Abstract
The subject of this study was an analysis of the role of active site residues in the phosphoryl transfer reaction catalyzed by 4-methyl-5-β-hydroxyethylthiazole kinase (ThiK). The ThiK-catalyzed reaction is of special interest due to the lack of a highly conserved aspartate residue serving as a catalytic base. ONIOM(B3LYP:PM3) models of stationary points along the reaction pathway consisted of reactants, two magnesium ions and several highly conserved ThiK active site residues. The results indicate that an SN2-like mechanism of ThiK, with γ-phosphate acting as an alcohol-activating base is reasonable. Geometries of substrates, transition state and products were utilized in the non-empirical analysis of the physical nature of catalytic interactions taking place in the ThiK active site. The role of particular residues was investigated in terms of their ability to preferentially stabilize the transition state relative to substrates (differential transition state stabilization, DTSS) or products (differential product stabilization, DPS). It seems that Mg2, Glu126 and Cys198 play a major catalytic role, whereas Mg1 and the same Cys198 are responsible for product release. It is remarkable that no dominant role of an electrostatic term in the interactions involved in catalytic activity is observed for product release. Determination of catalytic fields expressing differential electrostatic potential of the transition state with respect to substrates revealed the optimal electrostatic features of an ideal catalyst for the studied reaction. The predicted catalytic environment is in agreement with experimental data showing increased catalytic activity of ThiK upon mutation of Cys198 to aspartate.

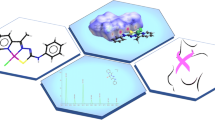
Catalytic fields for ThiK-catalyzed reaction juxtaposed with the positions of active site residues of a model system. Magnesium ions are considered part of the transition state/reactants. The surface of constant electronic density is colored according to differential electrostatic potential of transition state with respect to reactants. The sign of the differential potential reflects the electrostatic properties of a complementary molecular environment. Red (green) color denotes regions where a negative (positive) charge would be optimal for catalytic activity







Similar content being viewed by others
References
Cheek S, Zhang H, Grishin NV (2002) J Mol Biol 320:855–881
Adams JA (2001) Chem Rev 101:2271–2290
Bork P, Sander C, Valencia A (1993) Prot Sci 2:31–40
Campobasso N, Mathews II, Begley TP, Ealick SE (2000) Biochemistry 39:7868–7877
Sigrell JA, Cameron AD, Jones TA, Mowbray SL (1998) Structure 6:183–193
Mathews II, Erion MD, Ealick SE (1998) Biochemistry 37:15607–15620
Li MH, Kwok F, Chang WR, Lau CK, Zhang JP, Lo SCL, Jiang T, Liang DC (2002) J Biol Chem 277:46385–46390
Matte A, Tari LW, Delbaere LTJ (1998) Structure 6:413–419
Becke AD (1993) J Chem Phys 98:5648–5652
Lee CT, Yang WT, Parr RG (1988) Phys Rev B 37:785–789
Stewart JJP (1989) J Comput Chem 10:209–220
Maseras F, Morokuma K (1995) J Comput Chem 16:1170–1179
