Abstract
Objective
Recent studies have demonstrated an altered expression of certain microRNAs in patients with rheumatoid arthritis (RA) as well as their first-degree relatives (FDRs) compared to healthy controls (HCs), suggesting a role of microRNA in the progression of the disease. To corroborate this, a set of well-characterized RA families originating from northern Sweden were analyzed for differential expression of a selected set of microRNAs.
Method
MicroRNA was isolated from frozen peripheral blood cells obtained from 21 different families and included 26 RA patients, 22 FDRs, and 21 HCs. Expression of the selected microRNAs miR-22-3p, miR-26b-5p, miR-34a-3p, miR-103a-3p, miR-142-3p, miR-146a-5p, miR-155, miR-346, and miR-451a was determined by a two-step quantitative real-time polymerase chain reaction (qRT-PCR). Statistical analysis including clinical variables was applied.
Results
Out of the nine selected microRNAs that previously have been linked to RA, we confirmed four after adjusting for age and gender, i.e., miR-22-3p (p = 0.020), miR-26b-5p (p = 0.018), miR-142-3p (p = 0.005), and miR-155 (p = 0.033). Moreover, a significant trend with an intermediate microRNA expression in FDR was observed for the same four microRNAs. In addition, analysis of the effect of corticosteroid use showed modulation of miR-103a-3p expression.
Conclusions
We confirm that microRNAs seem to be involved in the development of RA, and that the expression pattern in FDR is partly overlapping with RA patients. The contribution of single microRNAs in relation to the complex network including all microRNAs and other molecules is still to be revealed.
Key Points • Expression levels of miR-22-3p, miR-26b-5p, miR-142-3p, and miR-155 were significantly altered in RA patients compared to those in controls. • In first-degree relatives, a significant trend with an intermediate microRNA expression in FDR was observed for the same four microRNAs. |
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Rheumatoid arthritis (RA) is a chronic autoimmune disease that affects 0.3–1% of the world population. RA is more common in northern Europe and North America, with predominance among women and elderly [1]. Previous studies suggest an interplay between genetic factors, environmental factors, and the immune system [2,3,4] although the pathogenesis of RA is still not fully understood. In families containing RA individuals, the majority of first-degree relatives (FDRs) never develop RA, even if they display risk factors such as smoking, anti-citrullinated protein antibodies (ACPAs), or carriage of HLA-shared epitope.
MicroRNA (miRNA or miR) is a non-coding sequence of single-stranded RNA that is approximately 18–25 nucleotides in length [5]. Its main function is to alter gene expression by post-transcriptional modifications and has shown to regulate about 30% of the protein coding genes. The mature microRNA will bind to a complementary sequence on mRNA that, depending on the grade of complementarity, will either lead to degradation of the mRNA or interfere with the translation process. Regarding this, microRNA primarily downregulates gene expression [2, 6]. A single microRNA can have the ability to regulate multiple genes, and it is also possible that one gene can be regulated by several microRNAs [2].
In a recent study based on indigenous North Americans, Anaparti et al. have shown an altered expression of several microRNAs in blood samples of RA patients as well as their FDRs, compared to healthy controls (HCs) [5]. Several other studies have similarly identified a dysregulated expression of microRNAs within the inflamed joints [7,8,9,10] and serum [11] as well as peripheral blood mononuclear cells (PBMCs) [12] from patients with RA, indicating that microRNA might have a role in the pathogenesis of the disease.
In a previous study of RA patients, their FDRs, and HCs originating from the four northernmost counties of Sweden [13], we have shown differences in the levels of IgG, IgA, and IgM ACPAs as well as of rheumatoid factor (RF) of IgM and IgA isotype. However, microRNA expression in peripheral blood cells has not been analyzed in this group. In this study, the expression of a number of carefully chosen microRNAs was studied. Thus, miR-22-3p, miR-26b-5p, miR-34a-3p, miR-103a-3p, miR-142-3p, miR-146a-5p, miR-155, miR-346, and miR-451a were examined in peripheral blood cells of our cohort comparing the three groups: patients with RA, their FDRs, and HCs.
