Abstract
The emergence of antibiotic resistance in Acinetobacter spp. is a rising public health concern worldwide. The objective of this study was to investigate the prevalence of antibiotic-resistance genes and the virulence of Acinetobacter spp. isolated from soil and crops obtained from agricultural fields in South Korea. Eight Acinetobacter spp. isolates carried various antibiotic resistance genes, such as emrAB (100%), cat/craA (100%), and aadA gene (87.5%). Minimum inhibitory concentration (MIC) analysis revealed that strains harboring antibiotic resistance genes exhibited high resistance to the respective antibiotics, such as colistin, chloramphenicol, and streptomycin. Interestingly, most of these isolates had high capability of biofilm formation and swarming motility, along with faster growth rates. Taken together, our study demonstrated that antibiotic-resistant Acinetobacter isolated from agricultural settings in South Korea not only frequently carries antibiotic resistance genes but also has virulence-related traits.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Acinetobacter spp. are Gram-negative, aerobic, opportunistic bacteria which are widely distributed in hospital environments or hospitalized patients, and also in soil and water (Cisneros and Rodríguez-Baño, 2002). Acinetobacter spp. can survive on either dry or moist surfaces for long periods and grow at a range of different temperatures and pH values. Acinetobacter spp. are also known to form biofilms, known as the most effective virulence factor, which further compromises the use of antibiotics in treating infections. Acinetobacter spp., especially A. baumannii, have been numerously reported for increasing involvement in outbreaks in clinical settings. Furthermore, the emergence of multidrug-resistant (MDR) Acinetobacter has led to a rise in public health concerns (Carvalheira et al., 2021). According to the World Health Organization (WHO), one of the most emphasized antibiotic-resistant “priority pathogens” for which new antibiotics are urgently needed is A. baumannii (Tacconelli et al., 2018). Antibiotic resistance genes are easily transferred in Acinetobacter especially by conjugation, leading to the increasing emergence of MDR Acinetobacter (Leungtongkam et al., 2018).
Recent studies report that Acinetobacter spp. can be introduced into the hospital environment through kitchens or by food (Campos et al., 2019). Several studies have also reported that MDR Acinetobacter spp. is found in agricultural products such as lettuce and fruits (Carvalheira et al., 2017). MDR Acinetobacter spp. poses a major threat to hospitals causing severe nosocomial infections that are related to high mortality rates. However, few studies have reported the prevalence of MDR Acinetobacter spp. in agricultural environments in South Korea (Ababneh et al., 2022; Kim et al., 2016). The purpose of this study was to investigate the prevalence of antimicrobial resistance of Acinetobacter spp. among soil and agricultural products from agricultural fields in South Korea and analyze their virulence-related traits.
Materials and methods
Isolation of Acinetobacter spp. from agricultural environments in South Korea
Soil and crops from agricultural environments were collected randomly from different local fields located in three Korean provinces, Seoul, Jeju, and Chungnam, using standard procedures in September 2022. Collected samples were enriched with Luria–Bertani (LB) broth (Difco, Detroit, MI, USA) at 37°C overnight and plated onto the selective agar medium of Acinetobacter, CHROMagar™ Acinetobacter (CHROMagar, Paris, France). The plates were incubated at 37°C overnight. Colonies on plates were picked and confirmed with PCR using specific primers for Acinetobacter and 16S rRNA sequencing (see supplementary data Table S1). Isolated Acinetobacter spp. were stored in glycerol stock at − 80°C. The Acinetobacter spp. isolates were grown on LB media at 37°C.
