Abstract
The regulatory role of protein cysteine phosphorylation is an under-researched area. The difficulty of accessing reference S-phosphorylated peptides (pCys-peptides) hampers progress in MS-driven cysteine phosphoproteomics, which requires targeted analytical procedures. This work describes an uncomplicated process for the conversion of disulfide-bridged protein into a complex model mixture of combinatorially modified peptides. Hen egg-white lysozyme was reduced with tris(2-carboxyethyl)phosphine (TCEP) followed by alkylation of cysteine with (3-acrylamidopropyl)trimethyl-ammonium chloride (APTA) and subsequent beta-elimination in aqueous Ba(OH)2 to yield modified polypeptides containing multiple dehydroalanine (Dha) residues. The conjugate addition of thiophosphoric acid to Dha residues followed by trypsinolysis led to numerous D/L phosphocysteine-containing peptides, which were identified by higher-energy collisional-dissociation tandem mass spectrometry (HCD-MS/MS). Our results show that some pCys-peptides produce prominent neutral losses of 80 Da, 98 Da and a weak 116 Da loss. These are similar to the neutral-loss triplets generated by phosphohistidine peptides.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Continuing progress in phosphoproteomics studies is elucidating cellular processes that are modulated by dynamic phosphorylation on hydroxyl (Duan and Walther 2015), carboxyl (Attwood et al. 2011; Lai et al. 2017) and amine groups (Marmelstein et al. 2017) in protein amino acid residues. Moderately susceptible to acid hydrolysis, S-linked phosphorylation is among the least explored modifications (Shannon and Weerapana 2013; Piggott and Attwood 2017) and it is discussed mainly in the context of transient cysteine phosphorylation of protein tyrosine phosphatases (Tonks 2014). In 2012, the reversible cysteine phosphorylation of several global transcriptional regulators was reported in Staphylococcus aureus (Sun et al. 2012). It was suggested that this modification plays a crucial role in bacterial signaling, and might be prevalent in various organisms. In 2016, Bertran-Vicente et al. described studies on chemoselective S-phosphorylation of cysteine-containing synthetic peptides. Using the Escherichia coli system, the authors overexpressed the IIBGlc domain of the glucose transporter IICBGlc, identified the tryptic peptide ENITNLDApCITR and independently synthesized it to verify the sequence (Bertran-Vicente et al. 2016). At the same time, Gehring’s group proved the appearance of S-phosphorylation on phosphatases of regenerating liver (PRLs), overexpressed in HeLa cells. This modification is located on the catalytic cysteine, and in vivo it is abundant and persistent (Gulerez et al. 2016; Zhang et al. 2017). Phosphatases of regenerating liver play a causative role in cancer progression and metastasis, and thus deeper insight into the function of S-phosphorylation could be scientifically valuable (Rubio and Köhn 2016; Yu and Zhang 2018). Transient catalytic cysteine phosphorylation has also been reported for the protein tyrosine phosphatase, non-receptor type 12 (PTPN12) (Dong et al. 2018) and the phosphoglucan phosphatase OsttaDSP from the green alga Ostreococcus tauri (Carrillo et al. 2018).
More work is needed to understand the major factors that influence the chemical stability (Buchowiecka 2014) and MS-fragmentation behavior of S-phosphopeptides. Thus, generating a reference MS-library of the model peptides, categorized by sequence, pCys-position, and pI values, would be a worthwhile effort. The methodological utility of spectral libraries has already been proven in detailed studies of phosphorylation events such as arginine phosphorylation in Staphylococcus aureus (Marx et al. 2013; Junker et al. 2018; Deutsch et al. 2019).
Currently, there are two general approaches to transforming cysteine-containing proteins into their respective S-phosphorylated derivatives: via dehydroalanine (Dha) intermediates (Chalker et al. 2012; Morrison et al. 2015), or through three-step transformation on cysteine sulphur involving the use of photo-labile thiophosphoric acid O,O-diesters (Bertran-Vicente et al. 2016). However, these methods are not readily applicable to the synthesis of a larger set of S-phosphorylated structures.
This work offers a robust method for converting a single disulfide-bonded protein into a very complex, model mixture of combinatorially modified peptides, comprising diverse S-phosphorylated sequences (pCys-peptides). An empirical mass spectra library of pCys-peptides and a larger reference library of Cys-containing peptides were collected and described. Finally, general advice for detecting intact S-phosphorylated peptides and their dephosphorylated analogs is proposed for consideration in the context of cysteine phosphoproteomics.
Materials and methods
Reagents
Water from a Simplicity® Water Purification System, Millipore (Darmstadt, Germany) was used throughout the experiments. Spectra/Por® 3 Float-A-Lyzer®, MWCO: 3500, Biotech Regenerated Cellulose Membranes were sourced from Spectrum Laboratories Inc. Apart from Seq. Grade Modified Trypsin (Promega, V511A) the following reagents and solvents were purchased from Sigma-Aldrich and were used as received: 2-aminopyrazine (AP); 3-(acrylamidopropyl)trimethyl-ammonium chloride (APTA); acetonitrile (ACN); ammonium bicarbonate; barium carbonate; barium hydroxide; 2,5-dihydroxybenzoic acid (DHB); formic acid (FA); guanidine hydrochloride (Gdn–HCl); HCl; lysozyme from hen egg-white; N-methylmaleimide; trans-3,5-dimethoxy-4-hydroxycinnamic acid–sinapinic acid (SA); solid carbon dioxide, trifluoroacetic acid (TFA); Tris; tris-carboxymethyl phosphine (TCEP); urea. Sodium thiophosphate tribasic hydrate (Na3PO3S∙12 H2O) was a gift from Professor Andrzej Okruszek, Institute of Technical Biochemistry, Lodz University of Technology, Poland.
Analytical methods
MALDI-TOF MS
MALDI-TOF mass spectra were recorded with an UltrafleXtreme mass spectrometer (Bruker) in the positive mode; laser frequency: 2000 Hz, voltage source 1: 24.80 kV, voltage source 2: 22.95 kV
Sample type: proteins (lyophilizate); double layer, ground steel targets.
Matrix solution A: a saturated solution of SA in ethanol; matrix solution B: a saturated solution of SA in (acetonitrile: 0.1% TFA in water [30:70 v/v]). Sample preparation: deposit 0.5 μL of matrix solution A onto the MALDI target and allow to dry; mix one part of matrix solution B with one part of analyte solution in 0.1% TFA; deposit 0.5 μL of matrix/analyte mixture onto a matrix spot and allow to dry.
