Abstract
Festuca is one of the most ecologically and economically important genera of temperate grasses. Species of its type subgenus, Festuca, are common components of cool-seasonal pastures and are especially frequent in mountainous areas, where there are often several morphologically similar species that grow in the same or comparable habitats and sometimes live in sympatry. Nuclear DNA assessments by flow cytometry can be used to identify taxa and uncover new ploidy levels in species complexes for which new chromosome data are provided or previous chromosome counts and genome sizes are known. Holoploid (2C) values of newly studied Pyrenean Festuca subgen. Festuca sects. Eskia, Festuca and Aulaxyper species fall within the expected ranges for these taxonomic groups and include 2x, 4x, 6x and 8x ploidy levels. Monoploid (1Cx) genome sizes of diploids and polyploids are larger in the species of the more ancestral F. sect. Eskia group showing a decreasing trend in the species of the more recently evolved F. sects. Festuca and Aulaxyper lineages. 1Cx values of high polyploid Aulaxyper taxa are among the smallest of the three Festuca sections, corroborating previous findings. Our analysis provides new genome size values and inferred ploidy levels for hexaploid F. heteromalla and octoploid F. trichophylla and highlights the genomic and ecological differentiation of tetraploid F. gautieri susbsp. gautieri from diploid F. gautieri subsp. scoparia.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Festuca L. (Poaceae) is one of the main genera of the temperate pooid grasses, both in number of species and in ecological and economic importance, as it encompasses some of the world’s best forages, pastures and lawn species. The genus includes more than 500 species (Festuca; Catalán 2006), considering both broad and fine-leaved Festuca taxa, and up to 644 accepted species, considering fine-leaved subtribe Loliinae taxa [Festuca and several minor subsumed genera (Plants of the World On-line http://www.plantsoftheworldonline.org/taxon/urn:lsid:ipni.org:names:328907-2, as per 5th April 2023]. Fescues are native to temperate zones and mountainous subtropical and tropical regions of the five continents (Catalán 2006; Minaya et al. 2017). Phylogenetic analyses have consistently detected two main lineages within Festuca and its close genera of subtribe Loliinae, the broad-leaved and the fine-leaved clades (Catalán et al. 2004, 2007; Inda et al. 2008; Minaya et al. 2017; Moreno-Aguilar et al. 2020, 2022a). Taxonomically, Festuca has been classified into 11 subgenera with several sections within each, according to Alexeev’s revision of world fescues (Catalán 2006; Moreno-Aguilar et al. 2022b). Nine of those subgenera were assigned to broad-leaved Festuca and two to fine-leaved Festuca, including the largest subgenus Festuca (Catalán 2006; Moreno-Aguilar et al. 2022b). Subgenus Festuca includes sections Eskia Willk., Festuca (F. ovina group) and Aulaxper Dumort. (F. rubra group) that contain taxa broadly distributed in Eurasia. Biogeographic studies have shown that the Loliinae originated in the Palearctic region and that the Mediterranean—SW Asia area has been the main center of diversification of both broad- and fine-leaved Loliinae (Inda et al. 2008, 2014; Minaya et al. 2017). The Iberian Peninsula and the Pyrenees are one of the richest regions in Festuca species, accounting for near 100 taxa of which 40% are endemics (Kerguélen and Plonka 1989; Devesa et al. 2013, 2020). Most of the Iberian and Pyrenean fescues grow in mountainous areas where there are often several different species living together.
