Abstract
In this study, we formulated a mathematical model of hypoxia-inducible factor 1 (HIF-1) mediated regulation of cellular energy metabolism describing the reprogramming of cell metabolic processes from oxidative phosphorylation to glycolysis under reduced oxygen levels. The model considers the dynamics of fifteen biochemical species and the proton concentration with the underlying reaction processes localized in three intracellular compartments, i.e. the cytoplasm, mitochondrion and nucleus. More than sixty parameters of the model were calibrated using both the published data and the system steady-state based identification procedure. The model was validated by generating predictions which could be compared to empirical observations. The model behaviors representing the cell metabolism switching over in response to transitioning from a normoxic to hypoxic environment are consistent with the current views of the role of HIF-1 in hypoxia.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
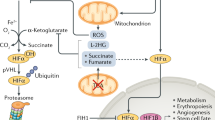
Energy metabolism drives the functioning of organs and individual cells. There is a growing need in controlling specific metabolic processes that are skewed in many pathologies, such as cancer, infectious diseases, and sepsis [9, 19, 25]. Hypoxia (low oxygen tension) is an essential hallmark of the pathological processes. The metabolic reprogramming of cells that occurs in hypoxia is regulated by hypoxia-inducible factor 1 (HIF-1) [17, 21,22,23]. The activity of HIF-1 is determined by the oxygen level. In turn, HIF-1 regulates the balance between oxidative and glycolytic metabolism [21]. With oxygen as the substrate, the respiratory chain paves the way for the efficient generation of ATP from citric acid cycle products. Lowered oxygen levels lead to increased production of reactive oxygen species by respiratory complexes and without further protection, this may lead to cellular dysfunction or cell death. HIF acts as a master regulator adapting the metabolism to such hypoxic conditions by initiating several mechanisms which reduce mitochondrial activity and amplify anaerobic ATP production [21]. Although ATP levels cannot be maintained at normoxic values, the cell benefits from this switch, because ROS production in the mitochondria is prevented [32].
Understanding the complex regulation of cellular energy metabolisms under normal and hypoxic settings requires the development of mechanistic mathematical models of cellular energetics. Some mathematical models of intracellular respiration have already been proposed [1, 2, 10, 15, 28]. These models mostly describe the biophysical processes of oxidative phosphorylation but do not touch upon HIF-mediated regulation. Recently, two studies addressed the transcriptional activity of the HIF-1 signaling network [16] and the interleukin-15 dependent regulation of HIF-1a in Natural Killer cells [6]. As a mathematical model of HIF-1 regulated reprogramming of cellular metabolism does not yet exist, we aim to formulate and calibrate such an integrative model.
In this study, we formulate and calibrate the mathematical model of HIF-1 mediated regulation of cellular energy metabolism describing the reprogramming of cell metabolic processes from oxidative phosphorylation to glycolysis under reduced oxygen levels. The model considers biochemical and biophysical processes underlying the dynamics of sixteen chemical species and the proton concentration localized in the cytoplasm, mitochondrion and nucleus compartments. We identify unknown parameters such that the system exhibits a stable stationary state under normoxic conditions (see Section 4.1). The stability of the steady state is shown by computer experiments (simulation of the evolution of the metabolites for perturbed (± 10%) steady states), see also Fig. 2, and by a linear stability analysis. The existence and uniqueness of a global solution to the model equations is shown in Section 3. Comparing numerical simulations under hypoxic conditions with and without HIF-mediated adaption confirms the ability of HIF to indirectly limit respiratory chain activity (by inhibiting the pyruvate dehydrogenase) and to increase glycolytic ATP production (by activating glycolytic enzymes), see Figs. 4, 5, and 6 in Section 4.
2 The Model
The mathematical model is given by a system of ordinary differential equations (ODEs) and consists of the main parts of the cellular energy metabolism, namely glycolysis and lactic acid fermentation in the cytoplasm as well as citric acid cycle and oxydative phosphorylation in the mitochondria, as summarized in Fig. 1. In addition, a HIF-mediated activation of the glycolysis and inactivation of the citric acid cycle at low oxygen concentrations is implemented. Reaction rates based on the law of mass action are preferably used to model the biochemical reactions. Oxydative phosphorylation is modeled using phenomenological rate equations. To keep the extent of the model to a minimum, properties of enzymatically driven reactions such as inhibition, saturation or other regulatory mechanisms are not considered for large parts of the model and subsequent reactions are often interpreted as one entity. Irreversibility of according reaction rates is often justified by the irreversibility of at least one of the underlying reactions. To make the paper more accessible for readers who are not familiar with the biomedical application, we described all species considered in our model together with the corresponding acronym in Table 1. All parameters of the model are listed with their physiological meaning in Table 2.
Biochemical scheme of the mathematical model of the cellular energy metabolism describing the species, metabolic reaction networks and corresponding regulatory mechanisms. Number i at reaction arrows refers to the corresponding rate equation vi. A simplified fatty acid catabolism is outlined in pale gray, but not yet considered in the model
2.1 The Cytoplasm
Glycolysis.
