Abstract
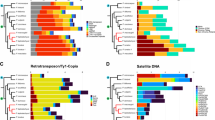
Alstroemeria species present a well-conserved and asymmetric karyotype. The genus is divided into a Chilean clade, rich in heterochromatin, and a Brazilian clade, poor in heterochromatin. We investigated the distribution of the main repetitive sequences in the chromosomes of the Brazilian species A. longistaminea (2n = 16 + 0-6B) aiming to evaluate the role played by these sequences on the structural organization of the karyotype. In situ hybridization of the three most abundant retrotransposons, corresponding to ~ 45% of the genome, was uniformly distributed. Three satellite DNA sequences, representing near half of the whole satellite fraction (1.93% of the genome), were mainly concentrated on the heterochromatin and one of them painted the whole B chromosome. Noteworthy, some satellites were located on euchromatin, either dispersed or concentrated in clusters along the chromosomes, revealing a G-band-like pattern. The two satellites that presented more C-band- and G-band-like labeling were also hybridized in situ in two other Alstroemeria species. They revealed astonishing similar patterns of distribution, indicating an unusually structural karyotype conservation among Brazilian species.





Similar content being viewed by others
References
Báez M, Vaio M, Dreissig S, Schubert V, Houben A, Pedrosa-Harand A (2019) Together but different: the subgenomes of the bimodal Eleutherine karyotypes are differentially organized. Front Plant Sci 10:1170. https://doi.org/10.3389/fpls.2019.01170
Baeza C, Schrader O, Budahn H (2007) Characterization of geographically isolated accessions in five Alstroemeria L. species (Chile) using FISH of tandemly repeated DNA sequences and RAPD analysis. Plant Syst Evol 269(1):1–14
Baeza C, Finot V, Ruiz E (2015) Comparative karyotype analysis of populations in the Alstroemeria presliana Herbert (Alstroemeriaceae) complex in Chile. Genet Mol Biol 38(2):199–204. https://doi.org/10.1590/S1415-4757382220140277
Baeza C, Finot V, Ruiz E, Carrasco P, Novoa P, Stuessy T, González A (2016) Comparative karyotypic analysis and cytotaxonomy in the Alstroemeria ligtu L. (Alstroemeriaceae) complex of Chile. Braz J Bot 39:305–313. https://doi.org/10.1590/S1415-47572010005000012
Barros e Silva AE, Marques A, dos Santos KGB, Guerra M (2010) The evolution of CMA bands in Citrus and related genera. Chromosome Research 18 (4):503-514. https://doi.org/10.1007/s10577-010-9130-2
Barros e Silva AE, Guerra M (2010) The meaning of DAPI bands observed after C-banding and FISH procedures. Biotech Histochem 85(2):115–125. https://doi.org/10.1080/10520290903149596
Belyayev A, Paštová L, Fehrer J, Josefiová J, Chrtek J, Mráz P (2018) Mapping of Hieracium (Asteraceae) chromosomes with genus-specific satDNA elements derived from next-generation sequencing data. Plant Syst Evol 304:387–396. https://doi.org/10.1007/s00606-017-1483-y
Buitendijk JH, Ramanna MS (1996) Giemsa C-banded karyotypes of eight species of Alstroemeria L. and some of their hybrids. Ann Bot 78(4):449–457
Chacón J, de Assis MC, Meerow AW, Renner SS (2012a) From East Gondwana to Central America: historical biogeography of the Alstroemeriaceae. J Biogeogr 39(10):1806–1818. https://doi.org/10.1111/j.1365-2699.2012.02749.x
Chacón J, Sousa A, Baeza CM, Renner SS (2012b) Ribosomal DNA distribution and a genus-wide phylogeny reveal patterns of chromosomal evolution in Alstroemeria (Alstroemeriaceae). Am J Bot 99(9):1501–1512. https://doi.org/10.3732/ajb.1200104
Chen R, Song W, Li X, An Z (1994) Chromosome G-banding in plants by inducing with trypsin and urea. Cell Res 4:79–87
de Assis R, Baba VY, Cintra LA, Gonçalves LSA, Rodrigues R, Vanzela ALL (2020) Genome relationships and LTR-retrotransposon diversity in three cultivated Capsicum L. (Solanaceae) species. BMC Genomics 21:237. https://doi.org/10.1186/s12864-020-6618-9
De Jeu MJ, Lasschuit J, Kuipers AGJ, Kamstra SA, Visser RGF (1997) Characterization and localization of repetitive DNA sequences in the ornamental Alstroemeria aurea Graham. Theor Appl Genet 94:982–990