Cytogenetic analyses have provided a cue for the analysis of evolutionary trends and speciation processes within subtribe Loliinae. Catalán (2006) showed the first comparative study of molecular phylogenetic inference and karyotype evolution in Festuca, a genus with a uniform chromosome base number of x = 7 and ploidy levels ranging from 2x to 12x. Recently, a new ploidy level of 14x was recorded within F. sect. Festuca (F. yvesii subsp. summilusitana 2n = 14x = 98; Martínez-Sagarra et al. 2021). Festuca exhibits striking differences in monoploid genome sizes (1Cx) between the broad- and the fine-leaved lineages, showing a 2.5-fold range decrease in chromosome size and C-values from more ancestral broad-leaved lineages (Drymanthele, Scariosa, Subbulbosae) to more recently evolved fine-leaved lineages (Festuca, Aulaxyper) (Catalán 2006). The early diverging fine-leaved Eskia lineage reveals an intermediate karyotype pattern between the broad- and fine-leaved groups (Catalán et al. 2006). Šmarda et al. (2008) further hypothesized that the Loliinae lineage probably experienced a two-fold genome and chromosome size enlargement and a GC enrichment with respect to its close relatives, which was subsequently followed by dramatic reductions, especially in the rapidly evolving fine-leaved group. Loureiro et al. (2007) and Šmarda et al. (2008) also detected genomic downsizing in the fine-leaved Loliinae and in the polyploids, with the genomic losses being more pronounced in allopolyploids with large progenitor genomes than in autopolyploids with small progenitor genomes. Moreno-Aguilar et al. (2022a) found a large and significant correlation between the sizes of the Loliinae genomes and their respective amounts of repetitive elements (especially Retand and Angela retrotransposons). These authors also showed that changes in the Loliinae genome sizes were likely caused by gains or losses in their repetitive elements. Their analysis of the evolution of the Loliinae repeatome suggested that the large genome sizes of broad-leaved diploid Loliinae were likely the result of past allopolyploidizations followed by diploidizations, while the massive repeatome and genome contractions observed in some polyploid lineages (e. g., F. sect. Aulaxyper, F. subgen. Schedonorus) were probably the consequence of large genomic rearrangements and losses in these highly hybridogenic lineages (Moreno-Aguilar et al. 2022a). Within Festuca subgen. Festuca, the two taxonomically complex and most broadly studied groups are F. sects. Festuca and Aulaxyper Dumort. (Catalán 2006; Catalán et al. 2004, 2006, 2007). A third section that contains a relatively high number of taxa distributed in mountainous areas of the circum-Mediterranean region is F. sect. Eskia Willk. (Torrecilla et al. 2003, 2013). Some lineages experienced complex reticulate evolutionary processes, including homoploid and heteroploid hybridizations and polyploidizations (Catalán 2006; Torrecilla et al. 2013; Marques et al. 2016). Some species of fine-leaved fescues are morphologically very similar, and their classification is still controversial.
Numerous karyological and cytogenetic studies have been conducted in Festuca subgen. Festuca in Europe (summarized in Kerguélen and Plonka 1989; Fuente and Ortúñez 1988, 2001; Fuente et al. 2001; Ortúñez and Fuente 1995, 2004; Ferrero and Fuente 1996; Loureiro et al. 2007; Šmarda et al. 2008; Devesa et al. 2013, 2020; Martínez-Sagarra et al. 2021, 2022), sometimes to identify the species or to clarify the relationships between these taxa growing in the mountains and to assign them to the species or complexes within a phylogenetic context. Nuclear DNA assessments by flow cytometry can be a useful tool to estimate the ploidy levels of the samples and to carry out an initial selection of the studied taxa, especially in areas where they coexist and in species complexes in which previous chromosome counts have already been carried out. Despite the numerous taxa treated in previous studies (https://cvalues.science.kew.org/search/angiosperm, accessed 5th April 2023), some of the species growing in the Pyrenees and other mountain systems have not been thoroughly surveyed yet.
This study deals with the cytogenetic analysis of taxonomically complex SW European Pyrenean Festuca subgen. Festuca plants from sections Eskia, Festuca and Aulaxyper. The understanding gained from genome size assessments, combined with careful observation regarding morphology and habitat, in addition to our present knowledge of the chromosome number, will contribute toward clarifying the identity of some of the more critical taxa. The aims of our research are: (i) to assess the genome size variation between and within some groups of Pyrenean fescues, and (ii) to test if the genome size assessments fit previous hypotheses on different evolutionary trends of monoploid, 1Cx, genomes for species of each section and if they could help discover new ploidy levels and distinguish or discriminate between some related species.
Material and methods
Plant material
Plant material studied together with the localities of provenance is summarized in Table 1. The herbarium vouchers of these taxa are deposited in the herbaria of the Botanical Institute of Barcelona (BC) and of the Escuela Politécnica Superior de Huesca (University of Zaragoza).
DNA content assessment and chromosome counting
Nuclear DNA content was assessed for 102 individuals of 33 populations belonging to 14 taxa.
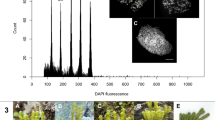
Fresh young leaves of the plants were co-chopped using a razor blade with an internal standard in the proportions 2:1 in 1200 μl of LB01 buffer (Doležel et al. 1989) with 0.5% Triton X-100 and supplemented with 100 µg/ml ribonuclease A (RNase A, Boehringer, Meylan, France) in a Petri dish. Pisum sativum L. ‘Express Long’ (2C = 8.37 pg; Marie and Brown 1993) was used as internal standard and was first analyzed separately in 600 μl of LB01 buffer in order to locate the peak position of the standard in the fluorescence histogram. Nuclei were filtered through a 70-μm nylon filter in order to eliminate cell debris before the addition of 36 μl of propidium iodide (1 mg/ml, solution in water; Invitrogen Eugene, Oregon, USA). From one to five individuals were used per population (Table 1), and two samples of each individual collected were extracted and measured independently. Fluorescence analysis was carried out using an Epics XL flow cytometer (Coulter Corporation, Hialeah, Florida, USA) at the Centres Científics i Tecnològics de la Universitat de Barcelona, with the standard configuration as described in Garnatje et al. (2006). Acquisition was stopped at 8000 nuclei. The DNA content was calculated assuming a linear correlation between the fluorescence signals (of the stained nuclei) and DNA amount. Mean and standard deviations were calculated for 2C values of each population. The mean of the half-peak coefficient of variation (HPCV) was also calculated for both target plant and internal standard. A nonparametric Kruskal–Wallis test was performed to compare the 1Cx values of the three studied sections after a Shapiro–Wilk test of normality.