Materials and methods
Study design
Participants within this study originate from the four most northern counties in Sweden. The RA families were identified by a questionnaire provided to patients with RA at their respective rheumatology clinic. A total of 194 patients with confirmed diagnosis of RA (1987 ARA criteria) [14] and 191 unaffected FDRs were recruited from northern Sweden. All FDRs were interviewed by a questionnaire and all of the relatives with symptoms or signs of joint disease were clinically assessed by a rheumatologist as previously reported [13]. For this study, members from the 21 families living closest to the University Hospital in Umeå participated, resulting in 26 patients with RA and 22 FDRs. For comparison, we included 21 HCs in the study (Table 1). The FDRs included in the study were given a second questionnaire about symptoms and signs of joint disease and were clinically evaluated by a rheumatologist if signs of joint disease were reported. Blood samples were collected in PAXgene Blood RNA tubes (PreAnalytiX, Hombrechtikon, Switzerland) and were kept in − 80 °C until microRNA isolation.
Ethical considerations
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards. Informed consent was obtained from all individual participants included in the study. Ethical permits for this study were from the Regional Ethics Committee, Umeå (Dnr 05-068M and Dnr 2016-216-32M).
Isolation of microRNA
The collected blood samples were defrosted in room temperature for 2 h and turned eight to ten times, then centrifuged for 10 min at 3000g in 4 °C using a centrifuge with a swing-out rotor (Allegra® X-15R Centrifuge, Beckman Coulter). The supernatant was removed and 1 mL Gibco™ PBS (pH 7.4, 1×, Thermo Fisher scientific, Waltham, MA, USA) was added to the pellet. The samples were transferred into 2-mL tubes and centrifuged for 10 min at 3000g in 4 °C and the supernatant was removed. The microRNA was isolated from the pellet using MirVana miRNA Isolation Kit (Life Technologies Europe B.V., Bleiswijk, Netherlands) according to the manufacturer’s instructions with the following changes: instead of acid-phenol:chloroform:isoamyl alcohol (125:24:1, pH 4.5 ± 0.2), phenol:chloroform:isoamyl alcohol (25:24:1, pH 6.7 ± 0.2) or phenol:chloroform:isoamyl alcohol (125:24:1, pH 4–5) with an additional volume of chloroform in a 2:1 ratio was used. Nuclease-free water was used as elution solution. The concentration of the isolated RNA was determined by a NanoDrop® ND-1000 Spectrophotometer with software NanoDrop 1000 3.7.1, or Thermo Scientific™ NanoDrop™ One C, where the absorbance at 230 nm, 260 nm, and 280 nm was measured. The samples were diluted with nuclease-free water to a final concentration of 2.0 ng/μL.
Reverse transcription PCR
TaqMan® MicroRNA Reverse Transcription Kit (Life Technologies Europe B.V.) was used in order to perform the reverse transcription. RT primers from TaqMan® MicroRNA assays for RNU48 (assay ID: 001006), U6 snRNA (assay ID: 001973), hsa-miR-22-3p (assay ID: 000398), hsa-miR-26b-5p (assay ID: 000407), hsa-miR-34a-3p (assay ID: 002316), hsa-miR-103a-3p (assay ID: 000439), hsa-miR-142-3p (assay ID: 000464), hsa-miR-146a-5p (assay ID: 000468), hsa-miR-155 (assay ID: 002623), hsa-miR-346 (assay ID: 000553), and mmu-miR-451a (assay ID: 001141) with a concentration of 5× were used (Life Technologies Europe B.V.). The reverse transcription reaction was performed according to the protocol of the manufacturer for TaqMan® Small RNA Assays. The cDNA was amplified in PTC-100™ Programmable Thermal Controller, and the cDNA was stored in − 20 °C until qPCR was performed.
Quantitative PCR
qPCR was performed using TaqMan™ Fast Universal PCR Master Mix (2×) no AmpErase™ UNG (Life Technologies Europe B.V.) and primers and probes from TaqMan® MicroRNA assays as mentioned above, with a concentration of 20× (Life Technologies Europe B.V.). The PCR reaction mix was prepared according to the manufacturer’s protocol for TaqMan® Small RNA Assays. Negative controls, without cDNA, were run on every plate. In the majority of cases, samples and negative controls were analyzed in triplicates. The qPCR was performed in QuantStudio5 and analyzed in QuantStudio™ Design and Analysis Software (Applied Biosystems, Foster City, CA, USA). For each of the nine microRNAs analyzed, the threshold value was adjusted to lay within the linear phase of the amplification curve and was held constant for all analyses of one particular microRNA. In initial analyses, two reference genes, i.e., RNU48 and U6 snRNA, were tested. The U6 snRNA was found to display the lowest deviation in expression in between the runs and was therefore selected as the reference gene.
