Abstract
Site-directed spin labeling electron paramagnetic resonance (SDSL-EPR) is an established tool for exploring protein structure and dynamics. Although nitroxide side chains attached to a single cysteine via a disulfide linkage are commonly employed in SDSL-EPR, their internal flexibility complicates applications to monitor slow internal motions in proteins and to structure determination by distance mapping. Moreover, the labile disulfide linkage prohibits the use of reducing agents often needed for protein stability. To enable the application of SDSL-EPR to the measurement of slow internal dynamics, new spin labels with hindered internal motion are desired. Here, we introduce a highly ordered nitroxide side chain, designated R9, attached at a single cysteine residue via a non-reducible thioether linkage. The reaction to introduce R9 is highly selective for solvent-exposed cysteine residues. Structures of R9 at two helical sites in T4 Lysozyme were determined by X-ray crystallography and the mobility in helical sequences was characterized by EPR spectral lineshape analysis, Saturation Transfer EPR, and Saturation Recovery EPR. In addition, interspin distance measurements between pairs of R9 residues are reported. Collectively, all data indicate that R9 will be useful for monitoring slow internal structural fluctuations, and applications to distance mapping via dipolar spectroscopy and relaxation enhancement methods are anticipated.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Methanethiosulfonate nitroxides are extensively used as reagents for spin labeling cysteines in proteins due to their high reactivity and specificity [1,2,3]. The most popular reagent (MTSSL) generates a disulfide-linked side chain designated R1 (Fig. 1a). The high reactivity of the methanethiosulfonate functionality allows facile derivatization of partially and even fully buried cysteine residues [3, 4]. The conformation and internal dynamics of R1 in proteins have been extensively characterized by crystallography [5,6,7,8,9], EPR spectral simulations [4, 10, 11], and density functional theory computations [12]. An important feature of R1 that has emerged from these studies is the interaction of the Sδ sulfur of the disulfide linkage with the Cα–H hydrogen of the residue in helical structures (Fig. 1a). This interaction constrains the conformational space and gives rise to an anisotropic motion on the ns time scale described by torsional oscillations about the χ4/χ5 dihedral angles [4, 8]. The anisotropic motion averages the magnetic tensor elements of the nitroxide in such a manner as to make the continuous-wave electron paramagnetic resonance (CW-EPR) spectral lineshape exquisitely sensitive to fast (ns) structural fluctuations, and to modulation of the side chain motion by local interactions. Thus, R1 has become a sensor of choice for monitoring fast backbone fluctuations and mapping secondary and tertiary structures of the protein [11, 13,14,15].
Structures of spin labeling reagents and reaction products with cysteine residues on proteins involved in this study, among which the reagent HO-4072/the side chain R9 is the focus. a Structure of the reagent 1-oxyl-2,2,5,5-tetramethypyrroline-3-methyl (MTSSL, HO-225), the R1 side chain with designations of atoms and dihedral angles used in the text, and the molecular model (from PDB code 2CUU). The Cα-H···Sδ interaction that restricts the motion of the disulfide linkage is shown. b Structure of the reagent 3,4-bis-(methanethiosulfonylmethyl)-2,2,5,5-tetramethyl-2,5-dihydro-1H-pyrrol-1-yloxy radical (HO-1944), the RX side chain, and the molecular model (from PDB code 5LWO). c Structure of the reagent bis (2,2,5,5-tetramethyl-3-imidazoline-1-oxyl-4-il)-disulfide (IDSL), the V1 side chain with designations of atoms and dihedral angles used in the text, and the molecular model (from PDB code 3K2R). The Cα-H···Sδ interaction and the N···Sγ interaction are shown. d Structure of the reagent methyl 4-fluoro-2,2,5,5-tetramethyl-2,5-dihydro-1H-pyrrole-3-carboxylate-1-yloxyl radical (HO-4072), the R9 side chain with designations of atoms and dihedral angles used in the text, and the molecular model (from PDB code 8TAT, see Result section of this article)
However, the internal motion of R1 limits its utility for monitoring slow protein internal motions (μs–ms), for example, using saturation transfer electron paramagnetic resonance (ST-EPR [16,17,18]) or pulsed electron–electron double resonance (ELDOR [19]) methods. Moreover, the existence of multiple rotamers [8] in addition to the internal motion leads to uncertainty in the location of the nitroxide and hence in the interpretation of interspin distance distributions determined from pulsed dipolar spectroscopy [20, 21].
To limit the conformational space and restrain the motion of the nitroxide, we have introduced two other disulfide-linked side chains. The first, designated RX [22], crosslinks pairs of suitably placed cysteine residues in α-helices and β-sheets (Fig. 1b) using a bi-functional methanethiosulfonate reagent (HO-1944, Fig. 1b). Due to constraints imposed by the two sites of attachment, the RX conformational space is highly constrained and the limited internal motion is strongly ordered [22,23,24]. Another, designated V1 [25], is introduced at a single cysteine using a disulfide-linked imidazoline nitroxide biradical (IDSL, Fig. 1c) via a thiol/disulfide exchange reaction [26, 27]. V1 has a highly restricted internal motion resulting from the same Cα-H···Sδ interaction as in R1, but in addition a unique N···Sγ intra-side chain interaction [25, 28] (Fig. 1c). The strong ordering and localization make RX and V1 attractive for use in distance measurements [22, 28] and for slow motion detection [22, 29,30,31].
Despite the positive features of these disulfide-linked spin label side chains, there are some disadvantages. First, the disulfide is readily cleaved by reducing agents that are necessary for the stability and activity of some proteins. In the case of V1, the reactive disulfide is relatively unstable and susceptible to spontaneous hydrolytic cleavage in aqueous solution. Moreover, the high reactivity of methanethiosulfonate and IDSL reagents to even buried cysteine residues requires the removal of at least some native cysteines [3, 25]. RX necessitates the introduction of two cysteine residues, and the cross-link can stabilize strained, non-native structures.
In this study, we introduce a new nitroxide reagent (HO-4072) that reacts exclusively with solvent-exposed cysteine residues (Fig. 1d). The nitroxide side chain formed by the reaction, designated R9, has a stable thioether linkage that tolerates the commonly used reducing reagents and has a short linker that positions the nitroxide close to the protein backbone. Crystal structures are presented that identify unique interactions that lead to a generally high ordering of R9.
2 Results
The R9 side chain has been introduced at solvent-exposed sites in T4 Lysozyme (T4L) carrying cysteine mutations in the pseudo wild-type background [32,33,34]. In the sections below, we characterize the reaction mechanism and rate of HO-4072 with cysteine residues, explore the internal dynamics of the R9 side chain using various EPR methods, and present X-ray crystal structures of R9 on helical sites and identify the structural origin of the strongly ordered motion. Applications to determine inter-residue distance with double electron–electron resonance (DEER), and to monitor slow protein dynamics with ST-EPR are presented. Spin–lattice relaxation times in T4L are measured via Saturation Recovery (SR-EPR [29, 35, 36]) to evaluate the potential of R9 for measuring distances via relaxation enhancement.
2.1 Reaction Mechanism and Kinetics
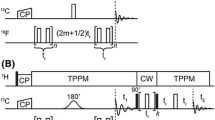
The reaction of HO-4072 with cysteine is anticipated to proceed via a Michael addition followed by E1 elimination (Fig. 2a). The Michael addition should proceed most rapidly in the alkaline solution with RS− as the nucleophile. Reaction progress is readily followed by the decrease in sharp spectral component corresponding to unreacted HO-4072 as it reacts under pseudo-first-order conditions with a large excess of protein (Fig. 2b; see Materials and Methods). Figure 2c shows the pseudo-first order rate constant determined in this manner as a function of pH for T4L 72C. The apparent pK determined by fitting is 8.35 ± 0.2, a reasonable value for the pKa of cysteine in a protein [37].
Characterization of the reaction of HO-4072 with cysteine. a Proposed spin-label reaction mechanism with HO-4072 to generate the R9 side chain. b The disappearance of the CW-EPR signal (I(t)/I(0), see Materials and Methods) for unreacted HO-4072 as a function of time in the presence of a limiting amount of HO-4072. This example is recorded at pH 8.41 in solution. Under these conditions, the reaction is pseudo-first order. The trace (red) fit to the data points (blue) is a monoexponential with rate constant \(k^{\prime}_{1}\) = 1.552 (± 0.01) × 10–4 s−1. c Plot of the reaction rate constant, determined as for (b), vs pH. The fit to the data is a Henderson–Hasselbalch equation and yields a pKa for cysteine of 8.35 ± 0.2. d Deconvoluted ESI mass spectra before and after the spin-label reaction between HO-4072 and T4L mutant with a single solvent-exposed cysteine
However, these data do not identify reaction product(s), and it could in principle be the intermediate preceding the elimination of the HF molecule (Fig. 2a). Moreover, the 4-ester could be partially or fully hydrolyzed at pH ≥ 8 in which the reaction is favorably carried out. Thus, a mixture of products is possible. However, mass spectrometry indicates a major species with a mass increase of 196 ± 1 Da corresponding to the addition of the R9 side chain (Fig. 2d), which is readily distinguishable from the intermediate (a theoretical 216 ± 1 Da increase). Moreover, the mass spectra recorded at different times during the reaction did not reveal any significant accumulation of reaction intermediates (Fig. S1), indicating that the final elimination step is rapid. In a separate experiment, it was found that ester hydrolysis of a thioether analog of R9 does not take place at a measurable rate up to a pH 8.7, T = 28 ℃, within 48 h (see details in SI Materials and Methods, Fig. S2).
