Abstract
Concept
MNDO-based semi-empirical methods in quantum chemistry have found widespread application in the modelling of large and complex systems. A method for the analytic evaluation of first and second derivatives of molecular properties against semi-empirical parameters in MNDO-based NDDO-descendant models is presented, and the resultant parameter Hessian is compared against the approximant currently used in parameterization for the PMx models.
Methods
As a proof of concept, the exact parameter Hessian is employed in a limited reparameterization of MNDO for the elements C, H, N, O and F using 1206 molecules for reference data (heats of formation, ionization energies, dipole moments and reference geometries). The correctness of our MNDO implementation was verified by comparing the calculated molecular properties with the MOPAC program.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In modern computational chemistry, semi-empirical methods based on the neglect of diatomic differential overlap (NDDO) [1,2,3,4,5,6,7,8,9,10,11,12] have found widespread applications in studies where more computationally intensive ab initio methods are unfeasible. While semi-empirical model Hamiltonians developed via machine-learning have gained significant attention [13, 14], recent work on the use of machine-learning techniques to construct correction terms for molecular properties calculated at lower levels of theory [15, 16] or develop semi-empirical molecular Hamiltonians [17] suggests that semi-empirical methods can achieve chemical accuracy. Essential to the success of a developed semi-empirical model is a robust parameterization procedure that allows the model to best reproduce experimental data; as such, parameter optimization is an important area of study for the development of effective and accurate semi-empirical models.
While alternative formulations based on the NDDO approximation [18,19,20,21,22] have been proposed in recent years, most NDDO-descendant semi-empirical models [1, 2, 5, 9] use an identical formalism for the construction of the one-electron matrix \(\mathbf{H}\) and two-electron matrix \(\mathbf{G}\); with the exception of minor corrections to the asymptotic behaviour of two-electron integrals in PM7 [10], these methods also employ the same approximations and semi-empirical expressions [23, 24] for the evaluation of relevant molecular integrals. The difference between most NDDO-descendant models hence lies only in the parameter values chosen as well as the empirical expressions for core-core repulsion terms, which have been modified significantly between models [2, 9,10,11]; as such, any parameterization scheme developed for one NDDO-descendant model can be readily applied with few modifications to other NDDO-descendant models.
As noted by Stewart when developing PM7 [10], the parameter Hessian can be used to determine the nature of stationary points detected during parameterization; likewise, accurate second-derivative information can be used to great effect in searching for minima on surfaces. Construction of the parameter Hessian via direct differentiation of the error function appears straightforward; in differentiating the error function \(\mathcal{S}\) with respect to parameters \({}^{{\mathrm Z}_{\mathrm A}}\mathrm p_{\mathrm i}\) and \({}^{{\mathrm Z}_{\mathrm B}}\mathrm p_{\mathrm i}\), we find that
It should be noted, however, that the parameter Hessian constructed in the development of PM7 appears to neglect second derivatives of the reference functions; as given in [10], the expression for \(\frac{{\partial }^{2}\mathcal{S}}{\partial {}{}^{{Z}_{A}}{p}_{i}\partial {}{}^{{Z}_{B}}{p}_{j}}\) appears to be
The neglect of the second-derivative term is expected to significantly affect the nature of the constructed Hessian matrix and its eigenvalues, impacting the quality of parameter optimization.
Evaluation of parameter derivatives of molecular properties via finite difference, as in standard procedure, results in numerical instability and may lead to an undesirable irreproducibility in results; thus, an analytical method for evaluation of parameter first and second derivatives is sought. This requires the evaluation of second derivatives of the density matrix as the idempotency condition [25] does not apply for derivatives of the ionization energy or dipole moment; an efficient method for solution of the second order coupled-perturbed Hartree–Fock (CPHF) equations is hence necessary for analytical derivative evaluation. NDDO methods formally operate in the Lowdin basis, where the overlap matrix between basis functions is substituted for the identity matrix; thus, the relevant equations for CPHF equation solution are greatly simplified and may be easily implemented.
