Abstract
Recombinant vaccinia viruses harboring the complete hemagglutinin (HA) or neuraminidase (NA) genes from the influenza A/Anhui/1/2013 (H7N9) virus were constructed (rVac-H7 HA and rVac-N9 NA viruses). The HA and NA proteins were expressed in the cytoplasm and on the plasma membrane of thymidine-kinase-negative (TK-) cells infected with these recombinant viruses. Only one form of the HA protein was expressed in infected TK- cells, with a molecular weight (MW) of 75 kDa, but three forms were found when the culture medium was supplemented with trypsin (MWs of 75, 50 and 27 kDa), which was similar to what was found in Madin-Darby canine kidney (MDCK) cells infected with reverse genetic (rg) influenza viruses carrying HA genes of H7N9 virus origin. One form of hyperglycosylated NA protein with a MW of 75 kDa was produced in rVac-N9-NA-virus-infected TK- or MDCK cells. The MW decreased to 55 kDa after deglycosylation. The hyperglycosylated recombinant NA protein demonstrated sialidase activity in a fetuin-based neuraminidase assay. The rVac-H7 HA and rVac-N9 NA viruses elicited significantly higher anti-HA and anti-NA antibody titers in BALB/c mice that were immunized once than in ICR mice. The anti-HA and anti-NA antibodies showed activity against homosubtypic HA or NA, but not against heterosubtypic HA or NA, as determined by hemagglutination-inhibition and microneutralization assays for anti-HA antibodies and neuraminidase-inhibition and replication-inhibition assays for anti-NA antibodies. Taken together, our data demonstrated immunobiological properties of recombinant HA and NA proteins that might be useful for vaccine development.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In March 2013, a human case of influenza A (H7N9) was first reported to the World Health Organization from the People’s Republic of China [1]. The event was followed by three outbreak waves, which occurred in spring 2013, winter 2013/14, and winter 2014/15 [2]. In addition, a rise in the number of cases in October 2015 predicted the occurrence of a fourth outbreak wave [3]. At present, the virus is confined solely to China with exceptions of travel-related cases in Malaysia [4] and Canada [5]. As of 29 November 2015, the cumulative number of confirmed human cases has reached 679, including 272 deaths in China [6].
Phylogenetic analysis suggested that this virus might be a reassortant with the haemagglutinin (HA) gene derived from an influenza H7N3 virus from Eurasian wild birds, the neuraminidase (NA) gene from either influenza H11N9 or H2N9 viruses from wild birds, and the six internal genes (PB2, PB1, PA, NP, M, and NS) from H9N2 viruses in poultry in China [7].
There is concern about the pandemic potential of the emerging H7N9 virus for several reasons. This avian-virus is able to bind both the avian type receptor α-2,3 galactose-linked sialic acid (SA α-2,3 gal) and the human-type receptor (SA α-2,6 gal) on epithelial cells lining the human upper respiratory tract [8]. The presence of the amino acid substitutions Glu627Lys and Asp701Asn in the PB2 protein suggested that the virus has adapted to grow in mammalian cells [9]. Moreover, pre-existing neutralizing antibodies specific for these viruses are absent in the general population [10], and current seasonal influenza vaccines cannot induce cross-protection [11].
In order to develop a vaccine, the HA protein of H7N9 viruses has been expressed in many virus vectors, including baculovirus [12], modified vaccinia Ankara (MVA) virus [13], Newcastle disease virus [14], and parainfluenza virus 5 [15]. These studies demonstrated high hemagglutination-inhibition (HAI) and/or neutralization antibody titers in immunized animals, and in particular, protection against lethal virus challenge in some models. Nevertheless, information about the biological properties of the H7 HA is limited [8, 11], and the immunological properties of N9 NA are not understood.
Here, we constructed recombinant vaccinia (rVac) viruses with the complete HA or NA genes derived from the H7N9 influenza virus and determined the immunobiological properties of the proteins expressed in infected thymidine-kinase-negative (TK-) cells. The antigenicity of the recombinant HA and NA proteins was determined by immunofluorescence (IF) and Western blot (WB) assays using reference sera, and their immunogenicity was assessed by inoculating two species of mice (inbred BALB/c and outbred ICR) with recombinant viruses and assaying for a specific antibody response. Various strains of wild-type and reverse genetics (rg)-derived influenza viruses belonging to different HA and NA genetic groups [16, 17] were employed as test viruses to demonstrate common and cross-reactive antibodies using various serological assays, including HAI and micro-neutralization (MN) assays for anti-HA antibody and neuraminidase-inhibition (NAI) assay and replication-inhibition (RI) assay for the anti-NA antibody.
Materials and methods
Viruses
Influenza viruses employed in this study included the following human viruses: the 2009, pandemic A/California/07/2009 virus (CA-07 virus) from the National Institute for Biological Standards and Control, UK, and A/Thailand/NMA-1/2011 (A/Texas/50/2012 (H3N2)-like virus), a human virus of avian origin A/Laos/Nong Khai 1/2007 (H5N1) virus clade 2.3.4, and the low-pathogenic avian influenza (LPAI) viruses: A/Chicken/HK/G9/1997 (H9N2), A/Ostrich/Zinbabwe/222/1996 (H7N1) and A/Duck/Shan Tou/461/2000 (H4N9), kindly provided by Prof. Robert G. Webster and Dr. Richard Webby, St. Jude Children’s Research Hospital, Memphis, USA. Human viruses were grown in Madin-Darby canine kidney (MDCK) cells maintained in Eagle’s minimum essential medium (EMEM) (Gibco, Grand Island, NY, USA) in the presence of trypsin treated with tosyl phenylalanyl chloromethyl ketone (TPCK), and avian viruses were grown in embryonated chicken eggs.
Vaccinia virus, Lister strain, was kindly provided by the Thai Government Pharmaceutical Organization. The virus was propagated in TK- cells maintained in Dulbecco’s modified Eagle medium (DMEM) (Gibco) supplemented with 2 % fetal bovine serum (Gibco).