Glucose uptake from the extracellular medium is modeled by a constant rate, assuming that the cell is always provided with sufficient extracellular glucose and intracellular processes do not affect its transport across the membrane:
Next, glucose is further processed at the expense of two molecules of ATP, in order to prepare it for the following degradation process. Repeated phosphorylation and isomerization on the way from glucose to fructose 1,6-bisphosphate is summarized as one mechanism in the second rate equation:
The subsequent cleavage of fructose 1,6-bisphosphate into two three-carbon units and their conversion into pyruvate is considered as the second and last step of glycolysis in the model. It is accompanied by the formation of four molecules ATP and two molecules NADH, making glycolysis an overall ATP-producing pathway.
To obtain this rate equation, two more assumptions are made: Firstly, the amount of adenosine bodies is a conserved quantity in the model and adenosine only exists as ATP or ADP (\(A_{cyt}^{tot} = [ATP]_{cyt} + [ADP]_{cyt}\)) and secondly, there is always a sufficient and constant amount of inorganic phosphate such that its concentration does not affect the velocity of the reaction.
Lactic Acid Fermentation.
Analogously to the adenosine bodies, the conservation \(N_{cyt}^{tot} = [NADH]_{cyt} + [NAD]_{cyt}\) is considered. While almost all energy consuming processes in the cell use ATP as an energy source and therefore regenerate ADP, the energy of NADH cannot be used directly and the cell would run out of NAD without further metabolic pathways. Therefore, lactic acid fermentation enables NAD to be regained in the cytoplasm, as pyruvate and NADH are converted into lactate and NAD.
Reversibility of the reaction is not taken into account here and so the only possible fate of lactate is to be transported out of the cell:
ATP Consumption.
All energy consuming processes in each compartment of the cell are represented by the following rate equation, which was adopted from a model of Koenig et al. [13]:
It consists of Michaelis–Menten type kinetics, accounting for basal life-sustaining processes and a linear term, which respects the ability of the cell to adjust its energy demand according to the current ATP supply.
2.2 Metabolite Transport Across the Inner Mitochondrial Membrane
Pyruvate Transport
An alternative fate of cytosolic pyruvate is to be transported into mitochondria, where it can fuel the citric acid cycle. The transport is assumed to be reversible in the model, because mitochondrial pyruvate could otherwise accumulate arbitrarily when the citric acid cycle is inhibited. The direction of the transport is prescribed by the difference in particle numbers directly at the membrane. As the model does not resolve the distribution of pyruvate spatially, the respective metabolite concentration is divided by the volume of the membrane and then compared. Depending on the compartment, the resulting value is converted into cytosolic or mitochondrial volume units:
with \(\tilde {k}_{7} := k_{7}V_{m}\), Vm: volume of the membrane, Vcyt: volume of the cytosol and Vmito := NmitoVmito,single: total mitochondrial volume.
ADP/ATP Transport
The adenine nucleotide translocator enables the exchange of ATP and ADP across the inner mitochondrial membrane. It is modeled as a reversible reaction, following the law of mass action. ADP and ATP amounts of both compartments are compared and then converted into the correct particle number per volume unit.
where \(\tilde {k}_{ANT}:= k_{ANT}{V_{m}^{2}}\). Keq,ANT is introduced such that ATP is transported out of the mitochondria under normal conditions.
2.3 The Mitochondrion
The model for the mitochondrial energy metabolism is adopted from Nazaret et al. [15] with extensions regarding the oxygen dependence of the oxydative phosphorylation. The conservations \(A_{mito}^{tot} = [ATP]_{mito} + [ADP]_{mito}\) and \(N_{mito}^{tot} = [NADH]_{mito} + [NAD]_{mito}\) are assumed.
Citric Acid Cycle
An additional preliminary step of the citric acid cycle is the oxydative decarboxylation of mitochondrial pyruvate into acetyl-CoA. It is the first occurrence of mitochondrial NADH production, one of the main objectives of the mitochondrial energy metabolism, and also an important regulatory point.
The concentration of Coenzyme A is assumed to be constant and without impact on the reaction rate.
A round of the citric acid cycle starts, when the acetyl group of acetyl-CoA is transferred into oxaloacetate, forming citrate.
The reaction from citrate to α-ketoglutarate via isocitrate with NADH production is simplified to:
α-ketoglutarate undergoes further transformations and finally, oxaloacetate is formed to complete the circle. During this reaction sequence, two molecules NADH, one molecule FADH2 and one molecule GTP are gained. For the model, only NADH is considered and GTP is replaced by ATP as they are energetically equivalent.
The model respects some additional features of the TCA cycle. α-ketoglutarate and oxaloacetate are also linked via a reversible reaction within the scope of the amino acid metabolism. For the sake of simplicity, additional reactants are omitted:
An output of the TCA cycle is modeled by the concentration dependent degradation of oxaloacetate in order that the system can reach a steady state.
Finally, the ATP dependent transformation of pyruvate into oxaloacetate complements the citric acid cycle by an replenishing reaction:
Oxydative Phosphorylation
Energy is now intermediately stored as NADH and the mitochondrial regeneration of NAD is the task of the respiratory chain. It comprises four enzyme complexes which are located in the impermeable inner mitochondrial membrane. The high energy electrons of NADH are transferred sequentially onto oxygen and the energy that is released during this process is used to translocate ten protons per NADH molecule from the mitochondrial matrix into the intermembrane space.
When the matrix and the intermembrane space, separated by the inner mitochondrial membrane, are interpreted as a capacitor, the resulting proton difference can be expressed as a potential. In accordance with Nazaret et al., the proton concentration outside the mitochondrial matrix is assumed to be constant and therefore the temporal change of the membrane potential is linearly linked to the change of the intramitochondrial proton concentration:
There are several processes which affect this concentration, firstly the respiratory chain and the ATPase. The latter couples the re-entrance of protons from the intermembrane space into the matrix to the synthesis of ATP.