The second-order CPHF equations under the NDDO approximation
The form of the first-order CPHF equations in the Lowdin basis for both closed- and open-shell species under the NDDO approximation has been well-documented [26] due to their implementation in geometry optimization routines [26, 27]; the second-order CPHF equations have been presented only in the MO basis [28] and are hence given in analogous form for the UHF case in the Lowdin basis.
All equations presented in this section are based on the Unrestricted Hartree–Fock (UHF) formalism; equations for the restricted case are obtained as a special case of these results. Matrices for a specific spin (e.g. the Fock matrices for alpha- and beta-spin) are denoted as \({}^{\sigma }\mathbf{M}\), where \(\sigma \in \left\{\alpha ,\beta \right\}\) is an arbitrary spin. \(\mathbf{P}={}{}^{\alpha }\mathbf{P}+{}{}^{\beta }\mathbf{P}\) represents the density matrix in the Lowdin basis, which is the sum of the alpha- and beta-spin density matrices.
We first define the generalized Coulomb and exchange matrices \(\mathbf{J}\left({\varvec{\Delta}}\right)\) and \(\mathbf{K}\left({\varvec{\Lambda}}\right)\) as well as their associated static derivatives as the contraction of the relevant 2-electron integrals with arbitrary matrices \({\varvec{\Delta}}\) and \({\varvec{\Lambda}}\):
In conventional NDDO methods [1,2,3,4,5,6,7,8,9,10,11], the core matrix is given by
In the above, \({V}_{\mu \nu ,B}\) represents the two-centre nuclear-electron attraction integral between basis functions \(\mu ,\nu\) and atom \(B\); \({\beta }_{\mu \nu }=\frac{{\beta }_{\mu }+{\beta }_{\nu }}{2}{S}_{\mu \nu }\) represents the resonance integral between two basis functions on different atoms.
The Fock matrix in the NDDO formalism can hence be given as follows:
In matrix form, the first-order orbital coefficients \({}^{\sigma }{\mathbf{x}}^{{q}_{1}}\) are related to the direct derivative of the (AO) coefficient matrix \({}^{\sigma }\mathbf{C}\) as follows:
Analogously, the second-order orbital coefficients can be obtained via differentiation of the first-order orbital coefficients:
Application of the orthonormality condition \({{}^{\sigma }\mathbf{C}}^{\mathrm{T}}{}{}^{\sigma }\mathbf{C}=1\) yields the relevant commutator relations for the first- and second-order orbital coefficients:
The derivatives of the off-diagonal elements of the MO-basis Fock matrix must be zero due to the variational condition:
The first-order CPHF equations are well-documented in the literature and are hence only presented for completeness.
Evaluation of the first derivative \(\frac{d{}{}^{\sigma }\mathfrak{F}}{d{q}_{1}}\) yields the following:
The “static” derivative term \(\frac{d\mathbf{H}}{d{q}_{1}}+{\mathbf{J}}^{{q}_{1}}\left(\mathbf{P}\right)-{\mathbf{K}}^{{q}_{1}}\left({}{}^{\sigma }\mathbf{P}\right)\) is referred to as \({}^{\sigma }{\mathbf{F}}^{{q}_{1}}\) and the response term \(\mathbf{J}\left(\frac{d\mathbf{P}}{d{q}_{1}}\right)-\mathbf{K}\left(\frac{d{}{}^{\sigma }\mathbf{P}}{d{q}_{1}}\right)\) is referred to as \({}^{\sigma }{\mathbf{R}}^{{q}_{1}}\); collection of terms yields
Thus, the first-order CPHF equations are given as follows, as reported in [26, 27]:
To evaluate \({{}^{\sigma }\mathfrak{R}}_{ij}^{{q}_{1}}\), \(\frac{d {}{}^{\sigma }\mathbf{P}}{d {q}_{1}}\) must be cast in terms of \({}^{\sigma }{\mathbf{x}}^{{q}_{1}}\); since \({}^{\sigma }\mathbf{P}={{}{}^{\sigma }\mathbf{C}}_{occ}^{\mathrm{T}}{}{}^{\sigma }{\mathbf{C}}_{occ}\),
Lastly, the expression for \(\frac{d\mathbf{H}}{d{q}_{1}}\) is of Eq. (10):
The first-order CPHF equations are hence linear in the first-order coefficients \({x}_{ij}^{{q}_{1}}\).