Chromosome counts were carried out according to the following protocol. Root tips from plants growing in pots were pretreated in distilled water for 24 h at 4 °C. This material was fixed in a solution of absolute ethanol and glacial acetic acid (3:1) for 30 min in the dark. Subsequently, the material was transferred to a new solution of absolute ethanol and glacial acetic acid (3:1) for 4 h in the dark. After this step, the material was conserved at − 20 °C in the same solution. For chromosome counts, root tip meristems were excised, hydrolized for 10–12 min on 1N HCl at 60 °C, washed with distilled water, stained with 1% aceto-orcein and squashed on a drop of 9:1 45% acetic acid: glycerol. Slides were observed with a Zeiss Axioplan microscope, and the best metaphase plates were photographed with an AxioCam HRm camera.
Results and discussion
Data on the genome size assessments (2C and 1Cx values), chromosome numbers (new data and previously published data), and estimated ploidy levels for the studied taxa are shown in Table 1. Some ploidy levels have been inferred from genome size values compared to genome sizes of other species of Festuca included in this study or analyzed by other authors (Kerguélen and Plonka 1989; Loureiro et al. 2007; Šmarda et al. 2008; Devesa et al. 2020; Martínez-Sagarra 2018; Martínez-Sagarra et al.2021, 2022; Moreno-Aguilar et al. 2022a) [Table 1. asterisk: estimations based on the congruence with chromosome number, “ploidy level”; no asterisk: estimations based solely on genome size, “DNA-ploidy level”]. Overall, the 2C values range from 4.41 pg of diploid F. marginata subsp. andresmolinae to 16.36 pg of octoploid F. laevigata sensu lato, both within F. sect. Festuca, and monoploid (1Cx) values range from 1.85 pg of F. nigrescens subsp. microphylla (F. sect. Aulaxyper) to 2.92 pg of F. gautieri subsp. scoparia (F. sect. Eskia). However, the 2C/1Cx values and inferred ploidy levels (2x, 4x, 6x, 8x) of the studied taxa vary considerably among sections (Table 1), confirming the previous findings of Catalán (2006), Šmarda et al. (2008) and Moreno-Aguilar et al. (2022a) on the decreasing genome-size trend from F. sect. Eskia, to F. sects. Festuca and Aulaxyper, and the larger 1Cx reduction observed for polyploid genomes in F. sect. Aulaxyper. The nonparametric Kruskal–Wallis test shows statistically significant differences (p < 0.0001) between 1Cx values of the sect. Eskia and those of the other two sections, Festuca and Aulaxyper. The average of HPCV (half peak coefficient of variation) of the investigated plant is 2.29 and that of the standard is 3.46, most values being below 5%, accounting for data reliability.
Festuca sect. Eskia. Within the F. sect. Eskia group, mean monoploid genome values (1Cx) of F. eskia (2.79 pg) and F. gautieri subsp. scoparia (2.85 pg) are highly similar; the two sister species constitute one of the earliest diverging lineages of the fine-leaved clade (Inda et al. 2008; Minaya et al. 2017; Moreno-Aguilar et al. 2020). 2C values for nuclear DNA amount of diploid F. eskia (5.45–5.69 pg; 2n = 2x = 14; Table 1) fall within those reported in the broadly sampled study of Marques et al. (2016). This subalpine species, endemic to the Pyrenees and the Cantabrian mountains, is only known to be diploid, though it participated in the origin of two recent homoploid hybrids after its interspecific crossings with its close F. gautieri subsp. scoparia (F. × picoeuropeana Nava) and F. quadriflora (F. × souliei St-Yves) in different parts of its distribution range (Torrecilla et al. 2013; Marques et al. 2016). Genome size values of F. gautieri correspond to two ploidy levels, which have been corroborated by chromosome counting (Table 1). 