Construction of recombinant vaccinia virus
Two kinds of recombinant vaccinia viruses (rVac-H7 HA or rVac-N9 NA virus) carrying HA or NA gene inserts derived from the H7N9 virus were constructed. The recombinant pCI plasmids harboring the complete H7 HA or N9 NA genes derived from A/Anhui/1/2013 (H7N9) virus were kindly provided by Dr. Yuelong Shu, Chinese National Influenza Center. The HA and NA genes were amplified by polymerase chain reaction (PCR) using the primer pairs SXm-HA-1 (5′-TAT TCC CGG GAG AGC AAA AGC AGG GG-3′) and SXm-NS-890R (5′-ATA TCC CGG GTA TTA GTA GAA ACA AGG GTG TTT T-3′) for the HA gene and SXm-NA-1 (5′-TAT TCC CGG GAG AGC AAA AGC AGG AGT-3′) and SXm-NA-1413R (5′-ATA TCC CGG GTA TTA GTA GAA ACA AGG AGT TTT TT-3′) for the NA gene. These two primer pairs were derived from those designed by Hoffmann, et al. [18] and modified in our laboratory by adding overhangs at both the 5′ and 3′ termini with XmaI restriction sites (C^CCGGG) to generate a sticky-end PCR product for an ease of DNA ligation. The amplified products were then cloned into the pSC11 plasmid vector (kindly provided by Prof. Bernard Moss, National Institute of Allergy and Infectious Diseases, Maryland, USA) under the 7.5 k promoter. The pSC11-HA or pSC11-NA recombinant plasmids were mixed with DMRIE-C transfection reagent (Invitrogen, Carlsbad, CA, USA) in DMEM and used to transfect TK-cells that had been pre-infected with wide-type vaccinia virus at a multiplicity of infection (MOI) of 0.01 plaque-forming units (PFU)/ml for 2 hours. The HA or NA gene flanked by TK R and TK L sequences in the vector plasmids was inserted into the parental vaccinia virus genome by homologous recombination with the viral TK gene. As a result, the recombinant virus harboring the HA or NA gene insert lost its ability to produce thymidine kinase (TK- phenotype) but gained the ability to produce β-galactosidase. The transfected culture was further incubated for 2 days to allow virus replication. The recombinant vaccinia virus was distinguished from the parental TK+ vaccinia virus by plaque selection on TK- cell monolayers in the presence of bromodeoxyuridine (BrdU) and X-gal, in which the plaques produced by cells infected with the recombinant vaccinia virus appeared blue. Plaque purification was performed three times using low-melting-point agarose containing BrdU and X-gal.
Construction of reverse genetic viruses
Reassortant influenza viruses were rescued by reverse genetics based on the pHW2000 plasmid vector system comprising eight recombinant plasmids carrying each gene of A/Puerto Rico/8/1934 (H1N1) virus (PR8 virus). These recombinant plasmids were kindly provided by Prof. Robert G. Webster.
In the construction process, the recombinant pCI plasmids carrying an HA or NA DNA insert were amplified by PCR using the universal primers designed by Hoffmann et al. [18], and the amplified products were cloned into the pHW2000 plasmid vector. Subsequently, eight recombinant plasmids carrying the HA and/or NA of the H7N9 virus and the other gene segments of the PR8 virus were used to co-transfect a co-culture of human embryonic kidney cells (HEK-293T) and MDCK cells. The transfection protocol has been described previously [19]. At 3 to 5 days post-transfection, the culture supernatants were inoculated into 10-day-old embryonated eggs and incubated at 33 °C for 72 hours for virus propagation.
A total of five reassortant influenza viruses used in this study included the rescued PR8 virus, and four rg viruses that contained an HA and/or NA gene from the donor viruses in the backbone of PR8 virus, i.e., rgH7N9 HANA (6 + 2), rgH7 HA (7 + 1), rgN9 NA (7 + 1) and the rgH5N1 HANA (6 + 2) virus in which the multiple basic amino acids RERRRK/R at the HA cleavage site were changed to IET/R, the monobasic amino acid sequence at the cleavage site of an avirulent H6 subtype.
Animals
Both inbred BALB/c and outbred ICR mice at five weeks of age were used in the study. Each strain of mice was divided into five groups, and each group comprising five mice was injected intraperitoneally with rVac-H7 HA, rVac-N9 NA, rVac-pSC11, or wild-type vaccinia virus at inoculum dose of 107 PFU/mouse, and the fifth group was for mock injection. PolyI:C at a concentration of 10 μg per mouse was used as an adjuvant. The animals were handled by experienced veterinarians. The physical condition of the animal was monitored once a day until the end of the experiment. All animals were healthy. None was ill or died prior to the experimental endpoint. The animals were anesthetized using isoflurane inhalation and bled by cardiac puncture at six weeks after immunization. Death of the animals was confirmed by physical examination.
Indirect immunofluorescence (IF) assay
TK- cells infected with rVac-HA or rVac-NA virus were investigated for expression and localization of the HA or NA protein by indirect IF. The slides of infected cell deposits were incubated with primary antibody (goat antiserum against H7 HA or N9 NA, kindly provided by Prof. Robert G. Webster) for one hour at 37 °C, followed by secondary antibody (fluorescein-isothiocyanate-conjugated rabbit anti-goat Ig) (DakoCytomation, Glostrup, Denmark) for one hour at 37 °C. The slides were counterstained with Hoechst 33342 trihydrochloride trihydrate (Invitrogen, Paisley, Scotland, UK) and examined for the presence of fluorescent cells under a laser scanning confocal microscope (LSM 510 Meta, Zeiss, Jena, Germany).