Since only the occupied-virtual block of \({}^{\sigma }{\mathbf{x}}^{{q}_{1}}\) is necessary to solve for \(\frac{d{}{}^{\sigma }\mathbf{P}}{d{q}_{1}}\), the remaining elements of \({}^{\sigma }{\mathbf{x}}^{{q}_{1}}\) are evaluated afterwards via direct substitution:
The full matrix \({}^{\sigma }{\mathbf{x}}^{{q}_{1}}\), not just the occupied-virtual block, is necessary for solution of the second-order CPHF equations. These are obtained via further differentiation of Eq. (18):
Simplification yields:
Application of the commutator relation further reduces the expression complexity:
Evaluation of \(\frac{{d}^{2}{}{}^{\sigma }\mathbf{F}}{d{q}_{1}d{q}_{2}}\) is performed analogously to Eq. (19), albeit with significantly more terms in the resultant expression:
Lastly, direct differentiation of the density matrix yields
Since the first three terms in \({\left(\frac{{d}^{2}{}{}^{\sigma }\mathbf{P}}{d{q}_{1}d{q}_{2}}\right)}_{\mu \nu }\) are independent of the second-order coefficients, we define
The static derivative term in \(\frac{{d}^{2}{}{}^{\sigma }\mathbf{F}}{d{q}_{1}d{q}_{2}}\) is hence termed \({{}^{\sigma }\mathbf{F}}^{{q}_{1}{q}_{2}}\), with the response term correspondingly referred to as \({{}^{\sigma }\mathbf{R}}^{{q}_{1}{q}_{2}}\):
Accordingly, defining \({{}^{\sigma }\mathfrak{F}}^{{q}_{1}{q}_{2}}\) and \({{}^{\sigma }\mathfrak{R}}^{{q}_{1}{q}_{2}}\) yields the second-order CPHF equations:
The form of the second-order CPHF equations has intentionally been cast into a form that resemble the first-order CPHF equations; as such, the same algorithms [26, 29] employed to solve the first-order CPHF equations may be applied for solution of the second-order CPHF equations.
Derivatives of molecular properties under the MNDO formalism
The first derivatives \(\frac{d\left(\Delta {H}_{f}\right)}{d{}{}^{{Z}_{A}}\alpha }\) and \(\frac{d\left(\Delta {H}_{f}\right)}{d{}{}^{{Z}_{A}}{E}_{eisol}}\) are easily evaluated:
If another NDDO-based semi-empirical method using the same formalism as MNDO for \(\mathbf{H}\) and \(\mathbf{G}\) [1,2,3,4,5,6,7,8,9,10,11] is required, the derivatives of \(\Delta {H}_{f}\) with respect to the additional core-repulsion function parameters will depend on the expression for \({V}_{BC}^{CRF}\) and can be easily obtained via direct differentiation.