2C values of diploid F. gautieri subsp. scoparia (5.50–5.84 pg; 2n = 2x = 14) also overlap with those reported in the large study of Marques et al. (2016); by contrast, 2C values of tetraploid F. gautieri subsp. gautieri (10.30–10.86 pg, 2n = 4x = 28; Table 1) are given here for the first time. Subalpine F. gautieri subsp. gautieri shows a restricted distribution on both sides of the Eastern Pyrenees (France: Canigó, Puigmal; Spain: Ribes de Freser (Núria, Toses)), and Montseny growing only on siliceous soils (Küpfer 1974; Kerguélen and Plonka 1989; this study), by contrast subalpine F. gautieri subsp. scoparia is largely distributed on mountain ranges of the eastern Iberian Peninsula and the Pyrenees, and grows preferentially on calcareous substrates, but also in cation-rich granites (Catalán et al. 2013; Torrecilla et al. 2013; Marques et al. 2016). Kerguélen and Plonka (1989) differentiated the two F. gautieri taxa based on the larger sizes of spikelets and lemmas of tetraploid F. gautieri subsp. gautieri compared to those of diploid F. gautieri subsp. scoparia, detecting a common feature of larger reproductive structures in polyploids than in their progenitor diploids also observed in other fescues (Catalán 2006). However, Fuente and Ortúñez (1988), Fuente et al. (2001) and Devesa et al. (2020) did not segregate them, as they could not separate them morphologically. Our study has shown that tetraploid F. gautieri subsp. gautieri occupies a slightly broader range in the Eastern Pyrenees than previously known (Montseny) and that its strong siliceous ecological niche does not overlap with the calcicolous or cation-rich niche of its diploid F. gautieri subsp. scoparia conspecific taxon. The almost duplicated genome size of F. gautieri subsp. gautieri 4x with respect to that of F. gautieri subsp. scoparia 2x (Table 1) and their close morphology may suggest that the tetraploid taxon may have originated from the diploid taxon recently and has not yet experienced severe genome losses. In addition, the mean monoploid genome value (1Cx) of the tetraploid (2.67 pg) is roughly similar to that of the diploid, fitting the proposed evolutionary scenario of a potential autopolyploid origin for these types of Festuca polyploids (Šmarda et al. 2008). Further investigations would be necessary, however, to corroborate the kind of polyploid nature of the tetraploid F. gautieri subsp. gautieri and the morphological differentiation of the two taxa.
Festuca sect. Festuca. Monoploid genome values (1Cx) of the F. sect. Festuca species show similar values for the diploids F. airoides (2.23 pg) and F. marginata subsp. andresmolinae (2.21 pg); by contrast, those of the polyploids vary from values close to those of the diploids, like that of tetraploid F. inops (2.27 pg), to others slightly lower, like those of hexaploid F. yvesii subsp. yvesii (1.98 pg), octoploid F. laevigata (1.95–2.05 pg), and hexaploid F. ochroleuca Timb.-Lagr. subsp. bigorronensis (St.-Yves) Kerguélen (2.02 pg) (Table 1). The genome size of Prepyrenean F. marginata subsp. andresmolinae matched its reported chromosome number of 2n = 2x = 14 (Fuente and Ortuñez 1993; Devesa et al. 2020). Alpine F. airoides has been the subject of some taxonomic debate in relation to its status alongside F. supina Schur, and its morphological similarity to F. niphobia (St.-Yves) Kerguélen, which may grow sympatrically in some parts of the Eastern Pyrenees (Kerguélen and Plonka 1989). Devesa et al. (2020) amalgamated these taxa into F. airoides s.l. differentiating two subspecies, F. airoides subsp. airoides 2x (including F. supina auct. pyr. non-Schur 1866) and subsp. molinieri (Litard.) Mart.-Segarra & Devesa 2x, 4x (including F. niphobia) in the Eastern Pyrenees, the second taxon being more robust and more broadly distributed than the former. Our F. airoides sample fits the morphology of the typical subspecies; its 2C and 1Cx values are in agreement with those reported by Šmarda et al. (2008) for French and Bulgarian samples. The 2C values of F. inops correspond to a tetraploid taxon (Table 1). Šmarda et al. (2008) only detected the diploid level in Italian and French populations of this species; however, Devesa et al. (2020) subsumed F. gracilior (Hack.) Markgr.-Dann. under F. inops subsp. inops. Although Kerguelen and Plonka (1989) considered F. gracilior to be a diploid species in France, Martínez-Sagarra (2018) and Devesa et al. (2020) also detected tetraploids in F. inops subsp. inops in Spain; our genome size value of this taxon from Montserrat corroborates the findings of the latter authors.