Western blot (WB) assay
A WB assay was performed following the Laemmli method [20] to determine the molecular weight (MW) of H7 HA and N9 NA proteins from various sources, i.e., TK- cells infected with rVac-H7 HA or rVac-N9 NA virus, MDCK cells infected with rgH7N9 HANA (6 + 2) virus, and also recombinant H7 HA (rBV-H7 HA, BEI Resources, USA, catalog no. NR-44081) and the N9 NA proteins (rBV-N9 NA, BEI Resources, catalog no. NR-44082) expressed in a baculovirus-insect cell system. The infected cell pellets were lysed with RIPA buffer containing 0.1 % sodium dodecyl sulphate (SDS), 1 % Triton X-100, 0.5 % sodium deoxycholate, 50 mM TrisCl, pH 7.5, and 150 mM NaCl. The solubilized proteins or reference antigens were mixed with 4× reducing sample buffer containing 8 % SDS, 8 % β-mercaptoethanol, 250 mM Tris Cl, pH 6.8, 0.4 % bromophenol blue and 40 % glycerol and boiled for 10 minutes prior to analysis by 10 % denaturing discontinuous sodium dodecyl sulphate-polyacrylamide gel electrophoresis (SDS-PAGE). The protein bands present in the gel were blotted onto nitrocellulose membranes (Protran, Whatman, GmbH, Germany) using a Trans-Blot SD semidry transfer cell (Bio-Rad, Hercules, CA, USA). The blotted membrane was blocked with 1 % bovine serum albumin (BSA) in Tris-buffered saline plus 0.1 % Tween-20 (Sigma-Aldrich, St. Louis, MO, USA) and stained with goat antiserum against H7 HA or N9 NA overnight at 4 °C. The membranes were treated with horseradish peroxidase (HRP)-conjugated-rabbit anti-goat Ig (Dako Corp., Santa Barbara, CA, USA) for 2 hours at room temperature, followed by 3′-diaminobenzidine (Sigma-Aldrich) as the chromogenic substrate.
Deglycosylation
A cell lysate containing NA protein from a different expression system was denatured with 0.1 volume of glycoprotein denaturing buffer and incubated for 10 minutes at 100 °C, followed by the addition of 0.1 volume of G7 reaction buffer, 0.1 volume of NP40, and 0.1 volume of peptide-N-glycosidase F (PNGase F; New England Biolabs, Herts, UK) and further incubated for 1 hour at 37 °C. The digested products were subjected to WB assay for comparison to the undigested cell lysate.
Hemagglutination-inhibition (HAI) assay
The HAI assay was conducted in duplicate in 96-well microtiter plates with “V” shaped wells, as described previously [21]. The test sera were mixed with receptor-destroying enzyme (RDE) (Denka Seiken, Tokyo, Japan) at a ratio of 1:3 for 18 hours at 37 °C, followed by heat inactivation for 45 minutes at 56 °C and absorption with goose erythrocyte suspension for 60 minutes at 4 °C. The treated sera were serially diluted twofold from 1:10 to 1:1280. In each well, a 25-μl volume of the diluted serum was incubated with a 25-μl volume containing 4 HA units of the test virus for 30 minutes at room temperature, followed by addition of a 50-μl volume of 0.5 % goose erythrocyte suspension. Virus back-titration was performed in parallel. The end result was read after the reaction plate was incubated for 30 minutes at 4 °C. The antibody titer was defined as the reciprocal of the highest serum dilution that completely inhibited the hemagglutination reaction.
Neuraminidase-inhibition (NAI) assay
The NAI assay was conducted following the protocol described by Couzens et al. [22]. The test viruses were titrated by neuraminidase assay prior to NAI antibody determination. The virus samples were serially diluted twofold with sample diluent containing 1 % BSA and 0.5 % Tween-20 in Dulbecco’s phosphate-buffered saline (Sigma-Aldrich) from 1:2 to 1:2048. Then, a 50-μl volume of the diluted virus or culture supernatants, together with a 50 μl volume of the sample diluent was transferred to a well of a 96-well plates, pre-coated with fetuin (Sigma-Aldrich) and incubated for 17 hours at 37 °C. Subsequently, peanut-agglutinin-conjugated horseradish peroxidase (PNA-HRP) (Sigma-Aldrich) was added, and the plate was further incubated at room temperature for 2 hours in the dark. The assay employed o-phenylenediamine dihydrochloride (OPD) as the chromogenic substrate. The optical density was determined by reading the reaction plate with a spectrophotometer at wavelength of 490 nm. The absorbance values corresponding to the NA enzyme activity were observed to form a sigmoidal regression curve. The virus dilution that yielded 90–95 % of the maximum signal was chosen for further use in the NAI assay.
To measure the NAI antibody titer, each serum sample was pretreated with RDE followed by heat inactivation as described for the HAI assay, and then serially diluted twofold with sample diluent from 1:10 to 1:20,480. A 50-μl volume of each diluted serum was added in duplicate wells of a fetuin-coated plate, followed by a 50-μl volume of the virus at the working dilution. At least four wells containing only virus in sample diluent were used as virus controls, and at least four wells containing only sample diluent were used as a background control. The reaction plates were incubated for 16-18 hours at 37 °C prior to washing and incubation with PNA-HRP for 2 hours at room temperature. OPD was used as the chromogenic substrate as described above. The end result was calculated by subtracting the mean O.D. value of the test wells from the mean O.D. value of the background wells. The NAI antibody titer was defined as the reciprocal of the highest serum dilution that yielded 50 % inhibition of the virus control.
Micro-neutralization (MN) assay
The protocol for the MN assay was identical to that described by the World Health Organization [23] with the modification that we did not remove the virus-serum mixture from the reaction well. Briefly, each serum sample was pre-treated with RDE, followed by heat inactivation as described above for the HAI assay. The treated serum was then serially diluted twofold in EMEM from 1:10 to 1:1280 in duplicate. Thereafter, a 60-µl volume of each dilution was mixed with a 60-µl volume of the test virus at the working concentration of 200 TCID50/100 μl and incubated for 2 hours at 37 °C. Then, 100 μl of serum-virus mixture was transferred to a well containing a MDCK cell monolayer and maintained in EMEM supplemented with TPCK-treated trypsin for 3 days at 37 °C. The presence of viruses in the culture supernatant was determined by hemagglutination assay using a 0.5 % goose erythrocyte suspension. The wells that showed no hemagglutination indicated that the test antiserum protected the cell monolayers from virus infection. The reciprocal of the highest serum dilution that completely inhibited hemagglutination was defined as the antibody titer.
Replication-inhibition (RI) assay
An RI assay was employed to measure anti-NA antibody that is able to block virus replication. The assay protocol and the endpoint determination method for anti-NA antibody titer was similar to that used in the MN assay described above.
Results
Expression and localization of rVac-H7 HA and rVac-N9 NA proteins
The expression of recombinant HA and NA proteins in TK- cells infected with rVac-H7 HA or rVac-N9 NA virus was examined by confocal IF assay using goat antiserum against H7 HA or N9 NA. TK- cells infected with rVac-pSC11 virus were used as a negative control. Both rVac-H7 HA and rVac-N9 NA proteins localized in the cytoplasm and on the plasma membrane of the infected cells as visualized by confocal microscopy. No fluorescent signal was observed in the rVac-pSC11-virus-infected TK- cell controls (Fig. 1).