Instead of modeling the complex mechanisms at the respiratory chain in detail, a phenomenological approach is chosen. The four enzyme complexes are assembled into one rate equation, which exhibits a Michaelis–Menten type dependence in the NADH concentration as well as a complete inhibition once the membrane potential becomes too high to translocate more protons. For details, see [15]. For our purposes, this reaction rate is extended by a Michaelis–Menten type term for the oxygen dependence of the respiratory chain:
The formation of ATP from ADP and Pi, together with the backflow of protons into the mitochondrial matrix, is reversible:
To gain a phenomenological, reversible rate equation, Nazaret et al. consider the overall Gibb’s free energy of the coupled reaction:
with \(K_{app} := \frac {[ATP]_{eq}}{[ADP]_{eq} [P_{i}]_{eq}}\). From the ansatz ΔG = 0, they derive a formula for the critical ATP concentration in the equilibrium, dependent on the membrane potential. The difference between the actual ATP concentration and the critical ATP concentration then prescribes the direction of the reaction in the following phenomenological rate:
For more details about the derivation, see again [15].
Proton Leak
Since the inner mitochondrial membrane is not completely impermeable for protons, some of them flow back into the matrix without contributing to the synthesis of ATP. This process is taken into account by a linear dependence on the membrane potential:
Glycerol Phosphate Shuttle
The glycerol phosphate shuttle for the feeding of high energy electrons from cytosolic NADH is assumed to have the same phenomenological properties as the respiratory chain: Michaelis–Menten like dependencies on the oxygen and cytosolic NADH concentration as well as an inhibition at high membrane potentials.
We do not model the transport of oxygen from the blood into the mitochondria but interpret [O2] as the mitochondrial oxygen concentration and set it manually to a constant value for the simulations.
2.4 Hypoxia Inducible Factor 1
The HIF Pathway.
HIF1, assembled from the two subunits HIF-1α and HIF-1β, acts as a master regulator for the adaption of cellular metabolism to low oxygen levels. It is a transcription factor and initiates the enhanced transcription of a large number of different genes. For this model, the increased transcription of glycolysis-related genes is of interest, as well as the HIF-mediated transcription of the pyruvate dehydrogenase kinase. The latter can phosphorylate and thus inactivate the enzyme which catalyzes the oxydative decarboxylation in the mitochondria, resulting in a lowered citric acid cycle activity.
While HIF-1β is an abundant protein, HIF-1α is not present in relevant amounts during normoxia. Instead, the HIF-1α-gene is constantly transcribed and translated, but subsequently degraded in an oxygen-dependent manner. Only when oxygen concentrations are sufficiently low, HIF-1α can accumulate and unite with HIF-1β to affect the metabolism.
The mathematical model for the HIF accumulation during hypoxia is partly based on the a dynamic model of the HIF-1α network by Nguyen et al. [16]. A first simplification is to merge HIF-1α and HIF-1β into one entity, HIF1. The permanent and oxygen-independent formation of HIF1 is modeled by a constant rate:
HIF1, like all proteins, is then subject to an unspecific protein turnover:
HIF1-prolyle-hydroxylases (HIF-PHDs), the primary oxygen sensors of the cell, mark the HIF1 protein for degradation by the proteasomal machinery, when the oxygen concentration is sufficiently high:
A negative feedback loop for HIF is incorporated via the modeling of the prolyl hydroxylase concentration. As the PHD-gene is also a target gene of HIF, the corresponding enzyme concentration is expected to rise during hypoxia:
This guarantees the dampening of the HIF concentration after some time and also promotes a fast return to the HIF-independent metabolism once the cell is re-oxygenated.
Updated Rate Equations For HIF1-Dependent Reactions
We model the effect of high HIF levels on the glycolytic rates and on the rate of oxadative decarboxylation by imposing HIF-dependent activation/inactivation terms in the following way:
For high HIF levels, the additional term approaches the value prescribed by parameter Pi.
2.5 The ODE System
For the ODE system, we use the following notation: x1 : [Gluc], x2 : [F1,6bP], x3 : [Pyr]cyt, x4 : [Lac], x5 : [ATP]cyt, x6 : [NAD]cyt, x7 : [Pyr]mito, x8 : [AcCoA], x9 : [Cit], x10 : [KG], x11 : [OAA], x12 : [ATP]mito, x13 : [NAD]mito, x14 : [ΔΨ], x15 : [HIF], x16 : [PHD]. \(g_{i}(x_{15}) := \frac {c + P_{i}(Dx_{15})^{4}}{c + (Dx_{15})^{4}}\), i ∈{1,2,3,4,8}.