The remaining derivatives (against \({}^{{Z}_{A}}{\beta }_{s},{}{}^{{Z}_{A}}{\beta }_{p}, {}{}^{{Z}_{A}}{U}_{ss},{}{}^{{Z}_{A}}{U}_{pp}, {}{}^{{Z}_{A}}{\zeta }_{s},{}{}^{{Z}_{A}}{\zeta }_{p}\)) are identical for [1,2,3,4,5,6,7,8,9,10,11] but will require modification for other methods (e.g. MNDO-F, OMx):
Likewise,
The expressions for \(\frac{d\mathbf{H}}{d{}{}^{{Z}_{A}}p}\), \(\frac{{d}^{2}\mathbf{H}}{d{}{}^{{Z}_{A}}{p}_{i}d{}{}^{{Z}_{B}}{p}_{j}}\), \({\mathbf{J}}^{{}{}^{{Z}_{A}}p}\left({\varvec{\Delta}}\right)\), \({\mathbf{J}}^{{}{}^{{Z}_{A}}{p}_{i}{}{}^{{Z}_{B}}{p}_{j}}\left({\varvec{\Delta}}\right)\), \({\mathbf{K}}^{{}{}^{{Z}_{A}}p}\left({\varvec{\Lambda}}\right)\) and \({\mathbf{K}}^{{}{}^{{Z}_{A}}{p}_{i}{}{}^{{Z}_{B}}{p}_{j}}\left({\varvec{\Lambda}}\right)\) for each of the parameters \({}^{{Z}_{A}}{\beta }_{s},{}{}^{{Z}_{A}}{\beta }_{p}, {}{}^{{Z}_{A}}{U}_{ss},{}{}^{{Z}_{A}}{U}_{pp}, {}{}^{{Z}_{A}}{\zeta }_{s},{}{}^{{Z}_{A}}{\zeta }_{p}\) are detailed in the Supplementary Information.
Direct differentiation yields the first and second derivatives of the (semi-empirical) dipole moment and the energy matrix \({}^{\sigma }{\varvec{\upepsilon}}\) via which the ionization energy can be obtained:
Derivatives of dipole moments are presented for the Restricted Hartree–Fock case, as no extension to UHF systems is necessary.
Lastly, the elements of \(\frac{d\mathbf{g}}{d{}{}^{{Z}_{C}}p}\) (derivative of the gradient vector in Cartesian coordinates against arbitrary parameter \({}^{{Z}_{C}}p\)) are given by
Elements of \(\frac{{d}^{2}\mathbf{g}}{d{}{}^{{Z}_{C}}{p}_{i}d{}{}^{{Z}_{D}}{p}_{j}}\) are likewise obtained by direct differentiation of the above expression and are omitted for brevity.
Further details on the equations in this section, where necessary, are provided in the Supplementary Information.
Nature of the Hessian approximant in PM7
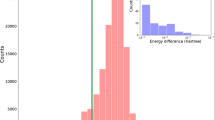
To illustrate the differences between the exact Hessian \(\mathbf{H}\) and the approximant \({}^{\mathrm{PM}7}\mathbf{H}\), a pictorial representation of the two matrices is given in Fig. 1. The matrices were computed using original MNDO parameters [1] for a chosen training set of 1206 molecules (see Supplementary Information).
A heatmap of the elements of the exact Hessian \(\mathbf{H}\) (left) and Hessian approximant \({}^{\mathrm{PM}7}\mathbf{H}\) (right), raised to the fifth root. The \(37\times 37\) Hessian contains second derivatives for the MNDO parameters \(\alpha ,{\beta }_{s},{\beta }_{p},{U}_{ss},{U}_{pp},{\zeta }_{s},{\zeta }_{p},{E}_{isol}\) for the elements H, C, N, O and F
Furthermore, the approximant \({}^{\mathrm{PM}7}\mathbf{H}\) is observed to be positive definite while \(\mathbf{H}\) reflects that the parameter surface is non-convex (Table 1).
Methods for parameter optimization
In PM7, parameter optimization is performed via an approximated line search, with the search direction obtained via a direct Hessian descent (HD) step on the approximated Hessian [5]:
This method of determining the step size shall be termed the approximated line search (ALS) method; an alternative would be a trust radius (TR), where the step size is constrained by a dynamic trust radius. The modification of the trust radius in our attempt is given by the following computation, with \(\mathbf{B}\) representing an arbitrary Hessian or Hessian approximant:
In addition to determination of \(\widehat{\mathbf{d}}\) via direct Hessian descent, a trust region optimization (TRO) was also attempted, where \(\mathbf{d}\) is computed via:
The shift parameter \(\lambda\) is computed iteratively, with the chosen numerical method detailed in the Supplementary Information.