Sample type: peptides (lyophilizate); dried droplet, ground steel targets; matrix DHB-AP (2:1 v/v)
Matrix solution: DHB-AP 10 mg/mL in the ratio 2:1 in 50% acetonitrile/0.1% TFA in water. Sample solvent: water of pH 7; 0.2% TFA in water added just before analysis to make a final concentration of 0.1% TFA. Sample preparation: one part of the analyte solution was mixed with one part of the matrix solution. 0.5 µL of the matrix/analyte mixture was transferred onto the target plate and left to dry.
LC ESI-MS/MS
Mass spectrometric analysis was performed on an LTQ Orbitrap Velos (Thermo Fisher Scientific, Bergen). The peptide mixture was loaded on an RP-18 pre-column (nano-ACQUITY Symmetry C18; Waters no. 186003514) using 0.1% aqueous TFA as the mobile phase and then transferred to a nano-HPLC RP-18 column (nanoACQUITY BEH C18; Waters no. 186003545) using an acetonitrile gradient (0–60% acetonitrile in 120 min) in the presence of 0.05% FA with a flow rate of 150 nL/min. The column outlet was directly coupled to the ion source of the spectrometer working in the regime of data-dependent mode and HCD fragmentation. The normalized collision energy was set to 30%. Dynamic exclusion was disabled. A blank run ensuring a lack of cross-contamination from previous samples preceded each analysis.
MS-data analysis: acquired raw data were processed using Byonic™ by Protein Metrics Inc, (v2.0–25) with UniProt-accession_p00698.decoys.fasta. Protein FDR was set to 1% FDR.
The presented searches used 6 ppm precursor ion tolerance, 20 ppm fragment ion tolerance, and considered tryptic peptides with a maximum of two missed cleavages. The searches allowed 14 variable modifications with a maximum number of common modifications (c) per peptide: 4, and rare modification (r) per peptide: 1, as listed below:
Qat/+ 170.1419 | C (c4) |
Dha/− 33.9877 | C (c4) |
Phosphorylation/+ 79.9663 | C (c4) |
Deamidation/+ 0.9840 | N (c3) |
Deamidation/+ 0.9840 | Q (c1) |
Carbamylation/+ 43.0058 | R (c3) |
Carbamylation/+ 43.0058 | K (c3) |
Carbamylation/+ 43.0058 | NTerm (c1) |
Oxidation/+ 15.9949 | M (c1) |
Dioxidation/+ 31.9898 | M (c1) |
Dioxidation/+ 31.9898 | W (c1) |
Trioxidation/+ 47.9847 | C (c1) |
Gln > pyro-Glu/− 17.0265 | NTerm Q (r1) |
Glu > pyro-Glu/− 18.0105 | NTerm E (r1) |
Synthetic procedures
Steps A and B: lysozyme reduction and alkylation with APTA reagent; L ~ [S-Qat]8 ~ [Side+]~COOH
Under a nitrogen atmosphere (ca. 1 mM protein solution), 150 g of lysozyme powder Lysozyme[S–S]4 was dissolved in 10 mL of denaturing buffer DB (0.1 M Tris–HCl in 8 M urea, pH 8.5) followed by the addition of 44 mg of crystalline TCEP (15 mM) to obtain pH 5.2–5.4. The mixture was incubated 1 h at 37 °C giving reduced Lysozyme[SH]8. Next, 3.4 mL of S-alkylating APTA reagent (commercial 40% water solution) was added and the pH of the solution was adjusted to 8.1 with a few crystals of Tris reagent. The mixture was incubated in the dark under a nitrogen atmosphere for 2 h at room temperature. The resulting lysozyme derivatives (L ~ [S-Qat]8 ~ [Side+]~COOH) were precipitated with isopropanol in the ratio of 5:1 (v/v) and after centrifugation (10 min., 10,000 g, rt.) the white precipitate was dissolved in 10 mL of DB buffer. The obtained product of lysozyme S-alkylation was analyzed by MALDI MS [S1].
Step C: β-elimination reaction; preparation of L ~ [Dha]x ~ [S-Qat]y ~ [Side+]~COOH
Procedure 1
Preparation of L ~ [Dha]5–7 ~ [S-Qat]1–3 ~ [Side+]~COOH: 10 mL of [S-Qat]8-derivatives of lysozyme in DB buffer were combined with 2.5 mL of 1.5% w/v Ba(OH)2 in 8 M urea to reach pH ~ 12.5. After 3 h of incubation at 50 °C the pH of the mixture dropped to 11.5. The reaction was terminated by carefully adding small pieces of solid carbon dioxide to obtain pH 7. The precipitated BaCO3 was separated by centrifugation (15 min., 10,000 g, rt.) and the supernatant (pH 8) was dialyzed (MWCO: 3500 kDa) against Milli-Q water, with several changes of water. A slightly opaque water solution of L ~ [Dha]x ~ [S-Qat]y ~ [Side+]~COOH was stored in a fridge. The approximate concentration of protein was 0.2 mM as determined spectrophotometrically at A280 using a lysozyme solution as the standard. The average degree of β-elimination x = 5–7 Dha residues per protein molecule was determined by MALDI MS [S2].
Procedure 2
Preparation of L ~ [Dha]6–8 ~ [S-Qat]0–2 ~ [Side+]~COOH:10 mL of [S-Qat]8-derivatives of lysozyme in DB buffer was combined with 2.5 mL of 1.5% Ba(OH)2 in 8 M urea to reach pH ~ 12.5. The β-elimination reaction was carried out at 50 °C with intermitted pH control. When needed, a few crystals of Ba(OH)2 were added to maintain the required pH range between 12.5 and 12.2. The reaction was terminated after 3 h of incubation and the product was processed as described above. The average degree of β-elimination x = 6–8 Dha residues per protein molecule was determined by MALDI MS [S3].
Step D: conjugate addition of thiophosphoric acid; preparation of L ~ [pCys]q ~ [Dha]p ~ [S-Qat]y ~ [Side+]~COOH
Conjugate addition of thiophosphate to L ~ [Dha]x ~ [S-Qat]y ~ [Side+]~COOH was tested under several reaction conditions: in the presence of urea (variant 1), in the presence of guanidine hydrochloride (variant 2) and in the absence of strong chaotropic agents (variant 3).