3 Existence And Uniqueness Of Global Solutions
Proposition 1
We consider the initial value problem (IVP) consisting of (18)–(33) together with initial values xi(0) = xi,0, i = 1, … ,16 which are non-negative except for component 14 (which represents the mitochondrial membrane potential) and satisfy
Then there exists a unique global solution \(x = (x_{1},\dots ,x_{16})^{T} \in C^{1}([0, \infty )\); \(\mathbb {R}^{16})\) of this IVP. The solution remains non-negative except for component x14 and for all \(t\in (0, \infty )\) it holds that
Proof
Existence and uniqueness of a local solution on the maximal time interval follows due to the local Lipschitz-continuity of the right-hand side of (18)–(33). Next, we prove that the solution exists globally to the right. Let us consider the set
with
First, we show that \(\overline {D}\) is positive invariant for the IVP corresponding to (18)–(33), i.e., for all \({x_{0}} \in \overline {D}\), the solution x to the IVP satisfies \(x(t) \in \overline {D}\), for all t ≥ 0 belonging to the maximal existence interval. This follows by using e.g., Appendix 1, Theorem in [24] or Theorem 5.6.1 in [14] (see also [20]) and the following properties of the right-hand side of the system (18)–(33):
-
For all x ∈ ∂D such that there exists xi = 0, i = 1, … , 16, i≠ 14, it holds that fi(x) ≥ 0.
-
For all x ∈ ∂D such that there exists xi = bi, i = 5,6,12,13, it holds that fi(x) ≤ 0.
To prove that the remaining species xi, i = 1, … , 16, i ≠ 5, 6, 12, 13, 14 are bounded from above on the maximal existence interval, we set
and obtain
Thus, Gronwall’s lemma implies that S cannot be infinite in finite time. Finally, we have to study the behavior of x14. We have
The estimates
and
together with Gronwall’s lemma imply the boundedness of x14 on any finite time interval. Thus the global existence of a unique solution to the IVP is proven. □
4 Numerical Results
Computer experiments allow simulating processes which are not (yet) accessible by biological experimental methods and may help to deepen the understanding—or to predict the behavior—of a system under certain conditions. Prior to that, the underlying model has to be calibrated, i.e. the parameters of the model have to be set to physically meaningful values. This is often an inverse problem, where parameters have to be determined such that the system reproduces some observed data. Unfortunately, appropriate data on the time course of metabolites, in particular under hypoxic conditions, is, to our knowledge, barely available. Therefore, we are restricted to stationary information and have to collect the metabolite concentrations from the literature with the aim of representing a metabolic equilibrium state (see Table 1). The parameters are then determined such that the model is able to reproduce this prescribed equilibrium.
Parameters which are not rate constants (such as for instance parameters of HIF-regulated activation/inactivation or KM-values) are either obtained from the literature and previous modeling studies or fixed manually to biologically reasonable values (see Table 2). Rate constants kATP and kresp are adopted from [15].
4.1 Parameter Identification
The identification of the unknown rate constant vector
of the system defined by (18)–(31) is based on the assumption that the metabolite concentrations are constant over time in the cell under normal conditions. Hence we look for a k such that the model reproduces the biological equilibrium which is prescribed by the steady state metabolite concentrations in Table 1. Thus, k has to be chosen in such a way that the right-hand side of the ODE system (18)–(31) vanishes when the metabolite concentrations are at their equilibrium values. This leads to the following constrained optimization problem:
where \(\tilde {\mathbf {x}} = (x_{1}(t),\dots ,x_{14}(t))^{T}\) and \(\tilde {f}\) is the right-hand side of the ODE system (18)–(31). Note that the parameters k15 − k19 of the HIF-subsystem are determined separately and that the HIF concentration is for now set constant to its basal value in Table 1. Constraint (35) guarantees non-negativity of the rate constants and constraints (36)–(37) lead to reaction rates, which are all within one order of magnitude. We use the default interior-point-algorithm of the MATLAB-function fmincon to solve the COP numerically and obtain the rate constants in Table 2.
It remains to determine the parameters of the HIF-subsystem, which is composed of (32) and (33). The KM-values for v17 are adopted directly from Nguyen et al. ([16]). The rate constants of v15 - v19 are chosen manually (cf. Table 2) such that the HIF-subsystem exhibits the following characteristic properties:
1) The system is in an equilibrium for the basal concentrations of HIF1 and PHD (see Table 1) during normoxia. 2) HIF1 protein accumulation during hypoxia takes place in the time range of hours ([11, 27]). 3) Hypoxia-induced increase of HIF1-levels is attenuated after some time and HIF1-levels eventually even decrease again during prolonged hypoxia ([11, 27]). 4) A negative feedback loop between HIF1 and PHD promotes rapid return to the basal HIF1-concentration after reoxygenation ([7, 16]).
4.2 Numerical Simulations Under Normoxic Conditions
In the following, all numerical simulations of the initial value problem for the ODE-system are performed with the MATLAB-function ode13s (https://www.mathworks.com/). Using the parameters identified in Section 4.1, see Table 2, we parameterize the system (18)–(33) and obtain the stationary state given in Table 1. To examine the local stability of this steady state, the Jacobian matrix of the system is computed by using symbolic differentiation in MATLAB (https://www.mathworks.com/). The eigenvalues of the Jacobian evaluated in the steady state are determined numerically and without exception exhibit a negative real part. Thus we conclude exponential stability. This stability property is visualized in Fig. 2, where the initial values of all species are chosen ± 10% away from the respective steady state values. The system eventually moves back to the equilibrium after some damped oscillations. Lactate takes the longest time to approach its steady state value, but as it does not interact with other metabolites, this does not influence the rest of the system. The system is robust to a severe drop in the ATP concentration. Indeed, when the ATP levels in the Cytosol and Mitochondrion compartments are reduced to 50% of their equilibrium values as shown in Fig. 3, the steady state of the metabolic system is also rapidly recovered. Mitochondrial ATP levels rise on the expense of the membrane potential ΔΨ and since ATP is transported from mitochondria into the cytosol during the simulations, the cytosolic ATP levels reach their steady state value even faster than the mitochondrial ATP levels. This is accompanied by some damped oscillations in the mitochondria and in the cytosol. Note that, if not indicated otherwise, the concentration curves in all plots are divided by their respective steady-state concentration (cf. Table 2) for the sake of a better visualization.