Whole genome sequencing and bioinformatics analysis
Genomic DNA (gDNA) was extracted using PureHelix Genomic DNA Prep Kit (Solution Type)-Bacteria (Nanohelix, Daejon, Korea). Then, gDNA was quantified and qualified by gel electrophoresis 260/230 nm and 260/280 nm absorbance ratio and Quant-iT™ PicoGreenR dsDNA Assay Kit (Invitrogen). Following the manufacturer’s guidelines, a Nanopore MinION long-read sequencing library was prepared using a ligation sequencing-native barcoding kit (SQK-NBD114.24; Oxford Nanopore Technologies, UK) and sequenced on a FLO-MIN114 (R10.4.1) flow cell with a MinION Mk1B device and MinKNOW software (22.10.7). De novo assembly were conducted using Flye version 2.9.1 (Kolmogorov et al., 2019). Pangenome analysis was performed using Roary Pangenome Pipeline (Page et al., 2015). Roary was used (with a 95% BLASTp percentage identity cut-off) to cluster the genes encoding complete protein sequences into core (hard core and soft core) and accessory (shell and cloud) genomes.
Growth curve
Overnight cultures were diluted 1:100 in fresh LB broth in a 24-well plate. Growth was monitored every 15 min for 12 h by measuring optical density at 600 nm (OD600) using the SpectraMax i3 Plus Microplate Spectrophotometer (Molecular Devices, USA). Experiments were conducted in triplicates.
Antimicrobial susceptibility testing
Antimicrobial susceptibility was determined using a broth dilution method according to the Clinical and Laboratory Standards Institute (CLSI) guideline (CLSI, 2020). The minimum inhibitory concentrations (MICs) of colistin (Sigma), chloramphenicol (Duchefa), streptomycin (Sigma), gentamycin (Sigma), piperacillin (Duchefa), cefotaxime (Sigma), tetracycline (Sigma), ciprofloxacin (Sigma), doripenem (Sigma), imipenem (Sigma), meropenem (Sigma, Saint Louis, MO, USA) were determined using E. coli ATCC 25922 (American Type Culture Collection; ATCC, Manassas, VA, USA) as a quality control strain (CLSI, 2020). The antibiotic resistance level of chloramphenicol and streptomycin was determined based on previous studies (Wei and Yang, 2017; Yang et al., 2019).
Biofilm formation assay
The biofilm formation capacity of each strain was estimated using the crystal violet staining method described previously (Cha et al., 2019). Briefly, strains were cultured in LB broth overnight and was diluted to reach OD600 = 0.1 with fresh LB broth in 96-well plates. After incubation at 37°C for 24 h in static conditions, the plates were washed three times with PBS (Dulbecco’s Phosphate-Buffered Saline, GenDEPOT), and each well was stained with 200 μl of 0.1% crystal violet (CV) for 20 min. The plates were again washed three times to remove excess stain, and solubilized with 200 μl of 33% acetic acid for 20 min. The OD570 was then measured using the microplate spectrophotometer, to calculate the biofilm formation capacity of the isolates. The extent of biofilm formation of each isolate was compared to the reference strain, A. baumannii CCARM 12001, which is known as a multidrug-resistant strain. A. baumannii CCARM 12001 was obtained from the Culture Collection of Antimicrobial Resistant Microbes, Korea. Un-inoculated LB broth was used as a negative control. All experiments were carried out in triplicates.
Swarming motility assay
Freshly grown cultures were stabbed to enable spread of bacteria on the surface of the medium (0.4% semisolid) as described previously (Choi et al., 2013). Modified LB broth (tryptone—10 g/l; NaCl—5 g/l; yeast extract—5 g/l; micro-agar—4 g/l) was used for the motility assay (Vijayakumar et al., 2016). Plates were prepared on the same day as the inoculation. A 2 μl aliquot of an overnight culture diluted to an OD600 of 1 was spotted in the middle of the plate and allowed to dry at room temperature. Then, plates were incubated at 37°C for 24 h. Experiments were performed at least three times.
Results and discussion
Isolation and identification of Acinetobacter isolates
Eight strains of Acinetobacter spp. were isolated from soil and crops from agricultural fields in South Korea. Various Acinetobacter spp. was identified including A. seifertii, A. vivianii, A. pittii, and A. baumannii isolates (Table 1). A. seifertii and A. pittii are close relatives of A. baumannii isolates, the control of which is critical in clinical settings (Towner, 2009). The phylogenetic tree of the 8 isolated strains was constructed, showing that the isolates could be divided into 2 big clades, where RAES06 is distinctly separated from the rest of the 7 isolates (Fig. 1).
