Abstract
We analyzed the population genetic pattern of 12 fragmented Geropogon hybridus ecological range edge populations in Israel along a steep precipitation gradient. In the investigation area (45 × 20 km2), the annual mean precipitation changes rapidly from 450 mm in the north (Mediterranean-influenced climate zone) to 300 mm in the south (semiarid climate zone) without significant temperature changes. Our analysis (91 individuals, 12 populations, 123 polymorphic loci) revealed strongly structured populations (AMOVA ΦST = 0.35; P < 0.001); however, differentiation did not change gradually toward range edge. IBD was significant (Mantel test r = 0.81; P = 0.001) and derived from sharply divided groups between the northernmost populations and the others further south, due to dispersal or environmental limitations. This was corroborated by the PCA and STRUCTURE analyses. IBD and IBE were significant despite the micro-geographic scale of the study area, which indicates that reduced precipitation toward range edge leads to population genetic divergence. However, this pattern diminished when the hypothesized gene flow barrier was taken into account. Applying the spatial analysis method revealed 11 outlier loci that were correlated to annual precipitation and, moreover, were indicative for putative precipitation-related adaptation (BAYESCAN, MCHEZA). The results suggest that even on micro-geographic scales, environmental factors play prominent roles in population divergence, genetic drift, and directional selection. The pattern is typical for strong environmental gradients, e.g., at species range edges and ecological limits, and if gene flow barriers and mosaic-like structures of fragmented habitats hamper dispersal.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Species’ distribution ranges are commonly defined by ecological limits, which are most often determined by ecological gradients. As species approach their range limits, their populations typically become smaller and more fragmented (Bridle and Vines 2007). As a consequence, range edge (margins or verge) populations often feature decreased genetic diversity due to reduced gene flow as result of the more fragmented distribution or random genetic drift effects enhance genetic diversity due to neutral, negative, or positive mutations which can appear and increase in frequency over time (e.g., Ellstrand and Elam 1993). If gene flow patterns align with geographic distance (isolation by distance, IBD; Wright 1943), this affects the distribution of variation within and among range edge populations (Eckert et al. 2008; Sexton et al. 2009). If ecological gradients cause range limits, populations are generally more likely to maintain specialized genotypes that are well adapted to particular ecological conditions (e.g., Rehm et al. 2015). This is especially the case when populations that are geographically closer together are genetically more similar than populations that are further apart (IBD) and where genetic and environmental differences among populations (isolation by environment, IBE) are positively correlated. Especially in smaller and more fragmented range edge populations, positive, advantageous mutations are more likely to become fixed, as directional selection within and among range edge populations might be stronger than in central populations, where stabilizing selection through higher rates of gene flow tend to oppose effects of local selection and therefore limits adaptation (Hoffmann and Blows 1994; Lenormand 2002; Bridle and Vines 2007; Sexton et al. 2009). Under an IBD and IBE pattern, range edge populations eventually diversify and undergo niche evolution during adaptation to novel environments, or alternatively may depauperate where adaptation is prevented by small population size (Sexton et al. 2009).
For decades, evolutionary ecologists have investigated local adaptation across different systems and scales, however, rarely on micro-geographic scales. This is due to the assumption that high rates of gene flow prevent adaptive divergence at fine spatial grains (Richardson et al. 2014). However, several investigations on small geographic scales suggest that micro-geographic divergence is more widespread than commonly assumed (Kettlewell 1955; Antonovics and Bradshaw 1970; Steiner and Berrang 1990; Kavanagh et al. 2010; Willi and Hoffmann 2012; Krueger-Hadfield et al. 2013; Richardson and Urban 2013; Richardson et al. 2014).
Environmental association studies allow the investigation of the nature of local adaptation by identifying its leading causes (Savolainen et al. 2013; Gray et al. 2014). This is achieved by linking genetic variation to environmental variables (Manel et al. 2012); however, in non-model species with limited or absent genomic information, the identification of adaptive genetic variation can only be achieved indirectly. By investigating anonymous loci in numerous individuals, it is possible to detect outlier loci of ecological relevance that may be linked to adaptive genes (Haldane 1948; Endler 1986; Schmidt et al. 2008; Stinchcombe and Hoekstra 2008; Manel et al. 2012). Their distribution differs from alleles at neutral loci, and their correlation to environmental influences reveals potential indications for adaptations (Holderegger et al. 2010).
Environmental association studies have formerly concentrated on the correlation between genetic and geographic distances to analyze IBD, while more recent studies also incorporate multiple environmental variables with a main focus on environmental gradients (e.g., climatic-, elevation-, environmental- and habitat gradients) with climatic variables being the most commonly used (Gerber et al. 2004; Nahum et al. 2008; Nakazato et al. 2008; Manel et al. 2012; Jones et al. 2013; Gray et al. 2014; Harter et al. 2015). Environmental association studies by means of climatic variables generally comprise temperature and precipitation that over large geographic distances normally change simultaneously. Thus, it is often impossible to detangle one factor from another to determine the exact driving forces for putative adaptation (Linhart and Grant 1996). As a consequence, only a few studies could detect outlier loci in non-model plant species. In Campanula (Jones et al. 2013) and Cotinus (Lei et al. 2015), these outlier loci are clearly associated with precipitation, while Manel et al. (2012) in alpine plant species, Gray et al. (2014) in Andropogon, and Hübner et al. (2009) in Hordeum showed that precipitations in combination with temperature were the best environmental predictors.
Here, we investigated the genetic diversity and structure of range edge populations of the annual Geropogon hybridus (L.) Sch.Bip. along a steep precipitation gradient (450–300 mm) on a micro-geographic scale (45 km) without any significant temperature change. Using amplified fragment length polymorphism (AFLP) analysis (Vos et al. 1995), we asked the following questions: (1) Does genetic diversity and differentiation change gradually toward range edge? (2) Can we identify significant IBD or IBE pattern despite the micro-geographic scale of the study area? (3) Is there putative precipitation-related adaptation among the surveyed populations?
Materials and methods
Study species
Geropogon hybridus is a diploid annual herbaceous Asteraceae species, 10–40–(80) cm high. The flowering stem is erect, usually glabrous with narrowly linear, grass-like leaves. The pedicule is hollow or swollen below the capitula. The linear, long involucral bracts often exceed the capitulum. The zygomorphic flowers (from March to May) have pink to violet corollas, with dark purple anthers. The species is self-compatible. It is pollinated by a variety of generalist insect families that are abundant in the landscape. Achenes are dimorphic, narrowly fusiform, or cylindrical in outline. The outer achenes are smooth with a long awened scabridulous pappus, most likely adapted to endozoochorous dispersal (mammalians). The inner achenes are prominently ribbed with a long, plumose pappus consisting of 20 unequal bristles, most likely adapted to anemochorous dispersal (Bobrov and Tzvelev 2000). The species is widely distributed throughout the Mediterranean region and toward southeast Asia (Danin 2016, ICN 2009+), but occurrences are restricted to semidesert or more humid environments.
Study region and sampling
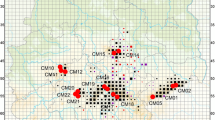
The study region is situated between the cities of Kirjat Gat and Be’er Sheva (31°24′00″–31°14′50″N, 34°48′30″–34°50′30″E) in the southern Judea Lowland, Israel (Fig. 1) and represents the southernmost range edge of G. hybridus in Israel (Giladi et al. 2011, Danin 2016). The northern part of the area is influenced by Mediterranean climate with an annual rainfall of 450 mm per year, whereas the southern part is situated in a much dryer, semiarid region with only 300 mm of annual rainfall (Ben-Gai et al. 1994; Goldreich 2003; Giladi et al. 2011). In contrast to the strong precipitation gradient, no significant temperature changes can be observed on this small micro-geographic scale (Goldreich 2003; Giladi et al. 2011).
