Abstract
Engineered endonucleases that digest the specific sequences can be used to modify target genomes precisely. This is called “genome editing” and is used widely to modify the genome of various organisms. The DNA-binding domains of zinc finger (ZF) proteins were the first to be used as a genome editing tool, in the form of designed ZF nucleases and, more recently, transcription activator-like effector as well as the clustered, regularly interspaced, short palindromic repeats and CRISPR-associated protein 9 (CRISPR/Cas9) system that targets RNA–DNA rather than protein–DNA interactions have been used successfully. These are powerful tools with which targeted gene modifications can be introduced in various organisms, including various plant species. A key step in genome editing is the generation of a double-stranded DNA break that is specific to the target gene. This is achieved using custom-designed endonucleases, which enable site-directed mutagenesis via a non-homologous end joining repair pathway and/or gene targeting via homologous recombination to occur efficiently at specific sites in the genome. This review provides an overview of the current status of genomics and genetic engineering of woody plants, and recent advances in genome editing technologies in plants as well as fungi. We also discuss how these strategies can provide insights into molecular breeding technology for woody plants.
Similar content being viewed by others
Introduction
Woody plants are important components of the global ecosystem; they play an important role in limiting emissions of carbon dioxide and other greenhouse gases, and the water-retaining capacity of forests is critical for flood control. In addition, woody plants are a major biomass resource and are gaining attention as a source of biofuel.
Various useful traits of woody plants have been investigated for sustainable production of biomass, bioremediation using trees, and improvements in the efficiency of energy production from woody plant materials. A good understanding of the key genetics factors regulating the phenotypes involved in those processes is crucial. Various genetic engineering tools have been used to analyze such factors, and important target genes have been manipulated. As a result, many genetically engineered trees with superior traits have been produced.
Recent advances in sequencing technologies have resulted in the availability of whole genome sequences of a number of industrially important woody plants. In addition, in so-called “multi-omics” analyses, data of useful traits have been accumulating rapidly in various woody plants. Such data are needed to develop breeding systems for the production of woody plants with novel favorable traits. These tools will contribute to a more detailed understanding of the gene expression involved in growth and development, environmental stress responses, and also the regulation of cell wall biosynthesis.
Although many instances of the genetic manipulation of important target genes have been reported in trees, the time and effort needed for the establishment of transgenic plants of this type still needs to be reduced by at least one order of magnitude in order for the technology to be cost effective. “Traditional” methods of genetic engineering require lengthy molecular or phenotypic screening to identify the desired characteristics. A recently emerged genetic engineering tool, so-called “genome editing” is now used widely to modify the genome of various organisms. Engineered endonucleases that digest specific sequences in the target genome are the key technology applied in genome editing. Currently, four types of systems are used for this purpose: zinc finger nucleases (ZFNs), transcription activator-like effector nucleases (TALENs), engineered mega-nucleases (EMNs), and the clustered, regularly interspaced, short palindromic repeats and CRISPR-associated protein 9 (CRISPR/Cas9) system. Of these, the CRISPR/Cas9 system has emerged as the most popular method of achieving target-specific manipulation of the gene of interest in the genomes of many organisms, including various plant species. Together with the accumulation of data on woody biomass production and conventional genome modification technologies, genome editing has the potential to produce novel tree species more rapidly than ever.
In this review article, we present a brief history of genetic engineering in woody plants, and then describe recent advances in this new technology, with an emphasis on genome editing to improve plant genomes via the generation of site-directed sequence modifications by engineered nucleases in model plant species. We also discuss the application and future prospects of these technologies in woody plant molecular breeding and biotechnology.
Molecular breeding: genomics and genetic engineering in woody plants
Molecular breeding, which has become an important approach used to accelerate the introduction of useful genetic traits into woody plant genomes, can be divided mainly into two distinct techniques: (1) marker-assisted selection/breeding and quantitative trait loci (QTLs) analysis, and (2) genetic engineering based on recombinant DNA and gene transfer techniques.
Marker-assisted breeding relies on genetic variations (most often DNA polymorphism) and requires many germplasm lines as a resource for selection. Currently, a number of genetic variations and germplasm lines are identified in both hardwoods and softwoods and information on genetic mapping and quantitative trait locus mapping for the valuable traits is available [1–6].
The availability of whole genome sequences of woody plants have helped molecular breeding. Recently, the whole genome sequences of Populus trichocarpa [7] (http://phytozome.jgi.doe.gov/pz/#!info?alias=Org_Ptrichocarpa) and Eucalyptus grandis [8] (http://phytozome.jgi.doe.gov/pz/;#!info?alias=Org_Egrandis) have been determined and published. Such data sets open up a wide range of research activities, such as gene discovery, gene function, gene expression, and comparative genomics as well as physical mapping via bio-informatics approaches. In addition, recent advance of “omics” techniques in woody plants have provided further valuable information.
The transcriptome and proteome provide a dynamic link between the genome and cellular characterization through gene/protein expression data. When initially introduced, microarray-based technology was applied to the transcriptome [9], and online tools comprising a microarray-based expression data (PoplarPLEX; http://www.plexdb.org/plex.php?database=Poplar) and Poplar eFP Browser (http://www.bar.utoronto.ca/efppop/cgi-bin/efpWeb.cgi) were available to better understanding the transcriptome and genome of Populus. Now, with development of the next-generation sequencing technologies, RNA-sequencing analysis is accelerating genome-wide expression studies [10] (http://www.eucgenie.org/).
The metabolome of woody plants also provides valuable resources for molecular breeding applications. The study of the “metabolome” is the study of all the collected end products of gene/protein expression. Therefore, “multi-omics” analyses, such as a combination of transcriptome and metabolome analyses, provide an interactive view of intracellular function [11, 12]. Metabolome analysis is also useful for screening in molecular breeding projects. Indeed, Eckert et al. [13] identified over 3000 associations for a total of 1617 unique single nucleotide polymorphisms (SNPs) associated with 255 metabolites.
Genetic engineering can produce novel traits that would not arise from natural variations in the target genome. In addition, genetic engineering techniques can reduce the breeding period of woody plants, which generally requires a long time when using molecular marker-assisted techniques. Although there are several techniques for genetic engineering for woody plants, Agrobacterium-mediated transformation is currently the method of choice for a wide variety of woody plant species.