The presence of F. laevigata Gaudin has not been recognized from within the Iberian Peninsula by Devesa et al. (2020), though it was recognized within the Pyrenees by Kerguélen and Plonka (1989) and Foggi and Müller (2009). Devesa et al. (2020) synonymised F. laevigata auct. p.p. non-Gaudin to F. yvesii subsp. yvesii. However, according to Arndt (2005) individuals showing preferentially an unbroken sclerenchyma ring and belonging to Pyrenean populations generally referred to F. cagiriensis Timb.-Lagr., could be united within a wider concept of F. laevigata. Following the classification system of Kerguélen and Plonka (1989), based on leaf blade anatomy and other morphometric traits, we have identified samples of Pyrenean and Prepyrenean populations as pertaining to F. laevigata s.l. and F. yvesii susp. yvesii (Table 1). We emphasize F. laevigata sensu lato because most populations from Spain correspond most closely to F. cagiriensis. Genome size values of Italian F. laevigata populations corresponded to octoploids (18.65 pg; Šmarda et al. 2008) although chromosome counts of Kerguélen and Plonka (1989) indicated the putative existence of additional hexaploids. The mean 2C values of our F. laevigata samples from the Central and Eastern Pyrenees and Prepyrenees (16.0 pg, Table 1) are between those expected for octoploids (~ 18 pg) and hexaploids (12–13 (14) pg) (Šmarda et al. 2008). Considering the hypothesized reduction in the genome size of the fine-leaved fescues high polyploids, those values may correspond to octoploids, thus allowing us to infer a mean monoploid genome 1Cx value of 1.95–2.0 pg for them. Interestingly, our F. yvesii subsp. yvesii sample from the Eastern Prepyrenees presents a genome size value of 11.89 pg, intermediate between that expected for hexaploids and tetraploids (9–9.5 pg) (Šmarda et al 2008) and octoploids (17.7–18.5 pg) (Martínez-Sagarra 2018). Both octoploid and hexaploid populations have previously been detected for this species in the Pyrenees (Kerguélen and Plonka 1989; Martínez-Sagarra 2018; Devesa et al. 2020); therefore, the 2C value of our Prepyrenean sample would probably correspond to a hexaploid, with an inferred small monoploid 1Cx value of 1.98 pg (Table 1). The presence in this same region of the hexaploid F. glauca Vill., which grows in siliceous terrain, mainly on the Catalonian coast but penetrating inland as, for example, in the Alberes, may well help to explain the nature of the apparently introgressive, hexaploid populations of this species that occur within Catalonia. In fact, when cultivated, plants from near the summit of Montseny become indistinguishable from the coastal plants referred to as F. glauca. F. ochroleuca subsp. bigorronensis is reported to be a tetraploid taxon (Kerguélen and Plonka 1989). Our genome size values (12.12–12.16 pg) correspond to a hexaploid, however. These high Tena Valley populations have been variously interpreted, since they are often not clearly identifiable with other populations which bear the characteristic striated and often pubescent distal culm portion, although their morphological and ecological features allow us to classify them as F. ochroleuca subsp. bigorronensis.
Festuca sect. Aulaxyper. The large variability of nuclear DNA contents found in the studied species of F. sect. Aulaxyper has allowed us to identify new ploidy levels (Table 1) and to confirm the dramatic reduction in monoploid 1Cx values from diploids to high polyploids (Šmarda et al. 2008; Moreno-Aguilar et al. 2022a). 2C values for F. rivularis (5.56 pg), the only known continental Europe diploid representative of this lineage (Díaz-Perez et al. 2008), are provided here for the first time, allowing us to infer a monoploid genome value of 2.78 pg, close to those of species of the more ancestral F. sect. Eskia lineage (Table 1). By contrast, 1Cx values of the F. sect. Aulaxyper polyploids are between one-third and one-fourth smaller than that of F. rivularis. Genome sizes of hexaploids F. nigrescens subsp. nigrescens (12.07–12.22 pg) and subsp. microphylla (11.10–12.48 pg) and their mean monoploid genome (1Cx) values (2.01–2.02 and 1.85–2.08 pg, respectively) are similar to each other and equivalent though slightly smaller than those reported by Šmarda et al. (2008) for French, Czech and Italian populations of these taxa. Devesa et al. (2020) did not differentiate the two close subspecies in the Iberian Peninsula though they have been recognized by other authors (Kerguélen and Plonka 1989; Šmarda et al. 2008). The leaf morphology of our Formigal samples is like F. bartherei Timb.