Map and sampling sites of the study region in Israel. In the magnified sections are the Isohyete (2011; worldclim.org), gray color depicts natural habitats and white color indicates agriculture fields. Cake diagrams at the right show the cluster of the STRUCTURE analysis (∆K = 2) for each population. In the left box, the distribution of Geropogon hybridus in Israel is shown [GBIF.org (29th July 2016) GBIF Occurrence Download http://doi.org/10.15468/dl.d2zwzr]. Map was prepared with ArcGIS Desktop (ArcGIS Desktop 10.2.2., Esri)
The area is characterized by a fragmented agroecosystem with scattered mosaic-like patches of natural vegetation, where insulated rock layers appear close to the topsoil (Svoray et al. 2007; Yaacobi et al. 2007; Gemeinholzer et al. 2012). On these patches, we sampled a total of 12 G. hybridus populations, with four populations each in the north (N1–N4), in the Center (C1–C4), and in the south (S1–S4) of the investigation area (Fig. 1; Table 1). The distance between the different regions are N to C ~12 km, C to S ~10 km, and N to S ~23 km (distances refer to the shortest distances between each region). Additionally, for some analyses, populations were grouped according to precipitation gradient (450 mm: N1–4; 400–350 mm: C1–4 + S1–2; 300 mm: S3–4). At each site, we randomly sampled 12 individuals; however, in some cases, evaluable data could only be obtained from less (Table 1; actual number of samples per population are provided). Samples of fresh and healthy leaves were randomly collected in the field and immediately dried in silica gel and stored at 4 °C until further processing. Total population size was estimated by counting the individuals in a plot of 150 × 150 m.
Laboratory work and data analyses
DNA isolation and subsequent AFLP analysis were conducted as described in (Lauterbach et al. 2011). After an initial primer screening on eight samples (N1_1; N3_1; C1_1; C1_3; C3_1; C3_3; S2_1; S4_1) from the three subgroups each to exclude ascertainment bias (N, C and S) testing 12 different primer combinations, we chose three selective primer combinations that proved to be informative and provided clear bands which were sufficiently polymorphic to show variation within and among populations: EcoRI_AGC/MseI_CCA, EcoRI_AGC/MseI_CTC, and EcoRI_ACA/MseI_CGA. The laboratory setup was designed in a 96-well format. Thus, each primer could be analyzed in one run, assuming that AFLP produces clear and reproducible bands (Jones et al. 1997); therefore, we did not add replicates for error rate estimation. AFLP data matrix establishment was carried out using Genographer, version 2.1.4 (Banks and Benham 2008). Rare bands were included in the analysis (Online Resources 1 and 2).
Genetic diversity of the surveyed populations was estimated as percentage of polymorphic loci (PLP) and as Nei’s gene diversity (H e ) (Nei 1987) using AFLPsurv 1.0 (Vekemans 2002). To test for significant differences of diversity estimates between groups of populations (i.e., N vs. C vs. S), we applied a one-way analysis of variance (ANOVA) in R 3.0.2 (R Development Core Team Team 2013) using the Shapiro–Wilk test to check for normality (P = 0.41) and the Tukey HSD post hoc test to explore significant ANOVA results. The Bartlett test (P = 0.21) indicated no evidence for non-constancy of variance (heteroscedasticity) in our data.
Patterns of genetic population structure were visualized with a principal component analysis (PCA) using the R package ADEGENET version 1.4-2 (Jombart 2008). Additionally, we explored individuals’ genetic affiliation to genetic clusters using STRUCTURE version 2.3.3 (Pritchard et al. 2000). We explored individuals’ genetic affiliation to genetic clusters by applying the admixture model, 100,000 MCMC replicates, with a burn-in period of 50,020 and repeats per run for each chosen cluster number (i.e., K = 1–12), PLOIDY = 2 and RECESSIVEALLELES = 1. For all other settings, the default options were used. To identify the most likely K modal distribution, delta K (Evanno et al. 2005) was determined using STRUCTURE HARVESTER (Earl and von Holdt 2012). Corresponding graphs were constructed with DISTRUCT (Rosenberg 2004).
Genetic variation among groups of populations (ΦCT), among populations within groups (ΦSC), and within populations (ΦST) was partitioned with hierarchical analyses of molecular variance (AMOVA) using ARLEQUIN 3.5.1.2 (Excoffier and Lischer 2010). Additionally, pairwise ΦST values were estimated among populations. Significance levels were determined after 9999 permutations.
To evaluate patterns of isolation by distance (IBD) and isolation by environment (IBE), we tested for the relationships between pairwise ΦST values and geographic distance, and pairwise ΦST values and difference in annual precipitation, respectively. Pairwise and partial Mantel tests were conducted using the ‘vegan’ library in R (Oksanen et al. 2007) with 999 permutations. For IBD analyses, we used Euclidian geographic distances, and for IBE analyses, we constructed a matrix displaying pairwise precipitation distances (i.e., the difference of annual rainfall in mm between pairs of populations). For both types of analyses, we always tested (i) the complete data set of all populations and (ii) the two pairs of adjacent groups of populations (i.e., N–C and C–S, respectively).
To detect signatures of selection that could indicate putative adaptations to particular conditions along the gradient, we applied two differentiation-based genome scan methods: BAYESCAN 2.1 (Foll and Gaggiotti 2008) and DFDIST (Beaumont and Balding 2004) as included in the workbench MCHEZA (Antao and Beaumont 2011). BAYESCAN analyses were run with a burn-in of 50,000 iterations, a sample size of 10,000, and a thinning interval of 50, resulting in a total of 550,000 iterations. An additional burn-in was carried out by 20 short pilot runs of 5000 iterations. DFDIST analyses were conducted with 50,000 simulations, using the combined ‘Neutral mean FST’ and ‘Force mean FST’ algorithms and correcting results for multiple comparisons by setting the false discovery rate to 0.1. To consider a locus under selection, we chose a conservative approach. In BAYESCAN, only loci with a posterior probability over 0.99 and that were detected with a false discovery rate (FDR) of 0.01 were considered to be putatively under divergent selection. Comparably, in MCHEZA, only outliers that were identified at the 99% confidence interval were treated as putative adaptive loci. To test if adaptations occur either gradual along the gradient or are linked to a specific precipitation threshold (or any other biological significant line) for each genome scan method, we conducted two types of analyses: (i) a ‘global analysis’ including all populations, and (ii) ‘N–C specific’ and ‘C–S specific’ analyses including either only northern and central or central and southern populations.
To further substantiate the results of the two differentiation-based genome scans and to more directly test for signatures of precipitation-related adaptation, we applied the spatial analysis method (SAM; Joost et al. 2007), which uses multiple univariate logistic regression to test for correlations between environmental variables and binary molecular data. In our case, the site-specific mean values of mean annual precipitation in mm (Table 1) were assigned to the AFLP data. SAM uses the individual as reference unit, functions independently of any presumed population structure and is largely assumption free (Joost et al. 2007). Only if the two statistical tests implemented in SAM (likelihood ratio G and Wald test) reject the null hypothesis, a model is considered as significant (Joost et al. 2007). Also in SAM, the Bonferroni correction of the significance level for multiple comparisons was applied, which corresponds to a 99% confidence interval.
Results
Genetic diversity and differentiation
AFLP analyses with three different primer combinations and 91 analyzed individuals resulted in 123 unambiguously scorable polymorphic loci, ranging from 50 to 450 base pairs. The mean genetic diversity of the investigated Geropogon hybridus populations was H e = 0.26, with genetic diversities ranging from H e = 0.21 (S4) to H e = 0.32 (C4, Table 1). The region specific mean values of diversity estimates (Table 1) varied from H eN = 0.26 to H eC = 0.29 with the lowest values in the south (H eS = 0.24). Genetic diversity among the three regions differed significantly: F = 4.33, P = 0.048. Post hoc tests showed a difference between central and south populations (P < 0.05). No differences in gene diversity estimates among precipitation specific groupings could be detected.
Principal component analysis depicted two large and distinct cluster that separated northern individuals from central and southern individuals along the first axes (Online Resource 3). Notably, one of the northern and three of the central individuals reflected contrary genotypic patterns, resembling those of the central or northern cluster, respectively. However, none of the southern individuals featured a northern-specific genotypic pattern. Overall, the first three principal components accounted for 31.4, 7.2, and 5.3% of the genetic variation.