Antimicrobial resistance profiles and pangenome analysis of Acinetobacter isolates
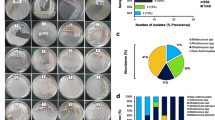
Antibiotic susceptibility results showed that Acinetobacter isolates were highly resistant to colistin (8/8, 100%), chloramphenicol (7/8, 87.5%), and streptomycin (4/8, 50%), whereas all isolates were susceptible to gentamycin, piperacillin, cefotaxime, tetracycline, ciprofloxacin, and carbapenems (Table 2). Since previous studies have reported that mcr-1 gene was detected in colistin-resistant Acinetobacter, we tested the presence of mcr-1 gene in the isolates (Hameed et al., 2019; Kalová et al., 2021). The mcr-1 gene was not detected in all isolates based on PCR results (data not shown). However, the emrA or emrB gene, which encodes efflux pumps associated with colistin resistance in Acinetobacter, was present in all isolates (see supplementary data Table S2) (Boinett et al., 2019; Lin et al., 2017). Based on the pangenome analysis, we discovered poor correlation between the aadA gene and streptomycin resistance in isolates (Fig. 1 and Table S2). The aadA gene, which encodes 3′′-adenylyltransferase, causes resistance to streptomycin and spectinomycin (Svab et al., 1990). All isolates, except RAES03, encoded the aadA gene, but only 50% of the isolates showed high resistance to streptomycin (see supplementary data Table S2). This result may be explained by poor expression or silencing of the aadA gene (Schmidt et al., 2018). All isolates also encoded the cat and the craA gene, which are both related to chloramphenicol resistance, and all isolates except RAES06 showed high resistance to chloramphenicol (see supplementary data Table S2) (Devaud et al., 1982; Roca et al., 2009). It is notable that Acinetobacter isolates from agricultural fields are mostly resistant to colistin, chloramphenicol, and especially to streptomycin, given the prevalent and abundant use of antibiotics as pesticides in South Korea (Lee et al., 2021; Sundin and Wang, 2018).
Growth rates of Acinetobacter isolates
The acquisition of antibiotic resistance is known to impair bacterial fitness resulting in a growth defect (Andersson and Hughes, 2010; Song et al., 2014). To estimate the growth rates of Acinetobacter spp. isolates, we measured bacterial growth with the MDR A. baumannii CCARM 12001 as a control strain. Most isolates exhibited significantly higher growth rates compared to the control strain (Fig. 2). Notably, RAES13, RAES19 and RAES20 showed a more rapid exponential growth phase compared to the other strains, with RAES06 exhibiting the slowest growth (Fig. 2). Interestingly, the fast-growing isolates showed the highest antibiotic resistance (27.3%) among the eight isolates, whereas RAES06 was the most susceptible (9.0%) to antibiotics (Table 2).
Growth curve of Acinetobacter isolates. Growth was monitored every 30 min for 12 h. The inset shows a magnified view at the end of the growth curve. Experiments were repeated three times and the data represent mean ± standard deviation. Results were analyzed based on CCARM 12001 results by the Student’s t-test. *P < 0.05; **P < 0.01; ns, not significant
Biofilm formation of Acinetobacter isolates
Previous study states that biofilm formation is one of the major virulence factors of Acinetobacter. (Dehbanipour and Ghalavand, 2022). To determine the extent of biofilm formation of each isolate, the biofilm formation assay was conducted and the biofilm formation ability of each isolate was compared to the MDR A. baumannii CCARM 12001 strain. RAES01, RAES06, RAES13, and RAES20 showed similar values with the control strain, exhibiting strong biofilm formation capability (Fig. 3). On the other hand, RAES03, RAES10, RAES11, and RAES19 showed low biofilm formation compared to the control strain (Fig. 3). Some reports claim the correlation between antibiotic resistance and biofilm formation in Acinetobacter (Babapour et al., 2016; Bardbari et al., 2017). However, our results do not show any correlation between antibiotic resistance and biofilm formation (Table 2 and Fig. 3). Considering that growth and motility affect biofilm formation of bacteria, further study is required for deeper analysis (Goller and Romeo, 2008). Our results support the idea that biofilm formation does not necessarily associate with antibiotic resistance in Acinetobacter.