Stability of the cellular energy metabolism under normoxic conditions. The dynamics of the species in Cytosol (left) and Mitochondrion (right) cell compartments is shown. The initial values of the model variables are randomly chosen ± 10% away from their respective steady state values to examine numerically the system’s stability
Robustness of the cellular energy metabolism under normoxic conditions. The dynamics of the species in Cytosol (left) and Mitochondrion (right) cell compartments is shown. The initial ATP levels are chosen at 50% of their steady state values in both compartments. All other species start at their original steady state value
4.3 Numerical Simulations Under Hypoxic Conditions
The following computer experiments simulate hypoxic conditions by imposing an oxygen concentration of 0.05μM to the system (the normoxic cellular oxygen concentration is set to be 100μM).
Computer Experiments Without HIF-Mediated Regulatory Mechanisms
In the simulation presented in Fig. 4, the HIF mechanism is not considered, i.e., meaning that there is no additional activation of the glycolysis or inhibition of oxydative decarboxylation due to HIF accumulation. After a transient phase, the system approaches a new steady-state (apparent for longer simulation times, not entirely shown) with decreased but still relatively high ATP values (47% of the equilibrium value in the cytosol, 15% of the equilibrium value in the mitochondria). Maintenance of such high ATP concentrations is possible, because the reaction rates of the respiratory chain and of the ATPase are still at 50% and 40% of their equilibrium activity. Although the Michaelis–Menten-like term for the oxygen dependence of the respiratory chain (see (16)) reduces from approximately 1 to 0.05, the only slightly decreased membrane potential ΔΨ and the increased NADH levels as seen in Fig. 4b) enable the relatively high value of the respiratory rate. This guarantees a sufficient membrane potential for the perpetual synthesis of ATP within the scope of oxydative phosphorylation. Accumulation of glucose can be observed in the cytosol (cf. Fig. 4a)), because its transformation to fructose 1,6-bP is limited due to the low ATP level. Increased cytosolic NADH concentration (follows from NAD decrease) amplifies the lactic acid fermentation, resulting in lactate accumulation and cytosolic pyruvate depletion. The latter in turn effects the time-delayed decline of mitochondrial pyruvate, after its concentration initially rises due to the lack of reaction partners for further breakdown.
Cellular metabolic system response to a severe drop in oxygen level in the absence of HIF-1 regulation. The dynamics of the species in Cytosol (left) and Mitochondrion (right) cell compartments is shown. All metabolites start at their steady state concentration, but the oxygen concentration is set to [O2] = 0.05μM. The HIF concentration is set to be constant on its low normoxic value during the simulations
Computer Experiments With HIF-mediated Regulatory Mechanisms
When the same experiment is repeated in Fig. 5, but this time with an active HIF-pathway, changes in the metabolite concentrations can be observed after 2–6 hours of simulated time. The HIF-mediated inhibition of the oxydative decarboxylation and amplification of lactic acid fermentation together re-establish high NAD concentrations in both compartments. Therefore the respiratory chain remains at less than 1% of its equilibrium activity and subsequently, the ATPase, which was modeled as a reversible mechanism in (17), now actually catalyses the backreaction from ATP to ADP. This leads to a ATP concentration close to zero in the mitochondria (cf. Fig. 5a)). In contrast, the cytosolic ATP levels only drop to approximately 15% of their equilibrium value due to the increased glycolytic flux. HIF-mediated increase of glycolytic reaction rates apparently leads to the accumulation of glycolytic metabolites (see Fig. 5b)) and eventually, the system reaches a new steady-state (for longer simulation times, not shown). These results are consistent with experiments from [32], where wildtype mouse embryo fibroblasts (MEFs) possessed lower cellular ATP levels than HIF1α−/− knock-out MEFs under hypoxic conditions.
Cellular metabolic system response to a severe drop in oxygen level subject to HIF-1 regulation. The dynamics of the species in Cytosol (left) and Mitochondrion (right) cell compartments is shown. All metabolites start at their steady state concentration, but the oxygen concentration is set to [O2] = 0.05μM. HIF accumulation is considered and since its effect takes place on the time scale of hours, the simulated time is extended to 24 hours
Simulation results presented in Fig. 6 outline the advantage of HIF-driven adaptation to hypoxic conditions. The cytosolic ATP levels attained by the system after 24 hours of adaptation under different oxygen concentrations have been computed, together with the corresponding respiration rate vresp (see (16)) at that time. When HIF is not considered (blue curve), both, ATP levels (Fig. 6a)) and respiratory rate (Fig. 6b)) remain high even for very low oxygen concentrations and only drop once it is very close to zero. Such behavior is characteristic for HIF KO cells and associated with high ROS formation [21]. Inhibition of the oxydative decarboxylation in the simulations (orange curve) results in lowered ATP levels already for moderately low oxygen levels (Fig. 6a)), but comes with the advantage of decreased respiratory chain activity (Fig. 6)). Additional amplification of the glycolytic rates (yellow and purple curves) leads to comparably low respiratory chain activity at least for low oxygen concentrations and helps to maintain higher ATP levels in the cytosol.