The cysteine-less T4L was also incubated with the HO-4072 under the same reaction conditions. It was determined that background labeling of groups other than cysteine contributed less than 1% of the total EPR signal of the labeled cysteine mutants.
2.2 Internal Motion of R9 on a Solvent Exposed Helix Site
X-band CW-EPR spectral lineshapes of spin-labeled proteins encode information of the nitroxide dynamics on the ps-ns time scale [15]. The rotational diffusion of the entire protein, backbone fluctuations, and internal side chain motions that can be further modulated by local interactions [3, 10, 11] occur in this time range.
To isolate the side chain and protein dynamic modes, the spin-labeled protein is either investigated in a high viscosity medium or immobilized on a solid support to reduce or eliminate rotational diffusion of the entire protein [3, 38, 39]. Further, contributions from backbone fluctuations and local interactions can be minimized by selecting a reference site for labeling in rigid sequences where the side chain is solvent-exposed and non-interacting with other residues. Site 72 in T4L meets these criteria as it is located in the center of the long and rigid Helix C [3, 4].
Thus, the CW-EPR spectra of the side chains shown in Fig. 1 at site 72 should reveal the relative side chain internal motions on the ps-ns time scale, in the absence of backbone contributions. Figure 3a (black solid traces) shows the CW-EPR spectra for RX cross-linked between sites 68 and 72 (68RX72), and for 72R9, 72V1, and 72R1; in each case, the protein is immobilized on solid supports (CNBr-Sepharose [38, 39]). The spectrum of 68RX72 is striking with well-resolved parallel (A∥) and perpendicular (A⊥) components of the hyperfine tensor, narrow linewidths for the outer hyperfine extrema, and a large effective hyperfine splitting \(2A^{\prime}_{zz}\) [40]. In agreement with published results, the spectrum is well fit by a Microscopic Order Macroscopic Disorder model (MOMD [41]) (red dashed trace) corresponding to a highly ordered but rapid anisotropic motion [22].
X-band CW-EPR spectra and simulations for spin labels on T4L attached to Sepharose solid supports. a Experimental spectra of T4L 68RX72, 72R9, 72V1, and 72R1 at room temperature (black solid traces) and the corresponding simulated spectra (red dashed traces). The parallel (A∥) and perpendicular (A⊥) components of the axially symmetric hyperfine tensor, and the effective hyperfine splitting, \(2A^{\prime}_{zz}\) are shown. The vertical lines are drawn and fixed to the value corresponding to the \(2A^{\prime}_{zz}\) of the T4L 68RX72 spectrum for comparison. The corresponding values of the S20 are provided. b Ribbon diagram of T4L (PDB code 1L63) showing the sites where R9 is introduced and neighboring residues which are replaced with alanine (indicated by the respective Cα atom). The Cα atoms are colored corresponding to the spectra in c below. c Corresponding CW-EPR spectra of T4L 72R9 with background mutations indicated in b show the effect of alanine-replacement of neighboring side chains on R9 side chain dynamics. The vertical lines are drawn and fixed to the value corresponding to the \(2A^{\prime}_{zz}\) of the 72R9 spectrum for comparison. The magnetic field scan width for spectra in a and c is 100 G
In the current implementation of MOMD, the 2p orbital of the nitroxide is constrained to move within a cone and the amplitude of motion is measured by an order parameter \(S_{20} = 1/2\langle (3\cos^{2} \alpha - 1)\rangle\), where α is the half angle of the cone. The values of S20 lie between 0 and 1 and increase with increasing order. The rate of motion is measured by an effective correlation time, τR, values of which values increase with decreasing rate. For 68RX72, S20 = 0.93 and τR = 2.35 ns. The high order corresponding to α = 12° is due to both the backbone rigidity near site 72 and the highly constrained internal motion of the cross-linked RX side chain.
The spectra of 72R9, 72V1, and 72R1 each resemble that of 68RX72 but with decreasing \(2A^{\prime}_{zz}\) and increasing linewidth. Remarkably, the lineshapes of each side chain can be simulated reasonably well by the same MOMD model and correlation time as RX but with varying degrees of order (Fig. 3a, red dashed traces). While small variations in other simulation parameters may further improve the fit quality (see Table S1, legend), it is apparent that spectral changes are dominated by the variation of the order parameter.
Within the context of this motional model, RX clearly has the most constrained (ordered) motion, but R9 with S20 = 0.86 (α = 18°) is similar and the most highly ordered among the singly attached spin label side chains. Note from Fig. 3a that the value of \(2A^{\prime}_{zz}\) is a qualitative measure of the amplitude of motion, i.e., the order, and that there is also a systematic increase in spectral linewidths proceeding from RX to R1.
The short linkage of the R9 side chain positions the nitroxide close to the backbone, and it is possible that the high order of R9 may be the result of interactions with a neighboring residue. To test this, each nearest neighbor was mutated, one at a time, to alanine [3]. Figure 3b shows the locations of the five residues mutated (N68, Q69, V71, V75, and R76) and Fig. 3c provides the corresponding, essentially indistinguishable, EPR spectra. This finding shows that no single side chain interaction can account for the high degree of immobilization, but does not eliminate the unlikely possibility that multiple side chains act in a concerted fashion, while any single mutation has no effect. The methyl groups of alanine could interact with R9, a possibility that will be revisited below.
2.3 Dependence of R9 Side Chain Mobility on the Local Structure
To explore the generality of labeling and the information content of R9 EPR spectra in a helical protein, the side chains were introduced, one at a time, at sites in distinct structural elements of T4L [3], namely solvent-exposed sites in the interior of regular helices (sites 68, 72, 76, 131), at the N-termini (82, 109) and a C-terminus (80), at an interhelical loop site (81), and at tertiary contact sites (65, 115, 134). No significant labeling was observed for cysteines at any of the buried sites tested (99, 118, 133, 153). Figure 4a shows the location of the above sites in T4L with the backbone color-coded according to crystallographic B-factors. Corresponding EPR spectra (Fig. 4b) of R9 and R1 at the same sites in T4L are shown for comparison; R1 spectra have all been previously published [3, 6, 25].
CW-EPR spectra of R9 and R1 at various sites of T4L. a Ribbon diagram of T4L (PDB code 1L63) showing all the sites used in this study, of which each residue is colored based on the value of B-factor. The blue spheres (non-interacting helical sites), blue–white spheres (helical termini sites), red spheres (tertiary contact or loop sites), and black spheres (buried sites) at the Cα indicate where spin labels are introduced. b CW-EPR spectra of R9 and R1 at room temperature in 30% (wt/wt) sucrose solution. The vertical lines are drawn and fixed to the value corresponding to the \(2A^{\prime}_{zz}\) of the 72R9 or 72R1 spectrum as a reference for comparison. Spectra of buried sites are measured under the same experimental conditions to reveal how negligible the amount of a buried site can be labeled by R9
The EPR spectra of all labeled mutants were measured in a 30% (wt/wt) sucrose solution to minimize the rotational diffusion of the protein. Considering the variety of topographical regions involved, the solid-support attachment was not used to avoid potential interactions between nitroxide and the solid support which may differently contribute to the side chain dynamics [39].
First, the spectra reveal that the R9 side chain can be introduced with overall good yields at sites in any of the structural elements tested (Fig. 4b), except for the buried sites where there is essentially no labeling. On the other hand, the methanethiosulfonate R1 reagent (and disulfide V1 reagent) reacts readily with these sites and yields spectra reflecting strong immobilization of the side chain, as previously reported [3, 6, 25]. The selectivity for solvent-exposed sites is an important advantage of the R9 labeling scheme.
At all sites, the spectra of R9 reflect lower mobility compared to R1 as reflected in values of \(2A^{\prime}_{zz}\). In particular, R9 side chains at sites in the center of ordered helices are strongly immobilized. However, in other regions, a site-specific variation in mobility is evident that may encode, among other things, information on local dynamics and interactions. Although an analysis of the structural and dynamical origin for the site-dependent mobility is beyond the scope of this work, crystal structures presented below identify the unique interactions of R9 that give rise to relative immobilization and offer clues to the origin of the site-dependent variation.
2.4 Crystal Structure of T4L 65R9/76R9
To identify the interactions that constrain the internal motion of R9 at helical sites, a 1.60 Å crystal structure of T4L containing R9 side chains at positions 65 and 76 in Helix C (T4L 65R9/76R9) was solved and refined to an R-factor of 19.8%; refinement statistics are provided in SI (Table S2). A structure of R9 at reference site 72 could not be obtained, since this site is involved in crystal contact [42]. There is little difference between the backbone dihedral angles (Fig. S3) along Helix C in T4L 65R9/76R9 and the T4L structures, indicating that R9 does not distort the local helical geometry.
