Abstract
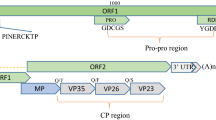
A novel virus infecting a Paris polyphylla var. yunnanensis plant, tentatively named "Paris polyphylla chlorotic mottle virus" (PpCMV), was discovered in the city of Lijiang, Yunnan Province, China. Its genome consists of 6384 nucleotides (nt), excluding the 3′-terminal poly(A) tail, and contains two open reading frames: ORF1 and ORF2. ORF1 is 6150 nt in length, encoding a large 2050-aa polyprotein with at least two conserved regions encoding a replication-associated protein and a coat protein, the latter of which is located at the 3′ end of ORF1. ORF2, consisting of 1185 nt, is located within ORF1 but has a different reading frame. It encodes a 394-aa-long putative movement protein. Phylogenetic analysis based on amino acid sequences revealed that the newly discovered virus exhibited the closest relationship to Hobart betaflexivirus 1 and rhodiola betaflexivirus 1, both of which belong to the genus Capillovirus, sharing 48.8% and 36.5% amino acid sequence identity, respectively, in the structural protein. This is the first report of the complete genome sequence of PpCMV in China.

Similar content being viewed by others
Availability of data and material
The nucleotide sequences reported in this manuscript have been deposited in the GenBank database under accession number MW822017 and are in the electronic supplementary material.
Bibliography
Mastuda H, Pongpiriyadacha Y, Morikawa T, Kishi A, Kataoka S, Yoshikawa M (2003) Protective effects of steroid saponins from paris polyphylla var. yunnanensis on ethanol- or indomethacin-induced gastric mucosal lesions in rats: structural requirement for activity and mode of action. Bioorg Med Chem Lett 13:1101–1106. https://doi.org/10.1016/S0960-894X(03)00052-0
He J, Zhang S, Wang H, Chen CX, Chen SF (2006) Advances in studies on and uses of paris polyphylla var. yunnanensis (Trilliaceae). Acta Bot Yunnanica 28(3):271–276. https://europepmc.org/article/CBA/619288
Li J, Zhao JL, Xu LJ, Zhou LG, Li XL, Wang JG (2008) Endophytic fungi from rhizomes of paris polyphylla var. yunnanensis. World J Microbiol Biotechnol 24(5):733–737. https://doi.org/10.1007/s11274-007-9531-3
Yang Y, Yang SC, Zhao J, Udikeri S, Liu T (2015) Microbial diversity in paris polyphylla var. yunnanensis rhizomes of varying ages. Genet Mol Res 14(4):17612–17621. https://doi.org/10.4238/2015.December.21.34
Wei ZX, Guo DQ, Li HF, Ding B, Zhang J, Zhou N, Yu J (2015) Photosynthetic parameters and physiological indexes of paris polyphylla var. yunnanensis influenced by arbuscular mycorrhizal fungi. Chin 40(20):3945–3952. https://doi.org/10.4268/cjcmm20152010
Zhou N, Ding B, Feng Y, Qi WH, Zhang H, Guo DQ, Xiang J (2015) Effects of mycorrhizal colonization and medicine quality of paris polyphylla var. yunnanensis inoculated by different foreign AM fungi species. Chin 40(16):3158–3167. https://doi.org/10.4268/cjcmm20151608
Liu T, Liao QH, Yu FQ, Zi SH, Tian SH, Fan LY (2021) Plant growth-promoting activities of bacterial endophytes isolated from the medicinal plant pairs polyphylla var. yunnanensis. World J Microbiol Biotechnol 38(1):15. https://doi.org/10.1007/s11274-021-03194-0
Du HH, Zhu FR, Yang M, Guo DQ, Zhao SX, Li QT, Zhou N (2021) Isolation and identification of phosphatolytic bacteria in paris polyphylla var. yunnanensis. Chin 46(4):915–922. https://doi.org/10.19540/j.cnki.cjcmm.20201120.102
Wen GS, Yang LY, Anane RF, Chen ZL, Yang YH, Chen L, Sun Y, Zhao MF (2019) First report of pepper mild mottle virus in paris polyphylla var. yunnanensis in china. Plant Dis 103(12):3289. https://doi.org/10.1094/PDIS-03-19-0445-PDN
Lan PX, Zhao JR, Zhou YL, Li YY, Shen DC, Liao QC, Li RH, Li F (2018) Complete genome sequence of paris mosaic necrosis virus, a distinct member of the genus potyvirus. Arch Virol 163(3):787–790. https://doi.org/10.1007/s00705-017-3649-x
Chen L, Anane RF, Wang Z, Yang LY, Chen ZL, Wen GS, Zhao MF (2020) Whole-genome sequence analysis of paris virus 1: a novel member of the genus Potyvirus infecting Paris polyphylla var. yunnanensis. Arch Virol 165(4):985–988. https://doi.org/10.1007/s00705-020-04560-3
Chen L, Anane RF, Wang Z, Chen ZL, Gao L, Wen GS, Zhao MF (2021) Characterization of a novel Tombusviridae species isolated from paris polyphylla var. yunnanensis. Arch Virol 166(11):3199–3205. https://doi.org/10.1007/s00705-021-05191-y
Chen ZL, Chen L, Anane RF, Wang Z, Gao L, Li SY, Wen GS, Yu DH, Zhao MF (2022) Complete genome sequence of a novel mitovirus detected in Paris polyphylla var. yunnanensis. Arch Virol 167(2):645–650. https://doi.org/10.1007/s00705-021-05339-w