Solution A: 0.4 g of Na3PO3S ∙ 12 H2O was dissolved in 0.6 mL of 2 M HCl to obtain 1 mL of solution at pH 8.3. Solution B: 4 mL of 9 M urea in water was mixed with 1 mL 0.2 M phosphate buffer at pH 8.3 to give 5 mL of 0.04 M phosphate buffer in 7.2 M urea, pH 8.8. Solution C: water solution of L ~ [Dha]5–7 ~ [S-Qat]1–3 ~ [Side +]~COOH with an approximate concentration of 0.2 mM; reaction mixture R1: 0.5 mL of solution A + 4.0 mL of solution B + 0.5 mL of solution C = 5 mL of mixture at pH 8.4.
The reaction mixture R1 was incubated for 2 h at 50 °C (final pH 8.1) then cooled to room temperature and dialyzed (MWCO 3500) against several changes of Milli-Q water, and finally against 10 mM ammonium bicarbonate (pH 7.3). The liquid product was freeze-dried and stored at 4 °C. The approximate yield of lysozyme transformation was 40–50%, i.e., from 150 mg of lysozyme powder about 60 mg of freeze-dried S-phosphorylated lysozyme derivatives was obtained. The sample of freeze-dried L1 was subjected to MALDI MS to estimate the average degree of S-phosphorylation q = 3 ± 1 pCys residues per protein molecule [S4].
Solution A: 0.4 g of Na3PO3S·12 H2O was dissolved in 7.5 ml of 8 M Gdn–HCl to obtain 8 ml of sodium thiophosphate (pH 8.6). Solution C: water solution of L ~ [Dha]6-8 ~ [S-Qat]0-2 ~ [Side+]~COOH of approximate concentration 0.2 mM. Reaction mixture R2: 8 ml of solution A + 1 ml of solution C = 9 ml mixture in 6.66 M Gdn–HCl (pH 8.4).
The reaction mixture R2 was incubated for 2 h at 50 °C (final pH 8.1) then cooled to room temperature and dialyzed (MWCO 3500) against several changes of Milli-Q water, and finally against 10 mM ammonium bicarbonate (pH 7.3). The solution of product L2 was freeze-dried and stored at 4 °C.
Solution A: 0.4 g of Na3PO3S · 12 H2O was dissolved in 0.6 mL of 2 M HCl to obtain 1 mL of solution at pH 8.3. Solution B: 0.2 M phosphate buffer of pH 8.3. Solution C: water solution of L ~ [Dha]6–8 ~ [S-Qat]0–2 ~ [Side+]~COOH with an approximate concentration of 0.2 mM. Reaction mixture R3: 0.5 mL of solution A + 5 mL of solution B + 0.5 mL of solution C = 6 mL of mixture at pH 8.4.
The reaction mixture R3 was incubated for 2 h at 50 °C (final pH 8.1) then cooled to room temperature and purified by dialysis as described above. The efficiency of the conjugate addition of thiophosphoric acid was noticeably lower in the absence of strong chaotropic agents in the reaction medium.
Step E: preparation of the peptide model mixtures Digest 1 and Digest 2
Peptide model mixtures were prepared from 2 mg of freeze-dried L1 or L2 products combined with 20 µg of trypsin, then re-suspended in 1 mL of 100 mM NH4HCO3 (pH 8.0). After incubating at 37 °C for 22 h the solution was aliquoted (50 µL) into individual Eppendorf tubes and freeze-dried. Individual 100 µg portions of Digest 1 and Digest 2 were used for mass spectrometric analysis. Digest 2 requires a separate discussion elsewhere.
Removal of S-linked phosphorylation from L2 sample by acidic hydrolysis
To remove S-linked phosphorylation from the L2 sample, 20 µL of 0.5% water solution of TFA was added to 70 µg of lyophilized L2 mixture. The solution was incubated overnight at room temperature. The chemically de-phosphorylated polypeptides were then subjected to MALDI MS analysis to indirectly estimate the average degree of S-phosphorylation q = 5 ± 1 pCys residues per protein molecule [S5].
Results
Transformation of lysozyme into S-phosphocysteine-containing peptides (pCys-peptides)
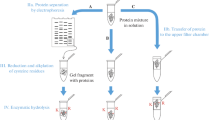
Hen egg-white lysozyme, a 129-amino acid polypeptide (14.3 kDa) containing four disulfide bridges, was selected for conversion into S-phosphorylated derivatives. The concept of transformation was inspired by an early work (Hiskey et al. 1970) reporting the partial β-elimination of benzyl thiolate from S-benzylated cysteine peptides. Thus, it was assumed that polypeptides bearing cysteine residue S-protected by other simple alkyl groups might also produce β-elimination products in amounts detectable by modern mass spectrometry. It was expected that conjugate addition of thiophosphoric acid to selectively generated Dha residues, followed by multiple proteases digestion, would create a diversified mixture of sequences with many bearing S-phosphocysteine residues. The transformation of lysozyme cysteines (Fig. 1, steps A–E) was accompanied by inherent chemical reactions involving other functional groups of the protein, in particular deamidation (Hao et al. 2011), carbamylation (Kollipara 2013), oxidations, and cysteine epimerization, as well as alkaline hydrolysis of peptide bonds, phosphorothiolate residue hydrolysis and several others. These seemingly unwelcome, additional modifications of peptide side chains [Side+] make the resulting model complex enough for testing various analytical procedures targeting pCys-peptides.
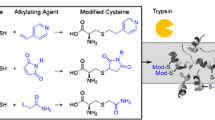
In preliminary experiments, reduced lysozyme was alkylated with several thiol-specific reagents to test the susceptibility of the resulting products to a β-elimination reaction. Ultimately, APTA was selected as the most convenient S-alkylating reagent [Electronic Supplementary Materials Fig. S1, ESM_1, S1].