The cycle threshold (CT) values received from the duplicates and triplicates were used to calculate a mean CT value for each sample. An inter-experimental reference sample was included in all runs, and the mean CT values for all microRNAs were adjusted to this reference sample. ΔCT was calculated as [CT (target) − CT (U6 snRNA)] and used in order to calculate ΔΔCT [ΔCT (RA) − ΔCT (HC)], [ΔCT (FDR) − ΔCT (HC)], or [ΔCT (RA) − ΔCT (FDR)]. Fold difference in expression was calculated as 2−ΔΔCT.
Statistical analysis and construction of figures
SPSS software version 26.0 (IBM, NY, USA) was used for statistical analyses. Comparisons between the groups of continuous variables were done using logistic regression presented with odds ratios (ORs) with 95% confidence intervals (CIs) or the Jonckheere-Terpstra test to analyze for the trend between the groups. Categorical variables were analyzed using the chi-square test. All tests were two-tailed. p values ≤ 0.05 were considered significant. The figures were created in GraphPad Prism 8 (version 8.4.3, GraphPad Software LLC, San Diego, CA, USA).
Results
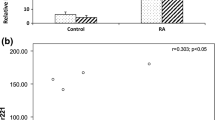
Analysis of the relative fold difference for the different microRNAs between the groups revealed that four of the nine microRNAs analyzed displayed significant differences in the comparison between RA and HCs (Table 2). In addition, difference in miR-155 expression was borderline significant in this comparison. After adjustments for age and gender, three microRNAs remained significant, i.e., miR-22-3p, miR-26b-5p, and miR-142-3p, and the p value of the miR-155 comparison became significant. Moreover, miR-26b-5p and miR-142-3p showed a significant difference comparing FDRs with HCs, where miR-26b-5p remained significant after adjustments. Notably, none of the analyzed microRNAs displayed significant difference between FDRs and RA after adjusting for gender and age (Table 2). A graphical overview of the findings is shown in Fig. 1.
Graphical representation of observed fold differences between RA patients, FDRs, and HCs respectively for indicated microRNAs. A star indicates significant difference for the corresponding logistic regression analysis as described in Table 2
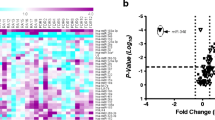
Next, we applied the Jonckheere-Terpstra test to compare the three groups regarding microRNA expression. We found significant differences between the three groups for miR-22-3p (p = 0.001), miR-26b-5p (p = 0.026), miR-142-3p (p = 0.001), and miR-451a (p = 0.045) as presented in Fig. 2. In addition, for miR-155, the difference between the three groups was borderline significant (p = 0.051). Principal component analysis based on relative expression level was applied to investigate potential clustering of individuals. No particular pattern was observed (data not shown).
MicroRNA expression for RA patients, FDRs, and HCs. ΔCT values for miR-22-3p, miR-26b-5p, miR-34a-3p, miR-103a-3p, miR-142-3p, miR-146a-5p, miR-155, miR-346, and miR-451a with mean values and standard deviation for RA patients, FDRs, and HCs noted out as solid lines. Expression of the miRNAs was adjusted to the reference gene U6 snRNA. The Jonckheere-Terpstra test was applied for each microRNA, where statistical significance between the three groups, p ≤ 0.05, is depicted in the figure
Next, influence of treatments of the RA patients was determined, and a significant effect of corticosteroids on miR-103a-3p levels was observed, with an upregulation of miR-103a-3p in the group treated with corticosteroids (p = 0.048).