Lastly, three different choices for the second-derivative matrix \(\mathbf{B}\) were investigated. In addition to the exact Hessian \(\mathbf{H}\) and the approximant \({}^{\mathrm{PM}7}\mathbf{H}\), a modified Hessian \({\mathbb{H}}\) was found to yield promising results:
The modified Hessian \({\mathbb{H}}\) preserves the eigenvectors of the exact Hessian while converting it to be positive (semi)definite, and ensures that a direct Hessian descent step will not traverse in the uphill direction given a surface of the wrong concavity.
It should be noted that \({}^{\mathrm{PM}7}\mathbf{H}\) is constructed without any second derivatives of molecular properties, and hence, optimization with \({}^{\mathrm{PM}7}\mathbf{H}\) is comparable to other methods using approximate Hessians that do not evaluate second derivative information (e.g. the BFGS or DFP schemes); however, \({}^{\mathrm{PM}7}\mathbf{H}\) appears to provide a reasonably good estimate for \(\mathbf{H}\) despite having incorrect eigenvalue information (as seen in Fig. 2, the percentage errors for specific elements in the Hessian matrix are at most 4–5%) and is thus expected to perform better than a regular quasi-Newton optimizer. We note in passing that PM3 and PM6 were optimized with the method outlined in [5] using a DFP update scheme while the more recent PM7 [10] was optimized using \({}^{\mathrm{PM}7}\mathbf{H}\), suggesting that \({}^{\mathrm{PM}7}\mathbf{H}\) better approximates the exact Hessian \(\mathbf{H}\).
The computation of the gradient \(\mathbf{g}\) is straightforward and may be obtained easily by both finite difference and analytic differentiation; this suggests that gradient-free optimization methods would not be necessary in optimizing parameters for NDDO-based methods. Such local optimizers can, however, be combined with other methods to identify global minima, e.g. genetic algorithms [30, 31], and may be indicative of possible avenues for further work on this subject. We defer a discussion of the nature of local and global minima on the parameter surface to the “Conclusion”.
Results of limited parameterisation
As a proof of concept, a limited parameterization of 1206 molecules consisting of the atom types C, H, N, O, F was performed using the MNDO formalism; the relevant routines for the semi-empirical evaluation of molecular properties using MNDO were correspondingly implemented and compared against MOPAC [32] for accuracy (see the Supplementary Information). Geometrical data, e.g., bond angles or bond lengths, are accounted for in our parameterization procedure by using the norm of the gradient vector \(\left|\mathbf{g}\right|\) at a reference geometry (either experimental or the result of high-level calculations), with the corresponding reference function set to zero to simulate a perfect correspondence between the semi-empirical and reference geometries; this was chosen to facilitate the ease of preparing the relevant inputs. The weighting functions \({\mathcal{C}}_{i}\) for the error function were chosen as specified in Table 2.
While additional properties such as reaction barriers were also viable choices for inclusion as reference data in our training set, such information was not used in our limited parameterization; nonetheless, the parameter derivatives of the molecular energy used in calculating derivatives of \(\Delta {H}_{f}\) can be applied to calculate the relevant parameter derivatives for the energies of transition states, allowing for an easy extension if necessary.
For all optimization methods, the graphs shown below are terminated once the decrease in \(\mathcal{S}\) is no longer appreciable; the optimization runs discussed in this section hence do not represent the identification of minima on the parameter surface.