Dependence of ATP levels and the respiration rates vresp on the degree of Hypoxia in Cytosol compartment. Cytosolic ATP levels (Left panel) and the value of vresp (Right panel) reached after 24 hours are predicted by simulations under different oxygen concentrations. The legends indicate whether the HIF-mediated regulation was considered and present the details of the specific choices of the activation/inactivation factors in the respective simulations
4.4 Sensitivity Analysis Under Normoxic And Hypoxic Conditions
We use the direct differential method to gain insights into the local sensitivity of the presented model with respect to 34 selected system parameters [26, 33]. In general, the system of sensitivity equations which is associated with some parameter of interest p, reads:
where we define
with n = 16 in our case. We also introduce the normalized sensitivity as
The solution of the sensitivity equations with respect to k1 in relative terms under normoxic conditions is shown in Fig. 7. There, the original system defined by (18)–(33) and parameters from Table 2 starts in the steady state given in Table 1 and remains there during the simulation while the sensitivity equations associated with parameter k1 are solved simultaneously.
Solution to the Sensitivity Equations for Parameter k1. The sensitivities of the metabolite concentrations with respect to the glucose uptake parameter k1 are solved numerically. Using the same numeration as in Section 2.5, the left panel shows the relative sensitivities for metabolites 1-8 and the right panel shows the relative sensitivities for metabolites 9-16
Analogous calculations are performed for 34 selected system parameters. To facilitate the evaluation and comparison of the results, we define a measure for the sensitivity of the system with respect to a certain parameter p as:
where ∥⋅∥2 is the euclidean norm. By tend we denote the endpoint of the simulated time interval, which is tend = 6 ⋅ 103s in the case of the sensitivity analysis under normoxic conditions. The outcome is compared to sensitivity results under hypoxic conditions in Fig. 7. The latter are obtained as follows: Again the system starts with the initial values from Table 1 and with parameters from Table 2, but this time the oxygen concentration is set to [O2] = 0.05μM, as already in the simulations for Fig. 5, and the simulated time interval is extended to tend = 24h. The sensitivity equations are solved simultaneously for all selected parameters and respective sensitivity measures are computed and depicted in Fig. 7.
These findings suggest that the system is dominated by the rate constant of the glucose uptake k1 and of the ATP consumption k6 during normoxia. The high influence of these two parameters is slightly dampened under hypoxic conditions and especially the HIF accumulation rate k15 as well as the HIF-dependent amplification factor of the glucose uptake P1 become more important.
Finally, the dependence of the steady-state concentration of cytosolic ATP on the most sensitive parameters from Fig. 8, k1, k6, k15, and P1, is investigated during normoxia and hypoxia. In Fig. 9, the cytosolic ATP concentration after 24h of simulated time is recorded, while varying the respective parameter over the interval from 50% to 200% of its baseline value from Table 2.
Sensitivity Analysis under Normoxic and Hypoxic conditions. Sensitivity measures as defined in (38) are calculated for simulations under normoxic conditions (blue bars) with tend = 6 ⋅ 103s and hypoxic conditions (red bars) with tend = 24h. During the hypoxic simulations, the oxygen concentration is set to the constant value [O2] = 0.05μM
Dependence of Cytosolic ATP on Variations of Selected Parameters. Parameters k1, k6, k15, and P1 are independently varied over the interval [0.5p∗,2p∗], where p∗ is the respective baseline parameter value from Table 2. In a), the steady-state cytosolic ATP concentrations at t = 24h are shown for simulations under normoxic conditions ([O2] = 100μM). In b), the calculations of a) are repeated for hypoxic conditions ([O2] = 0.05μM). Note that in a), the green and the red curve, and in b), the green and the black curve, coincide
As expected, the ATP concentration does not change with the HIF-associated parameters k15 and P1 during normoxia (cf. Fig. 9a) and therefore, the respective curves are constant and coincide. However, the ATP concentration increases with decreasing ATP consumption rate constant k6. Diminished glucose uptake, represented by the interval where k1 is below 100%, results in decreased ATP content, whereas increasing k1 beyond 100% has only a small effect, since the ATP concentration in the cytosol is bounded by the total concentration of adenosine, \(A_{cyt}^{tot}\).
In the case of simulated hypoxia (Fig. 9b), the trend of the ATP curves with respect to variations of parameters k1 and k6 is similar to the normoxic case. In contrast with Fig. 9a, the ATP curve maintains an almost constant slope and does not come close to its limit when increasing k1 beyond 100%, since the ATP level is generally lower in hypoxia. Not surprisingly, the ATP curve which is obtained by varying P1, is almost identical to the k1-dependent black curve. During hypoxia, the small constant c in the rate equation for the glucose uptake \(v_{1}^{\ast }\) can be neglected and hence the term scales linearly with both, k1 and P1. Apparently, the ATP concentration in the cytosol is not at all sensitive to the imposed changes of the HIF-accumulation rate k15 during hypoxia. In fact, the high sensitivity measure for k15 in Fig. 8 appears, since the HIF concentration itself is very sensitive to k15 and can reach very high levels for slightly increased k15, leading to a allegedly high sensitivity of the model to k15. But due to the saturation structure of the HIF-dependent activation/inactivation terms, a higher HIF concentration does not necessarily lead to a stronger response of the rest of the model.