Expression and localization of recombinant H7 HA and N9 NA proteins in TK- cells infected with rVac-H7 HA or rVac-N9 NA virus, detected by confocal microscopy. TK- cells infected with rVac-pSC11 virus vector were used as a negative control. Differential interference contrast images show the recombinant H7 HA or N9 NA proteins in fluorescent green. Nuclei were stained blue with Hoechst
Characterization of H7 HA protein produced from different virus-host cell systems
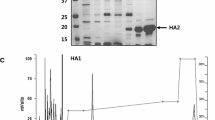
The expression of recombinant H7 HA in TK- cells was measured and compared to the expression in MDCK cells infected with rgH7N9 HANA (6 + 2) virus and to commercial HA protein expression in a baculovirus-insect cell system by WB assay. Uninfected MDCK and TK- cells were used as negative controls, as well as TK- cells infected with rVac-pSC11 virus or parental vaccinia virus. The result showed that HA protein expressed in infected TK- cells had a MW of approximately 75 kDa, corresponding to the HA0 precursor. When cells were cultured in medium containing TPCK-treated trypsin, two more proteins were detected at MWs of 50 and 27 kDa corresponding to the HA1 and HA2 cleavage products, respectively. Protein products at 75, 50 and 27 kDa were also found in MDCK cells infected with rgH7N9 HANA (6 + 2) virus. The recombinant H7 HA protein from baculovirus-insect cells had MW of 60 kDa, corresponding to HA0 precursor protein, as indicated in the product brochure. No band of specific protein was seen in the negative controls (Fig. 2).
Western blot assay displaying different MWs of H7 HA proteins expressed in various virus-host cell systems: 60 kDa for recombinant HA protein expressed in a baculovirus-insect cell system (rBV-H7 HA) and 75 kDa for recombinant protein expressed in TK- cells infected with rVac-H7 HA virus and maintained in culture medium without TPCK-treated trypsin. Three forms of HA proteins at MWs of 75, 50 and 27 kDa are expressed in TK- cells infected with rVac-H7 HA virus and maintained in culture medium supplemented with trypsin-TPCK. Similarly, those three forms of native H7 HA proteins are produced in MDCK cells infected with rgH7N9 HANA (6 + 2) virus
Characterization of N9 NA protein produced from different virus-host cell systems
When the N9 NA protein was expressed in TK- cells, MDCK cells infected with rVac-N9 NA virus, MDCK cells infected with rgH7N9 HANA (6 + 2) virus, or that from a baculovirus-insect cell system, a protein with a MW of 75, 75, 55 and 50 kDa, respectively, was detected (Fig. 3 a and b). The glycosylation pattern of these NA proteins was investigated by treatment with PNGase F glycosidase enzyme. There was no change in the MW of the 55-kDa NA protein produced in the rgH7N9 HANA (6 + 2)-infected MDCK cells after PNGase F digestion (Fig. 3a), while the MW of NA protein expressed in the rVac-N9 NA virus-infected TK- and MDCK cells decreased from 75 to 55 (Fig. 3a and b), and the MW of NA protein expressed in the baculovirus-insect cell system decreased slightly from 50 to 46 kDa (Fig. 3c). These results suggested that N9 NA proteins expressed in different expression systems differed in their glycosylation patterns.
Western blot assay demonstrating different MWs of N9 NA proteins expressed in various virus-host cell systems. A MW of 50 kDa for recombinant NA protein expressed in baculovirus-insect cell system (rBV-N9 NA), 75 kDa for recombinant protein expressed in TK- cells infected with rVac-N9 NA virus, and 55 kDa for N9 NA protein produced in MDCK cells infected with rgH7N9 HANA (6 + 2) virus are shown. After treatment with PNGase F, the MW of 75 kDa of N9 NA expressed in infected TK- cells decreased to 55 kDa, while there was no change in MW of N9 NA protein produced in infected MDCK cells (a). The MW of 75 kDa for recombinant protein expressed in MDCK cells infected with rVac-N9 NA virus decreased to 55 kDa after deglycosylation (b), and the MW of 50 kDa for rBV-N9 NA decreased to 46 kDa after deglycosylation (c)
Assay for sialidase activity of the recombinant NA protein
Culture supernatants of TK- cells infected with rVac-N9 NA virus were titrated for sialidase activity of the expressed NA protein using a fetuin-based enzyme-linked lectin assay (ELLA). The rgH7N9 HANA (6 + 2) virus was included as a positive control, and the supernatant from rVac-pSC11-infected TK- cell cultures was used in parallel as a negative control. The absorbance values (corresponding to sialidase activities) were plotted against the virus concentrations. NA enzymatic activity was absent in the culture supernatant of the negative control (Fig. 4).
Measurement of anti-H7 HA antibody in immune mouse sera by HAI assay
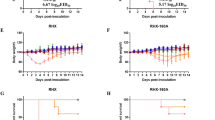
Immune sera collected from ICR or BALB/c mice immunized with rVac-H7 HA virus were assayed for anti-HA antibody by HAI assay using rgH7N9 HANA (6 + 2) virus as the test antigen. The geometric mean titer (GMT) of HAI antibody was 320 (range, 160-640) in BALB/c mice and 40 (range, 20-80) in ICR mice. The difference in HAI antibody titers in both mouse strains was statistically significant (P < 0.01). HAI antibody titers ≥ 40 are considered to provide 50 % protective level. Titers ≥ 40 were found in 100 % (5 of 5) of BALB/c mice, and 80 % (4 of 5) of ICR mice. These results suggested that BALB/c mice could mount a better immune response to influenza virus HA than ICR mice (Fig. 5).