-Lagr., a taxon now considered to be a mere form of F. nigrescens subsp. nigrescens. A species growing in the same subalpine peat bog population as F. rivularis has been identified as F. heteromalla Pourr.; this plant shows hexaploid 2C values of 13.61 pg and a monoploid genome size of 2.27 pg (Table 1). The chromosome counts and genome size values obtained so far for French and Austrian populations of F. heteromalla indicate that it is an octoploid species (Kerguélen and Plonka 1989; Šmarda et al. 2008), whereas Devesa et al. (2020) did not recognize the species and subordinated it to hexaploid F. rubra L. subsp. rubra. However, the morphology of the plant, and its mountain marsh ecology, clearly differentiate it from that of typical nemoral to prairie-dwelling F. rubra subsp. rubra, corresponding to that of a hexaploid F. heteromalla-type species, as indicated previously by Kerguélen and Plonka (1989). Our data confirm this new ploidy level and provide the first genome size value for 6x F. heteromalla. The largest genome size of the studied F. sect. Aulaxyper species is that of F. trichophylla (16.23 pg; Table 1). Šmarda et al. (2008) 2C values of Italian populations of F. trichophylla subsp. asperifolia (13.1 pg) and subsp. trichophylla (13.2 pg) corresponded to hexaploid individuals although Kerguélen and Plonka (1989) and Al-Bermani et al. (1992) indicated an additional decaploid ploidy level for F. trichophylla subsp. asperifolia and Devesa et al. (2020) a decaploid ploidy level for both subspecies. Our 2C data correspond to an octoploid genome within F. sect. Aulaxyper, similar to that reported for other octoploid taxa of this lineage (e.g., F. rubra subsp. juncea 8x, 16.97 pg; Šmarda et al. 2008) thus being the first record of an octoploid ploidy level within F. trichophylla s.l. A monoploid genome (1Cx) value of 2.03 pg calculated for this octoploid species (Table 1) is similar to those inferred for hexaploids F. heteromalla and F. nigrescens subsp. nigrescens and microphylla, supporting the extreme reduction and downsizing of monoploid genomes in high F. sect. Aulaxyper polyploids, compared to diploid F. rivularis, and their potential allopolyploid origins (Šmarda et al. 2008; Moreno-Aguilar et al. 2022a).
References
Al-Bermani AK, Catalán P, Stace CA (1992) A new circumscription of Festuca trichophylla (Gaudin) K. Richter (Gramineae). Anales Jard Bot Madrid 52:209–220
Arndt S (2005) Systematik der Festuca valesiaca und Festuca laevigata-Gruppe in den Westalpen. PhD Thesis, Friedrich-Schiller-Universität Jena, Jena
Catalán P (2006) Phylogeny and evolution of Festuca L. and related genera of the subtribe Loliinae (Poeae, Poaceae). In: Sharma AK, Sharma A (eds) Plant genome. Biodiversity and evolution, vol. 1(D). Science Publishers, Enfield, pp 255–303
Catalán P, Torrecilla P, López JA, Olmstead RG (2004) Phylogeny of the festucoid grasses of subtribe Loliinae and allies (Poeae, Pooideae) inferred from ITS and trnL-F sequences. Molec Phyl Evol 31:517–541. https://doi.org/10.1016/j.ympev.2003.08.025
Catalán P, Torrecilla P, López-Rodríguez JA, Müller J (2006) Molecular evolutionary rates shed new lights on the relationships of Festuca, Lolium, Vulpia and related grasses (Loliinae, Pooideae, Poaceae). In: Bailey JP, Ellis RG (eds) Current Taxonomic Research on the British and European Flora. Botanical Society of the British Isles, London, pp 45–70
Catalán P, Torrecilla P, López JA, Müller J, Stace CA (2007) A systematic approach to subtribe Loliinae (Poaceae, Pooideae) based on phylogenetic evidence. Aliso 23:380–405. https://doi.org/10.5642/aliso.20072301.31
Catalán P, Acedo C, Marques I, Llamas F, Alonso A, Pérez-Collazos E, Viruel J, Sahuquillo E, Sancho MC, Komac B, Manso JA, Segarra-Moragues JG, Draper D, Villar L (2013) Genética del paisaje y ecología de pastos subalpinos de Festuca eskia, F. gautieri y F. ×picoeuropeana en los Parques Nacionales de Ordesa-Monte Perdido, Aigüestortes y Estany de Sant Maurici, y Picos de Europa. In: Ramírez L, Asensio B (eds) Proyectos de investigación en Parques Nacionales: 2009–2012. Organismo Autónomo Parques Nacionales, Ministerio de Medio Ambiente, Madrid, pp 39–70
