bioingeniøren 6:20–26
Moen AE, Tannaes TM, Vatn S, Ricanek P, Vatn MH, Jahnsen J et al (2016) Simultaneous purification of DNA and RNA from microbiota in a single colonic mucosal biopsy. BMC research notes 9:328
Muller D, Greune L, Heusipp G, Karch H, Fruth A, Tschape H et al (2007) Identification of unconventional intestinal pathogenic Escherichia coli isolates expressing intermediate virulence factor profiles by using a novel single-step multiplex PCR. Appl Environ Microbiol 73(10):3380–3390
Brukner I, Longtin Y, Oughton M, Forgetta V, Dascal A (2015) Assay for estimating total bacterial load: relative qPCR normalisation of bacterial load with associated clinical implications. Diagn Microbiol Infect Dis 83(1):1–6
Saiki RK, Scharf S, Faloona F, Mullis KB, Horn GT, Erlich HA et al (1985) Enzymatic amplification of beta-globin genomic sequences and restriction site analysis for diagnosis of sickle cell anemia. Science 230(4732):1350–1354
Herlemann DP, Labrenz M, Jurgens K, Bertilsson S, Waniek JJ, Andersson AF (2011) Transitions in bacterial communities along the 2000 km salinity gradient of the Baltic Sea. The ISME journal 5(10):1571–1579
Wong SH, Kwong TNY, Chow TC, Luk AKC, Dai RZW, Nakatsu G et al (2017) Quantitation of faecal Fusobacterium improves faecal immunochemical test in detecting advanced colorectal neoplasia. Gut. 66(8):1441–1448
Mira-Pascual L, Cabrera-Rubio R, Ocon S, Costales P, Parra A, Suarez A et al (2015) Microbial mucosal colonic shifts associated with the development of colorectal cancer reveal the presence of different bacterial and archaeal biomarkers. J Gastroenterol 50(2):167–179
Iacopetta B (2001) Are there two sides to colorectal cancer? Int J Cancer 101(5):403–408
Koi M, Okita Y, Carethers JM (2018) Fusobacterium nucleatum infection in colorectal cancer: linking inflammation, DNA mismatch repair and genetic and epigenetic alterations. J anus, rectum and colon 2(2):37–46
Mima K, Cao Y, Chan AT, Qian ZR, Nowak JA, Masugi Y et al (2016) Fusobacterium nucleatum in colorectal carcinoma tissue according to tumor location. Clin Transl Gastroenterol 7(11):200
Arthur JC, Perez-Chanona E, Muhlbauer M, Tomkovich S, Uronis JM, Fan TJ et al (2012) Intestinal inflammation targets cancer-inducing activity of the microbiota (2012). Science 338(6103):120–123
Dutilh BE, Backus L, van Hijum SA, Tjalsma H (2013) Screening metatranscriptomes for toxin genes as functional drivers of human colorectal cancer. Best Pract Res Cl Ga 27(1):85–99
Acknowledgments
Students at Oslo Metropolitan University (Mia Sandgren et al., Sammy Ramstock et al., and Silje Mathiassen et al.) have contributed to parts of the contents in this manuscript. We thank the Pathology Dept. at Akershus University Hospital for characterization of tumors. We also express our gratitude to biomedical laboratory scientist Eva Smedsrud, Dept. of Multidisciplinary Laboratory Science and Medical Biochemistry, Genetic Unit, for all help with patient inclusion and sample processing.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical considerations
The regional committee for medical and health-related research ethics (REK 2012/1944) and the data protection manager at Ahus have approved the study. Written informed consent was obtained from all participants.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Tunsjø, H.S., Gundersen, G., Rangnes, F. et al. Detection of Fusobacterium nucleatum in stool and colonic tissues from Norwegian colorectal cancer patients. Eur J Clin Microbiol Infect Dis 38, 1367–1376 (2019). https://doi.org/10.1007/s10096-019-03562-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10096-019-03562-7