Biofilm formation of Acinetobacter isolates. Biofilm was quantified utilizing the crystal violet staining assay and measuring optical density at 570 (OD570). Isolates were incubated at 37°C for 24 h in static conditions. Experiments were repeated three times and the data represent mean ± standard deviation. Results were analyzed based on CCARM 12001 results by the Student’s t-test. *P < 0.05; **P < 0.01; ns, not significant
Motility assay of Acinetobacter isolates
Motility is one of the important virulence factors of Acinetobacter (Dehbanipour and Ghalavand, 2022). To compare the motility of the Acinetobacter isolates, we investigated the swarming motility with comparison to A. baumannii CCARM 12001. All isolates were highly motile compared to the control strain (Fig. 4). RAES06 showed the lowest motility among the isolates, which may be associated with its slow growth rate (Fig. 2). Interestingly, isolates with high biofilm-producing ability were revealed to have relatively low motility than low biofilm-forming isolates in our study (Figs. 3 and 4). Low motility is suggested to be coupled with higher adherence to surfaces, enhancing the efficiency of biofilm formation. Our results demonstrated the strong correlation between low motility and high biofilm formation in Acinetobacter spp. isolated from agricultural environments.
Swarming motility of Acinetobacter isolates. (A) Bar diagram of bacterial motility. The diameter of bacterial cells present on the plate was each measured for quantitative analysis. (B) Motility agar plates 24 h post-inoculation of Acinetobacter isolates. Experiments were repeated in triplicates and the data represent mean ± standard deviation. Results were analyzed based on CCARM 12001 results by the Student’s t-test. *P < 0.05; **P < 0.01; ***P < 0.001
In conclusion, the findings in this study demonstrated that Acinetobacter isolates from agricultural fields in South Korea were resistant to many antibiotics and showed a high biofilm formation and motility capability. Since contaminated agricultural products can be a possible transmission route of MDR Acinetobacter spp., further investigation is needed to control and reduce MDR Acinetobacter spp. in agricultural environments.
References
Ababneh, Q., Al-Rousan, E., & Jaradat, Z. Fresh produce as a potential vehicle for transmission of Acinetobacter baumannii. International Journal of Food Contamination. 9(1): 5 (2022)
Andersson, D. I., & Hughes, D. Antibiotic resistance and its cost: is it possible to reverse resistance? Nature Reviews Microbiology. 8: 260-271 (2010)
Babapour, E., Haddadi, A., Mirnejad, R., Angaji, S.-A., & Amirmozafari, N. Biofilm formation in clinical isolates of nosocomial Acinetobacter baumannii and its relationship with multidrug resistance. Asian Pacific Journal of Tropical Biomedicine. 6: 528-533 (2016)
Bardbari, A. M., Arabestani, M. R., Karami, M., Keramat, F., Alikhani, M. Y., & Bagheri, K. P. Correlation between ability of biofilm formation with their responsible genes and MDR patterns in clinical and environmental Acinetobacter baumannii isolates. Microbial Pathogenesis. 108: 122-128 (2017)
Boinett CJ, Cain AK, Hawkey J, Do Hoang NT, Khanh NNT, Thanh DP, Dordel J, Campbell JI, Lan NPH, Mayho M, Langridge GC, Hadfield J, Chau NVV, Thwaites GE, Parkhill J, Thomson NR, Holt KE, Baker S. Clinical and laboratory-induced colistin-resistance mechanisms in Acinetobacter baumannii. Microbial Genomics. 5: e000246 (2019)