Populus species were one of the first hardwood species to undergo genetic transformation, which was established in 1987 to produce herbicide resistance plants [14]. Since then, many transgenic Populus plants have been produced via Agrobacterium-mediated transformation, altering growth and wood characteristics [15], nitrogen metabolism [16], lignin content and character [17, 18] and improving salt tolerance [19, 20].
Eucalyptus species are another main target hardwood species for genetic engineering. Although Eucalyptus plants are relatively difficult to regenerate in tissue culture compared to Populus plants, several research groups have developed improved tissue culture and transformation protocols [21, 22], and both endogenous and exogenous genes have been introduced into Eucalyptus cells to produce Eucalyptus plants with modified secondary cell wall [23] and improved salt tolerance [24, 25].
Although softwoods are more recalcitrant to Agrobacterium-mediated transformation, improvements in tissue culture systems have allowed the production of transgenic softwoods, Pinus radiata [26], Pinus taeda [27, 28], and Picea abies [27]. As in the case of Populus and Eucalyptus plants, the introduction and overexpression of genes of commercial interest have also been demonstrated, e.g., production of salt-tolerant Pinus taeda [29]. A combination of the next-generation sequencing technology, multi-omics, and genetic engineering will also accelerate the molecular breeding of softwoods of commercial interest.
Genome editing: a modern tool for precise genome engineering of woody plants
Natural germplasm variations and mutated germplasm lines obtained using exogenous mutagens have been used in recent decades for breeding to increase quality and yield. Screening of the mutant population to identify individual plants that have the desired phenotype is needed to identify mutant lines of interest. This procedure is quite laborious, and genetic engineering has been used widely to reduce the time required for molecular breeding. However, conventional genetic engineering techniques rely on random gene transfer mechanisms. New methods of targeted gene modification are thus a key technology for improvement of woody plants as well as for analysis of gene function.
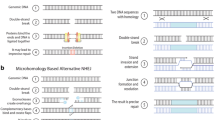
The recent development of engineered nucleases has allowed more precisely targeted gene engineering, namely genome editing. Four types of engineered nuclease systems are currently in use: EMNs, ZFNs, TALENs, and the CRISPR/Cas9 system; all these systems rely on the induction of double-stranded breaks (DSBs) in the target genome DNA [30, 31]. Creating site-directed DSBs in genomic DNA results in gene modifications through either non-homologous end joining (NHEJ) in the case of site-directed mutagenesis or homologous recombination repair (HR) in HR-mediated gene targeting [32].
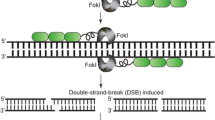
EMNs, ZFNs and TALENs are based on protein–DNA interactions. ZFNs and TALENs utilize the restriction enzyme Fok I [33, 34]. These latter two engineered nucleases comprise a DNA-binding domain (DBD) and a separate nuclease domain, and thus ZFNs and TALENs are easier to customize than EMNs [33, 34]. In the case of ZFNs, the ZF DBD comes from ZF transcription factor [35, 36]. The DBD of ZFNs is typically composed of three-to-four ZF arrays, and each array recognizes three bases. Similar to ZFNs, TALENs use the TALE domain as DBD; this DBD is composed of 34–35 amino acids, and each TALE DBD domain recognizes one base of the target DNA sequence [37]. In both ZFNs and TALENs, the identification and construction of highly specific DBDs are key steps for subsequent genome editing, and these steps are relatively laborious.
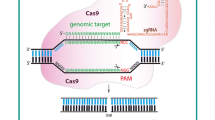
Unlike ZFNs and TALENs, the CRISPR/Cas9 system uses RNA–DNA recognition [38, 39]. The CRISPR/Cas9 system consists of two components: a so-called guide RNA (gRNA) for target recognition; and the RNA-binding/endonuclease protein, Cas9. The gRNA in the current system is a chimeric hybrid of an endogenous bacterial crRNA and tracrRNA, namely single guide RNA (sgRNA) [38, 39]. The base-pairing of part of the gRNA and its complementary DNA sequence on the target genome can guide the complex of sgRNA-Cas9 to the target DNA region, where Cas9 can then introduce a DSB at the target DNA sequence. For correct binding of the sgRNA-Cas9 complex onto the target DNA sequence, a so-called protospacer adjacent motif (PAM) sequence immediately following the target sequence is required. In the most widely used the CRISPR/Cas9 system, derived from Streptococcus pyogenes, the sequence “NGG” (N = A, T, G, C) for the PAM and a 20-nucleotide sequence complementary to the target at the 5′-end of the gRNA as guide is required for correct target recognition [38–40]. These features make the CRISPR/Cas9 system quite simple and easy to design, as well as being highly effective, as proven in human [38] and mouse [39] cells. Following successful site-directed mutagenesis in mammalian cells, the CRISPR/Cas9 system has been used widely in a variety organisms from bacteria to higher eukaryotes: Candida albicans [41], Caenorhabditis elegans [42], fruit fly [43], zebrafish [44], rodents [45], and cattle [46].
In higher plants, genome editing technologies exploiting ZFNs and TALENs have been utilized for site-directed mutagenesis and/or gene targeting in Arabidopsis [47–49], maize [50], and tobacco [51]. Since the appearance of the CRISPR/Cas9 system, the efficacy of genome editing has improved rapidly in a wide variety of plant species.
The first reports using the CRISPR/Cas9 system in plants confirmed that the transient expression of gRNA and Cas9 in Arabidopsis protoplasts, tobacco cells, and rice plants introduced mutations in the target gene of interest [52–54]. To establish knockout model plants lines, Arabidopsis [55] and liverwort [56] were subjected to Agrobacterium-mediated transformation. Fauser et al. [55] demonstrated the stable inheritance of nuclease-induced targeted mutagenesis events in the Arabidopsis ADH1 and TT4 genes at frequencies from 2.5 up to 70.0 % in the T3 generation. Sugano et al. [56] demonstrated the production of liverwort knockout plants for the ARF1 gene using the CRISPR/Cas9 system. It is interesting to note that liverwort is amenable to genome editing using haploid generation to obtain knockout plants in the parent generation of gene transfer (T0 generation). Other than model plants, genome editing with the CRISPR/Cas9 system has been performed in many crop plants, e.g., maize [57], rice [54, 58], sorghum [59], soybean [60, 61], tomato [62], and wheat [54, 63]. These reports demonstrated (1) a high-frequency of mutation following gene transfer of sgRNA/Cas9 via Agrobacterium-mediated transformation, with a variety of mutation rates depending on species; in rice, mutation efficiency reached 100 % [58], (2) the achievement of biallelic mutations after gene transfer of sgRNA/Cas9 in the T0 generation in several crop plants [58, 60–62], and (3) that multiplex gRNA expression can lead to simultaneous mutation of multiple target genes [54, 58, 60–63]. Biallelic mutations produced by the introduction of gRNA/Cas9 genes will have the advantage of obtaining plants with a knockout phenotype at the T0 generation, which will reduce the breeding period. In addition, the successive genome engineering of multiple genes will be of great value in breeding programs where the goal is to disrupt many target genes as well as the knockout of redundant genes.