β-Elimination reactions, generating Dha-residues, were carried out in two procedural variants that yielded two pools of derivatives with a lower and higher average degree of β-elimination x (i.e., x = 5–7 and x = 6–8). The resulting products were denoted, respectively, as L ~ [Dha]5–7[S-Qat]1–3[Side +]~COOH and L ~ [Dha]6–8[S-Qat]0–2[Side+]~COOH. Subsequent conjugate addition of thiophosphoric acid to Dha led to L1 and L2 collections of polypeptides substoichiometrically S-phosphorylated at epimerized cysteine residues. MALDI MS profiles revealed enormous complexity in the L1 and L2 mixtures, due to a random-like distribution of multiple modifications of the polypeptides chains [S2, S3]. The estimated average degree of polypeptide S-phosphorylation q ranged from q = 3 ± 1 pCys to q = 5 ± 1 pCys, for L1 [S4] and L2 [S5–S7] mixtures, respectively. Finally, L1 and L2 were subjected to trypsin digestion to yield Digest 1 and Digest 2 containing a complex mixture of modified peptides. Theoretically, each of the tryptic sequences incorporating a single cysteine residue could exist in six structural variants having at Cys position, respectively, S-Qat, Dha, D/L-pCys, as well as D/L-Cys originating from partial hydrolysis of D/L-pCys. For comparison, the exemplary sequence NLCNIPCSALLSSDTITASVNCAK incorporating three cysteine positions can hold as many as 63 = 216 permutations of the same six modifications. Moreover, a combinatorial deamidation of three asparagine residues raises the number of possible structures eightfold, to 1728, which does not yet comprise other probable modifications such as D-Asp, L-isoAsp, or D-isoAsp (Riggs et al. 2017). The MALDI spectra of Digest 1 show a surprisingly uncomplicated picture with one hundred peptide peaks [S8–S11], while HCD-MS/MS analyses routinely identify over 700 mono- and multiply modified unique sequences [ESM_2, S26]. The collection of peptides identified from Digest 1 poses a challenge for LC–MS/MS analysis. Using Byonic software tolerating multiple, variable modifications (Bern et al. 2012; Chalkley et al. 2014; Jiang et al. 2016), deamidation was detected for all fourteen Asn and three Gln sites, and carbamylation for all six Lys residues in the lysozyme sequence. Dha and S-Qat were spotted in all Cys positions. In addition to MS/MS spectra of pCys-peptides, many spectra of Cys-peptides were also registered; these provide evidence for hydrolytic S-dephosphorylation occurring in the course of the analysis [S13–S24]. The intact pCys-residues were identified at positions 6, 64, 94, 115, and 127, while freed Cys-residues were identified at positions 6, 30, 76, 80, 94, and 115. In summary, S-phosphorylation was substoichiometrically linked to all eight cysteine positions, as confirmed directly at five sites and revealed indirectly at six sites by detecting thiol groups.
Characteristics of HCD-type fragmentation of pCys-peptides
CID/HCD fragmentation of phosphopeptides has been the subject of extensive research, which has recently been the subject of a review article (Potel et al. 2018a). Typically, the resulting fragmentation spectra are dominated by a sequence-dependent loss of phosphoric acid H3PO4 (98 Da) or meta-phosphoric acid HPO3 (80 Da), with parallel peptide backbone fragmentation. In contrast, NL-events from pCys-peptides are poorly understood, as only a few HCD-fragmentation spectra have been published so far (Bertran-Vicente et al. 2016).
The pCys-peptides identified from Digest 1 are listed in Table 1, which shows the peptide sequence, the calculated value for the peptide pI, the charge state (z) of the peptide molecular ion (M) and the presence/absence of neutral loss ions (NL). The electronic supplementary materials contain HCD-spectra of the identified pCys-peptides [S13–S24].
As well as a significant level of backbone fragmentation, several HCD spectra of the pCys-peptides display M-80 Da (i.e., HPO3) and M-98 Da (i.e., H3PO4 or combine HPO3 + H2O) neutral losses (NL) from the molecular ion (M) and weak M-116 Da loss (i.e., H3PO4 + H2O). In the HCD spectra of the + 2 charged peptides, the M-80 ions are strong (up to 100%) and more intense than the M-98 ions, with some exceptions which will be discussed later. Phosphate neutral losses are also evident for peptides in the + 3 charge state, although these NL-signals are not dominant. Often, the major NL-triplet is accompanied by additional NL-triplet(s) with lower intensities. These additional triplets occur in the spectra of + 2 and + 3 charged peptides, shifted up by 0.492 Da and 0.328 Da, respectively. Whenever confirmed by the respective y- and/or b-fragment ions, the accompanying triplets were attributed to the co-fragmenting S-phosphorylated sequences carrying additional deamidations.
Intriguing differences were noticed in the case of fragmentation of the DRpCKGTDVQAWIR peptide in the mass spectra registered at 38.25 min, 38.81 and 40.12 min. In the spectrum scanned at 40.12 min, the intensity ratio of the two main NL-ions was as follows [M-80]INT > [M-98]INT (Fig. 2a). In the spectrum scanned at 38.25 min, this proportion was reversed, i.e., [M-80]INT < [M-98]INT (Fig. 2b). Notably, the intensities of both NL-signals were almost equal [M-80]INT ≈ [M-98]INT in the spectrum acquired at scan time 38.81 min [S17]. This observation may indicate the presence of phosphocysteine D/L-epimers of the peptide. The absolute configuration of a phosphorothiomethyl group affects interactions between the phosphate residue and peptide side chains. It is plausible that the conformation-dependent properties of D/L-epimers may be manifested by differences in their LC-retention times, as well as by clear differences in their patterns of phosphate neutral loss. At the same time, both epimers displayed almost identical MS backbone fragmentation, which correlates with the MS/MS-fragmentation feature described for other epimeric peptides (Serafin et al. 2005; Tao and Julian 2014) (Fig. 2). Presumably, the phosphoryl transfer (Oslund et al. 2014; Potel et al. 2018b) within L-pCys-peptide conformers to their carboxyl groups is not energetically equivalent to a similar transfer in the conformers of the respective D-pCys-epimer. This energetic inequivalence of the phosphate transfers may explain the observed changes in the relative intensity of the [M-98] ion. Clarification of the phenomenon would require experiments with the use of pure D and L-epimers of S-phosphopeptides displaying the NL-feature, yet such structures are not easily obtainable (Bertran-Vicente et al. 2016). Furthermore, common sequence-informative y- and/or b-ions may coexist with many dephosphorylated fragment ions, i.e., y-80, y-98, b-80, and b-98. Losses of 80 Da and/or 98 Da were recorded from the doubly charged x-, y-, b- as well as (b n–1 + H2O) fragment ions. The formation of a M-98 ion by a phosphoryl transfer mechanism (Cui et al. 2014) correlates with the neutral gain fragment ions y + 80 registered in several spectra of S-phosphopeptides [S14, S15, S17–19]. The phenomenon of intramolecular phosphoryl transfer during gas-phase collision-induced dissociation (CID) had previously been observed for O- and N-phosphopeptides and was interpreted as lowering confidence in phospho-site localization (Cui and Reid 2013; Schmidt et al. 2013).