The STRUCTURE analysis clearly confirmed the PCA results depicted two large and distinct clusters that separated northern individuals from central and southern individuals, detecting an optimal cluster number of ΔK = 2 (Fig. 2, Online Resource 4) that separated populations between the northern and central–south region (Fig. 1). Only few individuals of the northern and central site deviate from their respective group specific genetic pattern (i.e., in N2, C1 and C4).
Analysis of molecular variance (AMOVA) resulted in a global ΦST of 0.29 (Table 2) and population pairwise ΦST values between 0.000 and 0.406 (Online Resource 5), indicating strong genetic differentiation between some of the populations (i.e., N1–4 vs. C1–4, and S1–4). Hierarchical AMOVA (Table 2) showed that 33.1% of genetic variance resided between study regions, whereas only 1.9% resided among populations within study regions. However, most variation was partitioned within populations (65%). When separating populations only in two regional groups, for ‘N versus C + S’ a total of 42% and for ‘N + C versus S’ only 13.5% of variance resided between groups (Online Resource 6), further depicting that the main population differentiation is located between northern and central sites. Simple Mantel tests revealed strong and significant IBD and IBE patterns (Table 3) for both, the ‘complete’ and the ‘north-central’ data set (Mantel r values between 0.76 and 0.86) but not for the ‘central–south’ data set (Mantel r = 0.29 and −0.02). To further unravel the correlated effects of IBD and IBE, we applied partial Mantel tests. These still revealed significant relationships for the complete data set (IBD controlling for precipitation: r = 0.54; IBE controlling for geographic distances: r = 0.37) but not for the two reduced data sets (Table 3).
Outlier detection
The two differentiation-based genome scan approaches detected 15 (DFDIST) and 12 (BAYESCAN) putatively adaptive outlier loci, or loci which might be in the same linkage group as loci under adaptation, for the complete data set, respectively. Of these, 11 were similarly identified by both approaches (Table 4, Online Resource 7). Whereas, for the ‘central–south’ data set, none of the two approaches detected any outliers, for the ‘north-central’ data set, at least BAYESCAN identified nine loci to be potentially under divergent selection. Almost, all of these ‘north-central’-specific outliers (8/9) were similarly detected by DFDIST and BAYESCAN in the complete data set (Online Resource 7), indicating that putative differences in the selection regime are located between the north and central sites and do not gradually occur over the complete north–south gradient.
For correlation with mean annual precipitation data, SAM analysis revealed a total of 39 AFLP loci that showed a significant association at the 99% confidence interval. Interestingly, all 11 outliers that were similarly detected by the two differentiation-based genome scan approaches likewise were identified by SAM to be associated with the precipitation level (Table 4, Online Resource 7).
Discussion
Isolation by distance
Our population genetic investigation in fragmented range edge Geropogon hybridus populations along a steep precipitation gradient supports theory that range edge populations often feature reduced gene flow as result of the more fragmented distributions (Sexton et al. 2009). Other potential drivers explaining this pattern could be genetic bottlenecks (i.e., small population sizes), or selection differences. We detected potential IBD with the simple Mantel test (N–C–S r = 0.81; P = 0.001) and ΦST values (0.35; P < 0.001) indicative of a very strong differentiation along a short geographic distance. The findings are similar to the ones in Catananche lutea L., another annual Asteraceae species surveyed in the same investigation area (Gemeinholzer et al. 2012). This species co-occurs in the same vegetation along the same ecologic gradient under similar influences of fragmentation and also revealed variation to be greatest at largest distances, featuring an IBD scenario. Our analysis also supports the results of the summary analysis conducted by Sexton et al. (2014), who assessed gene flow with respect to environmental gradients in 110 published investigations. They found IBD to be the strongest pattern observed among plant studies.
IBD demands that the distribution of plots follows a linear model. The design of our sample setup in a natural environment of island like patches of natural vegetation in an agricultural surrounding takes this into account as far as possible. However, this strong overall population differentiation in our investigation was mainly driven by differences within northern populations, whereas central and southern populations alone showed no obvious IBD and only very weak population differentiation. Generally, significant IBD values can either be the result of continuous spatial variation in allele frequencies across the landscape, or they may derive from sharply divided groups where allele frequencies change rapidly at the point of division, due to environmental or dispersal limitations (Lindström et al. 2013) which may result in unaccounted overlapping effects of geographic barriers or potential bias connected to IBD (Meirmans 2012). Our results of the simple Mantel test (N–C) indicate significant barriers to gene flow and confirm restricted dissemination between northern and central populations, but not between central and southern populations, which is further corroborated by the STRUCTURE analyses. Meirmans (2012) argued, that by inducing allelic alterations in the sampling, the effect of geographic barriers may erroneously imply the occurrence of IBD (Silva et al. 2016) and IBD interpretations than, should be treated with caution.
Gene flow barriers may arise due to (i) factors restricting survival or persistence, or (ii) vector limitations in diaspore dispersal (pollen or seeds) as result of biotic or abiotic (physical or ecological) alterations. While dispersal limitation has been shown to limit species’ distributions at larger spatial scales (Eriksson 1998), its role in structuring communities at smaller scales has received less attention (Emery et al. 2009). No specific information about G. hybridus vector limitations in the investigation area is available, and the specific groups of insect pollinators are not known. However, composites are commonly visited by a wide variety of hymenoptera and coleoptera (Cheplick 1987; Bosch et al. 1997; Kaul et al. 2000) for many of which the distances on our micro-geographic mosaic-like vegetation patches are still in effective reach (e.g., Kwak et al. 1998; Morandin and Winston 2006). In addition, there are no seed dispersal distances for G. hybridus available; however, they are likely similar to the closely related and comparatively similar seed dispersed Hypochaeris radicata L. that has shown most seeds to spread within a distance of 100 m and rarely reaching distances up to 400 m (Soons et al. 2004). A population genetic-based maximization-of-explained-variance procedure resulted in a threshold distance of 3.5 km above which H. radicata populations were effectively genetically isolated (Mix et al. 2006). G. hybridus dispersal distances are probably quiet alike; however, isolation between fragmented habitat patches additionally aggravates the problem, not only because dispersal distance influences gene flow, but also due to the reduced likelihood that propagules reach suitable environments for establishment. The gradient-like structure of the project set-up with more or less evenly distributed mosaic-like populations in three regions across the investigation area was meant to account for that and clearly indicates that the statistically significant barrier of gene flow identified by both the simple and partial Mantel tests between N1–N4 and C1–S4 cannot solely be explained by geographic distances limiting dispersal. We found no explanation for the observed split into a northern and a southern gene pool based on soil type. Both locations, Galon in the north and Dvir in the south are characterized by grumosolic soil, with a higher content of loess in Dvir, whereas the central collecting point Lachish is situated on Redzina soil (Dan and Raz 1970).
In conclusion, other vector limitations or environmental influences must have contributed to the detected barrier between northern and central populations, either due to alterations in pollen or seed dispersal capacities or modifications in recruitment rates (Primack and Miao 1992; Nathan and Muller-Landau 2000; Clarke and Davison 2001; Givnish 2010). A possible hypothesis for the coherence of the central population Lachish with the southern population of Dvir might be the cultural history of Lachish. Situated on a strategic important hill, the area of Lachish was colonized around 6500 BC. Between 900 and 800 BC, Lachish developed into the second important garrison and residence town after Jerusalem (Utzschneider 2005: 46; Velikovsky 2009: 53). Located at the old caravan route between Syria and Egypt (Salvator 1879), Lachish was a center for trading with an own caravansary where most of the transport between Hebron and Gaza in south-east direction happened. In the 14th century, the large amounts of pilgrims from the Sinai region to Jerusalem also followed the old caravan route via Be’er Sheva to Hebron, laterthey made the pilgrimage by the old coast-way to Gaza and then north-east to Jerusalem (Robinson and Smith 1841: 736). In none of the records, we found any evidence for a transport route from the Lachish area to the north via Galon or Kiryat Gat. Thus, a possible explanation for the observed split into two genetic detectable entities might be an anthropogenic land use scenario, by which a dispersal limitation of the up to 3000 years isolation of the central and southern population (which for an annual plant is equal to 3000 generations) has led to a genetic drift at the edge of the distribution range. Caravanning from Lachish south to Be’er Sheva might have expanded the range of an edge distributed genotype to the southern area of Dvir, the opposite movement kept these populations in contact with the Lachish populations. Since no further route into the northern part of the species range existed, a further exchange between the larger northern populations and the smaller southern populations was hampered, leading to the now observable split within G. hybridus. In a further step, adaptation of the central and southern population to the low amount of precipitation might has occurred.