De la Herrán R, Robles F, Cuñado N, Santos JL, Ruiz Rejón M, Garrido-Ramos MA, Rejón CR (2001) A heterochromatic satellite DNA is highly amplified in a single chromosome of Muscari (Hyacinthaceae). Chromosoma 110(3):197–202. https://doi.org/10.1007/s004120000115
Fillion GJ, van Bemmel JG, Braunschweig U, Talhout W, Kind J, Ward LD, Brugman W, de Castro IJ, Kerkhoven RM, Bussemaker HJ, van Steensel B (2010) Systematic protein location mapping reveals five principal chromatin types in Drosophila cells. Cell 143(2):212–224. https://doi.org/10.1016/j.cell.2010.09.009
Fonsêca A, Ferreira J, Santos TRB, Mosiolek M, Belluci E, Kami J, Gepts P, Geffroy V, Schweizer D, dos Santos KGB, Pedrosa-Harand A (2010) Cytogenetic map of common bean (Phaseolus vulgaris L.). Chromosom Res 18(4):487–502. https://doi.org/10.1007/s10577-010-9129-8
Fransz PF, Armstrong S, Hans de Jong J, Parnell LD, van Drunen C, Dean C, Zabel P, Bisseling T, Jones GH (2000) Integrated cytogenetic map of chromosome arm 4S of A. thaliana: structural organization of heterochromatic knob and centromere region. Cell 100(3):367–376. https://doi.org/10.1016/s0092-8674(00)80672-8
Gaiero P, Vaio M, Peters SA, Schranz ME, de Jong H, Speranza PR (2019) Comparative analysis of repetitive sequences among species from the potato and the tomato clades. Ann Bot 123(3):521–532. https://doi.org/10.1093/aob/mcy186
Grewal SIS, Jia S (2007) Heterochromatin revisited. Nat Rev Genet 8(1):35–46. https://doi.org/10.1038/nrg2008
Gross DS (2015) Heterochromatin: dark matter or variation on a theme? Curr Biol 25(11):R448–R469. https://doi.org/10.1016/j.cub.2015.04.003
Heitz E (1928) Das Heterochromatin der Moose. Jahrb Wiss Bot 69:762–818
Kamstra SA, Kuipers AGJ, De De Jeu MJ, Ramanna MS, Jacobsen J (1997) Physical localisation of repetitive DNA sequences in Alstroemeria: karyotyping of two species with species-specific and ribosomal DNA. Genome 40(5):652–658. https://doi.org/10.1139/g97-086
Kamstra S, Ramanna M, De Jeu M, Kuipers A, Jacobsen E (1999) Homoeologous chromosome pairing in the distant hybrid Alstroemeria aurea x A. inodora and the genome composition of its backcross derivatives determined by fluorescence in situ hybridization with species-specific probes. Heredity 82(1):69–78. https://doi.org/10.1038/sj.hdy.6884650
Kelly LJ, Renny-Byfield S, Pellicer J, Macas J, Novák P, Neumann P, Lysak MA, Day PD, Berger M, Fay MF, Nichols RA, Nichols RA, Leitch AR, Leitch IJ (2015) Analysis of the giant genomes of Fritillaria (Liliaceae) indicates that a lack of DNA removal characterizes extreme expansions in genome size. New Phytol 208(2):596–607. https://doi.org/10.1111/nph.13471
Kuipers AG, van Os DP, de Jong JH, Ramanna MS (1997) Molecular cytogenetics of Alstroemeria: identification of parental genomes in interspecific hybrids and characterization of repetitive DNA families in constitutive heterochromatin. Chromosom Res 5(1):31–39. https://doi.org/10.1023/a:1018489318300
Kuipers AG, Heslop-Harrison JS, Jacobsen E (1998) Characterisation and physical localisation of Ty1-copia-like retrotransposons in four Alstroemeria species. Genome 41(3):357–367. https://doi.org/10.1139/g98-048
Kuipers GJ, Kamstra SA, de Jeu MJ, Visser RGF (2002) Molecular characterization and physical localization of highly repetitive DNA from Brazilian Alstroemeria species. Chromosom Res 10(5):389–398. https://doi.org/10.1023/a:1016801702777
Lee Y-I, Yap JW, Izan S, Leitch IJ, Fay MF, Lee Y-C, Hidalgo O, Dodsworth S, Smulders MJM, Gravendeel B, Leitch AR (2018) Satellite DNA in Paphiopedilum subgenus Parvisepalum as revealed by high-throughput sequencing and fluorescent in situ hybridization. BMC Genomics 19(1):578. https://doi.org/10.1186/s12864-018-4956-7
Marques A, Klemme S, Houben A (2018) Evolution of plant B chromosome enriched sequences. Genes 9(10):515. https://doi.org/10.3390/genes9100515
Mata-Sucre Y, Sader M, Van-Lume B, Gagnon E, Pedrosa-Harand A, Leitch IJ, Lewis GP, Souza G (2020) How diverse is heterochromatin in the Caesalpinia group? Cytogenomic characterization of Erythrostemon hughesii Gagnon & G.P. Lewis (Leguminosae: Caesalpinioideae). Planta 252(4):49. https://doi.org/10.1007/s00425-020-03453-8