Discussion
miRNAs are involved in RA pathogenesis
Familial studies provide a unique possibility to evaluate the impact of potential contributing factors in the at-risk FDR group. Taking this approach, we have previously shown that the frequency of HLA-DRB1*0401/0404/0408 and the T variant of PTPN22, which are risk factors of RA development, are significantly increased in RA and FDRs compared to those in HCs [13]. Moreover, levels of ACPAs and RF in plasma, in particular of the IgA and IgM isotypes, were significantly different between the three groups, where FDRs showed an intermediate phenotype [13]. Despite the presence of risk factors in the FDRs for RA development, regulatory mechanisms appear to still be in place. MicroRNAs could potentially be involved in this process. Indeed, several recent studies [5, 11, 12, 15] have demonstrated both overlaps and differences in the expression of multiple microRNAs comparing RA patients, FDRs, and HCs.
Thus, in the current study, we have analyzed the expression levels of nine different microRNAs in peripheral blood cells of well-characterized RA, FDR, and HC individuals. Expression of miR-26b-5p and miR-142-3p was downregulated in FDRs compared to that in HCs, and miR-26b-5p remained significant after adjustment for age and gender. Comparison of FDR to RA patients showed that FDR and RA patients display similar levels of these microRNAs, suggesting that FDR display this disease-associated feature. Our observations are in line with Zhu et al. where downregulation of miR-26b-5p and miR-142-3p was observed in PBMC of RA patients compared with controls [12]. MiR-26b has been associated to RA and inflammation where inhibition of miR-26b in rheumatoid arthritis fibroblast-like synoviocytes (RAFLS) causes increased levels of TNF-α, IL-1β, and IL-6, while miR-26b mimics mediate downregulation of the same proinflammatory cytokines [16]. Chen et al. have found that miR-26 is able to downregulate IL-6 expression as well as TNF-α/NF-κB signaling, by targeting mucosa-associated lymphoid tissue lymphoma translocation protein 1 (MALT1) and high-mobility group AT-hook 1 (HMGA1) [17]. Moreover, levels of miR-26b are downregulated in the cartilage of osteoarthritis (OA) patients compared to those in controls [18], where decreased levels of miR-26b also have been suggested to contribute to disease progression by upregulation of the NF-κB signaling pathway. In our analysis, however, we did not observe a correlation between miR-26b and IL-6 or IL-1β in RA and FDR (data not shown), which indicates that regulation of the expression of these cytokines depends on multiple influencing factors. Our result also deviates from the study by Anaparti et al., where significant decreased levels of miR-26b-5p were observed in FDRs compared to RA, and when compared to HCs, no significant difference was noted [5].
In this study, miR-142-3p was shown to be significantly downregulated in RA (and FDRs before adjustment) compared to that in HCs, which is in line with the study by Zhu et al., comparing PBMCs in RA and HC [12]. However, in serum and RAFLS, miR-142-3p is upregulated in patients with very early RA and CCP-positive at-risk individuals [11], as well as in RA patients [19] compared to HCs. This corroborates the complexity of microRNA function and regulation including that miR-142-3p seems to play various roles in various tissues.
An extensively studied microRNA in the development of RA is miR-155, which has been found to be upregulated in RA patients and in FDRs compared to HCs or OA patients [5, 20, 21]. MiR-155 is associated with inflammation and has been associated with erythrocyte sedimentation rate (ESR), disease activity score (DAS-28), and plasma levels of proinflammatory cytokines [22] in RA patients. MiR-155 has several targets, including suppressor of cytokine signaling 1 (SOCS1) and SH-2 containing inositol 5′ polyphosphatase 1 (SHIP1). Inhibition of SOCS1 and SHIP1 by miR-155 results in an enhancement of inflammatory cytokine production, i.e., TNF, IL-1, and IL-6 [23], thus contributing to an inflammatory milieu. Other reported targets are the Fas-associated death domain (FADD) protein and the IκB kinase (IKK), which both are anti-inflammatory molecules [22]. This further supports the proinflammatory role of miR-155. Unexpectedly, in the current study, we found a decrease of miR-155 expression in RA patients compared to HCs after adjustment for age and gender, which deviate from previous findings. In addition, miR-146 that also has been associated with inflammation did not differ in our cohort. The underlying cause for this is still to be revealed, but one possibility is that the RA patients are undergoing treatments of various kinds and the disease was well controlled at the time of sampling.