Theory predicts that IBD pattern often coincide with pattern indicative for niche evolution as result of strong selection and micro-geographic adaptation, or genetic impoverishment due to small population size and reduced gene flow from central populations (Ehrlich and Raven 1969; Nosil et al. 2005; Edelaar et al. 2008; Sexton et al. 2009; Urban 2011; Richardson and Urban 2013). IBD can especially be found in populations of fragmented habitats toward range edges (van Treuren et al. 1994; Young et al. 1996; Griffin and Barrett 2004). Eckert et al. (2008) reviewed 134 investigations to evaluate population structures across species’ geographic ranges by comparing central and range edge genetic population structures and found in 64% within-population genetic diversity declined toward range edges. In G. hybridus, however, nonsignificant lower H e values and equally high population sizes at range edge populations (Table 1) render genetic impoverishment unlikely and rather points to inter-population genetic divergence as result of increased random genetic drift, selection or adaptation (e.g., Young et al. 1996; Sork et al. 1999; Lowe et al. 2005; Aguilar et al. 2008). The mean genetic diversity of G. hybridus (Ø H e = 0.262) over the whole study region was almost twice as high as in the comparable annual C. lutea (Gemeinholzer et al. 2012; Ø H e = 0.136) in the same study area, but was comparatively low to investigations in fragmented solely out crossing H. radicata populations (Mix et al. 2006; Ø H e = 0.88). Our values are similar to findings in other Asteraceae species across their whole distribution range, e.g., Leucochrysum albicans var. tricolor (DC.) Paul G.Wilson (Morgan et al. 2013, H e = 0.13–0.39, with allozyme markers), Ixeridium dentatum subsp. nipponicum (Nakai) J.H.Pak & Kawano (Tanaka et al. 2014), Centaurea cineraria L. (Synonym: C. gymnocarpa Moris & De Not., Guarino et al. 2013; H e = 0.027–0.567, with SSR markers) and Hieracium eriophorum St.-Amans (Frey et al. 2012), thus are comparatively high on this rather local scale. Similar high levels of genetic diversity were also found in Grevillea barklyana F.Muell. ex Benth (Hogbin et al. 1998) and Primula (P. veris L. and P. vulgaris Huds; van Rossum et al. 2004) in comparative analyses between central and range edge populations.
Isolation by environment
High genetic diversity in combination with IBD at species’ distribution ranges may be indicative of diversifying areas where ecological niche evolution can occur. This may happen in the course of adaptations to environmental alterations (Sexton et al. 2009; Mallet et al. 2014). Especially, at strong climatic gradients, environmental variability can contribute to patterns of interspecific genetic variation and can act as ecological driver promoting population divergence (Huang et al. 2015). The positive IBE pattern in G. hybridus indicates genetic variation to be correlated with reduced precipitation. However, as shown by partial Mantel tests, both (‘potential,’ see “Discussion” above) IBD and IBE are present in the population genetic data (Maassen and Bakker 2001; Shafer and Wolf 2013). Such patterns may occur when IBD is really strong and environmental spatial autocorrelation is low, as geographic distance then functions as suppressor to the IBE correlation. By conducting partial Mantel tests controlling for either geography or precipitation, we could demonstrate that both factors significantly contributed to the population genetic data. This pattern changed however when we analyzed the populations at both sides of the hypothesized gene flow barrier separately. Toward range edge, we found low IBD and no IBE correlations, thus the vast amount of unaccounted variation there must largely be due to other non-spatially structured biological or unmeasured environmental variables, and/or random processes triggered by ecological drift and dispersal (Legendre et al. 2009; Huang et al. 2015). In contrast, the northern populations featured highly positive IBD and IBE values with clear indications that the enhanced geographic distance here is more strongly affecting the population genetic pattern than precipitation thus IBD may drive IBE.
Several investigations confirmed high levels of divergence or local adaptation for populations distributed across climatic gradients, and particularly, steep environmental gradients can affect gene flow and local adaptation (Aitken et al. 2008; Leimu and Fischer 2008; Hereford 2009; Alberto et al. 2013; Savolainen et al. 2013; Rehm et al. 2015). Rundle and Nosil (2005) stated that divergent selections between distinct environments are the best understood drivers for speciation and population differentiation (see also Mallet et al. 2014). However, this mainly accounts for population genetic investigations across large geographic scales, while on micro-geographic scales, this is still relatively unexploited.
Outlier loci
By identifying outlier loci, we sought to determine how genetic differentiation is affected by sharp environmental gradients (Gray et al. 2014). We could identify 11 loci which putatively undergo diversifying selection and which moreover could be associated with precipitation levels. In the literature review about genomic heterogeneities, Nosil et al. (2009) found that many studies report 5–10% of loci to be outlier. This is in concordance with our analysis where 8.9% variation significantly correlates to the precipitation-related environmental gradient and which suggests that rainfall may put considerable spatially divergent pressure on the G. hybridus genotypes even on micro-geographic scales. However, unexpected was that potentially IBE associated outliers were found in the N populations instead of the marginal populations, indicative of the unexpected pattern that core populations may drive a potential IBE associated pattern.
Due to the applied anonymous marker technique AFLP, it is still an open question if the loci—or some of them—are indeed indicative of natural selection, or other adaptive and non-adaptive processes leading to non-random gene flow, or rather if they account for false-positive observations. Even though AFLP markers are frequently used for outlier detection, it is well known that the anonymous marker technique may cause statistical bias due to homoplastic and non-independent characters, especially for smaller fragments (Vekemans 2002; Bonin et al. 2007; Caballero et al. 2008). As AFLP markers typically are <500 bp, they most likely fall in noncoding regions; thus, markers with potential outlier behavior will be in close linkage with the potential genes under selection rather than inside the gene sequences itself, or they may act as regulatory elements (Stinchcombe and Hoekstra 2008; Butlin 2010; Nunes et al. 2012). AFLP genome scans of potential outlier loci mostly detect noncoding fragments, often with repetitive or transposable elements, or the characters derive from areas of low recombination which are often the result of non-adaptive processes (Minder and Widmer 2008; Wood et al. 2008; Nunes et al. 2012; Paris and Despres 2012; Cruickshank and Hahn 2014). In such cases, the outlier regions could simply reflect demography, spatially heterogeneous selection (Guttman et al. 2009), background selection in areas containing genomic features (i.e., centromeres), or sequence assembly artifacts (Shafer et al. 2015) and thus are indicative for standing variation, sometimes referred to as soft selective sweeps due to contemporary disruptive natural selection. On the other hand, outliers could be the results of new variants (hard selective sweeps), e.g., by chromosomal regions with reduced recombination (Strasburg et al. 2012). Linkage as result of background or purifying selection or genetic hitchhiking can additionally affect differentiation which is known to be stronger in coding than in noncoding regions and might account for false-positive or false-negative observations (Lotterhos and Whitlock 2015). The influence of genetic hitchhiking is dependent on multiple evolutionary parameters, such as recombination rate, population size, the strength and mode of selection, loci number, and analytical methods (Hermisson and Pennings 2005; Oleksyk et al. 2010; Pritchard and Di Rienzo 2010; Tiffin and Ross-Ibarra 2014; Lotterhos and Whitlock 2015). In our analysis, several indications point to selective sweeps derived from standing variation, due to the comparatively high mutation rates on this micro-geographic scale, and as theory predicts standing variation to support mutations to sweep rapidly to fixation (Hermisson and Pennings 2005). Selective sweeps result in comparatively low variation at linked sites and high divergence between populations of different selective environments (Strasburg et al. 2012), which is in concordance to the pattern found here in the range edge populations of G. hybridus. To minimize false-positive observations, researchers often rely on very high thresholds which increase the false-negative rate but strongly reduce the probability to wrongly identify loci in linkage with selective sweeps (Thompson et al. 2003).