According to the classification proposed by Everley, all of the protonated pCys-peptides shown in Table 1 belong to a low proton mobility class of peptides, as their charge state z is equal to or lower than the sum of basic centers: (z ≤ Lys + Arg + His + 1) (Everley et al. 2017). For this set of pCys-peptides, an interesting correlation was found between the calculated pI value (Pirmoradian et al. 2014; Kozlowski 2016) of a plain peptide sequence and its NL-attribute. The pI criterion was used to test the set of low proton mobility peptides and revealed that NL-signals appeared in the HCD-spectra of the basic peptides incorporating the Arg-pCys unit but not in the spectra of acidic peptides which lacked this structural element (Table 1). The acidic peptide NLC[S-Qat]NIPpCSALLSSDITASVNC[S-Qat]AK and its deamidated variants were interesting exceptions. Although lacking Arg-pCys motifs, all of them displayed NL-triplets from + 3 charged molecular ions. It is likely that two incorporated residues of S-Qat formed a sequence environment of limited charge mobility, thereby promoting the neutral loss of phosphate from a stabilized molecular ion [S21]. These results cannot be generalized, due to the limited number of examined pCys-peptides.
Triplet-like phosphate losses, generated by HCD-fragmentation from molecular ions of pCys-peptides, are similar to the neutral-loss triplets (or missing M-116 signal NL-doublets) observed for many phosphohistidine peptides (Oslund et al. 2014; Potel et al. 2018a). Earlier, Hardman et al. had thoroughly evaluated identification of pHis-peptides by the triplet-like loss pattern (Hardman et al. 2017) and concluded that this trait was not a general hallmark of histidine phosphorylation.
The results show that, besides S-phosphorylated peptides, numerous Cys-containing peptides were identified from Digest 1 [S26]. Generated in the course of acidic LC-separation, Cys-peptides do not experience oxidation and are detectable by mass spectrometry. It is important to mention that proteomic samples may contain traces of thiol groups that emerged from various precursors. In general, the presence of free thiols is mainly associated with incomplete alkylation of thiol groups from reducible sulfhydryl modifications (Dong et al. 2014; Muller and Winter 2017; Suttapitugsakul et al. 2017). Moreover, even traces of S-acylated peptides can generate Cys-peptides as a result of acyl transfer events (Ji et al. 2013, 2016).
Discussion
Recent studies aimed at systematic large-scale detection of six acid-labile protein phosphorylation in human cell extracts have exposed the difficulty of confident localization of phosphate groups on Arg, His, Lys, Asp, Glu, and Cys residues. Although phosphorothiol ester linkage is moderately acid-stable and highly base-stable, protein cysteine phosphorylation was not detected in these studies (Hardman et al. 2017). The lack of basic information on the prevalence, properties, and behavior of this modification under analytical circumstances explains this lack of success.
This paper aims to provide new information that may help detect protein phosphocysteine in different proteomes. A new approach, imitating a proteomic workflow, was designed to obtain S-phosphorylated peptides from a cysteine-containing protein of a known sequence. The novelty of the method (Fig. 1A–E) is based on the incorporation into a sample preparation procedure of two additional, consecutive steps: the unusual β-elimination of alkyl thiolate from cysteine positions (Fig. 1, step C) and conjugate addition of thiophosphoric acid to the generated Dha residues (Fig. 1, step D). Subsequent standard proteolysis predictably converts the protein derivatives into a mixture of modified, MS-amenable peptides (Digest 1). Hence, the proposed approach contrasts with alternative de novo synthesis of pCys-peptides (Bertran-Vicente et al. 2016), in that the pCys-peptides are not synthesized sensu stricto by consecutive peptide bond formation.
Digest 1 exhibits enormous complexity due to the presence of numerous variable modifications in Cys positions and other peptide amino acids residues. Byonic software was applied to search for five modification types per a single peptide at a time. Searches were performed against the protein reference database being restricted to a single lysozyme sequence. Therefore, even poorly fragmented peptides were confidently identified, as generated fragment ions can be determined with high accuracy. However, for longer and poorly fragmented peptides it was not always possible to locate multiple modifications unambiguously. More detailed analysis of the model Digest 1 would certainly be possible, by additional HPLC-runs and alternative MS-fragmentation strategies (Dong et al. 2014).
Direct detection of S-linked phosphorylation
Several articles referring to pCys-containing proteins (Sun et al. 2012; Chooi et al. 2014; Bertran-Vicente et al. 2016; Gulerez et al. 2016; Zhang et al. 2017) have reported evidence of S-linked phosphorylation survival throughout various proteomic workflows. The HCD spectra of the model mixture assembled in this study demonstrate that direct detection of pCys-peptides is feasible, especially if their spectra exhibit NL-triplets, 80 Da losses from fragment ions, or ideally fragment ions carrying the pCys-residue. The protonated pCys peptides were found to produce triplet-like NL-pattern if their plain sequences manifested basicity, while those with the respective acidic sequences did not display this feature. Although this observation cannot be generalized, such interpretations may help detect S-phosphorylation of natural origins.
In real-world proteomics, cysteine residues of proteins are usually S-carbamidomethylated (Cys→camCys) (Muller and Winter 2017; Suttapitugsakul et al. 2017; Svozil and Baerenfaller 2017) and considered to have been permanently modified. Therefore, phosphocysteine targeting re-analysis of mass spectra should apply a non-routine bioinformatic approach, as MS/MS search settings affect the outcome of intact pCys-peptide identification. Allowing for variable camCys→pCys substitution, i.e., by setting [+ 22.94486 Da] on camCys, may enable detection of phosphocysteine residues. The [+ 22.94486 Da] mass gain is unique and distinguishable from a potential two-point modification, i.e., sodiation [+ 21.98195 Da] and deamidation [+ 0.98402 Da], which collectively yield a total mass gain of [+ 22.96597 Da]. This search variant is analogous to Bern’s “small trick” which allows variable de-carbamidomethylation [− 57.02146 Da] on permanently modified camCys-peptides (Bern 2014).