First, application of the optimization method used in the PMx models (Hessian descent with step size determined by the approximate line search) leads to surprisingly similar behaviour when both \({\mathbb{H}}\) and \({}^{\mathrm{PM}7}\mathbf{H}\) are used; nonetheless, there is an appreciable difference in the final parameter values for the two methods. Direct employment of the Hessian matrix \(\mathbf{H}\) results in a poor optimisation procedure as the evaluated step size shrinks significantly around \(\mathcal{S}=575000\); this is expected as the line-search procedure seeks to minimise \(\mathcal{S}\) even as the Newton–Raphson step traverses uphill for nonconvex regions. The optimization curves for \(\mathbf{H}\) are hence omitted in Figs. 3, 4 and 5.
To compare the line-search method used in the PMx models with a trust radius method, optimization using direct Hessian descent but with the step size determined via a dynamic trust radius was attempted. We note in passing that the resultant optimization curve strongly resembles that of the according line-search method (cf. Figure 3); once again, direct employment of \(\mathbf{H}\) leads to very poor optimisation, and the resultant curve has been omitted:
Lastly, optimization was attempted with a trust region optimizer; this led to similar performance for \({\mathbb{H}}\) while \({}^{\mathrm{PM}7}\mathbf{H}\) performed markedly poorer. Optimization with \(\mathbf{H}\) led to suboptimal parameters that encountered problems with SCF convergence and geometry optimization and was hence abandoned.
Optimization with \({}^{\mathrm{PM}7}\mathbf{H}\) appears to result in the identification of a saddle point, which is incorrectly identified as a minimum due to the positive-definite nature of \({}^{\mathrm{PM}7}\mathbf{H}\); the Hessian eigenvalues (from the exact Hessian \(\mathbf{H}\)) for the resultant parameters obtained via optimization with \({}^{\mathrm{PM}7}\mathbf{H}\) are given in Table 3.
In all methods (Hessian descent with both line search and trust radius determination of step size as well as the trust-region optimizer), the step size is constrained to a maximum of \({\left|\mathbf{d}\right|}_{max}=2\) to ensure that there are no significant and unexpected increases in the error function. We note in passing that the choice of \({\left|\mathbf{d}\right|}_{max}=2\), while arbitrary, does not play a significant role in the optimization of parameters; choices of \({\left|\mathbf{d}\right|}_{max}=1\) and \({\left|\mathbf{d}\right|}_{max}=3\) lead to very similar optimization curves (Fig. 6).
In the course of parameterization, it was realized that identification of a true local minimum would be difficult and require manual intervention in the parameterization procedure; thus, any set of parameters with a reasonably small gradient vector magnitude, a positive definite (exact) Hessian matrix and a marked resistance to further reduction in the error function should be accepted as a reasonable result from parameter optimization.
In this work, we report the identification of a local minimum from our limited parameterization, obtained with the imposition of no constraints on the parameters. The parameters, as well as the values of the associated gradient element, for an identified minimum with \(\mathcal{S}=261996\) are given in Table 4; the associated Hessian eigenvalues are presented in Table 5.
While clearly unphysical, these parameters indicate a position close to a local minimum on the parameter surface and are an indication of the significant deficiencies of the MNDO model in modelling charged or radical species.
Notably, ammonia is predicted to be planar using these parameters; as the non-planarity of ammonia was not a constraint imposed upon the system for parameterization, this is not an erroneous result. A detailed tabulation of the predicted molecular properties at the given parameters is provided in the Supplementary Information.
In Fig. 7, optimization using \({\mathbb{H}}\) with initial step size 0.1 and \({\left|\mathbf{d}\right|}_{max}=2\) is compared for three sets of starting parameters: the original MNDO parameters, the MNDO parameters reported for PDDG/MNDO [11] and the parameters for the elements C, H, N and O reported in NO-MNDO [22] alongside original MNDO parameters for fluorine (see Tables 6 and 7). Since the formalism used remains that of MNDO and not PDDG/MNDO or NO-MNDO, these additional sets of parameters are expected to perform worse than the original MNDO parameters; however, they provide an interesting case study for how our local optimizer might behave when starting from different initial parameter values. The graph is terminated after 150 optimization runs, and the final error function values obtained are hence not reflective of the identification of a local minimum; nonetheless, the very similar error function values despite the significant differences in final parameters (as reported in Table 8) seem to indicate that there may be numerous local minima with similar error function values.