Several other outlier tests have been developed beyond the outlier tests applied here (Beaumont and Nichols 1996; Vitalis et al. 2001; Beaumont and Balding 2004; Foll and Gaggiotti 2008; Excoffier et al. 2009; Bonhomme et al. 2010; Duforet-Frebourg et al. 2014) with several of them also accounting for genotype and environment associations by considering relations between allele frequencies and environmental parameters (Joost et al. 2007; Günther and Coop 2013; Gautier 2015; Lotterhos and Whitlock 2015). However, these analyses often suffer from high false-positive rates as they do not explicitly control for population structure in the test statistics (Sternberg et al. 2009; Meirmans 2012; De Mita et al. 2013; Lotterhos and Whitlock 2015). This might also be the case in our analysis, as populations in a landscape typically exhibit isolation by distance, and spatial autocorrelation in allele frequencies can cause false-positive associations between gene frequencies and the environment by chance. Natural populations would adapt to spatially heterogeneous environments at different spatial scales; therefore, geographically close populations in different selective environments would probably provide less false-positive hits than from distant populations where neutral demographic history might influence the pattern. Lotterhos and Whitlock (2015) state that the informative value of outlier-detections by associating genotype and environment associations is higher when populations physically near to each other (and therefore genetically similar for neutral genes) and yet in different environments (and therefore differentiated by selection) are being compared, like in our investigation.
Conclusions
The population genetic analysis of the annual Asteraceae Geropogon hybridus along a strong environmental gradient in a mosaic-like fragmented habitat revealed no significant population genetic diversity decline toward range edges. Contrary to expectations, population differentiation did not change gradually from the Mediterranean-influenced climate zone to the central and southern populations toward range edge in a more semiarid climatic region along a raster like sampling scheme. IBD and IBE were significant, despite the micro-geographic scale of the study area and partial Mantel tests controlling for either geography or precipitation showed that IBD and IBE significantly contribute to the population genetic divergent pattern over the whole investigation area. This pattern diminished when the hypothesized gene flow barrier was taken into account. The non-verge populations (N1–N4) featured highly positive IBD and IBE values with clear indications that the enhanced geographic distance here has more strongly affected the population genetic pattern than precipitation. Toward range edge (C1–S4), we found low IBD and no IBE correlations; thus, variation here must largely be due to other non-spatially structured biological or unmeasured environmental variables, and/or drift and dispersal.
By conducting environmental association studies, we could detect 11 outlier loci toward range edges which potentially evolved under selection, and which clearly indicate intraspecific population adaptive responses correlated to ecological drivers, in this case, precipitation. The results suggest that environmental factors can play prominent roles in population divergence, genetic drift, and directional selection, even on micro-geographic scales. This is especially the case along strong environmental gradients at species range edges when gene flow barriers and mosaic-like structures of fragmented habitats hamper dispersal. The results provide indications about potential niche evolution during adaptation. This highlights the significance of gene flow barriers, especially along fragmented range edge populations toward species’ ecological limits.
References
Aguilar R, Quesada M, Ashworth L, Herrerias-Diego Y, Lobo J (2008) Genetic consequences of habitat fragmentation in plant populations: susceptible signals in plant traits and methodological approaches. Molec Ecol 17:5177–5188. doi:10.1111/j.1365-294X.2008.03971.x
Aitken SN, Yeaman S, Holliday JA, Wang T, Curtis-McLane S (2008) Adaptation, migration or extirpation: climate change outcomes for tree populations. Evol Appl 1:95–111. doi:10.1111/j.1752-4571.2007.00013.x
Alberto FJ, Aitken SN, Alía R, González-Martínez SC, Hänninen H, Kremer A, Lefèvre F, Lenormand T, Yeaman S, Whetten R, Savolainen O (2013) Potential for evolutionary responses to climate change—evidence from tree populations. Glob Chang Biol 19:1645–1661. doi:10.1111/gcb.12181
Antao T, Beaumont MA (2011) Mcheza: a workbench to detect selection using dominant markers. Bioinformatics (Oxford, England) 27:1717–1718. doi:10.1093/bioinformatics/btr253
Antonovics J, Bradshaw AD (1970) Evolution in closely adjacent plant populations VIII. Clinal patterns at a mine boundary. Heredity 25:349–362. doi:10.1038/hdy.1970.36
Banks TW, Benham JJ (2008) Genographer Version: 2.1.4. Montana State University, Bozeman. Available at: http://sourceforge.net/projects/genographer
Beaumont MA, Balding DJ (2004) Identifying adaptive genetic divergence among populations from genome scans. Molec Ecol 13:969–980. doi:10.1111/j.1365-294X.2004.02125.x
Beaumont MA, Nichols RA (1996) Evaluating loci for use in the genetic analysis of population structure. Proc R Soc B 263:1619–1626. doi:10.1098/rspb.1996.0237
Ben-Gai T, Bitan A, Manes A, Alpert P (1994) Long-term changes in annual rainfall patterns in southern Israel. Theor Appl Climatol 49:59–67. doi:10.1007/BF00868190
Bobrov EG, Tzvelev NN (eds) (2000) Flora of the USSR, vol. 29. Compositae. Tribe Cichorieae. Smithsonian Institution Libraries, Washington, DC
Bonhomme M, Chevalet C, Servin B, Boitard S, Abdallah JM, Blott S, San Cristobal M (2010) Detecting selection in population trees: the Lewontin and Krakauer test extended. Genetics. doi:10.1534/genetics.110.117275
Bonin A, Ehrich D, Manel S (2007) Statistical analysis of amplified fragment length polymorphism data: a toolbox for molecular ecologists and evolutionists. Molec Ecol 16:3737–3758. doi:10.1111/j.1365-294X.2007.03435.x
Bosch J, Retana J, Xim C (1997) Flowering phenology, floral traits and pollinator composition in a herbaceous Mediterranean plant community. Oecologia 109:583–591. doi:10.1007/s004420050120
Bridle JR, Vines TH (2007) Limits to evolution at range margins: when and why does adaptation fail? Trends Ecol Evol 22:140–147. doi:10.1016/j.tree.2006.11.002
Butlin RK (2010) Population genomics and speciation. Genetica 138:409–418. doi:10.1007/s10709-008-9321-3
Caballero A, Quesada H, Rolan-Alvarez E (2008) Impact of amplified fragment length polymorphism size homoplasy on the estimation of population genetic diversity and the detection of selective loci. Genetics 179:539–554. doi:10.1534/genetics.107.083246
Cheplick GP (1987) The ecology of amphicarpic plants. Trends Ecol Evol 2:97–101. doi:10.1016/0169-5347(87)90166-2
Clarke PJ, Davison EA (2001) Experiments on the mechanism of tree and shrub establishment in temperate grassy woodlands: seedling emergence. Austral Ecol 26:400–412. doi:10.1046/j.1442-9993.2001.01124.x