As mentioned above, genome editing tools, such as ZFNs, TALENs, and the CRISPR/Cas9 system have been developed as site-specific endonucleases to introduce site-specific mutations. Recently, genome editing tools are also used for gene activation/repression and DNA/chromatin modification technologies. ZFs and TALEs were originally identified as transcription factors; therefore, the use of target-specific transcriptional activator/repressor with custom-designed ZFs and TALEs is quite straight forward. In the case of Cas9, the catalytically inactive mutant form of Cas9 (referred to as dCas9 [40]), which contained two mutations of the RuvC1 and HNH nuclease domains can be used for RNA-mediated custom-designed transcriptional factors. Currently, gene fusions of TALEs/dCas9 to several types of activation/repression domains are available for gene regulation tools in many organisms including mammalians and plants [64–66]. Piatek et al. [67] demonstrated the target-specific gene regulation using dCas9-based transcriptional activator and repressor in tobacco. They used the EDLL activation domain [68] and the TAL activation domain [64] for transcriptional activation, and the synthetic SRDX domain [69] for transcriptional repressor.
Effector domains of DNA-/chromatin-modifying enzyme are also used for the genome editing tool. Hilton et al. [70] reported the efficacy of dCas9-based histone acetyltransferase to fuse dCas9 with the catalytic core of the human acetyltransferase p300 in human cells. They demonstrated that the dCas9-based histone acetyltransferase catalyzed acetylation of histone H3 lysine 27 at its target sites, and this acetylation led transcriptional activation of target genes from promoters and enhancers. Establishment of dCas9-based DNA-/chromatin-modifying enzyme other than histone acetyltransferase, such as DNA methyltransferase, methylcytosine deoxygenase, ubiquitin ligase, and poly-ADP ribosyltransferase, will enable to perform precise epigenome editing to control genome-wide gene regulation and chromatin status.
Challenges for genome engineering of woody plants and wood decay basidiomycetes
There are a number of studies of genome editing in higher plants; however, reports of genome editing in tree species are limited. Trees are subject to the difficulties in application of genome editing technologies common to all higher plants: low transformation efficiencies, lack of information for optimal expression cassettes for expressing engineered nucleases, and difficulty of isolation of clonal engineered plants. This is compounded by the slower growth speeds of perennial trees compared to those of grass species. Therefore, the development of genome editing tools for tree species needs to be done in a limited number of trial-and-error cycles.
Jia and Wang [71, 72] reported CRISPR/Cas9-based genome editing in the sweet orange, Citrus sinensis. In this latter report, transient expression of plant-codon optimized Cas9 and CsPDS-targeted gRNA disrupted the endogenous CsPDS locus. Notably, the low transformation efficiency in sweet orange was overcome using Agrobacterium infection facilitated by pre-infection with Xanthomonas citri subsp. citri (Xcc). Xcc improves infection efficiencies of Agrobacterium to citrus [72]. In addition, the cauliflower mosaic virus (CaMV) 35S promoter, which is transcribed by RNA polymerase II, was used for the expression of gRNA in sweet orange [71]. RNA polymerase III-transcribed promoters, such as U3 and U6, have more commonly been used to express gRNA. However, in tree species, information on U3/U6-snRNA expression is somewhat lacking. RNA polymerase II-based gRNA expression is thus one of the possible approaches currently being followed up to design predictable CRISPR/Cas9-based expression in trees.
Peer et al. [73] utilized ZFN targeting of the uidA transgene, which expresses β-glucuronidase (GUS), and demonstrated ZFN-based site-directed mutagenesis in apple, Malus domestica, and fig, Ficus carica. Similar to the work in Arabidopsis [31], a heat-shock promoter was used to express ZFN cassettes to avoid toxicity of ZFN [73]. In this report, individual plants were produced and cloned after a tissue culture period of almost a year. Since Agrobacterium-mediated transformation of engineered nucleases produces NHEJ-based mutagenesis only in transfected cells, isolation of transfected cells and their regeneration is an unavoidable step in the cloning of engineered plants [73]. As another approach, systemic infection with a virus harboring genome editing vectors, which would not require regeneration from mutated cells if genome editing occurred in shoot apical meristems, was proposed by Peer et al. [73]. Combination of a systemic virus and an engineered nuclease is one possible approach to rapid breeding in woody plants.
Most recently, Zhou et al. [74] demonstrated the successful gene knockout of 4-coumarate:CoA ligase, the 4CL1 gene in Populus tremula × alba clone 717-1B4. The 4CL1 gene is a key gene in lignin biosynthesis [17, 75]. Although an RNA polymerase III-transcribed promoter has not been cloned so far in Populus, Zhou et al. used the Medicago U6.6 snRNA gene promoter [60] for gRNA expression, and highly efficient biallelic mutation of the 4CL1 gene was achieved. Mutant lines with these biallelic mutations showed similar phenotypes as in transgenic poplar plants expressing antisense 4CL1 RNA [75]. As mentioned above, highly efficient regeneration and transformation procedures have been developed for Populus plants. The establishment of efficient transformation methods, optimization of expression cassettes, and isolation methodologies of clonal plants will be helpful in developing genome editing tools in woody plants.
For efficient usage of woody plants biomass, it is necessary not only to breed woody plants themselves but also to utilize fungal and bacterial species that decay difficult-to-use biomass. Reports on the molecular breeding of wood decay basidiomycetes have been limited; however, there are some examples of molecular breeding in basidiomycete mushrooms and cellulose-degrading bacteria.