Campos, A., Lopes, M. S., Carvalheira, A., Barbosa, J., & Teixeira, P. Survival of clinical and food Acinetobacter spp. isolates exposed to different stress conditions. Food Microbiology. 77: 202-207 (2019)
Carvalheira, A., Silva, J., & Teixeira, P. Lettuce and fruits as a source of multidrug resistant Acinetobacter spp. Food Microbiology. 64: 119-125 (2017)
Carvalheira, A., Silva, J., & Teixeira, P. Acinetobacter spp. in food and drinking water—a review. Food Microbiology. 95: 103675 (2021)
Cha, Y., Son, B., & Ryu, S. Effective removal of staphylococcal biofilms on various food contact surfaces by Staphylococcus aureus phage endolysin LysCSA13. Food Microbiology. 84: 103245 (2019)
Choi, Y., Shin, H., Lee, J.-H., & Ryu, S. Identification and characterization of a novel flagellum-dependent Salmonella-infecting bacteriophage, iEPS5. Applied and Environmental Microbiology. 79: 4829-4837 (2013)
Cisneros, J. M., & Rodríguez-Baño, J. Nosocomial bacteremia due to Acinetobacter baumannii: epidemiology, clinical features and treatment. Clinical Microbiology and Infection. 8: 687-693 (2002)
CLSI. CLSI M100-ED30:2020 Performance Standards for Antimicrobial Susceptibility Testing, 30th Edition (2020)
Dehbanipour, R., & Ghalavand, Z. Acinetobacter baumannii: Pathogenesis, virulence factors, novel therapeutic options and mechanisms of resistance to antimicrobial agents with emphasis on tigecycline. Journal of Clinical Pharmacy and Therapeutics. 47: 1875-1884 (2022)
Devaud, M., Kayser, F. H., & Bächi, B. Transposon-mediated multiple antibiotic resistance in Acinetobacter strains. Antimicrobial Agents and Chemotherapy. 22: 323-329 (1982)
Goller, C., & Romeo, T. Environmental influences on biofilm development. Bacterial Biofilms. 322: 37-66 (2008)
Hameed, F., Khan, M. A., Muhammad, H., Sarwar, T., Bilal, H., & Rehman, T. U. Plasmid-mediated mcr-1 gene in Acinetobacter baumannii and Pseudomonas aeruginosa: first report from Pakistan. Revista Da Sociedade Brasileira De Medicina Tropical. 52: e20190237 (2019)
Kalová, A., Gelbíčová, T., Overballe-Petersen, S., Litrup, E., & Karpíšková, R. Characterisation of colistin-resistant enterobacterales and Acinetobacter strains carrying mcr genes from asian aquaculture products. Antibiotics (Basel). 10: 838 (2021)
Kim, S. Y., Kuppusamy, S., Kim, J. H., Yoon, Y.-E., Kim, K.-R., & Lee, Y. B. Occurrence and diversity of tetracycline resistance genes in the agricultural soils of South Korea. Environmental Science and Pollution Research. 23: 22190-22196 (2016)
Kolmogorov, M., Yuan, J., Lin, Y., & Pevzner, P. A. Assembly of long, error-prone reads using repeat graphs. Nature Biotechnology. 37: 540-546 (2019)
Lee, Y. S., Kim, G. H., Koh, Y. J., & Jung, J. S. Identification of strA-strB genes in streptomycin-resistant Pseudomonas syringae pv. actinidiae Biovar 2 strains isolated in Korea. The Plant Pathology Journal. 37: 489-493 (2021)
Leungtongkam, U., Thummeepak, R., Tasanapak, K., & Sitthisak, S. Acquisition and transfer of antibiotic resistance genes in association with conjugative plasmid or class 1 integrons of Acinetobacter baumannii. PLoS ONE. 13: e0208468 (2018)
Lin, M.-F., Lin, Y.-Y., & Lan, C.-Y. Contribution of EmrAB efflux pumps to colistin resistance in Acinetobacter baumannii. Journal of Microbiology. 55: 130-136 (2017)
Page AJ, Cummins CA, Hunt M, Wong VK, Reuter S, Holden MT, Fookes M, Falush D, Keane JA, Parkhill J. Roary: rapid large-scale prokaryote pan genome analysis. Bioinformatics. 31: 3691-3693 (2015)