Cruickshank TE, Hahn MW (2014) Reanalysis suggests that genomic islands of speciation are due to reduced diversity, not reduced gene flow. Molec Ecol 23:3133–3157. doi:10.1111/mec.12796
Dan J, Raz Z (1970) Israël. Soil association map. North and south. 1:250.000. Survey of Israel, Tel Aviv
De Mita S, Thuillet A, Gay L, Ahmadi N, Manel S, Ronfort J, Vigouroux Y (2013) Detecting selection along environmental gradients: analysis of eight methods and their effectiveness for outbreeding and selfing populations. Molec Ecol 22:1383–1399. doi:10.1111/mec.12182
Duforet-Frebourg N, Bazin E, Blum MGB (2014) Genome scans for detecting footprints of local adaptation using a Bayesian factor model. Molec Biol Evol 31:2483–2495. doi:10.1093/molbev/msu182
Earl DA, von Holdt BM (2012) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conservation Genet Resources 4:359–361. doi:10.1007/s12686-011-9548-7
Eckert CG, Samis KE, Lougheed SC (2008) Genetic variation across species’ geographical ranges: the central-marginal hypothesis and beyond. Molec Ecol 17:1170–1188. doi:10.1111/j.1365-294X.2007.03659.x
Edelaar P, Siepielski AM, Clobert J (2008) Matching habitat choice causes directed gene flow: a neglected dimension in evolution and ecology. Evolution 62:2462–2472. doi:10.1111/j.1558-5646.2008.00459.x
Ehrlich PR, Raven PH (1969) Differentiation of populations. Science 165:1228–1232
Ellstrand NC, Elam DR (1993) Population genetic consequences of small population size: Implications for plant conservation. Annual Rev Ecol Syst 24:217–242. doi:10.1146/annurev.es.24.110193.001245
Emery NC, Stanton ML, Rice KJ (2009) Factors driving distribution limits in an annual plant community. New Phytol 181:734–747. doi:10.1111/j.1469-8137.2008.02676.x
Endler JA (1986) Natural selection in the wild. Princeton University Press, Princeton
Eriksson A (1998) Regional distribution of Thymus serpyllum: management history and dispersal limitation. Ecography 21:35–43. doi:10.1111/j.1600-0587.1998.tb00392.x
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Molec Ecol 14:2611–2620. doi:10.1111/j.1365-294X.2005.02553.x
Excoffier L, Lischer HEL (2010) Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Molec Ecol Resources 10:564–567. doi:10.1111/j.1755-0998.2010.02847.x
Excoffier L, Hofer T, Foll M (2009) Detecting loci under selection in a hierarchically structured population. Heredity 103:285–298. doi:10.1038/hdy.2009.74
Danin A (2016) Flora of Israle Online. Available at: http://flora.org.il/en/plants. Accessed 1 Jan 2016
Foll M, Gaggiotti O (2008) A genome-scan method to identify selected loci appropriate for both dominant and codominant markers: a Bayesian perspective. Genetics 180:977–993. doi:10.1534/genetics.108.092221
Frey DJ, Haag CR, Kozlowski G, Tison J, Mráz P (2012) High genetic and morphological diversity despite range contraction in the diploid Hieracium eriophorum (Asteraceae) endemic to the coastal sand dunes of south-west France. Bot J Linn Soc 169:365–377. doi:10.1111/j.1095-8339.2012.01215.x
Gautier M (2015) Genome-wide scan for adaptive divergence and association with population-specific covariates. Genetics 201:1555–1579. doi:10.1534/genetics.115.181453
Gemeinholzer B, May F, Ristow M, Batsch C, Lauterbach D (2012) Strong genetic differentiation on a fragmentation gradient among populations of the heterocarpic annual Catananche lutea L. (Asteraceae). Pl Syst Evol 298:1585–1596. doi:10.1007/s00606-012-0661-1
Gerber J, Baltisberger M, Leuchtmann A (2004) Effects of a snowmelt gradient on the population structure of Ranunculus alpestris (Ranunculaceae). Bot Helv 114:67–78. doi:10.5169/seals-74720
Giladi I, Ziv Y, May F, Jeltsch F (2011) Scale-dependent determinants of plant species richness in a semi-arid fragmented agro-ecosystem. J Veg Sci 22:983–996
Givnish TJ (2010) Ecology of plant speciation. Taxon 59:1326–1366
Goldreich Y (2003) The climate of Israel: observation, research and application. Kluwer Academic Plenum Publishers, New York
Gray MM, St Amand P, Bello NM, Galliart MB, Knapp M, Garrett KA, Morgan TJ, Baer SG, Maricle BR, Akhunov ED, Johnson LC (2014) Ecotypes of an ecologically dominant prairie grass (Andropogon gerardii) exhibit genetic divergence across the U.S. Midwest grasslands’ environmental gradient. Molec Ecol 23:6011–6028. doi:10.1111/mec.12993
Griffin SR, Barrett SCH (2004) Genetic variation in Trillium erectum (Melanthiaceae), a widespread forest herb in eastern North America. Canad J Bot 82:316–321. doi:10.1139/b04-036
Guarino C, Salerno G, Cipriani G (2013) Effects of fragmentation phenomena on the genetic structure and gene flow in Centaurea cineraria group (Asteraceae) in the Mediterranean Basin. Pl Biosyst 147:996–1005. doi:10.1080/11263504.2013.855276
Günther T, Coop G (2013) Robust identification of local adaptation from allele frequencies. Genetics 195:205–220. doi:10.1534/genetics.113.152462
Guttman M, Amit I, Garber M, French C, Lin MF, Feldser D, Huarte M, Zuk O, Carey BW, Cassady JP, Cabili MN, Jaenisch R, Mikkelsen TS, Jacks T, Hacohen N, Bernstein BE, Kellis M, Regev A, Rinn JL, Lander ES (2009) Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature 458:223–227. doi:10.1038/nature07672
Haldane JBS (1948) The theory of a cline. J Genet 48:277–284. doi:10.1007/BF02986626
Harter DEV, Thiv M, Weig A, Jentsch A, Beierkuhnlein C (2015) Spatial and ecological population genetic structures within two island-endemic Aeonium species of different niche width. Ecol Evol 5:4327–4344. doi:10.1002/ece3.1682
Hereford J (2009) A quantitative survey of local adaptation and fitness trade-offs. Amer Naturalist 173:579–588. doi:10.1086/597611
Hermisson J, Pennings PS (2005) Soft sweeps: molecular population genetics of adaptation from standing genetic variation. Genetics 169:2335–2352. doi:10.1534/genetics.104.036947
Hoffmann AA, Blows MW (1994) Species borders: ecological and evolutionary perspectives. Trends Ecol Evol 9:223–227. doi:10.1016/0169-5347(94)90248-8
Hogbin PM, Ayre DJ, Whelan RJ (1998) Genetic variation and reproductive success of road verge populations of the rare shrub Grevillea barklyana (Proteaceae). Heredity 80:180–186. doi:10.1046/j.1365-2540.1998.00271.x
Holderegger R, Buehler D, Gugerli F, Manel S (2010) Landscape genetics of plants. Trends Plant Sci 15:675–683. doi:10.1016/j.tplants.2010.09.002
Huang C, Chang C, Huang B, Chung J, Chen J, Chiang Y, Hwang S (2015) Genetic relationships and ecological divergence in Salix species and populations in Taiwan. Tree Genet Genomes. doi:10.1007/s11295-015-0862-1
Hübner S, Höffken M, Oren E, Haseneyer G, Stein N, Graner A, Schmid K, Fridman E (2009) Strong correlation of wild barley (Hordeum spontaneum) population structure with temperature and precipitation variation. Molec Ecol 18:1523–1536. doi:10.1111/j.1365-294X.2009.04106.x
ICN (Hand R, Kilian N, Raab-Straube E von; eds) (2009+) International Cichorieae Network: Cichorieae Portal. Available at: http://wp6-cichorieae.e-taxonomy.eu/portal/. Accessed 4 Dec 2015
Jombart T (2008) Adegenet: a R package for the multivariate analysis of genetic markers. Bioinformatics 24:1403–1405