Hemagglutination-inhibition assay for anti-HA antibody in sera from mice immunized with rVac-H7 HA virus. BALB/c mice produced significantly higher antibody titers than ICR mice (one-tailed, unpaired Student t-test; P < 0.01, **), while no antibody was detected in mice immunized with rVac-pSC11 virus, wild-type vaccinia virus, or mock control
Reactivity of anti-H7 HA antibody against viruses with homologous and heterologous HA genes
Sera from BALB/c mice immunized with rVac-H7 HA virus were pooled and assayed for anti-HA antibody against viruses with HA genes from homologous or heterologous strains or subtypes by HAI and MN assays. Sera from mock-immunized mice were used as negative controls. The test viruses from HA genetic group 1 (H1, H5 and H9) included CA-07 and reassorted PR8 human viruses, H9N2 LPAI virus, rgH5N1 HANA (6 + 2) clade 2.3.4 virus and rgN9 NA (7 + 1) viruses. Viruses from HA genetic group 2 (H3, H4 and H7) included H3N2 human virus, H4N9 and H7N1 LPAI viruses, rgH7 HA (7 + 1) and rgH7N9 HANA (6 + 2) viruses (Fig. 6).
Reactivity of anti-H7 HA antibody in mouse sera against the test viruses carrying an HA gene from a homologous or heterologous strain of the H7 subtype or from heterologous subtypes belonging to HA group 1 and group 2 as demonstrated by A) HI assay and B) MN assay. A significant increase in antibody titer over the mock control was observed only with viruses belonging to the H7 subtype (one-tailed, Wilcoxon rank sum test; 0.01 < P < 0.05, *)
Both HAI and MN assay showed that the anti-H7 HA antibody only reacted with viruses with an H7 subtype, regardless of whether they were native virus or rg viruses (Fig. 6a and b). The titers against H7 viruses with a homologous HA gene were higher than that against H7 virus with a heterologous HA gene. Similarly, MN assay also demonstrated that anti-H7 HA antibody neutralized the viruses with a homologous H7 HA gene (rgH7 HA (7 + 1) and rgH7N9 HANA (6 + 2) viruses) to a higher degree than the virus with a heterologous HA gene (H7N1 LPAI virus).
Determination for anti-NA antibody in immune mouse sera by NAI assay
Immune sera collected from ICR or BALB/c mice immunized with rVac-N9 NA virus were assayed for anti-NA antibody by NAI assay using rgH7N9 HANA (6 + 2) virus as the test antigen. The GMT of NAI antibody was 2228 (range 1280-2560) in BALB/c mice and 970 (range 320-2560) in ICR mice. The difference in NAI antibody titers in these two mouse strains was statistically significant (P < 0.05). Anti-N9 antibody was not detected in mice immunized with rVac-pSC11 virus, wild-type vaccinia virus or mock control (Fig. 7).
Fetuin-based neuraminidase-inhibition assay for NAI antibody in BALB/c and ICR mice immunized with rVac-N9 NA. BALB/c mice produced a significantly higher antibody titer than ICR mice (one-tailed, unpaired Student t-test; 0.01 < P < 0.05, *), while no antibody was detected in mice immunized with rVac-pSC11 virus, wild-type vaccinia virus, or mock control
Reactivity of anti-N9 NA against viruses with homologous and heterologous NA genes
Sera from BALB/c mice immunized with rVac-N9 NA virus were pooled and assayed for anti-NA antibody against viruses with an NA gene from homologous and heterologous strains or subtypes by NAI and RI assay. The test viruses from NA genetic group 1 (N1) included CA-07 and reassorted PR8 human viruses, H7N1 LPAI virus, and rgH5N1 HANA (6 + 2) clade 2.3.4 and rgH7 HA (7 + 1) viruses. Viruses from the NA genetic group 2 (N2 and N9) included H3N2 human virus, H9N2 and H4N9 LPAI viruses, rgN9 NA (7 + 1) and rgH7N9 HANA (6 + 2) viruses, as shown in Fig. 8. The result showed that anti-N9 NA antisera were highly specific for viruses carrying the N9 subtype, as determined by both NAI and RI assays (Fig. 8a and b). Moreover, similar NAI antibody titers against N9 in various rg viruses with different HA subtypes were obtained, suggesting that reassortment with different HA subtypes did not affect the NA enzymatic activity of the test viruses.
Reactivity of anti-N9 NA antibody in mouse sera against the test viruses carrying an NA gene from homologous or heterologous strains of the N9 subtype or from heterologous subtypes belonging to NA group 1 and group 2, as demonstrated by A) NAI assay and B) RI assay. A significant increase in antibody titer over the mock control was observed with only viruses belonging to the N9 subtype (one-tailed, Wilcoxon rank sum test; 0.01 < P < 0.05, *)
Discussion
In this study, rVac-H7 HA and rVac-N9 NA viruses were constructed, and the immunobiological properties of recombinant HA and NA proteins expressed in infected TK- cells were characterized without interference from other influenza virus genes. Moreover, rgH7N9 HANA (6 + 2), rgH7 HA (7 + 1), rgN9 NA (7 + 1) viruses were also generated, and the properties of HA and NA proteins produced in different virus-host systems were compared. These rg viruses were also used as the test viruses in serological assays.
Our study showed good expression of HA and NA proteins in the cytoplasm and on the surface of TK- cells infected with recombinant viruses. One form of H7 HA with a MW of 75 kDa was produced in TK- cells infected with rVac-H7 HA virus. This MW was slightly higher than that of the recombinant H7 HA protein expressed in a baculovirus-insect cell system, i.e., 75 versus 60 kDa. In general, mammalian cells (TK- cells and MDCK cells) produce compositionally more complex N-glycans than insect cells [24]. Furthermore, TK- cells infected with the rVac-H7 HA virus and maintained in culture medium supplemented with TPCK-treated trypsin yielded three forms of HA proteins with a MW of 75, 50 and 27 kDa, corresponding to HA0, HA1 and HA2 domains. This pattern was also observed in MDCK cells infected with rgH7N9 virus in the presence of TPCK-treated trypsin. Similar HA patterns and sizes have also been observed with recombinant H1 HA protein expressed in the chicken cells infected with recombinant MVA virus carrying an HA gene derived from CA-07 virus [25] and recombinant H5 HA protein in TK- cells infected with recombinant vaccinia viruses harboring the HA gene of KAN-1 (H5N1) virus [26].