Devesa J, Catalán P, Müller J, Cebolla C, Ortuñez E (2013) Checklist de Festuca L. (Poaceae) en la Península Ibérica. Lagascalia 33:183–274
Devesa JA, Martínez-Sagarra G, López-Nieto E, Muñoz-Rodríguez A, Cebolla C, Ortúñez E (2020) Festuca L. In: Devesa JA, Romero-Zarco C, Buira A, Quintanar A, Aedo C (eds) Flora iberica, vol. XIX. I. Real Jardín Botánico, CSIC, Madrid, pp 200–373
Diaz-Pérez A, Sequeira M, Santos-Guerra A, Catalán P (2008) Multiple colonizations, in-situ speciation, and volcanism-associated stepping-stone dispersals shaped the phylogeography of the Macaronesian red fescues (Festuca L., Gramineae). Syst Biol 57:732–749. https://doi.org/10.1080/10635150802302450
Doležel J, Binarová P, Lucretti S (1989) Analysis of nuclear DNA content in plant cells by flow cytometry. Biol Pl 31:113–120. https://doi.org/10.1007/BF02907241
Doležel J, Bartoš J, Voglmayr H, Greilhuber J (2003) Nuclear DNA content and genome size of trout and human. Cytometry 51A:127–128. https://doi.org/10.1002/cyto.a.10013
Ferrero LM, Fuente V (1996) Aportaciones al estudio cariológico de algunas especies del género Festuca L. endémicas del Mediterráneo Occidental. Bol Soc Brot Ser 2:303–308
Foggi B, Müller J (2009) Festuca. In: Valdés B, Scholz H (eds) Poaceae. Euro+Med Plantbase - the information resource for Euro-Mediterranean plant diversity. Available at: http://ww2.bgbm.org/_GBIFComp/query.asp
Fuente V, Ortúñez E (1988) Datos corológicos de algunos taxones ibéricos del género Festuca L. Lagascalia 15:465–473
Fuente V, Ortúñez E (1993) Festuca marginata subsp. andres-molinae, subsp. nov.para la Península Ibérica. Bot Complutensis 18:105–112
Fuente V, Ortúñez E (2001) Festuca L. section Eskia Willk. subgenus Festuca in the Iberian Peninsula. Folia Geobot 36:385–421. https://doi.org/10.1007/BF02899988
Fuente V, Ortúñez E, Ferrero L (1997) Contribución al conocimiento del género Festuca L. (Poaceae) en el País Vasco y Sistema Ibérico septentrional (Península Ibérica). Itinera Geobot 10:317–351
Fuente V, Ferrero LM, Ortúñez E (2001) Chromosome counts in the genus Festuca L. section Festuca (Poaceae) in the Iberian Peninsula. Bot J Linn Soc 137:385–398. https://doi.org/10.1111/j.1095-8339.2001.tb02333.x
Garnatje T, Garcia S, Vilatersana R, Vallès J (2006) Genome size variation in the genus Carthamus (Asteraceae, Cardueae): systematic implications and additive changes during allopolyploidization. Ann Bot (Oxford) 97:461–467. https://doi.org/10.1093/aob/mcj050
Inda LA, Segarra-Moragues JG, Müller J, Peterson PM, Catalán P (2008) Dated historical biogeography of the temperate Loliinae (Poaceae, Pooideae) grasses in the northern and southern hemispheres. Molec Phyl Evol 46:932–957. https://doi.org/10.1016/j.ympev.2007.11.022
Inda LA, Sanmartin I, Buerki S, Catalán P (2014) Mediterranean origin and Miocene-Holocene Old World diversification of meadow fescues and ryegrasses (Festuca subgen. Schedonorus and Lolium). J Biogeog 41:600–614. https://doi.org/10.1111/jbi.12211
Kerguélen M, Plonka F (1989) Les Festuca de la flore de France (Corse comprise). Bull Soc Bot Centre-Ouest Sér 2:10
Kerguélen M (1975) Les Gramineae (Poaceae) de la Flore française. Essai de mise au point taxonomique et nomenclaturale. Lejeunia Nouv Ser 75:1–343
Küpfer P (1974) Recherches sur les liens de parenté entre la flore orophile des Alpes et celle des Pyrénées. Boissiera 23:1–322
Loureiro J, Kopecký D, Castro S, Santos C, Silveira P (2007) Flow cytometric and cytogenetic analyses of Iberian Peninsula Festuca spp. Pl Syst Evol 269:89–105. https://doi.org/10.1007/s00606-007-0564-8
Marie D, Brown SC (1993) A cytometric exercise in plant DNA histograms, with 2C values for 70 species. Biol Cell 78:41–51. https://doi.org/10.1016/0248-4900(93)90113-s
Marques I, Draper D, López-Herranz ML, Garnatje T, Segarra-Moragues JG, Catalán P (2016) Past climate changes facilitated homoploid speciation in three mountain spiny fescues (Festuca, Poaceae). Sci Rep 6:36283. https://doi.org/10.1038/srep36283
Martínez-Sagarra G, Castro S, Mota L, Loureiro J, Devesa JA (2021) Genome size, chromosome number and morphological data reveal unexpected infraspecific variability in Festuca (Poaceae). Genes 12:906. https://doi.org/10.3390/genes12060906
Martínez-Sagarra G, Casimiro-Soriguer F, Castro S, Loureiro J, Devesa JA (2022) Cytogenetic, morphometric, and ecological characterization of Festuca indigesta Boiss. in the Southeast of Spain. Plants 11:693. https://doi.org/10.3390/plants11050693