This study found a significant downregulation of miR-22-3p in RA compared to both FDRs and HCs, and the significant difference remained between RA and HCs after adjustment for age and gender. Lin et al. found that miR-22 was downregulated in synovial tissue of RA patients compared to that of OA patients, and that the levels of miR-22 were negatively correlated to the levels of Cyr61 overexpression and the promotion of IL-6 production [24]. Similar results have been reported by Yu et al., where inhibition of p53 in subarachnoid hemorrhage (SAH) mice led to decreased levels of miR-22 and upregulation of Cyr61, whereas knock down of p53 in HEB cells (human normal glial cell line) suppressed the expression of miR-22 [25]. Decreased levels of miR-22 in synovial tissue of RA patients compared to HCs have also been reported and this is negatively correlated to sirtuin 1 (SIRT1) levels [26]. Interestingly, addition of a miR-22 mimic in this context leads to decreased levels of TNF-α, IL-1β, and IL-6, supporting the role of miR-22 in inflammation and RA pathogenesis [26].
The levels of miR-451a were in this study significantly decreased in RA compared to those in HCs, and borderline significant comparing FDRs and HCs (p = 0.054). However, after adjustment for age and gender, no significant difference remained, indicating that levels of miR-451a are highly influenced by these parameters. Meta-analysis of likely targets of miR-451a [27] includes macrophage migration inhibitory factor (MIF) that is negatively modulated by miR-451a [28] and IL-6R that is also negatively regulated by miR-451a [29]. As we observed a downregulation of miR-451a in RA patients compared to HC, we speculate that this could represent an underlying inflammation and thus upregulation of IL-6R and MIF.
In contrast to Anaparti et al. [5], we did not observe any upregulation of miR-103a-3p expression in RA patients and FDRs compared to HCs. However, we observed downregulation of miR-103a-3p in RA patients using corticosteroids, which suggests a modulating effect of this treatment in line with a beneficial effect of miR-103a-3p suppression. The role of miR-103a-3p in bone remodulation has been extensively studied in both animal models and in various rheumatoid diseases [30,31,32]. In addition, the influence of corticosteroids on both miR-103 and osteoporosis has demonstrated to be tightly connected [33,34,35,36].
When applying the Jonckheere-Terpstra test, four of the nine microRNAs analyzed displayed a significant trend comparing RA patients, FDRs, and HCs, where FDRs exhibited an intermediate phenotype. The intermediary pattern for the microRNAs in the FDRs resembles our previously published observations [13], where an intermediate level of ACPA and RF in FDRs of the same cohort was observed. Taken together, this suggests that a dysregulation of microRNAs may be involved in the increased risk of RA development in FDRs. However, for some microRNAs, the expression level did not differ when a group-wise comparison of FDRs and HCs was performed, suggesting that microRNAs could also have a protective role, preventing FDRs to develop the disease. Overall, regulation of inflammation and other biological mechanisms by microRNAs and other factors is highly complex, and cannot be expected to be explained by a single microRNA.
In summary, we confirm that microRNAs seem to play a role in promotion and protection of RA development. Strength of this study is the usage of a well-characterized cohort, as well as the familial analysis approach where the participants originate from a restricted geographic area, and display a genetic and environmental homogeneity.
References