Schizophyllum commune is a wood-rotting fungus that has also found a use as one of the model organisms of mushrooms [76, 77]. Polyethylene glycol (PEG)-based transformation methods and HR-based gene targeting has already been developed in S. commune [77], although the HR rate was relatively low and selection markers should be used [77]. Using disruption of transcription factors fst3 and fst4, artificial regulation of mushroom development in S. commune has already been reported [76]. Higher HR rates could be achieved using artificially engineered nucleases, and other genetic factors regulating mushroom development and lignin-degradation may be uncovered.
Coprinopsis cinerea is another model organism of mushroom, and its genome had already been uncovered [78]. Some tools for molecular biology of this organism have been developed, such as PEG-based transformation, RNAi, fluorescent reporters, and HR-based gene targeting using selection markers [79]. C. cinerea generally grows on dung—a non-woody substrate—and is thought to be a non-wood-rotting fungus. However, heterologous expressions of lignin-degradation enzymes in Coprinopsis had been shown to increase lignin-decolorization activities [80]. Recently, electroporation-based gene delivery has been developed in Coprinopsis (Sugano et al. unpublished). Electroporation might possibly allow mRNA transfection. Like mRNA-injection-based genome editing in mouse and zebrafishes, transfection of engineered nuclease mRNA to basidiomycetes would open up marker-free genome manipulation in wood decay fungi. Without the need for foreign selection markers, fungal species derived from molecular breeding could rapidly be brought to market.
To date, there are no reports of genome editing in basidiomycetes; however, demonstration of efficient Platinum-TALEN-based genome editing (nearly 100 % gene targeting) in a filamentous fungus, Pyricularia oryzae, has been reported [81], so it is thought to be only a matter of time until genome editing in basidiomycetes is accomplished. Besides demonstration of genome editing per se, actual utilization of genome editing in basidiomycetes might be subject to a specific fungal problem. Basidiomycetes live dominantly as heterokaryotes; therefore, establishment of clonal individuals is more difficult than in plants. Cloning technologies using monokaryotic stages of fungal cells would be required.
Recently, CRISPR/Cas9-based genome editing in the wood decay bacteria, Clostridium cellulolyticum, has been reported: Xu et al. [82] reported that Cas9 expression with gRNA, which induces DSB in vivo, was lethal in C. cellulolyticum. It was assumed that C. cellulolyticum might have less active NHEJ-based repair pathways. On the other hand, “nickase type” Cas9n had been shown to induce highly efficient (>95 %) HR-based gene targeting at the targeted locus [74]. This high gene targeting efficiency will open the door to high-throughput molecular genetics in wood decay bacteria.
Concerning the molecular tools that have been developed in basidiomycetes, and the recent demonstration of highly efficient genome editing in filamentous fungi and wood decay bacteria, molecular breeding of wood decay fungi and bacteria look to be accelerated in the near future.
Concluding remarks and future prospects
Targeted gene engineering has been achieved effectively by custom-designed engineered nucleases. In gene modification, these “targetable” nucleases have the potential to become alternatives to standard breeding methods to identify novel traits in economically important plants, especially once the efficacy of the CRISPR/Cas9 system has been improved. In crop plants, several features of genome editing have been reported: (1) highly efficient gene disruption mediated by the coupling of the CRISPR/Cas9 system and Agrobacterium-mediated transformation, (2) multiple-gene disruption can be achieved using multiplex gRNA expression, and (3) biallelic mutations often occur.
However, to extend genome engineering technologies to make them more applicable and useful for woody plant species, further improvements are required to overcome their limitations. Highly specific and efficient genome editing systems will be required in woody plants, because most woody plant species exhibit lower transformation efficiencies compared to annual crop plant species tested with the CRISPR/Cas9 system.
The generation of off-target mutations is a problem encountered with the CRISPR/Cas9 system that will also need to be overcome in plant genome editing. One solution for the off-target problem was demonstrated recently using a double-nicking CRISPR/Cas9 system [83]. This system has already been tested in plants [55] and, therefore, could also be valuable for woody plant species. In addition to the double-nicking CRISPR/Cas9 system, a Fok I-based Cas9 nuclease system [84] and a novel gRNA design system with a 17- to 18-nt target sequence [85] have also been reported as a means of drastically reducing off-target mutation efficiency. A combination of these techniques will be required for precise genome engineering in woody plant species.
As new plant-breeding techniques develop, these efforts, together with a deeper understanding of the whole genome structure and function of wide variety of woody plants, will enable the development of future technologies in breeding novel and important traits in woody plants as well as fungal and bacterial species for efficient usage of woody plants biomass.