Novák P, Neumann P, Pech J, Steinhaisl J, Macas J (2013) RepeatExplorer: a Galaxy-based web server for genome-wide characterization of eukaryotic repetitive elements from next-generation sequence reads. Bioinformatics 29(6):792–793. https://doi.org/10.1093/bioinformatics/btt054
Novák P, Robledillo LA, Koblížková A, Vrbová I, Neumann P, Macas J (2017) TAREAN: a computational tool for identification and characterization of satellite DNA from unassembled short reads. Nucleic Acids Res 45(12):e111. https://doi.org/10.1093/nar/gkx257
Novák P, Neumann P, Macas J (2020) Global analysis of repetitive DNA from unassembled sequence reads using RepeatExplorer2. Nat Protoc 15(11):3745–3776. https://doi.org/10.1038/s41596-020-0400-y
Ostromyshenskii DI, Chernyaeva EN, Kuznetsova IS, Podgornaya OI (2018) Mouse chromocenters DNA content: sequencing and in silico analysis. BMC Genomics 19(1):151. https://doi.org/10.1186/s12864-018-4534-z
Park M, Park J, Kim S, Kwon J-K, Park HM, Bae IH, Yang T-J, Lee Y-H, Kang B-C, Choi D (2012) Evolution of the large genome in Capsicum annuum occurred through accumulation of single-type long terminal repeat retrotransposons and their derivatives. Plant J 69(6):1018–1029. https://doi.org/10.1111/j.1365-313X.2011.04851.x
Peters SA, Datema E, Szinay D, van Staveren MJ, Schijlen EGWM, van Haarst JC, Hesselink T, Abma-Henkens MHC, Bai Y, de Jong H, Stiekema WJ, Klein Lankhorst RM, van Ham RCHJ (2009) Solanum lycopersicum cv. Heinz 1706 chromosome 6: distribution and abundance of genes and retrotransposable elements. Plant J 58(5):857–869. https://doi.org/10.1111/j.1365-313X.2009.03822.x
Ribeiro T, Marques A, Novák P, Schubert V, Vanzela ALL, Macas J, Houben A, Pedrosa-Harand A (2017) Centromeric and non-centromeric satellite DNA organization differs in holocentric Rhynchospora species. Chromosoma 126(2):325–335. https://doi.org/10.1007/s00412-016-0616-3
Ribeiro T, Vasconcelos E, dos Santos KGB, Vaio M, Brasileiro-Vidal AC, Pedrosa-Harand A (2020) Diversity of repetitive sequences within compact genomes of Phaseolus L. beans and allied genera Cajanus L. and Vigna Savi. Chromosom Res 28(2):139–153. https://doi.org/10.1007/s10577-019-09618-w
Ribeiro T, Nascimento J, Santos A, Félix LP, Guerra M (2021) Origin and evolution of highly polymorphic rDNA sites in Alstroemeria longistaminea (Alstroemeriaceae) and related species. Genome. https://doi.org/10.1139/gen-2020-0159
Robledillo LA, Koblížková A, Novák P, Böttinger K, Vrbová I, Neumman P, Schubert I, Macas J (2018) Satellite DNA in Vicia faba is characterized by remarkable diversity in its sequence composition, association with centromeres, and replication timing. Sci Rep 8(1):5838. https://doi.org/10.1038/s41598-018-24196-3
Ruiz-Ruano FJ, López-Léon MD, Cabrero J, Camacho JPM (2016) High-throughput analysis of the satellitome illuminates satellite DNA evolution. Sci Rep 6:28333. https://doi.org/10.1038/srep28333
Samoluk SS, Chalup LM, Chavarro C, Robledo G, Bertioli DJ, Jackson SA, Seijo G (2019) Heterochromatin evolution in Arachis investigates through genome-wide analysis of repetitive DNA. Planta 249(5):1405–1415. https://doi.org/10.1007/s00425-019-03096-4
Sanso AM (2002) Chromosome studies in Andean taxa of Alstroemeria (Alstroemeriaceae). Bot J Linn Soc 138(4):451–459. https://doi.org/10.1046/j.1095-8339.2002.00019.x
Schimidt T, Heitkam T, Leidtke S, Schubert V, Gerhard M (2019) Adding color to a century-old enigma: multi-color chromosome identification unravels the autotriploid nature of saffron (Crocus sativus) as a hybrid of wild Crocus cartwrightianus cytotypes. New Phytol 222(4):1965–1980. https://doi.org/10.1111/nph.15715
Sonnhammer ELL, Durbin R (1995) A dot-matrix program with dynamic threshold control suited for genomic DNA and protein sequence analysis. Gene 167(1-2):GC1–G10. https://doi.org/10.1016/0378-1119(95)00714-8