Scott DL, Wolfe F, Huizinga TW (2010) Rheumatoid arthritis. Lancet 376(9746):1094–1108. https://doi.org/10.1016/S0140-6736(10)60826-4
Ammari M, Jorgensen C, Apparailly F (2013) Impact of microRNAs on the understanding and treatment of rheumatoid arthritis. Curr Opin Rheumatol 25(2):225–233. https://doi.org/10.1097/BOR.0b013e32835d8385
Karami J, Aslani S, Tahmasebi MN, Mousavi MJ, Sharafat Vaziri A, Jamshidi A, Farhadi E, Mahmoudi M (2020) Epigenetics in rheumatoid arthritis; fibroblast-like synoviocytes as an emerging paradigm in the pathogenesis of the disease. Immunol Cell Biol 98(3):171–186. https://doi.org/10.1111/imcb.12311
Scherer HU, Haupl T, Burmester GR (2020) The etiology of rheumatoid arthritis. J Autoimmun 110:102400. https://doi.org/10.1016/j.jaut.2019.102400
Anaparti V, Smolik I, Meng X, Spicer V, Mookherjee N, El-Gabalawy H (2017) Whole blood microRNA expression pattern differentiates patients with rheumatoid arthritis, their seropositive first-degree relatives, and healthy unrelated control subjects. Arthritis Res Ther 19(1):249. https://doi.org/10.1186/s13075-017-1459-x
Churov AV, Oleinik EK, Knip M (2015) MicroRNAs in rheumatoid arthritis: altered expression and diagnostic potential. Autoimmun Rev 14(11):1029–1037. https://doi.org/10.1016/j.autrev.2015.07.005
Gao J, Zhou XL, Kong RN, Ji LM, He LL, Zhao DB (2016) microRNA-126 targeting PIK3R2 promotes rheumatoid arthritis synovial fibro-blasts proliferation and resistance to apoptosis by regulating PI3K/AKT pathway. Exp Mol Pathol 100(1):192–198. https://doi.org/10.1016/j.yexmp.2015.12.015
Liu L, Zuo Y, Xu Y, Zhang Z, Li Y, Pang J (2019) MiR-613 inhibits proliferation and invasion and induces apoptosis of rheumatoid arthritis synovial fibroblasts by direct down-regulation of DKK1. Cell Mol Biol Lett 24:8. https://doi.org/10.1186/s11658-018-0130-0
Nakasa T, Miyaki S, Okubo A, Hashimoto M, Nishida K, Ochi M, Asahara H (2008) Expression of microRNA-146 in rheumatoid arthritis synovial tissue. Arthritis Rheum 58(5):1284–1292. https://doi.org/10.1002/art.23429
Stanczyk J, Pedrioli DM, Brentano F, Sanchez-Pernaute O, Kolling C, Gay RE, Detmar M, Gay S, Kyburz D (2008) Altered expression of microRNA in synovial fibroblasts and synovial tissue in rheumatoid arthritis. Arthritis Rheum 58(4):1001–1009. https://doi.org/10.1002/art.23386
Ouboussad L, Hunt L, Hensor EMA, Nam JL, Barnes NA, Emery P, McDermott MF, Buch MH (2017) Profiling microRNAs in individuals at risk of progression to rheumatoid arthritis. Arthritis Res Ther 19(1):288. https://doi.org/10.1186/s13075-017-1492-9
Zhu X, Wu L, Mo X, Xia W, Guo Y, Wang M, Zeng K, Wu J, Qiu Y, Lin X, Lu X, Deng F, Lei S (2020) Identification of PBMC-expressed miRNAs for rheumatoid arthritis. Epigenetics 15(4):386–397. https://doi.org/10.1080/15592294.2019.1676613
Arlestig L, Mullazehi M, Kokkonen H, Rocklov J, Ronnelid J, Dahlqvist SR (2012) Antibodies against cyclic citrullinated peptides of IgG, IgA and IgM isotype and rheumatoid factor of IgM and IgA isotype are increased in unaffected members of multicase rheumatoid arthritis families from northern Sweden. Ann Rheum Dis 71(6):825–829. https://doi.org/10.1136/annrheumdis-2011-200668
Arnett FC, Edworthy SM, Bloch DA, McShane DJ, Fries JF, Cooper NS, Healey LA, Kaplan SR, Liang MH, Luthra HS et al (1988) The American Rheumatism Association 1987 revised criteria for the classification of rheumatoid arthritis. Arthritis Rheum 31(3):315–324. https://doi.org/10.1002/art.1780310302
Ormseth MJ, Solus JF, Sheng Q, Ye F, Wu Q, Guo Y, Oeser AM, Allen RM, Vickers KC, Stein CM (2020) Development and validation of a microRNA panel to differentiate between patients with rheumatoid arthritis or systemic lupus erythematosus and controls. J Rheumatol 47(2):188–196. https://doi.org/10.3899/jrheum.181029