Jones CJ, Edwards KJ, Castaglione S, Winfield MO, Sala F, van de Wiel C, Bredemeijer G, Vosman B, Matthes M, Daly A, Brettschneider R, Bettini P, Buiatti M, Maestri E, Malcevschi A, Marmiroli N, Aert R, Volckaert G, Rueda J, Linacero R, Vazquez A, Karp A (1997) Reproducibility testing of RAPD, AFLP and SSR markers in plants by a network of European laboratories. Molec Breed 3:381–390. doi:10.1023/A:1009612517139
Jones MR, Forester BR, Teufel AI, Adams RV, Anstett DN, Goodrich BA, Landguth EL, Joost S, Manel S (2013) Integrating landscape genomics and spatially explicit approaches to detect loci under selection in clinal populations. Evolution 67:3455–3468. doi:10.1111/evo.12237
Joost S, Bonin A, Bruford MW, Després L, Conord C, Erhardt G, Taberlet P (2007) A spatial analysis method (SAM) to detect candidate loci for selection: towards a landscape genomics approach to adaptation. Molec Ecol 16:3955–3969. doi:10.1111/j.1365-294X.2007.03442.x
Kaul V, Koul A, Sharma C (2000) The underground flower. Curr Sci 78:39–44
Kavanagh KD, Haugen TO, Gregersen F, Jernvall J, Vøllestad LA (2010) Contemporary temperature-driven divergence in a Nordic freshwater fish under conditions commonly thought to hinder adaptation. Evol Biol 10:350. doi:10.1186/1471-2148-10-350
Kettlewell HBD (1955) Selection experiments on industrial melanism in the Lepidoptera. Heredity 9:323–342. doi:10.1038/hdy.1955.36
Krueger-Hadfield SA, Roze D, Mauger S, Valero M (2013) Intergametophytic selfing and microgeographic genetic structure shape populations of the intertidal red seaweed Chondrus crispus. Molec Ecol 22:3242–3260. doi:10.1111/mec.12191
Kwak MM, Velterop O, van Andel J (1998) Pollen and gene flow in fragmented habitats. Appl Veg Sci 1:37–54. doi:10.2307/1479084
Lauterbach D, Ristow M, Gemeinholzer B (2011) Genetic population structure, fitness variation and the importance of population history in remnant populations of the endangered plant Silene chlorantha (Willd.) Ehrh. (Caryophyllaceae). Plant Biol 13:667–777. doi:10.1111/j.1438-8677.2010.00418.x
Legendre P, Mi X, Ren H, Ma K, Yu M, Sun I, He F (2009) Partitioning beta diversity in a subtropical broad-leaved forest of China. Ecology 90:663–674. doi:10.1890/07-1880.1
Lei Y, Wang W, Liu Y, He D, Li Y (2015) Adaptive genetic variation in the smoke tree (Cotinus coggygria Scop.) is driven by precipitation. Biochem Syst Ecol 59:63–69. doi:10.1016/j.bse.2015.01.009
Leimu R, Fischer M (2008) A meta-analysis of local adaptation in plants. PLoS ONE 3(e4010):1–8. doi:10.1371/journal.pone.0004010
Lenormand T (2002) Gene flow and the limits to natural selection. Trends Ecol Evol 17:183–189. doi:10.1016/S0169-5347(02)02497-7
Lindström T, Brown GP, Sisson SA, Phillips BL, Shine R (2013) Rapid shifts in dispersal behavior on an expanding range edge. Proc Natl Acad Sci USA 110:13452–13456. doi:10.1073/pnas.1303157110
Linhart YB, Grant MC (1996) Evelutionary significance of local genetic differentiation in plants. Annual Rev Ecol Syst 27:237–277. doi:10.1146/annurev.ecolsys.27.1.237
Lotterhos KE, Whitlock MC (2015) The relative power of genome scans to detect local adaptation depends on sampling design and statistical method. Molec Ecol 24:1031–1046. doi:10.1111/mec.13100
Lowe AJ, Boshier D, Ward M, Bacles CFE, Navarro C (2005) Genetic resource impacts of habitat loss and degradation; reconciling empirical evidence and predicted theory for neotropical trees. Heredity 95:255–273. doi:10.1038/sj.hdy.6800725
Maassen GH, Bakker AB (2001) Suppressor variables in path models: definitions and interpretations. Sociol Methods Res 30:241–270. doi:10.1177/0049124101030002004
Mallet B, Martos F, Blambert L, Pailler T, Humeau L (2014) Evidence for isolation-by-habitat among populations of an epiphytic orchid species on a small oceanic island. PLoS ONE 9:e87469. doi:10.1371/journal.pone.0087469
Manel S, Gugerli F, Thuiller W, Alvarez N, Legendre P, Holderegger R, Gielly L, Taberlet P (2012) Broad-scale adaptive genetic variation in alpine plants is driven by temperature and precipitation. Molec Ecol 21:3729–3738. doi:10.1111/j.1365-294X.2012.05656.x
Meirmans PG (2012) The trouble with isolation by distance. Molec Ecol 21:2839–2846. doi:10.1111/j.1365-294X.2012.05578.x
Minder AM, Widmer A (2008) A population genomic analysis of species boundaries: neutral processes, adaptive divergence and introgression between two hybridizing plant species. Molec Ecol 17:1552–1563. doi:10.1111/j.1365-294X.2008.03709.x
Mix C, Arens PFP, Rengelink R, Smulders MJM, van Groenendael JM, Ouborg NJ (2006) Regional gene flow and population structure of the wind-dispersed plant species Hypochaeris radicata (Asteraceae) in an agricultural landscape. Molec Ecol 15:1749–1758. doi:10.1111/j.1365-294X.2006.02887.x
Morandin LA, Winston ML (2006) Pollinators provide economic incentive to preserve natural land in agroecosystems. Agric Eco-Syst Environm 116:289–292. doi:10.1016/j.agee.2006.02.012
Morgan JW, Meyer MJ, Young AG (2013) Severe habitat fragmentation leads to declines in genetic variation, mate availability, and reproductive success in small populations of a once-common Australian grassland daisy. Int J Pl Sci 174:1209–1218. doi:10.1086/673242
Nahum S, Inbar M, Ne’eman G, Ben-Shlomo R (2008) Phenotypic plasticity and gene diversity in Pistacia lentiscus L. along environmental gradients in Israel. Tree Genet Genomes 4:777–785. doi:10.1007/s11295-008-0150-4
Nakazato T, Bogonovich M, Moyle LC (2008) Environmental factors predict adaptive phenotypic differentiation within and between two wild Andean tomatoes. Evolution 62:774–792. doi:10.1111/j.1558-5646.2008.00332.x
Nathan R, Muller-Landau HC (2000) Spatial patterns of seed dispersal, their determinants and consequences for recruitment. Trends Ecol Evol 15:278–285. doi:10.1016/S0169-5347(00)01874-7
Nei M (1987) Molecular evolutionary genetics. Columbia University Press, New York
Nosil P, Vines TH, Funk DJ (2005) Perspective: reproductive isolation caused by natural selection against immigrants from divergent habitats. Evolution 59:705. doi:10.1554/04-428
Nosil P, Funk DJ, Ortiz-Barrientos D (2009) Divergent selection and heterogeneous genomic divergence. Molec Ecol 18:375–402. doi:10.1111/j.1365-294X.2008.03946.x
Nunes VL, Beaumont MA, Butlin RK, Paulo OS (2012) Challenges and pitfalls in the characterization of anonymous outlier AFLP markers in non-model species: lessons from an ocellated lizard genome scan. Heredity 109:340–348. doi:10.1038/hdy.2012.48
Oksanen J, Blanchet FG, Friendly M, Kindt R, Legendre P, McGlinn D, Minchin PR, O'Hara RB, Simpson GL, Solymos P, Stevens MHH, Szoecs E, Wagner H (2007) Vegan: community ecology package. R package version 1.15-1. Available at: http://www.r-project.org
Oleksyk TK, Smith MW, O’Brien SJ (2010) Genome-wide scans for footprints of natural selection. Philos Trans Roy Soc B Biol Sci 365:185–205. doi:10.1098/rstb.2009.0219
Paris M, Despres L (2012) Identifying insecticide resistance genes in mosquito by combining AFLP genome scans and 454 pyrosequencing. Molec Ecol 21:1672–1686. doi:10.1111/j.1365-294X.2012.05499.x
Primack RB, Miao S (1992) Dispersal can limit local plant distribution. Conservation Biol 6:513–519. doi:10.1046/j.1523-1739.1992.06040513.x