Such direct detection of S-linked phosphorylation seems suitable for less complex analytes, such as in the case of off-line fractionation (Ruprecht et al. 2017) or affinity enrichment procedures. In preliminary experiments, S-phosphopeptides from a sample of Digest 1 were enriched using a hydrophilic Ti-IMAC (He et al. 2015) prepared in-house from cotton–wool. The enriched analyte was subjected to a series of tandem MS applying complementary fragmentation methods CID, HCD, ETD, ETciD, and EThcD [S12]. In all, 12 additional pCys-peptides as well as numerous cysteine-containing peptides, were identified (data not shown). It is worth noting that recent studies of Gehrig’s group have demonstrated the utility of Phos-tag gels for the electrophoretic separation of S-phosphorylated PRLs and their non-phosphorylated counterparts (Gulerez et al. 2016; Zhang et al. 2017). Currently, Phos-tag affinity material, being a type of Zn-IMAC, is also available in micropipette-tip format (Yuan et al. 2017), which is suitable for testing phosphopeptide co-enrichment procedures.
Detection of S-linked phosphorylation by cognate Cys-peptides
The second approach to detecting S-phosphorylation is complementary to the first, and targets those pCys-peptides that went undetected in the first analytical run due to low concentrations, poor fragmentation, or limitations in the automated analysis of MS data. To some extent, this approach is comparable to the detection strategies of protein S-nitrosylation (Ju et al. 2015) and S-acylation (Ji et al. 2013, 2016) which exploit a specific split in these modifications, leading to detectable thiol groups. Overnight incubation of the thiol-deprived analyte at a low pH results in a total loss of acid-labile phosphorylations, including those that are S-linked. Therefore, any reappearance of thiol groups may be attributed to hydrolytically released Cys-peptides, which can be effectively extracted on thiol-affinity materials (Gu et al. 2015; Duan et al. 2017).
A library of HCD-spectra was compiled from the many tens of individual Cys-peptides [S26] identified from the Digest 1 analyte. Significant similarity was noted between the HCD-fragmentation features of the exemplary spectral pair DRpCKGTDVQAWIR and DRCKGTDVQAWIR [SI, Fig. S28], which clearly suggests that the spectrum of a non-phosphorylated peptide can help to predict the fragmentation pattern for the cognate pCys-peptide. Plausibly, a predictive spectra library of pCys-peptides could be constructed using an experimental spectral library of Cys-peptides as the reference. Applying the predictive library and appropriate spectrum-to-spectrum matching search engines (Hu and Lam 2013; Degroeve et al. 2015; Suni et al. 2015; Wang et al. 2015; Griss 2016; Shao and Lam 2017) could help to detect S-phosphorylated peptides which survived the proteomic workflow.
Conclusions
This paper has presented a procedure for transforming a single, disulfide-bridged protein with a known sequence into a complex mixture comprising modified, phosphocysteine-containing peptides. The resulting pCys-peptides could serve as standards for mass spectrometry. Typical LC–ESI-MS/MS analysis of the model mixture succeeded in identifying S-phosphorylated peptides, as well as numerous Cys-peptides that had lost the phosphate group during the workflow. Some features of the HCD-fragmentation spectra of the low proton mobility pCys-peptides were revealed. They exhibited strong M-80 and M-98 neutral loss ions, and weak M-116 neutral loss ions, if these were basic in the dephosphorylated form. They did not exhibit such losses if they were acidic. The identified Cys-peptides and their pCys-precursors exhibited similar backbone HCD-fragmentation patterns. Therefore, a spectral library of the Cys-peptides could be useful when searching in MS/MS data for intact S-phosphorylation using the spectra-matching approach.
References
Attwood PV, Besant PG, Piggott MJ (2011) Focus on phosphoaspartate and phosphoglutamate. Amino Acids 40:1035–1051. https://doi.org/10.1007/s00726-010-0738-5
Bern M (2014) Byonic™ : Focused protein databases. In: Appl. Note. https://www.proteinmetrics.com/wp-content/uploads/2014/10/AppNote-ByonicFocusedDatabase.pdf
Bern M, Kil Y, Becker C (2012) Byonic: advanced peptide and protein identification software. Curr Protoc Bioinform. https://doi.org/10.1002/0471250953.bi1320s40
Bertran-Vicente J, Penkert M, Nieto-Garcia O et al (2016) Chemoselective synthesis and analysis of naturally occurring phosphorylated cysteine peptides. Nat Commun 7:1–9. https://doi.org/10.1038/ncomms12703
Buchowiecka AK (2014) Puzzling over protein cysteine phosphorylation-assessment of proteomic tools for S-phosphorylation profiling. Analyst 139:4118–4123. https://doi.org/10.1039/c4an00724g
Carrillo JB, Gomez-Casati DF, Martín M, Busi MV (2018) Identification and analysis of OsttaDSP, a phosphoglucan phosphatase from Ostreococcus tauri. PLoS One 13:1–18. https://doi.org/10.1371/journal.pone.0191621
Chalker JM, Lercher L, Rose NR et al (2012) Conversion of cysteine into dehydroalanine enables access to synthetic histones bearing diverse post-translational modifications. Angew Chemie Int Ed 51:1835–1839. https://doi.org/10.1002/anie.201106432
Chalkley RJ, Bandeira N, Chambers MC et al (2014) Proteome Informatics Research Group (iPRG)_2012: a study on detecting modified peptides in a complex mixture. Mol Cell Proteomics. https://doi.org/10.1074/mcp.M113.032813
Chooi PK, Galan SR, Raj R et al (2014) Synthetic phosphorylation of p38 α recapitulates protein kinase activity. J Am Chem Soc 136:1698–1701. https://doi.org/10.1021/ja4095318