Conclusion
In this paper, we report a fully analytic differentiation routine for the evaluation of parameter derivatives for construction of the parameter gradient and Hessian in MNDO-based semi-empirical methods and have applied these equations for a proof-of-concept optimization of the MNDO parameters for the elements C, H, N, O and F. While the PM7 Hessian \({}^{\mathrm{PM}7}\mathbf{H}\) appears to work remarkably well in parameterization schemes, it does not guarantee that the identified stationary point on the parameter surface will be a local minimum; furthermore, \({}^{\mathrm{PM}7}\mathbf{H}\) does appear to perform worse when the optimization nears a local minimum or stationary point. The full Hessian \(\mathbf{H}\) may be necessary for optimization near the stationary point, and the accurate eigenvalue information may also be helpful in ensuring that a local minimum is attained at the termination of parameterization.
We have placed significant emphasis on the identification of local minima on the parameter surface. While it is conceded that identification of global minima is of greater concern in parameterization, we note that parameter surfaces encountered in the optimization of neural networks often show multiple local minima, all with similar loss function values [33] and posit that a similar situation may be observed in NDDO-based methods.
Data Availability
The relevant data in this paper shall be provided upon request.
References
Dewar MJS, Thiel W (1977) Ground states of molecules. 38. The MNDO method. Approximations and parameters. J Am Chem Soc 99:4899–4907
Dewar MJ, Zoebisch EG, Healy EF, Stewart JJ (1985) Development and use of quantum mechanical molecular models. 76. AM1: a new general purpose quantum mechanical molecular model. J Am Chem Soc 107(13):3902–3909
Rocha GB, Freire RO, Simas AM, Stewart JJ (2006) RM1: a reparameterization of AM1 for H, C, N, O, P, S, F, Cl, Br, and I. J Comput Chem 27:1101–1111
Thiel W, Voityuk AA (1992) Extension of the MNDO formalism to d orbitals: integral approximations and preliminary numerical results. Theoret Chim Acta 81:391–404
Stewart JJP (1989) Optimization of parameters for semiempirical methods I Method. J Comput Chem 10:209–220
Stewart JJ (1989) Optimization of parameters for semiempirical methods II. Appl J Comput Chem 10:221–264
Stewart JJ (1991) Optimization of parameters for semiempirical methods III extension of PM3 to Be, Mg, Zn, Ga, Ge, as, SE, CD, in, Sn, Sb, Te, Hg, Tl, Pb, and Bi. J Comput Chem 12:320–341
Stewart JJ (2004) Optimization of parameters for semiempirical methods IV: extension of MNDO, AM1, and PM3 to more main group elements. J Mol Model 10:155–164
Stewart JJP (2007) Optimization of parameters for semiempirical methods V: modification of NDDO approximations and application to 70 elements. J Mol Model 13:1173–1213
Stewart J (2012) Optimization of parameters for semiempirical methods VI: more modifications to the NDDO approximations and re-optimization of parameters. J Mol Model 19(1):1–32
Repasky MP, Chandrasekhar J, Jorgensen WL (2002) PDDG/PM3 and PDDG/MNDO: improved semiempirical methods. J Comput Chem 23:1601–1622
Husch T, Vaucher AC, Reiher M (2018) Semiempirical molecular orbital models based on the neglect of diatomic differential overlap approximation. Int J Quantum Chem 118:e25799
Zhang H, Lau JWZ, Wan L, Shi L, Shi Y, Cai H, Luo X, Lo GQ, Lee CK, Kwek LC, Liu AQ, (2023) Molecular Property Prediction with Photonic Chip-Based Machine Learning. Laser Photonics Rev 17:2200698