Li J, Zhang MQ, Du YN, Chen S, Zhang ZK, Zhao LH (2022) First report of tobacco mosaic virus infection in paris polyphylla var. yunnanensis in China. Plant Dis PMID 36018555. https://doi.org/10.1094/pdis-07-22-1539-pdn
Wang Z, Anane RF, Chen ZL, Gao L, Li SY, Chu BF, Wen GS, Zhao MF (2022) Complete genome sequence analysis of paris alphapartitivirus 1: a novel member of the genus Alphapartitivirus infecting paris polyphylla var. yunnanensis. Arch Virol 167(11):2365–2370. https://doi.org/10.1007/s00705-022-05531-6
Zhang BX, Li QN, Hu JY, Zhang L, Dong X, Ji PZ, Dong JH (2023) Complete genome sequence analysis of a new potyvirus isolated from paris polyphylla var. yunnanensis. Arch Virol 168(2):43. https://doi.org/10.1007/s00705-022-05655-9
Wei CB, Chen J, Zhang QY, Shi YH, Lin L, Zheng HY, Adams MJ, Chen JP (2005) A new potyvirus from thunberg fritillary (Fritillaria thunbergii Miq.) in zhejiang, china. Arch Virol 150(7):1271–1280. https://doi.org/10.1007/s00705-005-0519-8
Chen BH, Wu DL, Zheng HY, Li GZ, Cao YH, Chen JP, Yan F, Song XM, Lin L (2021) Complete genome sequence of passiflora virus Y infecting passion fruit in China. Arch Virol 166(5):1489–1493. https://doi.org/10.1007/s00705-021-05013-1
Kumar S, Stecher G, Tamura K (2016) MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33(7):1870–1874. https://doi.org/10.1093/molbev/msw054
Liu QY, Yang L, Xuan ZY, Wu JX, Qiu YJ, Zhang S, Wu D, Zhou CY, Cao MJ (2020) Complete nucleotide sequence of loquat virus A, a member of the family Betaflexiviridae with a novel genome organization. Arch Virol 165(1):223–226. https://doi.org/10.1007/s00705-019-04444-1
Li ZT, Wang HX, Zhao RB, Zhang Z, Xia ZH, Zhai JL, Huang X (2020) Complete genome sequence of a novel capillovirus infecting hevea brasiliensis in China. Arch Virol 165(1):249–252. https://doi.org/10.1007/s00705-019-04459-8
Bak SM, Jeong WY, Kim MH, Lee SH, Kim ST, Lee ES, Kim CK, Kim SY (2023) Complete genome sequence of a tentative novel capillovirus isolated from gerbera jamesonii. Arch Virol 168(4):117. https://doi.org/10.1007/s00705-023-05730-9
Alas T, Baloglu S, Caglar BK, Gunes A (2019) Detection and characterization of citrus tatter leaf virus (CTLV) and citrus yellow vein clearing virus (CYVCV) in citrus trees from Cyprus. Saudi J Biol Sci 26(5):995–998. https://doi.org/10.1016/j.sjbs.2019.02.001\
Marais A, Faure C, Theil S, Candresse T (2018) Molecular characterization of a novel species of capillovirus from Japanese Apricot (Prunus mume). Viruses 10(4):144. https://doi.org/10.3390/v10040144
Acknowledgements
We thank all the reviewers who participated in the review, as well as MJEditor (www.mjeditor.com) for providing English editing services during the preparation of this manuscript. This work was financially supported by the National Key Research and Development Program of China (2019YFD1001800), the Science and Technology Agency Program of Yunnan Province (202101BC070003-51), and the K. C. Wong Magna Fund of Ningbo University.
Funding
This work was financially supported by the National Key Research and Development Program of China (2019YFD1001800), the Science and Technology Agency Program of Yunnan Province (202101BC070003-51), and the K. C. Wong Magna Fund of Ningbo University.
Author information
Authors and Affiliations
Contributions
Conceptualization: Lin Lin, Xuemei Song. Methodology: Binghua Chen, Yuhao Cao, Xuemei Song. Data analysis: Jiejun Peng, Junmin, Li. Formal analysis and investigation: Binghua Chen, Xuemei Song, Lin Lin. Writing – original draft preparation: Binghua Chen, Hongying Zheng, Jianping Chen, Fei Yan. Writing – review and editing: Binghua Chen, Xuemei Song, Lin Lin. Funding acquisition: Jianping Chen. Resources: Lin Lin, Qiongji He, Mengying Hua, Yueyan Yin, Jiejun Peng. Supervision: Jianping Chen, Fei Yan
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest/competing interests.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Communicated by Jesús Navas-Castillo
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic Supplementary Material
Below is the link to the electronic supplementary material
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
He, Q., Chen, B., Zheng, H. et al. Complete genome sequence of Paris polyphylla chlorotic mottle virus infecting Paris polyphylla var. yunnanensis in southwest China. Arch Virol 168, 292 (2023). https://doi.org/10.1007/s00705-023-05896-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00705-023-05896-2