Roca, I., Marti, S., Espinal, P., Martínez, P., Gibert, I., & Vila, J. CraA, a major facilitator superfamily efflux pump associated with chloramphenicol resistance in Acinetobacter baumannii. Antimicrobial Agents and Chemotherapy. 53: 4013-4014 (2009)
Schmidt, K., Stanley, K. K., Hale, R., Smith, L., Wain, J., O'Grady, J., & Livermore, D. M. Evaluation of multiplex tandem PCR (MT-PCR) assays for the detection of bacterial resistance genes among Enterobacteriaceae in clinical urines. Journal of Antimicrobial Chemotherapy. 74: 349-356 (2018)
Song T, Park Y, Shamputa IC, Seo S, Lee SY, Jeon HS, Choi H, Lee M, Glynne RJ, Barnes SW, Walker JR, Batalov S, Yusim K, Feng S, Tung CS, Theiler J, Via LE, Boshoff HI, Murakami KS, Korber B, Barry CE 3rd, Cho SN. Fitness costs of rifampicin resistance in Mycobacterium tuberculosis are amplified under conditions of nutrient starvation and compensated by mutation in the β' subunit of RNA polymerase. Molecular Microbiology. 91: 1106-1119 (2014)
Sundin, G. W., & Wang, N. Antibiotic resistance in plant-pathogenic bacteria. Annual Review of Phytopathology. 56(1): 161-180 (2018)
Svab, Z., Harper, E. C., Jones, J. D., & Maliga, P. Aminoglycoside-3′′-adenyltransferase confers resistance to spectinomycin and streptomycin in Nicotiana tabacum. Plant Molecular Biology. 14: 197-205 (1990)
Tacconelli E, Carrara E, Savoldi A, Harbarth S, Mendelson M, Monnet DL, Pulcini C, Kahlmeter G, Kluytmans J, Carmeli Y, Ouellette M, Outterson K, Patel J, Cavaleri M, Cox EM, Houchens CR, Grayson ML, Hansen P, Singh N, Theuretzbacher U, Magrini N; WHO Pathogens Priority List Working Group. Discovery, research, and development of new antibiotics: the WHO priority list of antibiotic-resistant bacteria and tuberculosis. The Lancet Infectious Diseases. 18: 318-327 (2018)
Towner, K. J. Acinetobacter: an old friend, but a new enemy. Journal of Hospital Infection. 73: 355-363 (2009).
Vijayakumar, S., Rajenderan, S., Laishram, S., Anandan, S., Balaji, V., & Biswas, I. Biofilm formation and motility depend on the nature of the Acinetobacter baumannii clinical isolates. Frontiers in Public Health. 4: 105 (2016)
Wei, W.-j., & Yang, H.-f. Synergy against extensively drug-resistant Acinetobacter baumannii in vitro by two old antibiotics: colistin and chloramphenicol. International Journal of Antimicrobial Agents. 49: 321-326 (2017)
Yang, C. H., Su, P. W., Moi, S. H., & Chuang, L. Y. Biofilm formation in Acinetobacter Baumannii: genotype-phenotype correlation. Molecules. 24: 1849 (2019)
Acknowledgments
S.M.S. and E.A. were supported by the BK21 Plus Program of the Department of Agricultural Biotechnology, Seoul National University, Seoul, Korea. This work was carried out with the support of “The Cooperative Research Program for Agricultural Science & Technology Development (Project No. PJ016120)” Rural Development Administration, Republic of Korea.
Funding
Open Access funding enabled and organized by Seoul National University.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
We declare no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Son, S.M., Ahn, E., Ahn, S. et al. Prevalence of antibiotic-resistant Acinetobacter spp. on soil and crops collected from agricultural fields in South Korea. Food Sci Biotechnol 33, 1931–1937 (2024). https://doi.org/10.1007/s10068-023-01496-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10068-023-01496-7