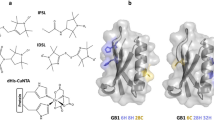
Figure 5 shows the electron density map for 65R9 and 76R9; electron densities of native side chains in Helix C are also shown to indicate the quality of the structure. Remarkably, the 2FO–FC electron density map reveals a single conformation of the side chain at both sites 65 and 76, with nearly all non-hydrogen atoms resolved. This is consistent with the high order of R9 reflected in CW-EPR spectra. In contrast, electron density maps of the more mobile R1 at solvent-exposed helical sites are not resolved for the nitroxide ring due to disorder about the final two single bonds (χ4 and χ5) [5, 8].
Crystal structure of T4L 65R9/76R9 at 100 K. a Electron density map for R9 and neighboring side chains are shown for the C helix. The electron density (cyan mesh) was calculated as an unweighted 2FO–FC map contoured at 1.0 σ. The experimental interspin distance between 65R9 {t, t} and 76R9 {t, t} is indicated. b and c Electron density maps of 65R9 and 76R9, respectively. The structure is deposited to the RCSB Protein Data Bank (PDB code: 8TAT)
Rotamers of a side chain with respect to dihedral angles {χ1, χ2} (Fig. 1) are designated t (trans), p (plus), or m (minus). For native amino acid side chains, nominal values are ± 180°, + 60° and − 60°, respectively, but variations of up to ± 30° are commonly observed [12, 43, 44]. For R1, V1, and RX, {m, m}, {t, p}, and {t, m}, are the predominant rotamers observed on α-helices in crystal structures [5, 8, 22, 25, 28, 45]. These rotamers are apparently stabilized by intra-residue interactions between the disulfide moiety and the peptide backbone, especially a Cα-H···Sδ interaction [8]. Instead, the R9 side chain adopts a {t, t} conformation at both sites 65 and 76 (Table 1) that has not so far been observed in any structures of disulfide-linked spin labels at α-helical sites.
Due to the short linkage of the R9 side chain, the nitroxide ring resides close to the backbone in the {t, t} rotamer and is sufficiently closely packed to exclude solvent on one side of the ring. This suggests that non-specific hydrophobic/van der Waals interactions [46] may play a role in determining the rotamer and the nitroxide immobilization (Fig. 6a). In addition are more specific interactions that may be grouped as intra-residue interactions within the R9, or inter-residue interactions with the i-3 and i-4 residues.
Interactions that stabilize the {t, t} rotamers and immobilize the nitroxides of R9. a CPK models of 65R9 {t, t} and 76R9 {t, t}. b Specific intra-residue, and c inter-residue interactions that stabilize the structure of 65R9 {t, t} and 76R9 {t, t} on Helix C determined from the crystal structure. In both panels, green dashes: interactions stabilizing the χ1 configuration as {t}; red dashes: interactions expected to hinder the motion of the nitroxide ring; black dashes: interactions restraining internally the configuration of the nitroxide ring
Specific interactions that stabilize the χ1 configuration as {t} in both 65R9 and 76R9 are shown as green dashed traces in Fig. 6b and c, including intra-residue C=O···Sγ interactions that meet the distance and geometry criteria for σ-hole interactions [47,48,49] and inter-residue Cβ-H···O=C hydrogen-bonding interactions [50, 51] with backbone residues i-3 and i-4. These interactions stabilize the {t, t} rotamer, but do not constrain the motion of the nitroxide ring.
Interactions expected to hinder the motion of the nitroxide ring involve both intra- and inter-residue interactions of the 4-ester substituent and are shown as red dashed traces in Fig. 6. The plane of the ester function is tilted approximately 30°–35° relative to the plane of the nitroxide ring, a configuration that optimizes intra-residue hydrogen-bonding between the ester oxygen and the C7–H hydrogen (black traces, Fig. 6b). In addition, the inter-residue interaction of the ester oxygen with Cα-H hydrogen of the i-3 residue is identified as potential Cα-H···O H-bonds [50,51,52,53,54,55] (Fig. 6c). Collectively, these attractive interactions stabilize the position of the nitroxide ring and are apparently involved in the relative immobilization of R9 compared to R1.
The interactions identified above for the {t, t} rotamer do not depend on the identity of the nearest neighbor side chains beyond the Cβ carbon. Thus, it is reasonable to assume that the above interactions, along with non-specific packing interactions, would be retained for R9 at helix surface sites in general. Under such conditions, the motion of the nitroxide would be modulated by collective motions of the helix, opening the possibility to utilize R9 as a sensor for slow protein dynamics, as discussed further below.
Although only the {t, t} rotamer is observed, other rotamers can be modeled with similar attractive interactions and without steric clashes (see below). Indeed, DEER distance measurements of the protein in frozen solution suggest the existence of at least one other preferred rotamer of R9. The results are presented in the following section.
2.5 R9 Interspin Distance Measurements by Pulsed Dipolar EPR Spectroscopy
Double electron–electron resonance (DEER) spectroscopy under cryogenic temperatures is well established for measuring long-range distance (20–80 Å) between pairs of spin labels selectively introduced in proteins [21, 56,57,58]. An important feature of DEER is that it provides analytical probability distributions, instead of average distances, making this technique a powerful tool for revealing structural heterogeneity.
While the position of a particular peak in a distance distribution contains structural information, the width reflects the spatial disorder in both the protein and nitroxide side chains. Disorder in the protein at cryogenic temperatures may reflect structural fluctuations at physiological temperature and is thus of significant interest. For a distribution width to reflect protein disorder, the disorder in the side chain must be minimized. As shown above, the motion of the R9 side chain exhibits the most ordered motion observed for singly attached spin labels at the T4L 72 reference site, making it a candidate for investigating protein structural heterogeneity.
To evaluate this point, and discover other possible rotamers of R9, DEER measurements were made on three pairs of R9 labels introduced at T4L sites in regular helices. Analogous R1 spin pairs were prepared for comparison. The Dipolar evolution functions and the corresponding fits (Fig. 7a), and derived distance distributions (Fig. 7b) are shown in Fig. 7. For each R9 pair, the distance distributions (blue traces) are dominated by a narrow peak with full width at half maximum (FWHM) of 3–3.5 Å contributing 60–80% of the total probability, while the FWHM of the corresponding R1 pairs is 5–6 Å. Although the distance distributions of R9 contain satellite peaks, the narrow width of the main peak is consistent with the high order observed for the R9 side chain.
Interspin distances measured by DEER. a Background-subtracted dipolar evolutions of the indicated T4L mutants bearing either two R9 side chains (blue) or two R1 side chains (red). b Corresponding probability distance distributions obtained from fits using a Gaussian model (see SI). Similar distance distributions were obtained with model-free analysis using optimized Tikhonov regularization. Validations of the distance distributions are shown in Fig. S4
Interestingly, the most probable distance for the 65R9/76R9 (20.08 Å) is significantly longer than the interspin distance determined from the crystal structure (Fig. 5a, 17.56 Å). Assuming that the protein structure in the crystal and solution are the same, a rotamer of R9 other than {t, t} must dominate at one or both sites in the solution. The rotamer at 65R9 is the more likely candidate since R9 makes direct contact with a symmetry-related molecule in the crystal lattice that may stabilize {t, t} (Fig. S5), and such interactions would be absent in the solution.
Modeling indicates that the {m, m} rotamer, which can account for the observed 20.0 Å DEER distance between 65R9/76R9 (Fig. 8a), can be accommodated at site 65 without steric clashes (Fig. 8b). In modeling, the {m, m} rotamer would be stabilized by favorable attractive interactions, both intra- and inter-residue (Fig. 8c, d), including a unique H-bond interaction between the ester carbonyl oxygen and the acidic Cα–H hydrogen [50,51,52,53,54]. For 72R9/131R9 and 109R9/131R9, the main distance peaks can be accounted for by sets of {m, m} and {t, t} rotamers (Fig. S6), but not with pairs of {t, t} rotamer. The minor distances observed in their distributions presumably arise from yet other rotamers.
Models of the {m, m} rotamer at site 65 that account for experimental DEER interspin distance measurements. a Model of the Helix C with 65R9 {m, m} and 76R9 {t, t} showing the interspin distance; b CPK model of 65R9 {m, m}; c intra-residue and d inter-residue interactions that stabilize 65R9 {m, m}, respectively. In both panels, green dashes: interactions stabilizing the χ1 configuration as {m}; red dashes: interactions expected to hinder the motion of the nitroxide ring
2.6 Spin–Lattice Relaxation Rate of R9
Nitroxide spin labels with highly ordered motions, such as R9, are expected to have long spin–lattice relaxation times (T1) [59,60,61], which are important for determining long-range distances at physiological temperatures using relaxation enhancement [62,63,64] or for measuring slow motion using ST-EPR [17, 65,66,67]. Based on CW-EPR lineshapes, it is anticipated that R9 should have a long T1 and therefore serve as a new probe for slow protein dynamics. To explore the suitability of R9 for those potential applications, T1 was measured using pulsed SR-EPR [29, 35, 36].