Bhat HS, Ranka K, Isborn CM (2020) Machine learning a molecular Hamiltonian for predicting electron dynamics. Int J Dyn Control 8(4):1089–1101
Ramakrishnan R, Dral PO, Rupp M, von Lilienfeld OA (2015) Big Data meets quantum chemistry approximations: the Δ-machine learning approach. J Chem Theory Comput 11(5):2087–2096
Chen J, Xu W, Zhang R (2021) A machine learning approach using frequency descriptor for molecular property predictions. New J Chem 45(44):20672–20680
Dral PO, von Lilienfeld OA, Thiel W (2015) Machine learning of parameters for accurate semiempirical quantum chemical calculations. J Chem Theory Comput 11(5):2120–2125
Kolb M, Thiel W (1993) Beyond the MNDO model: methodical considerations and numerical results. J Comput Chem 14:775–789
Weber W, Thiel W (2000) Orthogonalization corrections for semiempirical methods. Theor Chem Acc 103:495–506
Scholten M (2003) Semiemirische Verfahren mit Orthogonalisierungskorrek- turen: Die OM3 Methode. Heinrich-Heine-Universität Düsseldorf, Thesis
Dral PO, Wu X, Spörkel L, Koslowski A, Weber W, Steiger R, Scholten M, Thiel W (2016) Semiempirical quantum-chemical orthogonalization-corrected methods: theory, implementation, and parameters. J Chem Theory Comput 12:1082–1096
Sattelmeyer KW, Tubert-Brohman I, Jorgensen WL (2006) NO-MNDO: reintroduction of the overlap matrix into MNDO. J Chem Theory Comput 2:413–419
Goeppert-Mayer M, Sklar AL (1938) Calculations of the lower excited levels of benzene. J Chem Phys 6:645–652
Dewar MJS, Thiel W (1976) A semiempirical model for the two-center repulsion integrals in the NDDO approximation. Theor Chim Acta 46:89–104
Pulay P (1969) Ab initio calculation of force constants and equilibrium geometries in polyatomic molecules. Molecular Physics 17(2):197–204
Patchokovskii S, Thiel W (1996) Analytical second derivatives of the energy in MNDO methods. J Comput Chem 17(11):1318–1327
Frisch M, Scalmani G, Vreven T, Zheng G (2009) Analytic second derivatives for semiempirical models based on MNDO. Mol Phys 107:881–887
Osamura Y, Yamaguchi Y, Schaefer HF (1986) Second-order coupled perturbed Hartree-Fock equations for closed-shell and open-shell self-consistent-field wavefunctions. Chem Phys 103:227–243
Pople JA, Krishnan R, Schelegel HB, Binkley JS (1979) Derivative studies in Hartree-Fock and Møller-Plesset theories. Intl J Quantum Chem 13:225
Rossi I, Truhlar DG (1995) Parameterization of NDDO wavefunctions using genetic algorithms an evolutionary approach to parameterizing potential energy surfaces and direct dynamics calculations for organic reactions. Chem Phys Letters 233(3):231–236
Brothers EN, Merz KM (2002) Sodium parameters for AM1 and PM3 optimized using a modified genetic algorithm. J Phys Chem B 106(10):2779–2785
Stewart JJ (1990) Mopac: A semiempirical molecular orbital program. J Comput Aided Mol Des 4:1–103
Choromańska A, Henaff M, Mathieu M, Arous GB, LeCun Y (2014) The loss surfaces of multilayer networks. International Conference on Artificial Intelligence and Statistics
Author information
Authors and Affiliations
Contributions
A.O. wrote the main manuscript text and S.C. prepared all figures and tables in the manuscript. K.L.C. provided guidance and supervised the research work. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Ethical approval
Not applicable.
Competing interests
Not applicable.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ong, A.W.W., Cao, S.Y. & Kwek, L.C. An improved parameterization procedure for NDDO-descendant semi-empirical methods. J Mol Model 29, 118 (2023). https://doi.org/10.1007/s00894-023-05499-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00894-023-05499-3