This study showed that the NA protein expressed in TK- and MDCK cells infected with rVac-N9 NA virus was hyperglycosylated, and its MW was higher than those produced in MDCK cells infected with rgH7N9 HANA (6 + 2) virus and in insect cells infected with the recombinant baculovirus, i.e., 75, 75, 55 and 50 kDa, respectively. The glycan moiety of the hyperglycosylated NA expressed in TK- and MDCK cells infected with rVac-N9 NA virus could be removed by PNGase F enzyme, and a cleavage product with a MW of 55 kDa, which was the same as that of NA protein produced in rg-virus-infected MDCK cells, was obtained. Nevertheless, treatment with PNGase F enzyme could not reduce the MW of the NA protein produced in rg-virus-infected MDCK cells and only slightly decreased the MW of rBv-N9 NA. Our result suggested that recombinant NA proteins produced in various expression systems contained different glycans that could not be completely digested by PNGase F enzyme. Generally, PNGase F cleaves most, but not all kinds of glycans [27]. Previous investigators also observed hyperglycosylation of N1 NA protein in chicken cells infected with recombinant MVA virus carrying an NA gene derived from CA-07 virus [25]. Our previous work demonstrated a MW of 56 kDa for NA proteins produced in TK- cells infected with recombinant vaccinia viruses harboring an NA gene from KAN-1 (H5N1) virus, the same MW as shown in MDCK cells infected with KAN-1 wild-type virus [26], and the N1 NA protein in MDCK cells infected with KAN-1 wild-type virus was not deglycosylated by PNGase F [28]. This finding suggested that the MW of a protein produced in mammalian cells could be either similar or different, depending on the type of protein molecule investigated and the expression system. Moreover, this study also demonstrated NA sialidase activity of the recombinant N9 NA protein expressed in TK- cells and also in rgH7N9 HANA (6 + 2) virus using a fetuin-based-ELLA assay.
rVac-H7 HA or rVac-N9 NA virus was injected at the same dose into BALB/c and ICR mice to raise antibodies to HA or NA protein. Nevertheless, it was shown that both HAI and NAI antibody titers produced in BALB/c mice were significantly higher than those produced in ICR mice. In a pathogenesis study, BALB/c mice were more sensitive to H7N9 infection than ICR or C57BL/6 mice, as evidenced by their body weight loss and higher viral load in the lungs [29]. Thus, we can conclude that BALB/c mice are a good model for evaluation of immunogenicity.
Currently, 18 HA and 11 NA subtypes have been identified. All 18 HA subtypes fall into two distinct groups by phylogenetic classification: group 1 (H1, H2, H5, H6, H8, H9, H11, H12, H13, H16, H17 and H18) and group 2 (H3, H4, H7, H10, H14, and H15). Similarly, all NA subtypes also can be classified into two groups: group 1 (N1, N4, N5, and N8) and group 2 (N2, N3, N6, N7, and N9). The bat-derived N10 and N11 do not belong to either group [16, 17]. We tested the specificity of the anti-HA and anti-NA antibody response in BALB/c pooled sera using wild-type or reverse genetic influenza viruses as test viruses using various serological methods. Anti-H7 HA antibodies in immune mouse sera showed concordant results by HAI and MN assays. A significant increase in antibody levels over the mock control sera could be obtained only when rg viruses of the same HA subtype, i.e., rgH7 HA (7 + 1) and rgH7N9 HANA (6 + 2) viruses or a wild-type virus with homologous HA subtype, A/Ostrich/Zinbabwe/222/1996 (H7N1) virus, were used as the test viruses. No cross-reaction was observed with other virus subtypes belonging to the heterosubtypic HA group (H1, H5 and H9 viruses for HA group 1; or H3 and H4 viruses for HA group 2) in our study. On the other hand, previous investigators showed that the neutralizing antibody against the HA stalk region was broadly reactive and cross-neutralized the other virus subtypes belonging to the same HA genetic group, as demonstrated in humans and in a mouse model [30–32]. However, the antibody to the H7 HA stalk region in our mice might not be sufficiently cross-reactive after only one immunization, in contrast to boosted immunization in mice or repeated natural infections in humans in other studies.
Using NAI and RI assays, this study showed that anti-N9 NA antibody in pooled immune mouse sera reacted only with viruses with homosubtypic NA, i.e., A/Duck/Shan Tou/461/2000 (H4N9), rgN9 NA (7 + 1) and rgH7N9 HANA (6 + 2) viruses. Other investigators reported that the peptide “ILRTQESEC”, which is present in the enzyme active site of the NA globular head in almost all influenza subtypes, could induce the antibody that was broadly cross-reactive among all influenza subtypes in a fetuin-based-NAI assay [33]. However, cross-reactive anti-NA antibody against viruses with heterosubtypic NA was not seen in this study.
In conclusion, our study demonstrated that both H7 HA and N9 NA expressed in mice immunized with recombinant vaccinia viruses were immunogenic and capable of inducing satisfactory levels of functional antibodies that were subtype specific after one immunization. The availability of anti-NA antibody in addition to anti-HA antibody is potentially useful for fighting against H7N9 virus infections and could potentially be applied to other influenza subtypes. Nevertheless, to induce broadly reactive anti-HA or anti-NA antibody in humans, the inoculum dose, number of immunizations, immunogen preparation, and use of adjuvant need to be optimized.
