Abstract
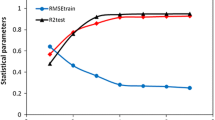
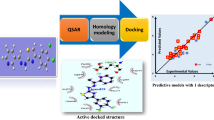
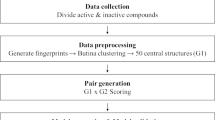
Studies on the interactions of antibiotic compounds with DNA can provide useful suggestions and guidance for the design of new and more efficient DNA-binding drugs. A quantitative structure–activity relationship (QSAR) study of the binding modes and binding affinities of the interactions between 30 antibiotic compounds and DNA was performed. A large number of descriptors that encode hydrophobic, topological, geometrical, and electronic properties were calculated to represent the structures of the antibiotic compounds. Aiming at a system with small, multidimensional samples, we utilized the genetic algorithm-support vector machine (GA-SVM) method to develop the QSAR, which can select an optimized feature subset and optimize SVM parameters simultaneously. A binary QSAR model for predicting binding mode and conventional QSAR models for predicting binding affinity were built based on the GA-SVM approach. The descriptors selected using GA-SVM represented the overall descriptor space and can account well for the binding nature of the considered dataset. The descriptors selected using the GA-SVM method were then used for developing conventional QSAR models by the artificial neural network (ANN) approach. A comparison between the conventional QSAR models using GA-SVM with those using ANN revealed that the former were much better. GA-SVM models can be useful for predicting binding modes and binding activities of the interactions of new antibiotic compounds with DNA.
Graphical abstract









Similar content being viewed by others
References
Haq I (2002) Arch Biochem Biophys 403:1
Wan KX, Shibue T, Gross ML (2000) J Am Chem Soc 122:300
Chaires JB (2006) Arch Biochem Biophys 453:26
Lerman LS (1961) J Mol Biol 3:18
Waring M (1970) J Mol Biol 54:247
Wartell RM, Larson JE, Wells RD (1974) J Biol Chem 249:6719
Kumar GS, He QY, Behr-Ventura D, Tomasz M (1995) Biochemistry 34:2662
Qu XG, Chaires JB (2001) J Am Chem Soc 123:1
Gopal M, Shenoy S (2003) J Photochem Photobiol B 72:69
Ibrahim MS (2001) Anal Chim Acta 443:63
Leng FF, Priebe W, Chaires JB (1998) Biochemistry 37:1743
Wang SF, Peng TZ, Yang CF (2003) J Biochem Biophys Methods 55:191
Wang J, Ozsoz M, Cai XH, Rivas G, Shiraishi H, Grant DH, Chicharro M, Fernandes J, Palecek E (1998) Bioelectrochem Bioenerg 45:33
Agrawal P, Barthwal SK, Barthwal R (2009) Eur J Med Chem 44:1437
Evstigneev MP, Mykhina VY, Davies DB (2005) Biophys Chem 118:118
Barthwal R, Sharma U, Srivastava N, Jain M, Awasthi P, Kaur M, Barthwal SK, Govil G (2006) Eur J Med Chem 41:27
Mallena S, Lee MPH, Bailly C, Neidle S, Kumar A, Boykin DW, Wilson WD (2004) J Am Chem Soc 126:13659
Haj HTB, Salerno M, Priebe W, Kozlowski H, Garnier-Suillerot A (2003) Chem Biol Interact 145:349
Miao Y, Lee MPH, Parkinson GN, Batista-Parra A, Ismail MA, Neidle S, Boykin DW, Wilson DW (2005) Biochemistry 44:14701
Teulade-Fichou MP, Carrasco C, Guittat L, Bailly C, Alberti P, Mergny GL, David A, Lehn JM, Wilson WD (2003) J Am Chem Soc 125:4732
Li VS, Choi D, Wang Z, Jimenez LS, Tang MS, Kohn H (1996) J Am Chem Soc 118:2326
Guan Y, Shi R, Li XM, Zhao MP, Li YZ (2007) J Phys Chem B 111:7336
Tian L, Wei WZ, Mao Y (2004) Clin Biochem 37:120
Lu XQ, Wang L, Liu HD, Chen J (2007) Talanta 73:444
Chen J, Lu XQ (2009) Talanta 79:129
Ismail I, Francois P, Elodie D, Olivier B, Andre M (2007) Bioorg Med Chem 15:4256
Menard PR, Lewis RA, Mason JS (1998) J Chem Inf Comput Sci 38:497
Menard PR, Mason JS, Morize I, Bauerschmidt S (1998) J Chem Inf Comput Sci 38:1204
Todeschini R, Consonni V (2000) Handbook of molecular descriptors. Wiley-VCH, Weinheim
Consonni C, Todeschini R, Pavan M (2002) J Chem Inf Comput Sci 42:682
Consonni V, Todeschini R, Pavan M, Gramatica P (2002) J Chem Inf Comput Sci 42:693
Kumar CV, Asuncion EH (1993) J Am Chem Soc 115:8541
Mazur S, Tanious FA, Ding D, Kumar A, Boykin DW, Simpson IJ, Neidle S, Wilson WD (2000) J Mol Biol 300:321
Rahimian M, Kumar A, Say M, Bakunov SA, Boykin DW, Tidwell RR, Wilson WD (2009) Biochemistry 48:1573
Banerjee D, Pal SK (2008) J Phys Chem B 112:1016
Athri P, Wilson WD (2009) J Am Chem Soc 131:7618
HyperChem Release 7, HyperCube Inc. (2002), http://www.hyper.com
Long W, Liu P, Li X, Xu Y, Yu J, Ma S, Yu L, Zou Z (2009) J Chemom 23:304
Li ZR, Han LY, Xue Y, Yap CW, Li H, Jiang L, Chen YZ (2007) Biotechnol Bioeng 97:389
Cortes C, Vapnik V (1995) Mach Learn 20:273
Wang WJ, Xu ZB, Lu WZ, Zhang XY (2003) Neurocomputing 55:643
Chang CC, Lin CJ (2001) http://www.csie.ntu.edu.tw/cjlin/libsvm
Fang SF, Wang MP, Qi WH, Zheng F (2008) Comput Mater Sci 44:647
Pai PF, Hong WC (2005) Electric Power Syst Res 74:417
Pourbasheer E, Riahi S, Ganjali MR, Norouzi P (2009) Eur J Med Chem 44:5023
Dolatabadi M, Nekoei M, Banaei A (2010) Monatsh Chem 141:577
Goodarzi M, Freitas MP, Wu CH, Duchowicz PR (2010) Chemometr Intell Lab Syst 101:102
Li ZC, Zhou XB, Lin YR, Zou XY (2008) Amino Acids 35:581
Li TH, Mei H, Cong PS (1999) Chemometr Intell Lab Syst 45:177
Habibi-Yangjeh A (2009) Monatsh Chem 140:523
Corwin H, Rajeshwar PV (2009) Mol Pharmaceutics 6:3849
Eduardo BM, Marcia MCF (2009) Eur J Med Chem 44:3577
Acknowledgments
This work was supported by the National Natural Science Foundation of China (Nos. 20945003, 21005063), and the Natural Science Foundation of Gansu (No. 096RJZA121).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zhou, X.B., Han, W.J., Chen, J. et al. QSAR study on the interactions between antibiotic compounds and DNA by a hybrid genetic-based support vector machine. Monatsh Chem 142, 949–959 (2011). https://doi.org/10.1007/s00706-011-0493-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00706-011-0493-7