References
Berlin S, Lagercrantz U, von Arnold S, Ost T, Rönnberg-Wästljung AC (2010) High-density linkage mapping and evolution of paralogs and orthologs in Salix and Populus. BMC Genom 11:129
Kang BY, Major JE, Rajora OP (2011) A high-density genetic linkage map of a black spruce (Picea mariana) × red spruce (Picea rubens) interspecific hybrid. Genome 54:128–143
Hudson CJ, Freeman JS, Kullan AR, Petroli CD, Sansaloni CP, Kilian A, Detering F, Grattapaglia D, Potts BM, Myburg AA, Vaillancourt RE (2012) A reference linkage map for Eucalyptus. BMC Genom 13:240
Neves LG, Davis JM, Barbazuk WB, Kirst M (2014) A high-density gene map of loblolly pine (Pinus taeda L.) based on exome sequence capture genotyping. G3 (Bethesda) 4:29–37
Freeman JS, Potts BM, Downes GM, Pilbeam D, Thavamanikumar S, Vaillancourt RE (2013) Stability of quantitative trait loci for growth and wood properties across multiple pedigrees and environments in Eucalyptus globulus. New Phytol 198:1121–1134
Muchero W, Guo J, DiFazio SP, Chen JG, Ranjan P, Slavov GT, Gunter LE, Jawdy S, Bryan AC, Sykes R, Ziebell A, Klápště J, Porth I, Skyba O, Unda F, El-Kassaby YA, Douglas CJ, Mansfield SD, Martin J, Schackwitz W, Evans LM, Czarnecki O, Tuskan GA (2015) High-resolution genetic mapping of allelic variants associated with cell wall chemistry in Populus. BMC Genom 16:24
Tuskan GA, Difazio S, Jansson S, Bohlmann J, Grigoriev I, Hellsten U, Putnam N, Ralph S, Rombauts S, Salamov A, Schein J, Sterck L, Aerts A, Bhalerao RR, Bhalerao RP, Blaudez D, Boerjan W, Brun A, Brunner A, Busov V, Campbell M, Carlson J, Chalot M, Chapman J, Chen GL, Cooper D, Coutinho PM, Couturier J, Covert S, Cronk Q, Cunningham R, Davis J, Degroeve S, Déjardin A, Depamphilis C, Detter J, Dirks B, Dubchak I, Duplessis S, Ehlting J, Ellis B, Gendler K, Goodstein D, Gribskov M, Grimwood J, Groover A, Gunter L, Hamberger B, Heinze B, Helariutta Y, Henrissat B, Holligan D, Holt R, Huang W, Islam-Faridi N, Jones S, Jones-Rhoades M, Jorgensen R, Joshi C, Kangasjärvi J, Karlsson J, Kelleher C, Kirkpatrick R, Kirst M, Kohler A, Kalluri U, Larimer F, Leebens-Mack J, Leplé JC, Locascio P, Lou Y, Lucas S, Martin F, Montanini B, Napoli C, Nelson DR, Nelson C, Nieminen K, Nilsson O, Pereda V, Peter G, Philippe R, Pilate G, Poliakov A, Razumovskaya J, Richardson P, Rinaldi C, Ritland K, Rouzé P, Ryaboy D, Schmutz J, Schrader J, Segerman B, Shin H, Siddiqui A, Sterky F, Terry A, Tsai CJ, Uberbacher E, Unneberg P, Vahala J, Wall K, Wessler S, Yang G, Yin T, Douglas C, Marra M, Sandberg G, Van de Peer Y, Rokhsar D (2006) The genome of black cottonwood, Populus trichocarpa (Torr. & Gray). Science 313:1596–1604
Myburg AA, Grattapaglia D, Tuskan GA, Hellsten U, Hayes RD, Grimwood J, Jenkins J, Lindquist E, Tice H, Bauer D, Goodstein DM, Dubchak I, Poliakov A, Mizrachi E, Kullan AR, Hussey SG, Pinard D, van der Merwe K, Singh P, van Jaarsveld I, Silva-Junior OB, Togawa RC, Pappas MR, Faria DA, Sansaloni CP, Petroli CD, Yang X, Ranjan P, Tschaplinski TJ, Ye CY, Li T, Sterck L, Vanneste K, Murat F, Soler M, Clemente HS, Saidi N, Cassan-Wang H, Dunand C, Hefer CA, Bornberg-Bauer E, Kersting AR, Vining K, Amarasinghe V, Ranik M, Naithani S, Elser J, Boyd AE, Liston A, Spatafora JW, Dharmwardhana P, Raja R, Sullivan C, Romanel E, Alves-Ferreira M, Külheim C, Foley W, Carocha V, Paiva J, Kudrna D, Brommonschenkel SH, Pasquali G, Byrne M, Rigault P, Tibbits J, Spokevicius A, Jones RC, Steane DA, Vaillancourt RE, Potts BM, Joubert F, Barry K, Pappas GJ, Strauss SH, Jaiswal P, Grima-Pettenati J, Salse J, Van de Peer Y, Rokhsar DS, Schmutz J (2014) The genome of Eucalyptus grandis. Nature 510:356–362
Wilkins O, Waldron L, Nahal H, Provart NJ, Campbell MM (2009) Genotype and time of day shape the Populus drought response. Plant J 60:703–715
Mizrachi E, Hefer CA, Ranik M, Joubert F, Myburg AA (2010) De novo assembled expressed gene catalog of a fast-growing Eucalyptus tree produced by Illumina mRNA-Seq. BMC Genom 11:681
Srivastava V, Obudulu O, Bygdell J, Löfstedt T, Rydén P, Nilsson R, Ahnlund M, Johansson A, Jonsson P, Freyhult E, Qvarnström J, Karlsson J, Melzer M, Moritz T, Trygg J, Hvidsten TR, Wingsle G (2013) OnPLS integration of transcriptomic, proteomic and metabolomic data shows multi-level oxidative stress responses in the cambium of transgenic hipI- superoxide dismutase Populus plants. BMC Genom 14:893
Hamanishi ET, Barchet GL, Dauwe R, Mansfield SD, Campbell MM (2015) Poplar trees reconfigure the transcriptome and metabolome in response to drought in a genotype- and time-of-day-dependent manner. BMC Genom 16:329
Eckert AJ, Wegrzyn JL, Cumbie WP, Goldfarb B, Huber DA, Tolstikov V, Fiehn O, Neale DB (2012) Association genetics of the loblolly pine (Pinus taeda, Pinaceae) metabolome. New Phytol 193:890–902
Fillatti JJ, Kiser J, Rose R, Comai L (1987) Efficient transfer of a glyphosate tolerance gene into tomato using a binary Agrobacterium tumefaciens vector. Bio Technol 5:726–730
Tuominen H, Sitbon F, Jacobsson C, Sandberg G, Olsson O, Sundberg B (1995) Altered growth and wood characteristics in transgenic hybrid aspen expressing Agrobacterium tumefaciens T-DNA indoleacetic acid-biosynthetic genes. Plant Physiol 109:1179–1189