Cui L, Reid GE (2013) Examining factors that influence erroneous phosphorylation site localization via competing fragmentation and rearrangement reactions during ion trap CID-MS/MS and -MS 3. Proteomics 13:964–973. https://doi.org/10.1002/pmic.201200384
Cui L, Yapici I, Borhan B, Reid GE (2014) Quantification of competing H3PO4 versus HPO3 + H2O neutral losses from regioselective 18O-Labeled Phosphopeptides. J Am Soc Mass Spectrom 25:141–148. https://doi.org/10.1007/s13361-013-0744-4
Degroeve S, Maddelein D, Martens L (2015) MS 2 PIP prediction server: compute and visualize MS 2 peak intensity predictions for CID and HCD fragmentation. Nucleic Acids Res 43:W326–W330. https://doi.org/10.1093/nar/gkv542
Deutsch EW, Perez-Riverol Y, Chalkley RJ, Wilhelm M, Tate S, Sachsenberg T, Walzer M, Käll L, Delanghe B, Böcker S, Schymanski EL, Wilmes P, Dorfer V, Kuster B, Volders P-J, Jehmlich N, Vissers JPC, Wolan DW, Wang AY, Mendoza L, Shofstahl J, Dowsey AW, Griss J, Salek RM, Neumann S, Binz P-A, Lam H, Vizcaíno JA, Bandeira N, Röst H (2019) Expanding the use of spectral libraries in proteomics. J Proteome Res 17(12):4051–4060. https://doi.org/10.1021/acs.jproteome.8b00485
Dong Q, Yan X, Kilpatrick LE et al (2014) Tandem mass spectral libraries of peptides in digests of individual proteins: Human Serum Albumin (HSA). Mol Cell Proteomics. https://doi.org/10.1074/mcp.O113.037135
Dong H, Zonta F, Wang S et al (2018) Structure and molecular dynamics simulations of protein tyrosine phosphatase non-receptor 12 provide insights into the catalytic mechanism of the enzyme. Int J Mol Sci 19:60. https://doi.org/10.3390/ijms19010060
Duan G, Walther D (2015) The roles of post-translational modifications in the context of protein interaction networks. PLoS Comput Biol 11:1–23. https://doi.org/10.1371/journal.pcbi.1004049
Duan J, Gaffrey MJ, Qian W-J (2017) Quantitative proteomic characterization of redox-dependent post-translational modifications on protein cysteines. Mol BioSyst 13:816–829. https://doi.org/10.1039/c6mb00861e
Everley RA, Huttlin EL, Erickson AR et al (2017) Neutral loss is a very common occurrence in phosphotyrosine—containing peptides labeled with isobaric tags. J Proteome Res 16:1069–1076. https://doi.org/10.1021/acs.jproteome.6b00487
Griss J (2016) Spectral library searching in proteomics. Proteomics 16:729–740. https://doi.org/10.1002/pmic.201500296
Gu L, Evans AR, Robinson RAS (2015) Sample multiplexing with cysteine-selective approaches: cysDML and cPILOT. J Am Soc Mass Spectrom 26:615–630. https://doi.org/10.1007/s13361-014-1059-9
Gulerez I, Funato Y, Wu H et al (2016) Phosphocysteine in the PRL-CNNM pathway mediates magnesium homeostasis. EMBO Rep 17:1890–1900
Hao P, Ren Y, Alpert AJ, Sze SK (2011) Detection, evaluation and minimization of nonenzymatic deamidation in proteomic sample preparation. Mol Cell Proteomics. https://doi.org/10.1074/mcp.O111.009381
Hardman G, Perkins S, Ruan Z et al (2017) Extensive non-canonical phosphorylation in human cells revealed using strong-anion exchange-mediated phosphoproteomics. bioRxiv. https://doi.org/10.1101/202820
He X, Chen X, Zhu G et al (2015) Hydrophilic carboxyl cotton chelator for titanium (IV) immobilization and its application as novel fibrous sorbent for rapid enrichment of phosphopeptides. ACS Appl Mater Interfaces 7:17356–17362. https://doi.org/10.1021/acsami.5b04572
Hiskey RG, Upham RA, Beverly GM, Jones WC (1970) Sulfur-containing polypeptides. X. A study of ß elimination of mercaptides from cysteine peptides. J Org Chem 35:2–4. https://doi.org/10.1021/jo00827a053
Hu Y, Lam H (2013) Expanding tandem mass spectral libraries of phosphorylated peptides: advances and applications. J Proteome Res 12:5971–5977. https://doi.org/10.1021/pr4007443
Ji Y, Leymarie N, Haeussler DJ et al (2013) Direct detection of S-palmitoylation by mass spectrometry. Anal Chem 85:11952–11959. https://doi.org/10.1021/ac402850s
Ji Y, Bachschmid MM, Costello CE, Lin C (2016) S- to N-Palmitoyl transfer during proteomic sample preparation. J Am Soc Mass Spectrom 27:677–685. https://doi.org/10.1007/s13361-015-1319-3
Jiang X, Horn D, Bomgarden R, et al (2016) Evaluation of search engines for phosphopeptide identification and quantitation. In: Poster Note 64793. 66TH ASMS CONFERENCE ON MASS SPECTROMETRY, San Diego
Ju Y, Fu M, Wu L, Yang G (2015) Strategies and tools for detection of protein S-Nitrosylation and S-Sulfhydration. Biochem Anal Biochem 4:1–7. https://doi.org/10.4172/2161-1009.1000224
Junker S, Maaß S, Otto A, Michalik S, Morgenroth F, Gerth U, Hecker M, Becher D (2018) Spectral library based analysis of arginine phosphorylations in staphylococcus aureus. Mol Cell Proteomics 17(2):335–348. https://doi.org/10.1074/mcp.RA117.000378
Kollipara L (2013) Protein carbamylation: in vivo modification or in vitro artefact? Proteomics 13:941–944. https://doi.org/10.1002/pmic.201200452
Kozlowski LP (2016) IPC—isoelectric point calculator. Biol Direct. https://doi.org/10.1186/s13062-016-0159-9
Lai S-J, Tu I-F, Wu W-L et al (2017) Site-specific His/Asp phosphoproteomic analysis of prokaryotes reveals putative targets for drug resistance. BMC Microbiol 17:1–10. https://doi.org/10.1186/s12866-017-1034-2
Marmelstein AM, Moreno J, Fiedler D (2017) Chemical approaches to studying labile amino acid phosphorylation. Top Curr Chem 375:1–32. https://doi.org/10.1007/s41061-017-0111-1
Marx H, Lemeer S, Schliep JE et al (2013) A large synthetic peptide and phosphopeptide reference library for mass spectrometry–based proteomics. Nat Biotechnol 31:557–564. https://doi.org/10.1038/nbt.2585