Martínez-Sagarra G (2018) Taxonomic study of Festuca L. sect. Festuca (Poaceae) in the Iberian Peninsula. PhD Thesis, University of Cordoba, UCO Press, Cordoba
Minaya M, Hackel J, Namaganda M, Brochmann C, Vorontsova MS, Besnard G, Catalán P (2017) Contrasting dispersal histories of broad- and fine-leaved temperate Loliinae grasses: range expansion, founder events, and the roles of distance and barriers. J Biogeog 44:1980–1993. https://doi.org/10.1111/jbi.13012
Moreno-Aguilar MF, Arnelas I, Sánchez A, Viruel J, Catalán P (2020) Museomics unveil the phylogeny and biogeography of the neglected Juan Fernandez archipelago Megalachne and Podophorus endemic grasses and their connection with relict Pampean–Ventanian fescues. Frontiers Pl Sci 11:819. https://doi.org/10.3389/fpls.2020.00819
Moreno-Aguilar MF, Inda LA, Sánchez A, Arnelas I, Catalán P (2022a) Evolutionary dynamics of the repeatome explains contrasting differences in genome sizes and hybrid and polyploid origins of grass Loliinae lineages. Frontiers Pl Sci 13:901733. https://doi.org/10.3389/fpls.2022.901733
Moreno-Aguilar MF, Inda LA, Sánchez A, Catalán P, Arnelas I (2022b) Phylogenomics and systematics of overlooked Mesoamerican and South American polyploid broad-leaved Festuca grasses differentiate F. sects. Glabricarpae and Ruprechtia and F. subgen. Asperifolia, Erosiflorae, Mallopetalon and Coironhuecu (subgen. nov.). Plants 11:2303. https://doi.org/10.3390/plants11172303
Ortúñez E, Fuente V (1995) Reports (394–400) In: Kamari G, Felber F, Garbari F (eds) Mediterranean chromosome number reports. 5. Fl Medit 5:261–373
Ortuñez E, Fuente V (2004) Chromosome counts in the genus Festuca section Eskia Willk. (Poaceae) in the Iberian Peninsula. Bot J Linn Soc 146:331–337. https://doi.org/10.1111/j.1095-8339.2004.00320.x
Šmarda P, Bureš P, Horová L, Foggi B, Rossi G (2008) Genome size and GC content evolution of Festuca: ancestral expansion and subsequent reduction. Ann Bot (Oxford) 101:421–433. https://doi.org/10.1093/aob/mcm307
Torrecilla P, López-Rodríguez JA, Stancik D, Catalán P (2003) Systematics of Festuca L. sects. Eskia Willk., Pseudatropis Kriv., Amphigenes (Janka) Tzvel., Pseudoscariosa Kriv. and Scariosae Hack. based on analysis of morphological characters and DNA sequences. Pl Syst Evol 239:113–139. https://doi.org/10.1007/s00606-002-0265-2
Torrecilla P, Acedo C, Marques I, Diaz-Pérez AJ, López-Rodríguez JA, Mirones V, Sus A, Llamas F, Alonso A, Pérez-Collazos E, Viruel J, Sahuquillo E, Sancho MC, Komac B, Manso JA, Segarra-Moragues JG, Draper D, Villar L, Catalán P (2013) Morphometric and molecular variation in concert: taxonomy and genetics of the reticulate Pyrenean and Iberian alpine spiny fescues (Festuca eskia complex, Poaceae). Bot J Linn Soc 173:676–706. https://doi.org/10.1111/boj.12103
Acknowledgements
We thank Jordi Luque, Beatriz Tenas and Iñigo Zabalgogeazcoa for their valuable help in the material collections, Miquel Veny for his technical support in the plant cultivation at the Barcelona Botanical Institute, Jaume Comas (Centres Científics i Tecnològics, Universitat de Barcelona) for the assistance given in flow cytometric measurements, and two anonymous reviewers for their valuable comments to an early version of the manuscript. This research was supported by projects 2017SGR1116 and 2021SGR00315 from the Generalitat de Catalunya and a LMP82-21 project from the Gobierno de Aragón.
Funding
Open Access funding provided thanks to the CRUE-CSIC agreement with Springer Nature.
Author information
Authors and Affiliations
Contributions
PC and TG collected the material, TG and JV carried out the experimental part, PC and SP wrote the main manuscript. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Ethical approval
This manuscript has been approved by all co-authors, as well as the responsible authorities at the Botanical Institute of Barcelona.
Additional information
Handling Editor: Karol Marhold.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Garnatje, T., Catalán, P., Inda, L.A. et al. Genome size of grass Festuca mountain species from the southwestern European Pyrenees: variation, evolution, and new assessments. Plant Syst Evol 309, 29 (2023). https://doi.org/10.1007/s00606-023-01867-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00606-023-01867-x