Figure 9 shows the recovery of the EPR signal from saturation for T4L 72R9 immobilized on CNBr-Sepharose. The recovery curve is reasonably well fit by a monoexponential function with a characteristic T1 = 8.3 μs, among the longest measured for a nitroxide spin-labeled protein at ambient temperatures [22]. With a T1 this long, it is predictable that R9 will be useful for relaxation enhancement experiments, extending the practical distance range up to ~ 45 Å [63, 64].
SR-EPR of T4L 72R9 attached to CNBr-Sepharose. The plot shows the signal intensity recorded at the nitroxide mI = 0 resonance maximum in the absorption spectrum as a function of time, following a saturating microwave pulse delivered at the same position. The Saturation Recovery signal (black), monoexponential fit (red), and residuals (gray) are indicated
2.7 Extending the Detection Range of Dynamics to the Microsecond Time Scale with R9
Although many functionally important processes of biological systems occur on the μs–ms time scale [68,69,70,71,72,73], a few spectroscopic techniques can offer direct dynamic information in this time domain. Conventional CW-EPR spectral lineshapes of nitroxides are insensitive to motions with correlation times longer than about 10–7 s. However, ST-EPR, a nonlinear continuous-wave EPR method introduced by Hyde and co-workers [16], extends the detection limit to motions with correlation times as long as milliseconds [17, 67, 74]. The technology has been used extensively to monitor the rotational diffusion of both soluble and membrane proteins [75,76,77,78,79].
To employ ST-EPR to observe slow internal modes of proteins, it is necessary to remove or minimize contributions from the overall Brownian protein diffusion and fast internal motions of the spin label. For this purpose, the protein is immobilized on a solid support [80, 81] and a spin label with restricted internal motion [81, 82] is needed. Here, we demonstrate that R9 is sufficiently immobilized on regular helix surface sites to detect microsecond dynamics in surface-attached proteins using ST-EPR.
Figure 10a and b (black traces) shows the CW-EPR and ST-EPR spectra, respectively, measured for T4L72R9 and T4L131R9 immobilized on CNBr-Sepharose at room temperature. Both CW-EPR lineshapes are similar and correspond to motion approaching the “rigid limit” for CW-EPR (τR ~ 100 ns). Information on motions in the μs and longer time domain is encoded in the line-height ratios [17, 74, 83, 84] H″’/H, C′/C, and L″’/L of ST-EPR spectra (Fig. 10b), using calibration plots [74] that relate the ratios to effective correlation times. The values of H″/H and L″’/L showing maximal sensitivity [17, 74] for motions in the time range of 10–5 ~ 10–3 s are used here (C′/C is sensitive when τR < 10–5 s).
ST-EPR of T4L 72R9 and 131R9. a CW-EPR and b ST-EPR spectra of T4L 72R9 and T4L 131R9 attached to CNBr-Sepharose at room temperature, before (black, solid), and after (red, dash) crosslinking using glutaraldehyde. The following field positions for ST-EPR analysis are indicated: L is the resolved peak in the low field. L″ is defined as the line height at the field position 10 Gauss upfield of L. H is the resolved peak in the high field, and H″ is 15 Gauss downfield of H. The C and C′ are the first peak and trough in the center. All line heights are measured with respect to the baseline
The ST-EPR of 72R9 and 131R9 are also qualitatively similar, and values of L″’/L indicate motions with correlation times around 23 μs for each. On the other hand, the H″’/H ratio, which has higher sensitivity to slower motions, corresponds to ~ 50 μs and ~ 100 μs for 72R9 and 131R9, respectively. The data show the utility of R9 in ST-EPR to reveal slow dynamics well beyond the detection limit of conventional CW-EPR. The different values of τR derived from different line-height ratios indicate the anisotropic nature of the motion [83,84,85,86,87].
To reveal the origin of microsecond dynamics, labeled proteins on CNBr-Sepharose were treated with glutaraldehyde under conditions that create crosslinks between lysine residues within a protein [88]. This strategy has been widely used in both solid-state NMR [89,90,91] and EPR [76, 92] studies to immobilize proteins via inter-molecular crosslinks, although intramolecular links are simultaneously formed. In the present case where each protein molecule is individually tethered to the Sepharose and widely separated, intramolecular crosslinks dominate and are expected to modulate internal fluctuations of the protein, while the internal motion of the R9 side chain is unlikely affected.
Crosslinking with glutaraldehyde has little effect on the CW-EPR spectra of both sites (red dashed traces, Fig. 10a), but pronounced changes corresponding to more than the doubling of correlation times are evident in the ST-EPR spectra (red dashed traces, Fig. 10b) of both 72R9 and 131R9 (> 100 μs, > 200 μs, respectively, using the H″/H ratio), suggesting that internal protein dynamics are the origin of the slow motions detected by R9.
3 Discussion
Elucidation of the sequence-specific nature of protein structure and dynamics is a primary goal of SDSL-EPR. Extracting such information from the EPR spectra of spin-labeled proteins requires knowledge of the configuration and motion of spin label side chain itself. Prior studies have thoroughly characterized the widely used nitroxide side chains R1, V1, and RX [3, 4, 8, 13, 15, 22, 25, 45, 93,94,95]. In this report, we provide a similar characterization of the new spin label side chain R9.
One important advantage of R9 is that it can be introduced with high selectivity to solvent-exposed cysteines. Presumably, this selectivity is due to the relatively slow reaction kinetics of HO-4072 with cysteine (Fig. 2b). Reaction with a buried cysteine can only occur during the lifetime of a protein fluctuation that exposes the residue to solvent, and the reaction rate of the Michael addition is too slow for a measurable product to form during the time required for reaction with a surface cysteine. On the other hand, the reactions of MTSSL and IDSL are extremely rapid, where quantitative modification with buried cysteines can occur [3].
An additional advantage of R9 is the thioether linkage that is not cleaved by the commonly employed reducing agents. Other spin label side chains linked to the protein via a thioether have been employed [81, 82, 96,97,98,99,100,101], but have not yet been characterized at the level required for interpreting EPR spectra in terms of protein structure and dynamics.
The CW-EPR lineshape of R9 at the T4L 72 reference site reflects a highly ordered anisotropic motion of the nitroxide (Fig. 3a), consistent with the very long T1 (Fig. 9) [61]. The high ordering of R9 is not unique to site 72 but is also observed at other sites in the interior of relatively rigid helical segments, including site 131, and sites 65 and 76 for which crystal structures were obtained (Fig. 5). The crystal structures suggest that the ordering arises from site-independent non-covalent bonding interactions with apparent contributions from non-specific packing interactions (Figs. 6, 8). On the other hand, the spectral lineshapes at most sites near helix termini and in connecting loops have increased mobility, suggesting that R9 may serve as a sensitive indicator of nanosecond motions in flexible regions of a protein (see below).
Highly constrained nitroxide side chains, such as RX and R9 with long T1, are of value in SDSL in part because they offer the potential for monitoring motions on the important μs–ms time scale, thus extending the detection range of conventional X-band CW-EPR lineshape analysis. To date, only methods relying on the saturation behavior of the nitroxide, determined by T1, have reported sensitivity to motions beyond the 100 ns time scale. Pulsed SR-EPR and ELDOR have been used to monitor exchange events in the range of 1 μs ~ 70 μs [29], while longer times are inaccessible. In principle, ST-EPR methods can extend the range farther into the ms domain [66, 67] provided that a highly constrained side chain is used and rotational diffusion of the whole protein can be minimized. Elegant studies by Thomas et al. have employed ST-EPR to monitor slow protein movements in muscle fibers as well as in lipid membranes, both of which provide a natural scaffolding on which labeled proteins are immobilized [18, 75, 76, 80, 81, 102,103,104]. Here, we extend this method to small (< 60 kDa), soluble proteins by immobilizing them on the Sepharose surface and employing R9 as the rigid probe.
The data in Fig. 10 demonstrate that ST-EPR using R9 can detect microsecond internal protein dynamics. The 20–100 μs motions detected for T4L 72R9 and T4L 131R9 attached on CNBr-Sepharose could in principle arise from motions of the protein relative to the solid support, R9 relative to the protein, and/or internal protein motions, i.e., structural fluctuations. The latter is clearly indicated by the strong effect of intramolecular crosslinking via glutaraldehyde. The ability to monitor site-specific internal motions might open the door to exploring the role of protein dynamics in ligand binding and protein–protein recognition, and applications will be presented elsewhere.
In addition to monitoring slow dynamics, R9 at highly ordered sites shows promise for distance measurement in proteins using DEER or T1-based relaxation enhancement methods (Fig. 7). For either method, an ideal label would exist in a single rotameric state to provide the highest distance resolution. It is unfortunate that multiple rotamers exist for any singly attached spin label published as yet, but due to the fewer number of rotatable bonds between the backbone and the nitroxide in R9 (3 bonds), as compared to R1 (5 bonds) and V1 (4 bonds) (Fig. 1), the rotameric space of R9 is more limited and the position of the EPR-active nitroxide is relatively localized.