Sun J, Yan P, Chen Y, Chen Y, Yang J, Xu G, Mao H, Qiu Y (2015) MicroRNA-26b inhibits cell proliferation and cytokine secretion in human RASF cells via the Wnt/GSK-3beta/beta-catenin pathway. Diagn Pathol 10:72. https://doi.org/10.1186/s13000-015-0309-x
Chen CY, Chang JT, Ho YF, Shyu AB (2016) MiR-26 down-regulates TNF-alpha/NF-kappaB signalling and IL-6 expression by silencing HMGA1 and MALT1. Nucleic Acids Res 44(8):3772–3787. https://doi.org/10.1093/nar/gkw205
Yin X, Wang JQ, Yan SY (2017) Reduced miR26a and miR26b expression contributes to the pathogenesis of osteoarthritis via the promotion of p65 translocation. Mol Med Rep 15(2):551–558. https://doi.org/10.3892/mmr.2016.6035
Qiang J, Lv T, Wu Z, Yang X (2019) Down-regulation of microRNA-142-3p inhibits the aggressive phenotypes of rheumatoid arthritis fibroblast-like synoviocytes through inhibiting nuclear factor-kappaB signaling. Biosci Rep 39(7). https://doi.org/10.1042/BSR20190700
Kurowska-Stolarska M, Alivernini S, Ballantine LE, Asquith DL, Millar NL, Gilchrist DS, Reilly J, Ierna M, Fraser AR, Stolarski B, McSharry C, Hueber AJ, Baxter D, Hunter J, Gay S, Liew FY, McInnes IB (2011) MicroRNA-155 as a proinflammatory regulator in clinical and experimental arthritis. Proc Natl Acad Sci U S A 108(27):11193–11198. https://doi.org/10.1073/pnas.1019536108
Pauley KM, Satoh M, Chan AL, Bubb MR, Reeves WH, Chan EK (2008) Upregulated miR-146a expression in peripheral blood mononuclear cells from rheumatoid arthritis patients. Arthritis Res Ther 10(4):R101. https://doi.org/10.1186/ar2493
Moran-Moguel MC, Petarra-Del Rio S, Mayorquin-Galvan EE, Zavala-Cerna MG (2018) Rheumatoid arthritis and miRNAs: a critical review through a functional view. J Immunol Res 2018:2474529–2474516. https://doi.org/10.1155/2018/2474529
Tavasolian F, Abdollahi E, Rezaei R, Momtazi-Borojeni AA, Henrotin Y, Sahebkar A (2018) Altered expression of microRNAs in rheumatoid arthritis. J Cell Biochem 119(1):478–487. https://doi.org/10.1002/jcb.26205
Lin J, Huo R, Xiao L, Zhu X, Xie J, Sun S, He Y, Zhang J, Sun Y, Zhou Z, Wu P, Shen B, Li D, Li N (2014) A novel p53/microRNA-22/Cyr61 axis in synovial cells regulates inflammation in rheumatoid arthritis. Arthritis Rheumatol 66(1):49–59. https://doi.org/10.1002/art.38142
Yu S, Zeng YJ, Sun XC (2018) Neuroprotective effects of p53/microRNA22 regulate inflammation and apoptosis in subarachnoid hemorrhage. Int J Mol Med 41(4):2406–2412. https://doi.org/10.3892/ijmm.2018.3392
Zhang C, Fang L, Liu X, Nie T, Li R, Cui L, Wang J, Ji Y (2020) miR-22 inhibits synovial fibroblasts proliferation and proinflammatory cytokine production in RASF via targeting SIRT1. Gene 724:144144. https://doi.org/10.1016/j.gene.2019.144144
Chou CH, Shrestha S, Yang CD, Chang NW, Lin YL, Liao KW, Huang WC, Sun TH, Tu SJ, Lee WH, Chiew MY, Tai CS, Wei TY, Tsai TR, Huang HT, Wang CY, Wu HY, Ho SY, Chen PR, Chuang CH, Hsieh PJ, Wu YS, Chen WL, Li MJ, Wu YC, Huang XY, Ng FL, Buddhakosai W, Huang PC, Lan KC, Huang CY, Weng SL, Cheng YN, Liang C, Hsu WL, Huang HD (2018) miRTarBase update 2018: a resource for experimentally validated microRNA-target interactions. Nucleic Acids Res 46(D1):D296–D302. https://doi.org/10.1093/nar/gkx1067
Bandres E, Bitarte N, Arias F, Agorreta J, Fortes P, Agirre X, Zarate R, Diaz-Gonzalez JA, Ramirez N, Sola JJ, Jimenez P, Rodriguez J, Garcia-Foncillas J (2009) microRNA-451 regulates macrophage migration inhibitory factor production and proliferation of gastrointestinal cancer cells. Clin Cancer Res 15(7):2281–2290. https://doi.org/10.1158/1078-0432.CCR-08-1818