5 Conclusions
In this study, we formulated the mathematical model of HIF-1 mediated regulation of cellular energy metabolism describing the reprogramming of cell metabolic processes from oxidative phosphorylation to glycolysis under reduced oxygen levels. The model considers the dynamics of sixteen biochemical species and the proton concentration with the underlying reaction processes localized in three intracellular compartments, i.e. the cytoplasm, mitochondrion and nucleus. More than sixty parameters of the model were calibrated using both the published data and the system steady-state based identification procedure. The model was validated by generating predictions which could be compared to empirical observations. The model behavior representing the cell metabolism switching over in response to transitioning from a normoxic to hypoxic environment is consistent with the current perception of the role of HIF-1 in hypoxia as detailed below:
-
Firstly, HIF-1 is known to prevent the persistence of potentially harmful ROS levels through the induction of pyruvate dehydrogenase kinase 1 (PDK1) [21, 22].
-
Secondly, glycolysis generates 2 molecules of ATP from one molecule of glucose whereas oxidative phosphorylation produces 36 ATP molecules per glucose molecule [17]. This difference in efficiency suggests that low oxygen tension will severely reduce cell energy availability [23].
The calibrated model presents a first step, i.e., a basic module for oxidative phosphorylation and glucolysis regulation, in the development an integrative multiscale model of cellular energetic metabolism. Such a model should consider the whole spectrum of metabolic fuels including the fatty acid metabolism, glutaminolysis. As glutamine is also required for the biosynthesis of proteins, it will be critical to identify an optimal physiological balance between its use for energy production and as a building block of proteins required for immune system function. Importantly, the immune cells metabolism is also regulated by multiple signaling networks, initiated by the formation of receptor synapses, determining the decisions on the cell fate under normal, hypoxic and inflammatory conditions. These also need to be considered on the way towards a comprehensive model of the cell.
There is growing need in developing pharmacological strategies for controlling specific metabolic processes that are skewed in many pathologies, such as autoimmune diseases, tumors, sepsis. Overall, we hope that our modeling approach should pave the way towards a systems level predictive model to be used for targeting the specific processes in the pathologies related to a skewed cellular metabolism [9].
References
Bazil, J.N., Buzzard, G.T., Rundell, A.E.: Modeling mitochondrial bioenergetics with integrated volume dynamics. PLos Comput. Biol. 6, e1000632 (2010)
Beard, D.A.: A biophysical model of the mitochondrial respiratory system and oxidative phosphorylation. PLos Comput. Biol. 1, e36 (2005)
Berg, J.M., Tymoczko, J.L., Stryer, L.: Biochemistry. W.H. Freeman and Company, New York (2012)
Bienfait, H.F., Jacobs, J.M., Slater, E.C.: Mitochondrial oxygen affinity as a function of redox and phosphate potentials. Biochim. Biophys. Acta 376, 446–457 (1975)
Chowdhury, H., Kreft, M., Jensen, J., Zorec, R.: Insulin induces an increase in cytosolic glucose levels in 3T3-L1 cells with inhibited glycogen synthase activation. Int. J. Mol. Sci. 15, 17827–17837 (2014)
Coulibaly, A., Bettendorf, A., Kostina, E., Figueiredo, A.S., Velasquez Giraldo, S.Y., Bock, H.-G., Thiel, M., Lindner, H.A., Barbarossa, M.V.: Interleukin-15 signaling in HIF-1α regulation in natural killer cells, insights through mathematical models. Front. Immunol. 10, 2401 (2019)
D’Angelo, G., Duplan, E., Boyer, N., Vigne, P., Frelin, C.: Hypoxia up-regulates prolyl hydroxylase activity. a feedback mechansim that limits HIF-1 responses during reoxygenation. J. Biol. Chem. 278, 38183–38187 (2003)
Degn, H., Wohlrab, H.: Measurement of steady-state values of respiration rate and oxidation levels of respiratory pigments at low oxygen tensions. a new technique. Biochim. Biophys. Acta (BBA)-Bioenerg. 245, 347–355 (1971)
Geltink, R.I.K., Kyle, R.L., Pearce, E.L.: Unraveling the complex interplay between t cell metabolism and function. Ann. Rev. Immunol. 36, 461–488 (2018)
Guillaud, F., Hannaert, P.: Dynamic simulation of mitochondrial respiration and oxidative phosphorylation: Comparison with experimental results. Acta Biotheor. 56, 157–172 (2008)
Jiang, B.-H., Semenza, G.L., Bauer, C., Marti, H.H.: Hypoxia-inducible factor 1 levels vary exponentially over a physiologically relevant range of O2 tension. Amer. J. Physiol. Cell Physiol. 271, C1172–C1180 (1996)