For high-resolution structure mapping by DEER, the existence of multiple rotamers is a limitation, but the very narrow dominant population of interspin distances for R9 pairs suggests that R9 will be useful for assessing structural heterogeneity, particularly when monitoring backbone flexibility or small changes in the global structure. In general, the elucidation of equilibrium structure and dynamics can be improved by comparing distance distributions obtained with R1, V1, and R9 using DEER.
4 Conclusions
Compared to the commonly used R1 side chain, the highly restricted internal motion of R9 makes measurement of slow motions within proteins possible, and allows for measurements of longer distances via relaxation enhancement at ambient temperatures and identification of structural heterogeneity via DEER. While the side chains V1 and RX have an ordering similar to R9, R9 has the advantages of labeling-selectivity at solvent-exposed surface sites, and singly attachment via a non-reducible thioether linker. Each of these side chains has advantages and disadvantages, but the unique properties of R9 make it a useful addition to the SDSL toolkit.
5 Materials and Methods
5.1 Construction, Purification, and Spin Labeling of Cysteine Mutants
T4L constructs were expressed, purified, and spin-labeled according to the general procedures previously described [3, 22, 25]. Reagents MTSSL and HO-1944 were generous gifts of Prof. Tamás Kálai (University of Pécs, Hungary). IDSL was purchased from Toronto Research Chemicals. Additional information is in SI Materials and Methods.
5.2 Synthesis, Characterization, and Reaction Kinetics of Reagent HO-4072

The synthesis of Methyl 4-fluoro-2,2,5,5-tetramethyl-2,5-dihydro-1H-pyrrole-3-carboxylate-1-yloxyl Radical (HO-4072): To a stirred solution of 2,2,6,6-Tetramethylpiperidine (TEMP) (1.51 g, 10.0 mmol) in anhydrous THF (10 mL), BuLi 4.0 mL (10.0 mmol, 2.5 M in hexanes) was added at 0 °C to generate the Li salt of TEMP (LiTEMP). After stirring at this temperature for 20 min. the solution was chilled to − 78 °C with dry ice/acetone and the solution of compound 1 [105] (1.98 g, 10.0 mmol) in anhydr. THF (10 mL) was added dropwise during 30 min. After stirring this solution at − 78 °C for 1 h, fluorobenzosulfonimide (NFSI) (3.15 g, 10.0 mmol) was added in anhydr. THF (20 mL) at − 78 °C during 30 min. Then, the mixture was allowed to warm to room temperature spontaneously (~ 1 h). The mixture was quenched with aq. sat. NH4Cl solution (20 mL) followed by adding EtOAc (20 mL). The organic phase was separated, then the aqueous phase was extracted with EtOAc (20 mL), and the combined organic phase was dried (MgSO4), filtered, and evaporated. The residue was purified with flash chromatography (hexane/ Et2O) to give compound 2 583 mg (27%) as orange crystals.
Compound 2 (reagent HO-4072) was characterized by chromatography and multiple spectroscopies. The kinetics of the reaction were determined using EPR spectroscopy, and the reaction products were characterized by Electrospray Ionization Mass Spectrometry (ESI–MS). Details are described in SI Materials and Methods.
5.3 EPR Spectroscopy
CW-EPR and ST-EPR experiments were performed on a Varian E-109 spectrometer fitted with a two-loop one-gap resonator [106, 107]. SR-EPR and Four-pulse DEER experiments were conducted on the Bruker ELEXSYS 580 spectrometer. Spectral simulations and DEER data analyses were carried out using the programs “MultiComponent” and “LongDistances”, respectively. The programs are available at https://www.biochemistry.ucla.edu/Faculty/Hubbell/software.html. Additional details in SI Materials and Methods.
5.4 X-Ray Crystallography
The diffraction quality crystal of T4L 65R9/76R9 was grown using the hanging drop vapor diffusion method. Crystals appeared in 1 week under 4℃. For the cryogenic diffraction study, a single crystal cryo-protected in mineral oil was flash-frozen, and data were collected at 100 K using an FR-E + SUPERBRIGHT X-ray Generator on a RIGAKU RAXIS HTC image plate detector. Data processing was performed as previously described [8] (SI Materials and Methods).
Availability of Data and Materials
The data that support the findings of this study are available from the corresponding author upon reasonable request.
References
L.J. Berliner, J. Grunwald, H.O. Hankovszky, K. Hideg, A novel reversible thiol-specific spin label: papain active site labeling and inhibition. Anal. Biochem. 119, 450–455 (1982). https://doi.org/10.1016/0003-2697(82)90612-1
A.P. Todd, J. Cong, F. Levinthal, C. Levinthal, W.L. Hubbell, Site-directed mutagenesis of colicin E1 provides specific attachment sites for spin labels whose spectra are sensitive to local conformation. Proteins Struct. Funct. Bioinform. 6, 294–305 (1989). https://doi.org/10.1002/prot.340060312
H.S. Mchaourab, M.A. Lietzow, K. Hideg, W.L. Hubbell, Motion of spin-labeled side chains in T4 lysozyme. Correlation with protein structure and dynamics. Biochemistry 35, 7692–7704 (1996). https://doi.org/10.1021/bi960482k
L. Columbus, T. Kálai, J. Jekö, K. Hideg, W.L. Hubbell, Molecular motion of spin labeled side chains in α-helices: analysis by variation of side chain structure. Biochemistry 40, 3828–3846 (2001). https://doi.org/10.1021/bi002645h
R. Langen, K.J. Oh, D. Cascio, W.L. Hubbell, Crystal Structures of spin labeled T4 lysozyme mutants: implications for the interpretation of EPR spectra in terms of structure. Biochemistry 39, 8396–8405 (2000). https://doi.org/10.1021/bi000604f
Z. Guo, D. Cascio, K. Hideg, T. Kálai, W.L. Hubbell, Structural determinants of nitroxide motion in spin-labeled proteins: tertiary contact and solvent-inaccessible sites in helix G of T4 lysozyme. Protein Sci. 16, 1069–1086 (2007). https://doi.org/10.1110/ps.062739107
Z. Guo, D. Cascio, K. Hideg, W.L. Hubbell, Structural determinants of nitroxide motion in spin-labeled proteins: solvent-exposed sites in helix B of T4 lysozyme. Protein Sci. 17, 228–239 (2008). https://doi.org/10.1110/ps.073174008
M.R. Fleissner, D. Cascio, W.L. Hubbell, Structural origin of weakly ordered nitroxide motion in spin-labeled proteins. Protein Sci. 18, 893–908 (2009). https://doi.org/10.1002/pro.96
P. Consentius, U. Gohlke, B. Loll, C. Alings, R. Müller, U. Heinemann, M. Kaupp, M. Wahl, T. Risse, Tracking transient conformational states of T4 lysozyme at room temperature combining X-ray crystallography and site-directed spin labeling. J. Am. Chem. Soc. 138, 12868–12875 (2016). https://doi.org/10.1021/jacs.6b05507
A.K. Kusnetzow, C. Altenbach, W.L. Hubbell, Conformational states and dynamics of rhodopsin in micelles and bilayers. Biochemistry 45, 5538–5550 (2006). https://doi.org/10.1021/bi060101v
C.J. López, S. Oga, W.L. Hubbell, Mapping molecular flexibility of proteins with site-directed spin labeling: a case study of myoglobin. Biochemistry 51, 6568–6583 (2012). https://doi.org/10.1021/bi3005686
D.T. Warshaviak, L. Serbulea, K.N. Houk, W.L. Hubbell, Conformational analysis of a nitroxide side chain in an α-helix with density functional theory. J. Phys. Chem. B. 115, 397–405 (2011). https://doi.org/10.1021/jp108871m