Morrison PM, Foley PJ, Warriner SL, Webb ME (2015) Chemical generation and modification of peptides containing multiple dehydroalanines. Chem Commun 51:13470–13473. https://doi.org/10.1039/c5cc05469a
Muller T, Winter D (2017) Systematic evaluation of protein reduction and alkylation reveals massive unspecific side effects by iodine-containing reagents. Mol Cell Proteomics 16(7):1173–1187
Oslund RC, Kee J-M, Couvillon AD et al (2014) A phosphohistidine proteomics strategy based on elucidation of a unique gas-phase phosphopeptide fragmentation mechanism. J Am Chem Soc 136:12899–12911. https://doi.org/10.1021/ja507614f
Piggott MJ, Attwood PV (2017) Focus on O-phosphohydroxylysine, O-phosphohydroxyproline, N1-phosphotryptophan and S-phosphocysteine. Amino Acids 49:1309–1323. https://doi.org/10.1007/s00726-017-2446-x
Pirmoradian M, Zhang B, Chingin K et al (2014) Membrane-assisted isoelectric focusing device as a micropreparative fractionator for two-dimensional shotgun proteomics. Anal Chem 86:5728–5732. https://doi.org/10.1021/ac404180e
Potel CM, Lemeer S, Orcid AJRH (2018a) Phosphopeptide fragmentation and site localization by mass spectrometry: an update. Anal Chem 91:126–141. https://doi.org/10.1021/acs.analchem.8b04746
Potel CM, Lin M-H, Heck AJ, Lemeer S (2018b) Widespread bacterial protein histidine phosphorylation revealed by mass spectrometry-based proteomics. Nat Methods 15:180–197. https://doi.org/10.1038/nmeth.4580
Riggs DL, Gomez SV, Julian RR (2017) Sequence and solution effects on the prevalence of D-Isomers produced by deamidation. ACS Chem Biol 12:2875–2882. https://doi.org/10.1021/acschembio.7b00686
Rubio T, Köhn M (2016) Regulatory mechanisms of phosphatase of regenerating liver (PRL)-3. Biochem Soc Trans 44:1305–1312. https://doi.org/10.1042/BST20160146
Ruprecht B, Zecha J, Zolg DPKB (2017) Methods in molecular biology. In: Comai L, Katz JMP (eds) Proteomics, vol 1550. Humana Press, New York, pp 83–98
Schmidt A, Ammerer G, Mechtler K (2013) Studying the fragmentation behavior of peptides with arginine phosphorylation and its influence on phospho-site localization. Proteomics 13:945–954. https://doi.org/10.1002/pmic.201200240
Serafin SV, Maranan R, Zhang K, Morton TH (2005) Mass spectrometric differentiation of linear peptides composed of L-amino acids from isomers containing one D-amino acid residue. Anal Chem 77:5480–5487. https://doi.org/10.1021/ac050490j
Shannon DA, Weerapana E (2013) Invited Review Orphan PTMs: rare, Yet Functionally Important Modifications of Cysteine. Biopolymers 101:156–164. https://doi.org/10.1002/bip.22252
Shao W, Lam H (2017) Tandem mass spectral libraries of peptides and their roles in proteomics research. Mass Spectrom Rev 36:634–648. https://doi.org/10.1002/mas
Sun F, Ding Y, Ji Q et al (2012) Protein cysteine phosphorylation of SarA/MgrA family transcriptional regulators mediates bacterial virulence and antibiotic resistance. PNAS 109:15461–15466. https://doi.org/10.1073/pnas.1205952109
Suni V, Imanishi SY, Maiolica A et al (2015) Confident site localization using a simulated phosphopeptide spectral library. J Proteome Res 14:2348–2359. https://doi.org/10.1021/acs.jproteome.5b00050
Suttapitugsakul S, Xiao H, Smeekens J, Wu R (2017) Molecular biosystems evaluation and optimization of reduction and alkylation methods to maximize peptide identification with MS-based proteomics. Mol BioSyst 13:2574–2582. https://doi.org/10.1039/c7mb00393e
Svozil J, Baerenfaller K (2017) A cautionary tale on the inclusion of variable posttranslational modifications in database-dependent searches of mass spectrometry data. In: Shukla AK (ed) Methods in enzymology. Academic Press Inc., Cambridge, pp 433–452
Tao Y, Julian RR (2014) Identification of amino acid epimerization and isomerization in crystallin proteins by tandem LC–MS. Anal Chem 86:9733–9741. https://doi.org/10.1021/ac502296c
Tonks NK (2014) Protein tyrosine phosphatases: from housekeeping enzymes to master-regulators of signal transduction. FEBS J 2(280):346–378. https://doi.org/10.1111/febs.12077
Wang Y, Yang F, Wu P et al (2015) OpenMS-Simulator: an open-source software for theoretical tandem mass spectrum prediction. BMC Bioinform 16:1–6. https://doi.org/10.1186/s12859-015-0540-1
Yu Z-H, Zhang Z-Y (2018) Regulatory mechanisms and novel therapeutic targeting strategies for protein tyrosine phosphatases. Chem Rev. https://doi.org/10.1021/acs.chemrev.7b00105
Yuan ET, Ino Y, Kawaguchi M et al (2017) A Phos-tag-based micropipette-tip method for rapid and selective enrichment of phosphopeptides. Electrophoresis 38:2447–2455. https://doi.org/10.1002/elps.201700175
Zhang H, Kozlov G, Li X et al (2017) PRL3 phosphatase active site is required for binding the putative magnesium transporter CNNM3. Sci Rep 7:1–9. https://doi.org/10.1038/s41598-017-00147-2
Acknowledgements
The author thanks Dr. Janusz Dębski, Laboratory of Mass Spectrometry, Institute of Biochemistry and Biophysics, Polish Academy of Science, Warsaw, for MS measurements and initial Mascot searches. Sincere appreciation is also given to Dr. Marshall Bern, Protein Metrics, for helpful discussions and access to Byonic software.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The author declares no conflict of interest.
Research involving human participants and/or animals
None.
Additional information
Handling Editor: J. Marshal.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Buchowiecka, A.K. Modified cysteine S-phosphopeptide standards for mass spectrometry-based proteomics. Amino Acids 51, 1365–1375 (2019). https://doi.org/10.1007/s00726-019-02773-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00726-019-02773-8