Joshi, A., Palsson, B.O.: Metabolic dynamics in the human red cell. Part III—metabolic reaction rates. J. Theor. Biol. 142, 41–68 (1990)
König, M., Holzhütter, H.-G., Berndt, N.: Metabolic gradients as key regulators in zonation of tumor energy metabolism: a tissue-scale model-based study. Biotechnol. J. 8, 1058–1069 (2013)
Ladas, G.E., Lakshmikantham, V.: Differential Equations in Abstract Spaces. Academy Press, New-York (1972)
Nazaret, C., Heiske, M., Thurley, K., Mazat, J.-P.: Mitochondrial energetic metabolism: a simplified model of TCA cycle with ATP production. J. Theor. Biol. 258, 455–464 (2009)
Nguyen, L.K., Cavadas, M.A.S., Scholz, C.C., Fitzpatrick, S.F., Bruning, U., Cummins, E.P., Tambuwala, M.M., Manresa, M.C., Kholodenko, B.N., Taylor, C.T., Cheong, A.: A dynamic model of the hypoxia-inducible factor 1α (HIF-1α) network. J. Cell Sci. 126, 1454–1463 (2013)
O’Neill, L.A.J., Kishton, R.J., Rathmell, J.: A guide to immunometabolism for immunologists. Nat. Rev. Immunol. 16, 553–565 (2016)
Park, J.O., Rubin, S.A., Xu, Y.-F., Amador-Noguez, D., Fan, J., Shlomi, T., Rabinowitz, J.D.: Metabolite concentrations, fluxes and free energies imply efficient enzyme usage. Nat. Chem. Biol. 12, 482–489 (2016)
Pearce, E.L., Poffenberger, M.C., Chang, C.-H., Jones, R.G.: Fueling immunity: insights into metabolism and lymphocyte function. Science 342 (6155):1242454 (2013)
Prüss, J.W., Wilke, M.: Gewöhnliche Differentialgleichungen Und Dynamische Systeme. Birkhäuser, Basel (2010)
Semenza, G.L.: Hypoxia-inducible factor 1: Regulator of mitochondrial metabolism and mediator of ischemic preconditioning. Biochim. Biophys. Acta (BBA)-mol. Cell Res. 1813, 1263–1268 (2011)
Simon, M.C.: Coming up for air: HIF-1 and mitochondrial oxygen consumption. Cell Metab. 3, 150–151 (2006)
Solaini, G., Baracca, A., Lenaz, G., Sgarbi, G.: Hypoxia and mitochondrial oxidative metabolism. Biochim. Biophys. Acta (BBA)-Bioenerg. 1797, 1171–1177 (2010)
Takeuchi, Y., Adachi, N., Tokumaru, H.: The stability of generalized Volterra equations. J. Math. Anal. Appl. 62, 453–473 (1978)
Taylor, C.T., Colgan, S.P.: Regulation of immunity and inflammation by hypoxia in immunological niches. Nat. Rev. Immunol. 17, 774–785 (2017)
Varma, A., Morbidelli, M., Wu, H.: Parametric Sensitivity in Chemical Systems. Cambridge University Press, Cambridge (2005)
Wang, G.L., Jiang, B.-H., Rue, E.A., Semenza, G.L.: Hypoxia- inducible factor 1 is a basic-helix-loop-helix-PAS heterodimer regulated by cellular O2 tension. Proc. Nat. Acad. Sci. 92, 5510–5514 (1995)
Wu, F., Yang, F., Vinnakota, K.C., Beard, D.A.: Computer modeling of mitochondrial tricarboxylic acid cycle, oxidative phosphorylation, metabolite transport, and electrophysiology. J. Biol. Chem. 282, 24525–24537 (2007)
Yamada, K., Hara, N., Shibata, T., Osago, H., Tsuchiya, M.: The simultaneous measurement of nicotinamide adenine dinucleotide and related compounds by liquid chromatography/electrospray ionization tandem mass spectrometry. Anal. Biochem. 352, 282–285 (2006)
Yang, H., Yang, T., Baur, J.A., Perez, E., Matsui, T., Carmona, J.J., Lamming, D.W., Souza-Pinto, N.C., Bohr, V.A., Rosenzweig, A., de Cabo, R., Sauve, A.A., Sinclair, D.A.: Nutrient-sensitive mitochondrial NAD+ levels dictate cell survival. Cell 130, 1095–1107 (2007)
Yu, Q., Heikal, A.A.: Two-photon autofluorescence dynamics imaging reveals sensitivity of intracellular NADH concentration and conformation to cell physiology at the single-cell level. J. Photochem. Photobiol. B: Biol. 95, 46–57 (2009)
Zhang, H., Bosch-Marce, M., Shimoda, L.A., Tan, Y.S., Baek, J.H., Wesley, J.B., Gonzalez, F.J., Semenza, G.L.: Mitochondrial autophagy is an HIF-1-dependent adaptive metabolic response to hypoxia. J. Biol. Chem. 283, 10892–10903 (2008)
Zi, Z.: Sensitivity analysis approaches applied to systems biology models. IET Syst. Biol. 5, 336–346 (2011)
Zupke, C., Sinskey, A.J., Stephanopoulos, G.: Intracellular flux analysis applied to the effect of dissolved oxygen on hybridomas. Appl. Microbiol. Biotechnol. 44, 27–36 (1995)
Acknowledgements
Open Access funding provided by Projekt DEAL. The research of G.B. was supported by the Russian Foundation for Basic Research (Grant no. 20-01-00352). The authors acknowledge the support by the Klaus Tschira Stiftung via the project SCIDATOS.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bocharov, G., Jäger, W., Knoch, J. et al. A Mathematical Model of HIF-1 Regulated Cellular Energy Metabolism. Vietnam J. Math. 49, 119–141 (2021). https://doi.org/10.1007/s10013-020-00426-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10013-020-00426-y
Keywords
- Cellular energy metabolism
- Oxidative phosphorylation
- Glycolysis
- Mathematical model
- Regulation
- Hypoxia-inducible factor 1