Gallardo F, Fu J, Canton FR, Garcia-Gutierrez A, Canovas FM, Kirby EG (1999) Expression of a conifer glutamine synthetase gene in transgenic poplar. Planta 210:19–26
Hu WJ, Kawaoka A, Tsai CJ, Lung JH, Osakabe K, Ebinuma H, Chiang VL (1998) Compartmentalized expression of two structurally and functionally distinct 4-coumarate:CoA ligase genes in aspen (Populus tremuloides). Proc Natl Acad Sci USA 95:5407–5412
Li L, Zhou Y, Cheng X, Sun J, Marita JM, Ralph J, Chiang VL (2013) Combinatorial modification of multiple lignin traits in trees through multigene cotransformation. Proc Natl Acad Sci USA 100:4939–4944
Hu L, Lu H, Liu Q, Chen X, Jiang X (2005) Overexpression of mtlD gene in transgenic Populus tomentosa improves salt tolerance through accumulation of mannitol. Tree Physiol 25:1273–1281
Li Y, Su X, Zhang B, Huang Q, Zhang X, Huang R (2009) Expression of jasmonic ethylene responsive factor gene in transgenic poplar tree leads to increased salt tolerance. Tree Physiol 29:273–279
Tournier V, Grat S, Marque C, El Kayal W, Penchel R, de Andrade G, Boudet AM, Teulières C (2003) An efficient procedure to stably introduce genes into an economically important pulp tree (Eucalyptus grandis × Eucalyptus urophylla). Transgenic Res 12:403–411
Chen ZZ, Ho CK, Ahn IS, Chiang VL (2006) Eucalyptus. Methods Mol Biol 344:125–134
Hussey SG, Mizrachi E, Spokevicius AV, Bossinger G, Berger DK, Myburg AA (2011) SND2, a NAC transcription factor gene, regulates genes involved in secondary cell wall development in Arabidopsis fibres and increases fibre cell area in Eucalyptus. BMC Plant Biol 11:173
Matsunaga E, Nanto K, Oishi M, Ebinuma H, Morishita Y, Sakurai N, Suzuki H, Shibata D, Shimada T (2012) Agrobacterium-mediated transformation of Eucalyptus globulus using explants with shoot apex with introduction of bacterial choline oxidase gene to enhance salt tolerance. Plant Cell Rep 31:225–235
Yu X, Kikuchi A, Shimazaki T, Yamada A, Ozeki Y, Matsunaga E, Ebinuma H, Watanabe KN (2013) Assessment of the salt tolerance and environmental biosafety of Eucalyptus camaldulensis harboring a mangrin transgene. J Plant Res 126:141–150
Grant JE, Cooper PA, Dale TM (2004) Transgenic Pinus radiata from Agrobacterium tumefaciens-mediated transformation of cotyledons. Plant Cell Rep 22:894–902
Wenck AR, Quinn M, Whetten RW, Pullman G, Sederoff R (1999) High-efficiency Agrobacterium-mediated transformation of Norway spruce (Picea abies) and loblolly pine (Pinus taeda). Plant Mol Biol 39:407–416
Tang W, Sederoff R, Whetten R (2001) Regeneration of transgenic loblolly pine (Pinus taeda L.) from zygotic embryos transformed with Agrobacterium tumefaciens. Planta 213:981–989
Tang W, Peng X, Newton RJ (2005) Enhanced tolerance to salt stress in transgenic loblolly pine simultaneously expressing two genes encoding mannitol-1-phosphate dehydrogenase and glucitol-6-phosphate dehydrogenase. Plant Physiol Biochem 43:139–146
Urnov FD, Rebar EJ, Holmes MC, Zhang HS, Gregory PD (2010) Genome editing with engineered zinc finger nucleases. Nat Rev Genet 11:636–646
Osakabe Y, Osakabe K (2015) Genome editing with engineered nucleases in plants. Plant Cell Physiol 56:389–400
Osakabe K, Endo M, Toki S (2012) Chapter I–1.6: DNA double-strand breaks and homologous recombination in higher plants. In: Quingyao S (ed) Plant mutagenesis—principles and applications. The Smiling Hippo, Greece, pp 71–80
Urnov FD, Miller JC, Lee YL, Beausejour CM, Rock JM, Augustus S, Jamieson AC, Porteus MH, Gregory PD, Holmes MC (2005) Highly efficient endogenous human gene correction using designed zinc-finger nucleases. Nature 435:646–651
Christian M, Cermak T, Doyle EL, Schmidt C, Zhang F, Hummel A, Bogdanove AJ, Voytas DF (2010) Targeting DNA double-strand breaks with TAL effector nucleases. Genetics 186:757–761
Mandell JG, Barbas CF III (2006) Zinc finger tools: custom DNA-binding domains for transcription factors and nucleases. Nucleic Acids Res 34:W516–W523 (Web Server issue)
Maeder ML, Thibodeau-Beganny S, Osiak A, Wright DA, Anthony RM, Eichtinger M, Jiang T, Foley JE, Winfrey RJ, Townsend JA, Unger-Wallace E, Sander JD, Müller-Lerch F, Fu F, Pearlberg J, Göbel C, Dassie JP, Pruett-Miller SM, Porteus MH, Sgroi DC, Iafrate AJ, Dobbs D, McCray PB Jr, Cathomen T, Voytas DF, Joung JK (2008) Rapid “open-source” engineering of customized zinc-finger nucleases for highly efficient gene modification. Mol Cell 31:294–301
Cermak T, Starker CG, Voytas DF (2015) Efficient design and assembly of custom TALENs using the Golden Gate platform. Methods Mol Biol 1239:133–159
Cong L, Ran FA, Cox D, Lin S, Barretto R, Habib N, Hsu PD, Wu X, Jiang W, Marraffini LA, Zhang F (2013) Multiplex genome engineering using CRISPR/Cas systems. Science 339:819–823
Mali P, Yang L, Esvelt KM, Aach J, Guell M, DiCarlo JE, Norville JE, Church GM (2013) RNA-guided human genome engineering via Cas9. Science 339:823–826
Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA, Charpentier E (2012) A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337:816–821
Vyas VK, Barrasa MI, Fink GR (2015) A Candida albicans CRISPR system permits genetic engineering of essential genes and gene families. Sci Adv 1:e1500248
Friedland AE, Tzur YB, Esvelt KM, Colaiácovo MP, Church GM, Calarco JA (2013) Heritable genome editing in C. elegans via a CRISPR-Cas9 system. Nat Methods 10:741–743