L. Columbus, W.L. Hubbell, A new spin on protein dynamics. Trends Biochem. Sci. 27, 288–295 (2002). https://doi.org/10.1016/S0968-0004(02)02095-9
L. Columbus, W.L. Hubbell, Mapping backbone dynamics in solution with site-directed spin labeling: GCN4−58 bZip free and bound to DNA. Biochemistry 43, 7273–7287 (2004). https://doi.org/10.1021/bi0497906
C. Altenbach, C.J. López, K. Hideg, W.L. Hubbell, Chapter three—exploring structure, dynamics, and topology of nitroxide spin-labeled proteins using continuous-wave electron paramagnetic resonance spectroscopy, in Methods in Enzymology. ed. by P.Z. Qin, K. Warncke (Academic Press, London, 2015), pp.59–100
J.S. Hyde, L. Dalton, Very slowly tumbling spin labels: adiabatic rapid passage. Chem. Phys. Lett. 16, 568–572 (1972). https://doi.org/10.1016/0009-2614(72)80426-3
D.D. Thomas, L.R. Dalton, J.S. Hyde, Rotational diffusion studied by passage saturation transfer electron paramagnetic resonance. J. Chem. Phys. 65, 3006–3024 (1976). https://doi.org/10.1063/1.433512
A.R. Thompson, N. Naber, C. Wilson, R. Cooke, D.D. Thomas, Structural dynamics of the actomyosin complex probed by a bifunctional spin label that cross-links SH1 and SH2. Biophys. J. 95, 5238–5246 (2008). https://doi.org/10.1529/biophysj.108.138982
D.A. Haas, T. Sugano, C. Mailer, B.H. Robinson, Motion in nitroxide spin labels: direct measurement of rotational correlation times by pulsed electron double resonance. J. Phys. Chem. 97, 2914–2921 (1993). https://doi.org/10.1021/j100114a014
P.P. Borbat, J.H. Freed, Double-quantum ESR and distance measurements, in Distance Measurements in Biological Systems by EPR. ed. by L.J. Berliner, G.R. Eaton, S.S. Eaton (Springer, Boston, 2000), pp.383–459
G. Jeschke, DEER distance measurements on proteins. Annu. Rev. Phys. Chem. 63, 419–446 (2012). https://doi.org/10.1146/annurev-physchem-032511-143716
M.R. Fleissner, M.D. Bridges, E.K. Brooks, D. Cascio, T. Kálai, K. Hideg, W.L. Hubbell, Structure and dynamics of a conformationally constrained nitroxide side chain and applications in EPR spectroscopy. Proc. Natl. Acad. Sci. 108, 16241–16246 (2011). https://doi.org/10.1073/pnas.1111420108
P. Consentius, U. Gohlke, B. Loll, C. Alings, U. Heinemann, M.C. Wahl, T. Risse, Combining EPR spectroscopy and X-ray crystallography to elucidate the structure and dynamics of conformationally constrained spin labels in T4 lysozyme single crystals. Phys. Chem. Chem. Phys. 19, 20723–20734 (2017). https://doi.org/10.1039/C7CP03144K
P. Consentius, B. Loll, U. Gohlke, C. Alings, C. Müller, R. Müller, C. Teutloff, U. Heinemann, M. Kaupp, M.C. Wahl, T. Risse, Internal dynamics of the 3-pyrroline-N-oxide ring in spin-labeled proteins. J. Phys. Chem. Lett. 8, 1113–1117 (2017). https://doi.org/10.1021/acs.jpclett.6b02971
D.T. Warshaviak, V.V. Khramtsov, D. Cascio, C. Altenbach, W.L. Hubbell, Structure and dynamics of an imidazoline nitroxide side chain with strongly hindered internal motion in proteins. J. Magn. Reson. 232, 53–61 (2013). https://doi.org/10.1016/j.jmr.2013.04.013
V.V. Khramtsov, V.I. Yelinova, L.M. Weiner, T.A. Berezina, V.V. Martin, L.B. Volodarsky, Quantitative determination of SH groups in low- and high-molecular-weight compounds by an electron spin resonance method. Anal. Biochem. 182, 58–63 (1989). https://doi.org/10.1016/0003-2697(89)90718-5
L. M. Weiner, Quantitative determination of thiol groups in low and high molecular weight compounds by electron paramagnetic resonance, in Methods in Enzymology. ed. by L. Packer (Academic Press, London, 1995), p. 87–105.
A.R. Balo, H. Feyrer, O.P. Ernst, Toward precise interpretation of DEER-based distance distributions: insights from structural characterization of V1 spin-labeled side chains. Biochemistry 55, 5256–5263 (2016). https://doi.org/10.1021/acs.biochem.6b00608
M.D. Bridges, K. Hideg, W.L. Hubbell, Resolving conformational and rotameric exchange in spin-labeled proteins using saturation recovery EPR. Appl. Magn. Reson. 37, 363–390 (2010). https://doi.org/10.1007/s00723-009-0079-2
R.F. Rayes, T. Kálai, K. Hideg, M.A. Geeves, P.G. Fajer, Dynamics of tropomyosin in muscle fibers as monitored by saturation transfer EPR of bi-functional probe. PLoS One 6, e21277 (2011). https://doi.org/10.1371/journal.pone.0021277
A.R. Thompson, B.P. Binder, J.E. McCaffrey, B. Svensson, D.D. Thomas, Chapter four—bifunctional spin labeling of muscle proteins: accurate rotational dynamics, orientation, and distance by EPR, in Methods in Enzymology. ed. by P.Z. Qin, K. Warncke (Academic Press, London, 2015), pp.101–123
L.J. Perry, R. Wetzel, Unpaired cysteine-54 interferes with the ability of an engineered disulfide to stabilize T4 lysozyme. Biochemistry 25, 733–739 (1986). https://doi.org/10.1021/bi00351a034
M. Matsumura, B.W. Matthews, Control of enzyme activity by an engineered disulfide bond. Science 243, 792–794 (1989). https://doi.org/10.1126/science.2916125
M. Matsumura, W.J. Becktel, M. Levitt, B.W. Matthews, Stabilization of phage T4 lysozyme by engineered disulfide bonds. Proc. Natl. Acad. Sci. 86, 6562–6566 (1989). https://doi.org/10.1073/pnas.86.17.6562
P.W. Percival, J.S. Hyde, Pulsed EPR spectrometer, II. Rev. Sci. Instrum. 46, 1522–1529 (2008). https://doi.org/10.1063/1.1134096
M.D. Bridges, Z. Yang, C. Altenbach, W.L. Hubbell, Analysis of saturation recovery amplitudes to characterize conformational exchange in spin-labeled proteins. Appl. Magn. Reson. 48, 1315–1340 (2017). https://doi.org/10.1007/s00723-017-0936-3
G.D. Fasman, Handbook of Biochemistry: Section D Physical Chemical Data, vol. I (CRC Press, Boca Raton, 2018)
C.J. López, M.R. Fleissner, Z. Guo, A.K. Kusnetzow, W.L. Hubbell, Osmolyte perturbation reveals conformational equilibria in spin-labeled proteins. Protein Sci. 18, 1637–1652 (2009). https://doi.org/10.1002/pro.180
C.J. López, M.R. Fleissner, E.K. Brooks, W.L. Hubbell, Stationary-phase EPR for exploring protein structure, conformation, and dynamics in spin-labeled proteins. Biochemistry 53, 7067–7075 (2014). https://doi.org/10.1021/bi5011128
J.H. Freed, Theory of slow tumbling ESR spectra for nitroxides, in Spin Labeling. ed. by L.J. Berliner (Academic Press, Amsterdam, 1976), pp.53–132
D.E. Budil, S. Lee, S. Saxena, J.H. Freed, Nonlinear-least-squares analysis of slow-motion EPR spectra in one and two dimensions using a modified Levenberg–Marquardt algorithm. J. Magn. Reson. 120, 155–189 (1996). https://doi.org/10.1006/jmra.1996.0113
D.R. Banatao, D. Cascio, C.S. Crowley, M.R. Fleissner, H.L. Tienson, T.O. Yeates, An approach to crystallizing proteins by synthetic symmetrization. Proc. Natl. Acad. Sci. 103, 16230–16235 (2006). https://doi.org/10.1073/pnas.0607674103
S.C. Lovell, J.M. Word, J.S. Richardson, D.C. Richardson, The penultimate rotamer library. Proteins Struct. Funct. Bioinform. 40, 389–408 (2000). https://doi.org/10.1002/1097-0134(20000815)40:3%3c389::AID-PROT50%3e3.0.CO;2-2
P. Gainza, K.E. Roberts, B.R. Donald, Protein design using continuous rotamers. PLoS Comput. Biol. 8, e1002335 (2012). https://doi.org/10.1371/journal.pcbi.1002335
D. Marsh, Site-directed spin labelling (SDSL), in Spin-Label Electron Paramagnetic Resonance Spectroscopy. ed. by D. Marsh (CRC Press, Boca Raton, 2019), p. 371–416.