Frisch MJ, Trucks GW, Schlegel HB, Scuseria GE, Robb MA, Cheeseman JR, Montgomery JR, Vreven T, Kudin KN, Burant JC, Millam JM, Iyengar SS, Tomasi J, Barone V, Mennucci B, Cossi M, Scalmani G, Rega N, Petersson GA, Nakatsuji H, Hada M, Ehara M, Toyota K, Fukuda R, Hasegawa J, Ishida M, Nakajima T, Honda Y, Kitao O, Nakai H, Klene M, Li X, Knox JE, Hratchian HP, Cross JB, Adamo C, Jaramillo J, Gomperts R, Stratmann RE, Yazyev O, Austin AJ, Cammi R, Pomelli C, Ochterski JW, Ayala PY, Morokuma K, Voth GA, Salvador P, Dannenberg JJ, Zakrzewski VG, Dapprich S, Daniels AD, Strain MC, Farkas O, Malick DK, Rabuck AD, Raghavachari K, Foresman JB, Ortiz JV, Cui Q, Baboul AG, Clifford S, Cioslowski J, Stefanov BB, Liu G, Liashenko A, Piskorz P, Komaromi I, Martin RL, Fox DJ, Keith T, Al-Laham MA, Peng CY, Nanayakkara A, Challacombe M, Gill PMW, Johnson B, Chen W, Wong MW, Gonzalez C, Pople JA (2004) Gaussian 03. Gaussian Inc, Wallingford CT
Dyguda E, Szefczyk B, Sokalski WA (2004) Int J Mol Sci 5:141–153
Bairoch A, Apweiler R, Wu CH, Barker WC, Boeckmann B, Ferro S, Gasteiger E, Huang H, Lopez R, Magrane M, Martin MJ, Natale DA, O’Donovan C, Redaschi N, Yeh LS (2005) Nucleic Acids Res 33:D154–D159
Thompson JD, Higgins DG, Gibson TJ (1994) Nucleic Acids Res 22:4673–4680
Gouet P, Courcelle E, Stuart DI, Metoz F (1999) Bioinformatics 15:305–308
Akola J, Jones RO (2003) J Phys Chem B 107:11774–11783
Diaz N, Field MJ (2004) J Am Chem Soc 126:529–542
Cheng Y, Zhang Y, McCammon JA (2005) J Am Chem Soc 127:1553–1562
Szefczyk B, Mulholland AJ, Ranaghan KE, Sokalski WA (2004) J Am Chem Soc 126:16148–16159
Dyguda E, Grembecka J, Sokalski WA, Leszczyński J (2005) J Am Chem Soc 127:1658–1659
Sokalski WA, Roszak S, Pecul K (1988) Chem Phys Lett 153:153–159
Boys FS, Bernardi D (1970) Mol Phys 19:553–566
Schmidt MW, Baldridge KK, Boatz JA, Elbert ST, Gordon MS, Jensen JH, Koseki S, Matsunaga N, Nguyen KA, Su SJ, Windus TL, Dupuis M, Montgomery JA (1993) J Comput Chem 14:1347–1363
Sokalski WA (1985) J Mol Catalysis 30:395–410
Cleland WW, Hengge AC (2006) Chem Rev 106:3252–3278
Brooks BR, Bruccoleri RD, Olafson BO, States DJ, Swaminathan S, Karplus M (1983) J Comput Chem 4:187–217
MacKerell AD Jr, Bashford D, Bellott M, Dunbrack RL Jr, Evanseck JD, Field MJ, Fischer S, Gao J, Guo H, Ha S, Joseph-McCarthy D, Kuchnir L, Kuczera K, Lau FTK, Mattos C, Michnick S, Ngo T, Nguyen DT, Prodhom B, Reiher WE III, Roux B, Schlenkrich M, Smith JC, Stote R, Straub J, Watanabe M, Wiorkiewicz-Kuczera J, Yin D, Karplus M (1998) J Phys Chem B 102:3586–3616
Grzywa R, Dyguda-Kazimierowicz E, Sieńczyk M, Feliks M, Sokalski WA, Oleksyszyn J (2007) J Mol Mod 13: DOI 10.1007/s00894-007-0193-8
Mildvan AS (1997) Proteins 29:401–416
Herberg FW, Doyle ML, Cox S, Taylor SS (1999) Biochemistry 38:6352–6360
Acknowledgments
This work is funded by the British–Polish Young Scientists Programme. The authors are also grateful for financial support from Wrocław University of Technology and Jackson State University subcontract #W912HZ-04-2-0002. Dr. Borys Szefczyk is acknowledged for the software for visualization of catalytic fields. Calculations were performed in Wrocław (WCSS) and Poznań (PCSS) Centers for Supercomputing and Networking as well as the Interdisciplinary Centre for Mathematical and Computational Modeling (ICM) in Warsaw.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Dyguda-Kazimierowicz, E., Sokalski, W.A. & Leszczyński, J. Non-empirical study of the phosphorylation reaction catalyzed by 4-methyl-5-β-hydroxyethylthiazole kinase: relevance of the theory of intermolecular interactions. J Mol Model 13, 839–849 (2007). https://doi.org/10.1007/s00894-007-0192-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00894-007-0192-9