Lee JS, Kwak SJ, Kim J, Kim AK, Noh HM, Kim JS, Yu K (2014) RNA-guided genome editing in Drosophila with the purified Cas9 protein. G3 (Bethesda) 4:1291–1295
Hwang WY, Fu Y, Reyon D, Maeder ML, Tsai SQ, Sander JD, Peterson RT, Yeh JR, Joung JK (2013) Efficient genome editing in zebrafish using a CRISPR-Cas system. Nat Biotechnol 31:227–229
Li D, Qiu Z, Shao Y, Chen Y, Guan Y, Liu M, Li Y, Gao N, Wang L, Lu X, Zhao Y, Liu M (2013) Heritable gene targeting in the mouse and rat using a CRISPR-Cas system. Nat Biotechnol 31:681–683
Heo YT, Quan X, Xu YN, Baek S, Choi H, Kim NH, Kim J (2015) CRISPR/Cas9 nuclease-mediated gene knock-in in bovine-induced pluripotent cells. Stem Cells Dev 24:393–402
Osakabe K, Osakabe Y, Toki S (2010) Site-directed mutagenesis in Arabidopsis using custom-designed zinc finger nucleases. Proc Natl Acad Sci USA 107:12034–12039
Zhang F, Maeder ML, Unger-Wallace E, Hoshaw JP, Reyon D, Christian M, Li X, Pierick CJ, Dobbs D, Peterson T, Joung JK, Voytas DF (2010) High frequency targeted mutagenesis in Arabidopsis thaliana using zinc finger nucleases. Proc Natl Acad Sci USA 107:12028–12033
Cermak T, Doyle EL, Christian M, Wang L, Zhang Y, Schmidt C, Baller JA, Somia NV, Bogdanove AJ, Voytas DF (2011) Efficient design and assembly of custom TALEN and other TAL effector-based constructs for DNA targeting. Nucleic Acids Res 39:e82
Shukla VK, Doyon Y, Miller JC, DeKelver RC, Moehle EA, Worden SE, Mitchell JC, Arnold NL, Gopalan S, Meng X, Choi VM, Rock JM, Wu YY, Katibah GE, Zhifang G, McCaskill D, Simpson MA, Blakeslee B, Greenwalt SA, Butler HJ, Hinkley SJ, Zhang L, Rebar EJ, Gregory PD, Urnov FD (2009) Precise genome modification in the crop species Zea mays using zinc-finger nucleases. Nature 459:437–441
Townsend JA, Wright DA, Winfrey RJ, Fu F, Maeder ML, Joung JK, Voytas DF (2009) High-frequency modification of plant genes using engineered zinc-finger nucleases. Nature 459:442–445
Li JF, Norville JE, Aach J, McCormack M, Zhang D, Bush J, Church GM, Sheen J (2013) Multiplex and homologous recombination–mediated genome editing in Arabidopsis and Nicotiana benthamiana using guide RNA and Cas9. Nat Biotechnol 31:688–691
Nekrasov V, Staskawicz B, Weigel D, Jones JD, Kamoun S (2013) Targeted mutagenesis in the model plant Nicotiana benthamiana using Cas9 RNA guided endonuclease. Nat Biotechnol 31:691–693
Shan Q, Wang Y, Li J, Zhang Y, Chen K, Liang Z, Zhang K, Liu J, Xi JJ, Qiu JL, Gao C (2013) Targeted genome modification of crop plants using a CRISPR-Cas system. Nat Biotechnol 31:686–688
Fauser F, Schiml S, Puchta H (2014) Both CRISPR/Cas-based nucleases and nickases can be used efficiently for genome engineering in Arabidopsis thaliana. Plant J 79:348–359
Sugano SS, Shirakawa M, Takagi J, Matsuda Y, Shimada T, Hara-Nishimura I, Kohchi T (2014) CRISPR/Cas9-mediated targeted mutagenesis in the liverwort Marchantia polymorpha L. Plant Cell Physiol 55:475–481
Liang Z, Zhang K, Chen K, Gao C (2014) Targeted mutagenesis in Zea mays using TALENs and the CRISPR/Cas system. J Genet Genomics 41:63–68
Xie K, Minkenberg B, Yang Y (2015) Boosting CRISPR/Cas9 multiplex editing capability with the endogenous tRNA-processing system. Proc Natl Acad Sci USA 112:3570–3575
Jiang W, Zhou H, Bi H, Fromm M, Yang B, Weeks DP (2013) Demonstration of CRISPR/Cas9/sgRNA-mediated targeted gene modification in Arabidopsis, tobacco, sorghum and rice. Nucleic Acids Res 41:e188
Jacobs TB, LaFayette PR, Schmitz RJ, Parrott W (2015) Targeted genome modifications in soybean with CRISPR/Cas9. BMC Biotechnol 15:16
Sun X, Hu Z, Chen R, Jiang Q, Song G, Zhang H, Xi Y (2015) Targeted mutagenesis in soybean using the CRISPR-Cas9 system. Sci Rep 5:10342
Brooks C, Nekrasov V, Lippman ZB, Van Eck J (2014) Efficient gene editing in tomato in the first generation using the clustered regularly interspaced short palindromic repeats/CRISPR-associated9 system. Plant Physiol 166:1292–1297
Upadhyay SK, Kumar J, Alok A, Tuli R (2013) RNA-guided genome editing for target gene mutations in wheat. G3 (Bethesda) 3:2233–2238
Mahfouz MM, Li L, Piatek M, Fang X, Mansour H, Bangarusamy DK, Zhu JK (2012) Targeted transcriptional repression using a chimeric TALE-SRDX repressor protein. Plant Mol Biol 78:311–321
Crocker J, Stern DL (2013) TALE-mediated modulation of transcriptional enhancers in vivo. Nat Methods 10:762–767
Maeder ML, Linder SJ, Cascio VM, Fu Y, Ho QH, Joung JK (2013) CRISPR RNA-guided activation of endogenous human genes. Nat Methods 10:977–979
Piatek A, Ali Z, Baazim H, Li L, Abulfaraj A, Al-Shareef S, Aouida M, Mahfouz MM (2015) RNA-guided transcriptional regulation in planta via synthetic dCas9-based transcription factors. Plant Biotechnol J 13:578–589
Tiwari SB, Belachew A, Ma SF, Young M, Ade J, Shen Y, Marion CM, Holtan HE, Bailey A, Stone JK, Edwards L, Wallace AD, Canales RD, Adam L, Ratcliffe OJ, Repetti PP (2012) The EDLL motif: a potent plant transcriptional activation domain from AP2/ERF transcription factors. Plant J 70:855–865
Hiratsu K, Matsui K, Koyama T, Ohme-Takagi M (2003) Dominant repression of target genes by chimeric repressors that include the EAR motif, a repression domain, in Arabidopsis. Plant J 34:733–739