References
Gao R, Cao B, Hu Y, Feng Z, Wang D, Hu W, Chen J, Jie Z, Qiu H, Xu K, Xu X, Lu H, Zhu W, Gao Z, Xiang N, Shen Y, He Z, Gu Y, Zhang Z, Yang Y, Zhao X, Zhou L, Li X, Zou S, Zhang Y, Li X, Yang L, Guo J, Dong J, Li Q, Dong L, Zhu Y, Bai T, Wang S, Hao P, Yang W, Zhang Y, Han J, Yu H, Li D, Gao GF, Wu G, Wang Y, Yuan Z, Shu Y (2013) Human infection with a novel avian-origin influenza A (H7N9) virus. N Engl J Med 368(20):1888–1897. doi:10.1056/NEJMoa1304459
Liu Y, Paquette SG, Zhang L, Leon AJ, Liu W, Xiuming W, Huang L, Wu S, Lin P, Chen W, Fang X, Zeng T, Kelvin N, Farooqui A, Kelvin DJ (2015) The third wave: H7N9 endemic reassortant viruses and patient clusters. J Infect Dev Ctries 9(2):122–127. doi:10.3855/jidc.6759
Food and Agriculture Organization of United Nations (2015) Fourth wave of H7N9 avian influenza threatens livelihoods, public health. http://www.fao.org/ag/againfo/programmes/en/empres/news_151015.html. Accessed 10 Nov 2015
William T, Thevarajah B, Lee SF, Suleiman M, Jeffree MS, Menon J, Saat Z, Thayan R, Tambyah PA (2014) Yeo TW (2015) Avian influenza (H7N9) virus infection in Chinese tourist in Malaysia. Emerg Infect Dis 21(1):142–145. doi:10.3201/eid2101.141092
Center for Disease Control and Prevention (2015) First Case of Avian Influenza A (H7N9) Virus Infection in a Human in North America. http://www.izsummitpartners.org/wp-content/uploads/2015/02/CDC-Key-Points-H7N9-Canada-Human-01-29-2015.pdf. Accessed 27 Jan 2015
Chinese National Influenza Center (2015) Influenza weekly report. http://cnic.org.cn/chn/. Accessed 8 Dec 2015
Lam TT, Wang J, Shen Y, Zhou B, Duan L, Cheung CL, Ma C, Lycett SJ, Leung CY, Chen X, Li L, Hong W, Chai Y, Zhou L, Liang H, Ou Z, Liu Y, Farooqui A, Kelvin DJ, Poon LL, Smith DK, Pybus OG, Leung GM, Shu Y, Webster RG, Webby RJ, Peiris JS, Rambaut A, Zhu H, Guan Y (2013) The genesis and source of the H7N9 influenza viruses causing human infections in China. Nature 502(7470):241–244. doi:10.1038/nature12515
Yang H, Carney PJ, Chang JC, Villanueva JM, Stevens J (2013) Structural analysis of the hemagglutinin from the recent 2013 H7N9 influenza virus. J Virol 87(22):12433–12446. doi:10.1128/JVI.01854-13
Zhu W, Li L, Yan Z, Gan T, Li L, Chen R, Chen R, Zheng Z, Hong W, Wang J, Smith DK, Guan Y, Zhu H, Shu Y (2015) Dual E627K and D701N mutations in the PB2 protein of A(H7N9) influenza virus increased its virulence in mammalian models. Sci Rep 5:14170. doi:10.1038/srep14170
Yang S, Chen Y, Cui D, Yao H, Lou J, Huo Z, Xie G, Yu F, Zheng S, Yang Y, Zhu Y, Lu X, Liu X, Lau SY, Chan JF, To KK, Yuen KY, Chen H, Li L (2014) Avian-origin influenza A(H7N9) infection in influenza A(H7N9)-affected areas of China: a serological study. J Infect Dis 209(2):265–269. doi:10.1093/infdis/jit430
Zhou J, Wang D, Gao R, Zhao B, Song J, Qi X, Zhang Y, Shi Y, Yang L, Zhu W, Bai T, Qin K, Lan Y, Zou S, Guo J, Dong J, Dong L, Zhang Y, Wei H, Li X, Lu J, Liu L, Zhao X, Li X, Huang W, Wen L, Bo H, Xin L, Chen Y, Xu C, Pei Y, Yang Y, Zhang X, Wang S, Feng Z, Han J, Yang W, Gao GF, Wu G, Li D, Wang Y, Shu Y (2013) Biological features of novel avian influenza A (H7N9) virus. Nature 499(7459):500–503. doi:10.1038/nature12379
Smith GE, Flyer DC, Raghunandan R, Liu Y, Wei Z, Wu Y, Kpamegan E, Courbron D, Fries LF 3rd, Glenn GM (2013) Development of influenza H7N9 virus like particle (VLP) vaccine: homologous A/Anhui/1/2013 (H7N9) protection and heterologous A/chicken/Jalisco/CPA1/2012 (H7N3) cross-protection in vaccinated mice challenged with H7N9 virus. Vaccine 31(40):4305–4313. doi:10.1016/j.vaccine.2013.07.043
Kreijtz JH, Wiersma LC, De Gruyter HL, Vogelzang-van Trierum SE, van Amerongen G, Stittelaar KJ, Fouchier RA, Osterhaus AD, Sutter G, Rimmelzwaan GF (2014) A single immunization with an MVA-based influenza virus H7 vaccine affords protection in the H7N9 pneumonia ferret model. J Infect Dis 211(5):791–800. doi:10.1093/infdis/jiu528
Liu Q, Mena I, Ma J, Bawa B, Krammer F, Lyoo YS, Lang Y, Morozov I, Mahardika GN, Ma W, Garcia-Sastre A, Richt JA (2015) Newcastle disease virus-vectored H7 and H5 live vaccines protect chickens from challenge with H7N9 or H5N1 avian influenza viruses. J Virol 89(14):7401–7408. doi:10.1128/JVI.00031-15
Li Z, Gabbard JD, Johnson S, Dlugolenski D, Phan S, Tompkins SM, He B (2015) Efficacy of a parainfluenza virus 5 (PIV5)-based H7N9 vaccine in mice and guinea pigs: antibody titer towards HA was not a good indicator for protection. PLoS One 10(3):e0120355. doi:10.1371/journal.pone.0120355
Gamblin SJ, Skehel JJ (2010) Influenza hemagglutinin and neuraminidase membrane glycoproteins. J Biol Chem 285(37):28403–28409. doi:10.1074/jbc.R110.129809
Wu Y, Wu Y, Tefsen B, Shi Y, Gao GF (2014) Bat-derived influenza-like viruses H17N10 and H18N11. Trends Microbiol 22(4):183–191. doi:10.1016/j.tim.2014.01.010
Hoffmann E, Stech J, Guan Y, Webster RG, Perez DR (2001) Universal primer set for the full-length amplification of all influenza A viruses. Arch Virol 146(12):2275–2289. doi:10.1007/s007050170002
Hoffmann E, Neumann G, Kawaoka Y, Hobom G, Webster RG (2000) A DNA transfection system for generation of influenza A virus from eight plasmids. Proc Natl Acad Sci USA 97(11):6108–6113. doi:10.1073/pnas.100133697
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning : a laboratory manual, 2nd edn. Cold Spring Harbor Laboratory, Cold Spring Harbor