A. Bondi, van der Waals volumes and Radii. J. Phys. Chem. 68, 441–451 (1964). https://doi.org/10.1021/j100785a001
C. Bleiholder, D.B. Werz, H. Köppel, R. Gleiter, Theoretical investigations on chalcogen−chalcogen interactions: what makes these nonbonded interactions bonding? J. Am. Chem. Soc. 128, 2666–2674 (2006). https://doi.org/10.1021/ja056827g
M.E. Brezgunova, J. Lieffrig, E. Aubert, S. Dahaoui, P. Fertey, S. Lebègue, J.G. Ángyán, M. Fourmigué, E. Espinosa, Chalcogen Bonding: experimental and theoretical determinations from electron density analysis. Geometrical preferences driven by electrophilic–nucleophilic interactions. Cryst. Growth Des. 13, 3283–3289 (2013). https://doi.org/10.1021/cg400683u
D.J. Pascoe, K.B. Ling, S.L. Cockroft, The origin of chalcogen-bonding interactions. J. Am. Chem. Soc. 139, 15160–15167 (2017). https://doi.org/10.1021/jacs.7b08511
K. Manikandan, S. Ramakumar, The occurrence of C-H…O hydrogen bonds in α-helices and helix termini in globular proteins. Proteins Struct. Funct. Bioinform. 56, 768–781 (2004). https://doi.org/10.1002/prot.20152
J.D. Yesselman, S. Horowitz, C.L. Brooks III., R.C. Trievel, Frequent side chain methyl carbon-oxygen hydrogen bonding in proteins revealed by computational and stereochemical analysis of neutron structures. Proteins Struct. Funct. Bioinform. 83, 403–410 (2015). https://doi.org/10.1002/prot.24724
Z.S. Derewenda, L. Lee, U. Derewenda, The occurence of C-H · · · O hydrogen bonds in proteins. J. Mol. Biol. 252, 248–262 (1995). https://doi.org/10.1006/jmbi.1995.0492
K. Molčanov, B. Kojić-Prodić, N. Raos, Analysis of the less common hydrogen bonds involving ester oxygen sp3 atoms as acceptors in the crystal structures of small organic molecules. Acta. Cryst. B 60, 424–432 (2004). https://doi.org/10.1107/S0108768104014442
S. Horowitz, R.C. Trievel, Carbon-oxygen hydrogen bonding in biological structure and function. J. Biol. Chem. 287, 41576–41582 (2012). https://doi.org/10.1074/jbc.R112.418574
G.J. Bartlett, D.N. Woolfson, On the satisfaction of backbone-carbonyl lone pairs of electrons in protein structures. Protein Sci. 25, 887–897 (2016). https://doi.org/10.1002/pro.2896
G. Jeschke, G. Panek, A. Godt, A. Bender, H. Paulsen, Data analysis procedures for pulse ELDOR measurements of broad distance distributions. Appl. Magn. Reson. 26, 223 (2004). https://doi.org/10.1007/BF03166574
M. Pannier, S. Veit, A. Godt, G. Jeschke, H.W. Spiess, Dead-time free measurement of dipole–dipole interactions between electron spins. J. Magn. Reson. 213, 316–325 (2011). https://doi.org/10.1016/j.jmr.2011.08.035
G. Jeschke, Conformational dynamics and distribution of nitroxide spin labels. Prog. Nucl. Magn. Reson. Spectrosc. 72, 42–60 (2013). https://doi.org/10.1016/j.pnmrs.2013.03.001
H. Sato, V. Kathirvelu, A. Fielding, J.P. Blinco, A.S. Micallef, S.E. Bottle, S.S. Eaton, G.R. Eaton, Impact of molecular size on electron spin relaxation rates of nitroxyl radicals in glassy solvents between 100 and 300 K. Mol. Phys. 105, 2137–2151 (2007). https://doi.org/10.1080/00268970701724966
H. Sato, S.E. Bottle, J.P. Blinco, A.S. Micallef, G.R. Eaton, S.S. Eaton, Electron spin–lattice relaxation of nitroxyl radicals in temperature ranges that span glassy solutions to low-viscosity liquids. J. Magn. Reson. 191, 66–77 (2008). https://doi.org/10.1016/j.jmr.2007.12.003
D. Marsh, Molecular order and T1-relaxation, cross-relaxation in nitroxide spin labels. J. Magn. Reson. 290, 38–45 (2018). https://doi.org/10.1016/j.jmr.2018.02.020
S. Jun, J.S. Becker, M. Yonkunas, R. Coalson, S. Saxena, Unfolding of alanine-based peptides using electron spin resonance distance measurements. Biochemistry 45, 11666–11673 (2006). https://doi.org/10.1021/bi061195b
Z. Yang, G. Jiménez-Osés, C.J. López, M.D. Bridges, K.N. Houk, W.L. Hubbell, Long-range distance measurements in proteins at physiological temperatures using saturation recovery EPR spectroscopy. J. Am. Chem. Soc. 136, 15356–15365 (2014). https://doi.org/10.1021/ja5083206
Z. Yang, M.D. Bridges, C.J. López, OYu. Rogozhnikova, D.V. Trukhin, E.K. Brooks, V. Tormyshev, H.J. Halpern, W.L. Hubbell, A triarylmethyl spin label for long-range distance measurement at physiological temperatures using T1 relaxation enhancement. J. Magn. Reson. 269, 50–54 (2016). https://doi.org/10.1016/j.jmr.2016.05.006
T. Páli, R. Bartucci, L.I. Horváth, D. Marsh, Distance measurements using paramagnetic ion-induced relaxation in the saturation transfer electron spin resonance of spin-labeled biomolecules: application to phospholipid bilayers and interdigitated gel phases. Biophys. J. 61, 1595–1602 (1992). https://doi.org/10.1016/S0006-3495(92)81963-X
T. Páli, V.A. Livshits, D. Marsh, Dependence of saturation-transfer EPR intensities on spin-lattice relaxation. J. Magn. Reson. 113, 151–159 (1996). https://doi.org/10.1006/jmrb.1996.0168
I.J. van den Dries, P.A. de Jager, M.A. Hemminga, Sensitivity of saturation transfer electron spin resonance extended to extremely slow mobility in glassy materials. J. Magn. Reson. 131, 241–247 (1998). https://doi.org/10.1006/jmre.1997.1354
R.H. Austin, K.W. Beeson, L. Eisenstein, H. Frauenfelder, I.C. Gunsalus, Dynamics of ligand binding to myoglobin. Biochemistry 14, 5355–5373 (1975). https://doi.org/10.1021/bi00695a021
A.G. Palmer, Probing molecular motion by NMR. Curr. Opin. Struct. Biol. 7, 732–737 (1997). https://doi.org/10.1016/S0959-440X(97)80085-1
M. Wolf-Watz, V. Thai, K. Henzler-Wildman, G. Hadjipavlou, E.Z. Eisenmesser, D. Kern, Linkage between dynamics and catalysis in a thermophilic-mesophilic enzyme pair. Nat. Struct. Mol. Biol. 11, 945–949 (2004). https://doi.org/10.1038/nsmb821
D.D. Boehr, H.J. Dyson, P.E. Wright, An NMR perspective on enzyme dynamics. Chem. Rev. 106, 3055–3079 (2006). https://doi.org/10.1021/cr050312q
K. Henzler-Wildman, D. Kern, Dynamic personalities of proteins. Nature 450, 964–972 (2007). https://doi.org/10.1038/nature06522
K.A. Henzler-Wildman, V. Thai, M. Lei, M. Ott, M. Wolf-Watz, T. Fenn, E. Pozharski, M.A. Wilson, G.A. Petsko, M. Karplus, C.G. Hübner, D. Kern, Intrinsic motions along an enzymatic reaction trajectory. Nature 450, 838–844 (2007). https://doi.org/10.1038/nature06410
T.C. Squier, D.D. Thomas, Methodology for increased precision in saturation transfer electron paramagnetic resonance studies of rotational dynamics. Biophys. J. 49, 921–935 (1986). https://doi.org/10.1016/S0006-3495(86)83720-1
D.D. Thomas, J.C. Seidel, J.S. Hyde, J. Gergely, Motion of subfragment-1 in myosin and its supramolecular complexes: saturation transfer electron paramagnetic resonance. Proc. Natl. Acad. Sci. 72, 1729–1733 (1975). https://doi.org/10.1073/pnas.72.5.1729
D.D. Thomas, C. Hidalgo, Rotational motion of the sarcoplasmic reticulum Ca2+-ATPase. Proc. Natl. Acad. Sci. 75, 5488–5492 (1978). https://doi.org/10.1073/pnas.75.11.5488
P. Thiyagarajan, M.E. Johnson, Saturation transfer electron paramagnetic resonance detection of sickle hemoglobin aggregation during deoxygenation. Biophys. J. 42, 269–274 (1983). https://doi.org/10.1016/S0006-3495(83)84394-X
N.J.P. Ryba, M.A. Hoon, J.B.C. Findlay, H.R. Saibil, J.R. Wilkinson, T. Heimburg, D. Marsh, Rhodopsin mobility, structure, and lipid–protein interaction in squid photoreceptor membranes. Biochemistry 32, 3298–3305 (1993). https://doi.org/10.1021/bi00064a012
Y.-H. Kuo, Y.-W. Chiang, Slow Dynamics around a protein and its coupling to solvent. ACS Cent. Sci. 4, 645–655 (2018). https://doi.org/10.1021/acscentsci.8b00139
D.D. Thomas, S. Ishiwata, J.C. Seidel, J. Gergely, Submillisecond rotational dynamics of spin-labeled myosin heads in myofibrils. Biophys. J. 32, 873–889 (1980). https://doi.org/10.1016/S0006-3495(80)85023-5
O. Roopnarine, K. Hideg, D.D. Thomas, Saturation transfer electron parametric resonance of an indane-dione spin-label. Calibration with hemoglobin and application to myosin rotational dynamics. Biophys. J. 64, 1896–1907 (1993). https://doi.org/10.1016/S0006-3495(93)81561-3