Hilton IB, D’Ippolito AM, Vockley CM, Thakore PI, Crawford GE, Reddy TE, Gersbach CA (2015) Epigenome editing by a CRISPR-Cas9-based acetyltransferase activates genes from promoters and enhancers. Nat Biotechnol 33:510–517
Jia H, Wang N (2014) Targeted genome editing of sweet orange using Cas9/sgRNA. PLoS One 9:e93806
Jia H, Wang N (2014) Xcc-facilitated agroinfiltration of citrus leaves: a tool for rapid functional analysis of transgenes in citrus leaves. Plant Cell Rep 33:1993–2001
Peer R, Rivlin G, Golobovitch S, Lapidot M, Gal-On A, Vainstein A, Tzfira T, Flaishman MA (2015) Targeted mutagenesis using zinc-finger nucleases in perennial fruit trees. Planta 241:941–951
Zhou X, Jacobs TB, Xue LJ, Harding SA, Tsai CJ (2015) Exploiting SNPs for biallelic CRISPR mutations in the outcrossing woody perennial Populus reveals 4-coumarate:CoA ligase specificity and redundancy. New Phytol 208:298–301
Hu WJ, Harding SA, Lung J, Popko JL, Ralph J, Stokke DD, Tsai CJ, Chiang VL (1999) Repression of lignin biosynthesis promotes cellulose accumulation and growth in transgenic trees. Nat Biotechnol 17:808–812
Ohm RA, de Jong JF, Lugones LG, Aerts A, Kothe E, Stajich JE, de Vries RP, Record E, Levasseur A, Baker SE, Bartholomew KA, Coutinho PM, Erdmann S, Fowler TJ, Gathman AC, Lombard V, Henrissat B, Knabe N, Kües U, Lilly WW, Lindquist E, Lucas S, Magnuson JK, Piumi F, Raudaskoski M, Salamov A, Schmutz J, Schwarze FW, van Kuyk PA, Horton JS, Grigoriev IV, Wösten HA (2010) Genome sequence of the model mushroom Schizophyllum commune. Nat Biotechnol 28:957–963
Ohm RA, de Jong JF, Berends E, Wang F, Wösten HA, Lugones LG (2010) An efficient gene deletion procedure for the mushroom-forming basidiomycete Schizophyllum commune. World J Microbiol Biotechnol 26:1919–1923
Stajich JE, Wilke SK, Ahrén D, Au CH, Birren BW, Borodovsky M, Burns C, Canbäck B, Casselton LA, Cheng CK, Deng J, Dietrich FS, Fargo DC, Farman ML, Gathman AC, Goldberg J, Guigó R, Hoegger PJ, Hooker JB, Huggins A, James TY, Kamada T, Kilaru S, Kodira C, Kües U, Kupfer D, Kwan HS, Lomsadze A, Li W, Lilly WW, Ma LJ, Mackey AJ, Manning G, Martin F, Muraguchi H, Natvig DO, Palmerini H, Ramesh MA, Rehmeyer CJ, Roe BA, Shenoy N, Stanke M, Ter-Hovhannisyan V, Tunlid A, Velagapudi R, Vision TJ, Zeng Q, Zolan ME, Pukkila PJ (2010) Insights into evolution of multicellular fungi from the assembled chromosomes of the mushroom Coprinopsis cinerea (Coprinus cinereus). Proc Natl Acad Sci USA 107:11889–11894
Ohm RA, Riley R, Salamov A, Min B, Choi IG, Grigoriev IV (2014) Genomics of wood-degrading fungi. Fungal Genet Biol 72:82–90
Ogawa K, Yamazaki T, Hasebe T, Kajiwara S, Watanabe A, Asada Y, Shishido K (1998) Molecular breeding of the basidiomycete Coprinus cinereus strains with high lignin-decolorization and -degradation activities using novel heterologous protein expression vectors. Appl Microbiol Biotechnol 49:285–289
Arazoe T, Ogawa T, Miyoshi K, Yamato T, Ohsato S, Sakuma T, Yamamoto T, Arie T, Kuwata S (2015) Tailor-made TALEN system for highly efficient targeted gene replacement in the rice blast fungus. Biotechnol Bioeng 112:1335–1342
Xu T, Li Y, Shi Z, Hemme CL, Li Y, Zhu Y, Van Nostrand JD, He Z, Zhou J (2015) Efficient genome editing in Clostridium cellulolyticum via CRISPR-Cas9 nickase. Appl Environ Microbiol 81:4423–4431
Ran FA, Hsu PD, Lin CY, Gootenberg JS, Konermann S, Trevino AE, Scott DA, Inoue A, Matoba S, Zhang Y, Zhang F (2013) Double nicking by RNA-guided CRISPR/Cas9 for enhanced genome editing specificity. Cell 154:1380–1389
Tsai SQ, Wyvekens N, Khayter C, Foden JA, Thapar V, Reyon D, Goodwin MJ, Aryee MJ, Joung JK (2014) Dimeric CRISPR RNA-guided FokI nucleases for highly specific genome editing. Nat Biotechnol 32:569–576
Fu Y, Sander JD, Reyon D, Cascio VM, Joung JK (2014) Improving CRISPR-Cas nuclease specificity using truncated guide RNAs. Nat Biotechnol 32:279–284
Acknowledgments
This work was supported by the Grant-in-Aid for Scientific Research C from the Japan Society for the Promotion of Science (21580125 to Y.O. and 25450002 to K.O.). This work was also supported by the Council for Science, Technology and Innovation (CSTI), Cross-ministerial Strategic Innovation Promotion Program (SIP), “Technologies for creating next-generation agriculture, forestry and fisheries” (funding agency: Bio-oriented Technology Research Advancement Institution, NARO).
Author information
Authors and Affiliations
Corresponding author
Additional information
This review article is published to coincide with the 60th anniversary of the Japan Wood Research Society.
Rights and permissions
This article is published under an open access license. Please check the 'Copyright Information' section either on this page or in the PDF for details of this license and what re-use is permitted. If your intended use exceeds what is permitted by the license or if you are unable to locate the licence and re-use information, please contact the Rights and Permissions team.
About this article
Cite this article
Osakabe, Y., Sugano, S.S. & Osakabe, K. Genome engineering of woody plants: past, present and future. J Wood Sci 62, 217–225 (2016). https://doi.org/10.1007/s10086-016-1548-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10086-016-1548-5