Liu X, Zhang A, Xiang J, Lv Y, Zhang X (2016) miR-451 acts as a suppressor of angiogenesis in hepatocellular carcinoma by targeting the IL-6R-STAT3 pathway. Oncol Rep 36(3):1385–1392. https://doi.org/10.3892/or.2016.4971
Chen J, Li K, Pang Q, Yang C, Zhang H, Wu F, Cao H, Liu H, Wan Y, Xia W, Wang J, Dai Z, Li Y (2016) Identification of suitable reference gene and biomarkers of serum miRNAs for osteoporosis. Sci Rep 6:36347. https://doi.org/10.1038/srep36347
Zhang H, Huang X, Ye L, Guo G, Li X, Chen C, Sun L, Li B, Chen N, Xue X (2018) B cell-related circulating microRNAs with the potential value of biomarkers in the differential diagnosis, and distinguishment between the disease activity and lupus nephritis for systemic lupus erythematosus. Front Immunol 9:1473. https://doi.org/10.3389/fimmu.2018.01473
Zuo B, Zhu J, Li J, Wang C, Zhao X, Cai G, Li Z, Peng J, Wang P, Shen C, Huang Y, Xu J, Zhang X, Chen X (2015) microRNA-103a functions as a mechanosensitive microRNA to inhibit bone formation through targeting Runx2. J Bone Miner Res 30(2):330–345. https://doi.org/10.1002/jbmr.2352
Kfir-Erenfeld S, Haggiag N, Biton M, Stepensky P, Assayag-Asherie N, Yefenof E (2017) miR-103 inhibits proliferation and sensitizes hemopoietic tumor cells for glucocorticoid-induced apoptosis. Oncotarget 8(1):472–489. https://doi.org/10.18632/oncotarget.13447
Adachi JD, Olszynski WP, Hanley DA, Hodsman AB, Kendler DL, Siminoski KG, Brown J, Cowden EA, Goltzman D, Ioannidis G, Josse RG, Ste-Marie LG, Tenenhouse AM, Davison KS, Blocka KL, Pollock AP, Sibley J (2000) Management of corticosteroid-induced osteoporosis. Semin Arthritis Rheum 29(4):228–251. https://doi.org/10.1016/s0049-0172(00)80011-6
Ringe JD (1989) Glucocorticoid-induced osteoporosis. Clin Rheumatol 8(Suppl 2):109–115. https://doi.org/10.1007/BF02207244
van Staa TP, Leufkens HG, Cooper C (2002) The epidemiology of corticosteroid-induced osteoporosis: a meta-analysis. Osteoporos Int 13(10):777–787. https://doi.org/10.1007/s001980200108
Acknowledgments
The authors would like to acknowledge Anne Israelsson for technical advice, Viktoria von Zweigbergk for blood sample collection, and the participants in the study.
Funding
Open access funding provided by Umea University. This work was supported by grants from the Swedish Research Council (Dnr: 2018-02551), King Gustaf V’s 80-Year Fund, the Swedish Rheumatism Association, and Umeå University, Umeå, Sweden.
Author information
Authors and Affiliations
Contributions
Study design was performed by Kristina Lejon, Solbritt Rantapää-Dahlqvist, Lisbeth Ärlestig, and Mikael Brink. Sample and patient-related data collection was done by Solbritt Rantapää-Dahlqvist, Lisbeth Ärlestig, and Mikael Brink. Laboratory work and data collection concerning the microRNAs were performed by Emma Renman. Data analysis was performed by Emma Renman, Mikael Brink, Solbritt Rantapää-Dahlqvist, and Kristina Lejon. Interpretation of data and writing of the first draft of the manuscript were done by Emma Renman and Kristina Lejon. Mikael Brink and Solbritt Rantapää-Dahlqvist reviewed and modified the manuscript, and all authors approved the final version.
Corresponding author
Ethics declarations
Disclosures
None.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Renman, E., Brink, M., Ärlestig, L. et al. Dysregulated microRNA expression in rheumatoid arthritis families—a comparison between rheumatoid arthritis patients, their first-degree relatives, and healthy controls. Clin Rheumatol 40, 2387–2394 (2021). https://doi.org/10.1007/s10067-020-05502-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10067-020-05502-9