M. Esmann, H.O. Hankovszky, K. Hideg, D. Marsh, A novel spin-label for study of membrane protein rotational diffusion using saturation transfer electron spin resonance. Application to selectively labelled Class I and Class II -SH groups of the shark rectal gland Na+/K+-ATPase. Biochim. Biophys. Acta Biomembr. 978, 209–215 (1989). https://doi.org/10.1016/0005-2736(89)90117-X
B.H. Robinson, L.R. Dalton, Anisotropic rotational diffusion studied by passage saturation transfer electron paramagnetic resonance. J. Chem. Phys. 72, 1312–1324 (1980). https://doi.org/10.1063/1.439194
E.C. Howard, K.M. Lindahl, C.F. Polnaszek, D.D. Thomas, Simulation of saturation transfer electron paramagnetic resonance spectra for rotational motion with restricted angular amplitude. Biophys. J. 64, 581–593 (1993). https://doi.org/10.1016/S0006-3495(93)81417-6
B. Jean, Gaffney, spin label-thiourea adducts. A model for saturation transfer ESR studies of slow anisotropic rotation. J. Phys. Chem. 83, 3345–3349 (1979). https://doi.org/10.1021/j100489a007
M.E. Johnson, L. Lee, L.W.M. Fung, Models for slow anisotropic rotational diffusion in saturation transfer electron paramagnetic resonance at 9 and 35 GHz. Biochemistry 21, 4459–4467 (1982). https://doi.org/10.1021/bi00261a041
P. Fajer, D. Marsh, Sensitivity of saturation transfer ESR spectra to anisotropic rotation. Application to membrane systems. J. Magn. Reson. 51, 446–459 (1983). https://doi.org/10.1016/0022-2364(83)90297-4
I. Migneault, C. Dartiguenave, M.J. Bertrand, K.C. Waldron, Glutaraldehyde: behavior in aqueous solution, reaction with proteins, and application to enzyme crosslinking. Biotechniques 37, 790–802 (2004). https://doi.org/10.2144/04375RV01
E. Persson, B. Halle, Nanosecond to microsecond protein dynamics probed by magnetic relaxation dispersion of buried water molecules. J. Am. Chem. Soc. 130, 1774–1787 (2008). https://doi.org/10.1021/ja0775873
E.P. Sunde, B. Halle, Slow internal protein dynamics from water 1H magnetic relaxation dispersion. J. Am. Chem. Soc. 131, 18214–18215 (2009). https://doi.org/10.1021/ja908144y
S. Kaieda, B. Halle, Internal water and microsecond dynamics in myoglobin. J. Phys. Chem. B. 117, 14676–14687 (2013). https://doi.org/10.1021/jp409234g
L.W. Fung, B.O. Kalaw, R.M. Hatfield, M.N. Dias, Erythrocyte spectrin maintains its segmental motions on oxidation: a spin-label EPR study. Biophys. J. 70, 841–851 (1996). https://doi.org/10.1016/S0006-3495(96)79626-1
W.L. Hubbell, C.J. López, C. Altenbach, Z. Yang, Technological advances in site-directed spin labeling of proteins. Curr. Opin. Struct. Biol. 23, 725–733 (2013). https://doi.org/10.1016/j.sbi.2013.06.008
Z. Yang, M. Bridges, M.T. Lerch, C. Altenbach, W.L. Hubbell, Chapter one—saturation recovery EPR and nitroxide spin labeling for exploring structure and dynamics in proteins, in Methods in Enzymology. ed. by P.Z. Qin, K. Warncke (Academic Press, London, 2015), pp.3–27
M.T. Lerch, Z. Yang, C. Altenbach, W.L. Hubbell, Chapter two—high-pressure EPR and site-directed spin labeling for mapping molecular flexibility in proteins, in Methods in Enzymology. ed. by P.Z. Qin, K. Warncke (Academic Press, London, 2015), pp.29–57
M. Esmann, H.O. Hankovszky, K. Hideg, J.A. Pedersen, D. Marsh, Vinyl ketone reagents for covalent protein modification. Nitroxide derivatives suited to rotational diffusion studies by saturation transfer electron spin resonance, using membrane-bound Na, K-ATPase as an example. Anal. Biochem. 189, 274–282 (1990). https://doi.org/10.1016/0003-2697(90)90120-X
L.I. Horváth, L. Dux, H.O. Hankovszky, K. Hideg, D. Marsh, Saturation transfer electron spin resonance of Ca2(+)-ATPase covalently spin-labeled with beta-substituted vinyl ketone- and maleimide-nitroxide derivatives. Effects of segmental motion and labeling levels. Biophys. J. 58, 231–241 (1990). https://doi.org/10.1016/S0006-3495(90)82368-7
M. Esmann, K. Hideg, D. Marsh, Conventional and saturation transfer EPR spectroscopy of Na+/K+-ATPase modified with different maleimide-nitroxide derivatives. Biochim. Biophys. Acta Protein Struct. Mol. Enzymol. 1159, 51–59 (1992). https://doi.org/10.1016/0167-4838(92)90074-N
M. Esmann, P.C. Sar, K. Hideg, D. Marsh, Maleimide, iodoacetamide, indanedione, and chloromercuric spin label reagents with derivatized nitroxide rings as ESR reporter groups for protein conformation and dynamics. Anal. Biochem. 213, 336–348 (1993). https://doi.org/10.1006/abio.1993.1430
M.A. Hollas, S.J. Webb, S.L. Flitsch, A.J. Fielding, A bifunctional spin label for ligand recognition on surfaces. Angew. Chem. Int. Ed. 56, 9449–9453 (2017). https://doi.org/10.1002/anie.201703929
G. Heaven, M.A. Hollas, L. Tabernero, A.J. Fielding, Spin labeling of surface cysteines using a bromoacrylaldehyde spin label. Appl. Magn. Reson. 52, 959–970 (2021). https://doi.org/10.1007/s00723-021-01350-1
D.D. Thomas, Rotational diffusion of membrane proteins, in Techniques for the Analysis of Membrane Proteins. ed. by C.I. Ragan, R.J. Cherry (Springer, Dordrecht, 1986), pp.377–431
Z.M. James, J.E. McCaffrey, K.D. Torgersen, C.B. Karim, D.D. Thomas, Protein–protein interactions in calcium transport regulation probed by saturation transfer electron paramagnetic resonance. Biophys. J. 103, 1370–1378 (2012). https://doi.org/10.1016/j.bpj.2012.08.032
R.N. Mello, D.D. Thomas, Three distinct actin-attached structural states of myosin in muscle fibers. Biophys. J. 102, 1088–1096 (2012). https://doi.org/10.1016/j.bpj.2011.11.4027
E.G. Rozantsev, H. Ulrich, Syntheses of some stable radicals and the most important intermediates, in Free Nitroxyl Radicals. ed. by E.G. Rozantsev, H. Ulrich (Springer, Boston, 1970), pp.203–246
W.L. Hubbell, W. Froncisz, J.S. Hyde, Continuous and stopped flow EPR spectrometer based on a loop gap resonator. Rev. Sci. Instrum. 58, 1879–1886 (1987). https://doi.org/10.1063/1.1139536
J.S. Hyde, W. Froncisz, Loop gap resonators. Adv. EPR Appl. Biol. Biochem. (1989). https://doi.org/10.1016/B978-0-444-88050-5.50012-4
Funding
This work was supported by an unrestricted grant from Research to Prevent Blindness USA to the UCLA-Department of Ophthalmology, an NEI Core Grant (P30) EY000311 to the Jules Stein Eye Institute, an R01GM137081 to M.E., and the Jules Stein Professorship Endowment to W.L.H. It was partly supported by the National Research, Development, and Innovation Fund of Hungary, NKFI K 137793. We are grateful to Dr. Christian Altenbach for providing EPR analysis programs. We thank Génesis Falcon and Mike Sawaya at the UCLA-DOE X-ray Crystallization and Crystallography Core Facilities, (supported by Department of Energy Grant DE-FC02-02ER63421) for assistance with crystallization screening, model building, and refinement. We also thank Dr. Mark Fleissner, Dr. Carlos López, and Evan Brooks for graciously providing some of the spin-labeled mutants reported herein.
Author information
Authors and Affiliations
Contributions
MC, ME, and WLH designed the research; MC, TK, DC, MDB, JPW, and ME performed the research; TK contributed new reagents; TK, DC, MDB, and JPW contributed analytic tools; MC, DC, MDB, JPW, ME, and WLH analyzed data; MC and WLH wrote the first draft of the manuscript, and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare no competing interests.
Ethical Approval
Not applicable.
Data Deposition
The atomic coordinates and structure factors for T4L 65R9/76R9 have been deposited in the Protein Data Bank, www.pdb.org (PDB ID code 8TAT).
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chen, M., Kálai, T., Cascio, D. et al. A Highly Ordered Nitroxide Side Chain for Distance Mapping and Monitoring Slow Structural Fluctuations in Proteins. Appl Magn Reson 55, 251–277 (2024). https://doi.org/10.1007/s00723-023-01618-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00723-023-01618-8