Steflova P, Tokan V, Vogel I, Lexa M, Macas J, Novak P, Hobza R, Vyskot B, Kejnovsky E (2013) Contrasting patterns of transposable element and satellite distribution on sex chromosomes (XY1Y2) in the Dioecious Plant Rumex acetosa. Genome Biol Evol 5(4):769–782. https://doi.org/10.1093/gbe/evt049
Summer AT (2003) Chromosomes Organization and Function. Blackwell, Oxford
Van-Lume B, Mata-Sucre Y, Báez M, Ribeiro T, Huettel B, Gagnon E, Leitch IJ, Pedrosa-Harand A, Lewis GP, Souza G (2019) Evolutionary convergence or homology? Comparative cytogenomics of Caesalpinia group species (Leguminosae) reveals diversification in the pericentromeric heterochromatic composition. Planta 250(6):2173–2186. https://doi.org/10.1007/s00425-019-03287-z
Waminal NE, Ryu KB, Park BR, Kim HH (2014) Phylogeny of Cucurbitaceae species in Korea based on 5S rDNA non-transcribed spacer. Genes Genom 36:57–64. https://doi.org/10.1007/s13258-013-0141-1
Zanela L (2009) Caracterização cariotípica de quatro espécies brasileiras de Alstroemeria (Alstroemeriaceae) com as técnicas de FISH, CMA, DAPI e AgNOR. M.Sc. Doctoral thesis, Agronomic Institute of Campinas (IAC), Campinas, Brazil
Availability of data and material
Not applicable
Code availability
Not applicable
Funding
The authors thank the Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq, Brazil) for the research fellowship to MG (PQ 311924/2016-6) and TR (PDJ 150912/2017-0).
Author information
Authors and Affiliations
Contributions
MG and TR conceived and designed the study. LPF and MG performed fieldwork and taxonomy. MV and TR performed the experiments. MG and TR carried out data analysis and draft the manuscript. All authors read and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Handling Editor: Handling Editor: Sergey Mursalimov
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Online Resource 1.
Primers used for amplification of repetitive sequences from A. logistaminea genome. (DOC 49 kb)
Online Resource 2.
Dotplot of satDNAs from Alstroemeria longistaminea identified using TAREAN tools. Contigs containing at least two monomers were extracted from clusters and compared to each other. Sequences which share similarity are bolded: AloSAT6A/6B vs. AloSAT6C and AloSAT7 vs. AloSAT10 (TIF 2212 kb)
Online Resource 3.
Monomers of satDNA reconstructed from A. longistaminea genomic reads (DOC 32 kb)
Online Resource 4.
Telocentric morphology of B chromosomes of A. longistaminea revealed by AloSAT7 in an individual with five Bs (arrows). In prophase (a), AloSAT7 labels only one chromosome end, indicating they are telocentrics. In highly condensed metaphase (b), they look like metacentrics or isotelocentrics because the interchromatid space looks like a primary constriction, but AloSAT7 signal reveal the real position of the centromere. Bar in b corresponds to 10 μm (TIF 2995 kb)
Online Resource 5.
In situ hybridization of AloSAT13 in A. longistaminea labelling the heterochromatic blocks and some additional sites on termini of all chromosomes. a, DAPI; a’, labelling with AloSAT13. Bar in a’ corresponds to 10 μm (TIF 2457 kb)
Online Resource 6.
In situ distribution of AloSAT4 (a, a’), AloSAT6B (b, b’), and AloSAT10 (c, c’). Metaphase in b was too spread and was mounted based in two photos. Arrowheads point to heterochromatic blocks. Inset in b’ shows amplified B chromosome. Bar in b’ corresponds to 10 μm (TIF 15035 kb)
Online Resource 7.
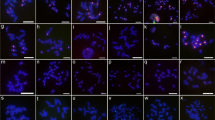
Profile of repetitive sequences mapped on B chromosomes of A. longistaminea. Chromovirus (a’), Ogre (b’), Tork (c’), AloSAT5 (d’), AloSAT6A (e’), AloSAT7 (f’), AloSAT11 (g’) and AloSAT13 (h’). Bar in h’ corresponds to 2.5 μm (TIF 902 kb)
Rights and permissions
About this article
Cite this article
Ribeiro, T., Vaio, M., Félix, L.P. et al. Satellite DNA probes of Alstroemeria longistaminea (Alstroemeriaceae) paint the heterochromatin and the B chromosome, reveal a G-like banding pattern, and point to a strong structural karyotype conservation. Protoplasma 259, 413–426 (2022). https://doi.org/10.1007/s00709-021-01681-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00709-021-01681-7