Louisirirotchanakul S, Lerdsamran H, Wiriyarat W, Sangsiriwut K, Chaichoune K, Pooruk P, Songserm T, Kitphati R, Sawanpanyalert P, Komoltri C, Auewarakul P, Puthavathana P (2007) Erythrocyte binding preference of avian influenza H5N1 viruses. J Clin Microbiol 45(7):2284–2286. doi:10.1128/JCM.00921-07
Couzens L, Gao J, Westgeest K, Sandbulte M, Lugovtsev V, Fouchier R, Eichelberger M (2014) An optimized enzyme-linked lectin assay to measure influenza A virus neuraminidase inhibition antibody titers in human sera. J Virol Methods 210C:7–14. doi:10.1016/j.jviromet.2014.09.003
World Health Organization (2002) WHO manual on animal influenza diagnosis and surveillance. WHO, Geneva
Harrison RL, Jarvis DL (2006) Protein N-glycosylation in the baculovirus-insect cell expression system and engineering of insect cells to produce “mammalianized” recombinant glycoproteins. Adv Virus Res 68:159–191. doi:10.1016/S0065-3527(06)68005-6
Hessel A, Schwendinger M, Fritz D, Coulibaly S, Holzer GW, Sabarth N, Kistner O, Wodal W, Kerschbaum A, Savidis-Dacho H, Crowe BA, Kreil TR, Barrett PN, Falkner FG (2010) A pandemic influenza H1N1 live vaccine based on modified vaccinia Ankara is highly immunogenic and protects mice in active and passive immunizations. PLoS One 5(8):e12217. doi:10.1371/journal.pone.0012217
Noisumdaeng P, Pooruk P, Prasertsopon J, Assanasen S, Kitphati R, Auewarakul P, Puthavathana P (2014) Homosubtypic and heterosubtypic antibodies against highly pathogenic avian influenza H5N1 recombinant proteins in H5N1 survivors and non-H5N1 subjects. Virology 454–455:254–262. doi:10.1016/j.virol.2014.02.024
Maley F, Trimble RB, Tarentino AL, Plummer TH Jr (1989) Characterization of glycoproteins and their associated oligosaccharides through the use of endoglycosidases. Anal Biochem 180(2):195–204. doi:10.1016/0003-2697(89)90115-2
Changsom D, Lerdsamran H, Wiriyarat W, Chakritbudsabong W, Siridechadilok B, Prasertsopon J, Noisumdaeng P, Masamae W, Puthavathana P (2016) Influenza neuraminidase subtype N1: immunobiological properties and functional assays for specific antibody response. PLoS One 11(4):e0153183. doi:10.1371/journal.pone.0153183
Zhu Z, Yang Y, Feng Y, Shi B, Chen L, Zheng Y, Tian D, Song Z, Xu C, Qin B, Zhang X, Guan W, Liu F, Yang T, Yang H, Zeng D, Zhou W, Hu Y, Zhou X (2013) Infection of inbred BALB/c and C57BL/6 and outbred Institute of Cancer Research mice with the emerging H7N9 avian influenza virus. Emerg Microbes Infect 2(8):e50. doi:10.1038/emi.2013.50
Ekiert DC, Bhabha G, Elsliger MA, Friesen RH, Jongeneelen M, Throsby M, Goudsmit J, Wilson IA (2009) Antibody recognition of a highly conserved influenza virus epitope. Science 324(5924):246–251. doi:10.1126/science.1171491
Ekiert DC, Friesen RH, Bhabha G, Kwaks T, Jongeneelen M, Yu W, Ophorst C, Cox F, Korse HJ, Brandenburg B, Vogels R, Brakenhoff JP, Kompier R, Koldijk MH, Cornelissen LA, Poon LL, Peiris M, Koudstaal W, Wilson IA, Goudsmit J (2011) A highly conserved neutralizing epitope on group 2 influenza A viruses. Science 333(6044):843–850. doi:10.1126/science.1204839
Krammer F, Margine I, Hai R, Flood A, Hirsh A, Tsvetnitsky V, Chen D, Palese P (2014) H3 stalk-based chimeric hemagglutinin influenza virus constructs protect mice from H7N9 challenge. J Virol 88(4):2340–2343. doi:10.1128/JVI.03183-13
Doyle TM, Hashem AM, Li C, Van Domselaar G, Larocque L, Wang J, Smith D, Cyr T, Farnsworth A, He R, Hurt AC, Brown EG, Li X (2013) Universal anti-neuraminidase antibody inhibiting all influenza A subtypes. Antiviral Res 100(2):567–574. doi:10.1016/j.antiviral.2013.09.018
Acknowledgments
This study was partially supported by the Office of the Higher Education Commission and Mahidol University under the National Research Universities Initiative, and also Siriraj Graduate Thesis Scholarship Program for International Students from Neighboring Countries. We thank Prof. Robert G. Webster and Dr. Richard Webbey, St. Jude Children’s Research Hospital, Memphis, USA, for the recombinant pHW2000 reverse genetic plasmids and the goat antisera against H7 HA and N9 NA; Dr. Yuelong Shu, Chinese National Influenza Center for providing the recombinant plasmids containing influenza H7 or N9 insert sequence; Prof. Bernard Moss, the National Institution of Allergy and Infection Disease, Maryland, USA, for the pSC11 plasmid vector; BRI resources, USA, for the recombinant HA and NA proteins; the Division of Medical Molecular Biology, Department of Research and Development, Faculty of Medicine Siriraj Hospital, Mahidol University, for the laser scanning confocal microscope facilities, and Dr. Andrew Broadbent, The Pirbright Institute, UK for useful edits and comments on the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
Li Jiang has received salaries and research support from the Office of the Higher Education Commission and Mahidol University under the National Research Universities Initiative, and Siriraj Graduate Thesis Scholarship Program for International Students from Neighboring Countries.
Conflict of interest
We have no competing interests.
Ethical approval
The use of animals in this study was approved by Animal Care and Use Committee of Veterinary Science, Mahidol University, Thailand (Permit Number: MUVS-2013-37).
Rights and permissions
About this article
Cite this article
Jiang, L., Changsom, D., Lerdsamran, H. et al. Immunobiological properties of influenza A (H7N9) hemagglutinin and neuraminidase proteins. Arch Virol 161, 2693–2704 (2016). https://doi.org/10.1007/s00705-016-2968-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-016-2968-7