Pritchard JK, Di Rienzo A (2010) Adaptation—not by sweeps alone. Nat Rev Genet 11:665–667. doi:10.1038/nrg2880
Pritchard JK, Stephens M, Donnelly Peter (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Rehm EM, Olivas P, Stroud J, Feeley KJ (2015) Losing your edge: climate change and the conservation value of range-edge populations. Ecol Evol 5:4315–4326. doi:10.1002/ece3.1645
Richardson JL, Urban MC (2013) Strong selection barriers explain microgeographic adaptation in wild salamander populations. Evolution 67:1729–1740. doi:10.1111/evo.12052
Richardson JL, Urban MC, Bolnick DI, Skelly DK (2014) Microgeographic adaptation and the spatial scale of evolution. Trends Ecol Evol 29:165–176. doi:10.1016/j.tree.2014.01.002
Robinson E, Smith E (1841) Palästina und die südlich angrenzenden Länder. Tagebuch einer Reise im Jahr 1838 in Bezug auf die biblische Geographie unternommen, vol 2. Verlag der Buchhandlung des Waisenhauses, Halle
Rosenberg NA (2004) DISTRUCT: a program for the graphical display of population structure. Molec Ecol Notes 4:137–138. doi:10.1046/j.1471-8286.2003.00566.x
Rundle HD, Nosil P (2005) Ecological speciation. Ecol Lett 8:336–352. doi:10.1111/j.1461-0248.2004.00715.x
Salvator L (1879) Die Karawanenstrasse von Aegypten nach Syrien. Verlag H. Mercy, Prag
Savolainen O, Lascoux M, Merilä J (2013) Ecological genomics of local adaptation. Nat Rev Genet 14:807–820. doi:10.1038/nrg3522
Schmidt PS, Serrão EA, Pearson GA, Riginos C, Rawson PD, Hilbish TJ, Brawley SH, Trussell GC, Carrington E, Wethey DS, Grahame JW, Bonhomme F, Rand DM (2008) Ecological genetics in the North Atlantic: environmental gradients and adaptation at specific loci. Ecology 89:S91–S107. doi:10.1890/07-1162.1
Sexton JP, McIntyre PJ, Angert AL, Rice KJ (2009) Evolution and ecology of species range limits. Annual Rev Ecol Evol Syst 40:415–436. doi:10.1146/annurev.ecolsys.110308.120317
Sexton JP, Hangartner SB, Hoffmann AA (2014) Genetic isolation by environment or distance: which pattern of gene flow is most common? Evolution 68:1–15. doi:10.1111/evo.12258
Shafer ABA, Wolf JBW (2013) Widespread evidence for incipient ecological speciation: a meta-analysis of isolation-by-ecology. Ecol Lett 16:940–950. doi:10.1111/ele.12120
Shafer ABA, Wolf JBW, Alves PC, Bergstrom L, Bruford MW, Brannstrom I, Colling G, Dalen L, de Meester L, Ekblom R, Fawcett KD, Fior S, Hajibabaei M, Hill JA, Hoezel AR, Hoglund J, Jensen EL, Krause J, Kristensen TN, Krutzen M, McKay JK, Norman AJ, Ogden R, Osterling EM, Ouborg NJ, Piccolo J, Popovic D, Primmer CR, Reed FA, Roumet M, Salmona J, Schenekar T, Schwartz MK, Segelbacher G, Senn H, Thaulow J, Valtonen M, Veale A, Vergeer P, Vijay N, Vila C, Weissensteiner M, Wennerstrom L, Wheat CW, Zielinski P (2015) Genomics and the challenging translation into conservation practice. Trends Ecol Evol 30:78–87. doi:10.1016/j.tree.2014.11.009
Silva LB, Sardos J, de Sequeira MM, Silva L, Crawford D, Moura M (2016) Understanding intra and inter-archipelago population genetic patterns within a recently evolved insular endemic lineage. Pl Syst Evol 302:367–384. doi:10.1007/s00606-015-1267-1
Soons MB, Heil GW, Nathan R, Katul GG (2004) Determinants of long-distance seed dispersal by wind in grasslands. Ecology 85:3056–3068. doi:10.1890/03-0522
Sork VL, Nason J, Campbell DR, Fernandez JF (1999) Landscape approaches to historical and contemporary gene flow in plants. Trends Ecol Evol 14:219–224. doi:10.1016/S0169-5347(98)01585-7
Steiner KC, Berrang PC (1990) Microgeographic adaptation to temperature in pitch pine progenies. Amer Midl Naturalist 123:292. doi:10.2307/2426557
Sternberg M, Holzapfel C, Tielbörger K, Sarah P, Kigel J, Lavee H, Fleischer A, Jeltsch F, Köchy M (2009) The use and misuse of climatic gradients for evaluating climate impact on dryland ecosystems—an example for the solution of conceptual problems. In: Blanco J, Kheradmand H (eds) Climate change—geophysical foundations and ecological effects. InTech Open Access Publisher, Rijeka, pp 361–374
Stinchcombe JR, Hoekstra HE (2008) Combining population genomics and quantitative genetics: finding the genes underlying ecologically important traits. Heredity 100:158–170. doi:10.1038/sj.hdy.6800937
Strasburg JL, Sherman NA, Wright KM, Moyle LC, Willis JH, Rieseberg LH (2012) What can patterns of differentiation across plant genomes tell us about adaptation and speciation? Philos Trans Roy Soc B 367:364–373. doi:10.1098/rstb.2011.0199
Svoray T, Mazor S, Bar P (2007) How is shrub cover related to soil moisture and patch geometry in the fragmented landscape of the Northern Negev desert? Landscape Ecol 22:105–116. doi:10.1007/s10980-006-9004-3
Tanaka K, Oshio A, Enoki S, Kubo A, Takahara Y (2014) Population genetic diversity of the endemic plant subspecies Ixeridium dentatum subsp. nipponicum (Asteraceae). Biochem Syst Ecol 55:73–78. doi:10.1016/j.bse.2014.02.014
R Development Core Team (2013) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna. Available at: http://www.R-project.org
Thompson WC, Taroni F, Aitken CGG (2003) How the probability of a false positive affects the value of DNA evidence. J Forensic Sci 48:2001171. doi:10.1520/JFS2001171
Tiffin P, Ross-Ibarra J (2014) Advances and limits of using population genetics to understand local adaptation. Trends Ecol Evol 29:673–680. doi:10.1016/j.tree.2014.10.004
Urban MC (2011) The evolution of species interactions across natural landscapes. Ecol Lett 14:723–732. doi:10.1111/j.1461-0248.2011.01632.x
Utzschneider H (2005) Micha. Teilband 3. Zürcher Bibelkommentare. TVZ, Zürich
van Rossum F, Campos De Sousa S, Triest L (2004) Genetic consequences of habitat fragmentation in an agricultural landscape on the common Primula veris, and comparison with its rare congener P. vulgaris. Conservation Genet 5:231–245. doi:10.1023/B:COGE.0000030007.85492.70
van Treuren R, Bijlsma R, Ouborg NJ, Kwak MM (1994) Relationships between plant density, outcrossing rates and seed set in natural and experimental populations of Scabiosa columbaria. J Evol Biol 7:287–302. doi:10.1046/j.1420-9101.1994.7030287.x
Vekemans X (2002) AFLP-SURV version 1.0. Laboratoire de Génétique et Evolution des Populations Végétales. Department of Biology, University of Lille 1, Lille
Velikovsky J (2009) Zeitalter im Chaos. Vol. II: Ramses II und seine Zeit. Verlag Julia White, Doebeln
Vitalis R, Dawson K, Boursot P (2001) Interpretation of variation across marker loci as evidence of selection. Genetics 158:1811–1823
Vos P, Hogers R, Bleeker M, Reijans M, van de Lee T, Homes M, Frijters A, Pot J, Peleman J, Kuiper M, Zabeau M (1995) AFLP: a new technique for DNA fingerprinting. Nucl Acids Res 23:4407–4414
Willi Y, Hoffmann AA (2012) Microgeographic adaptation linked to forest fragmentation and habitat quality in the tropical fruit fly Drosophila birchii. Oikos 121:1627–1637. doi:10.1111/j.1600-0706.2011.20156.x
Wood HM, Grahame JW, Humphray S, Rogers J, Butlin RK (2008) Sequence differentiation in regions identified by a genome scan for local adaptation. Molec Ecol 17:3123–3135. doi:10.1111/j.1365-294X.2008.03755.x
Wright S (1943) Isolation by distance. Genetics 28:114–138
Yaacobi G, Ziv Y, Rosenzweig ML (2007) Habitat fragmentation may not matter to species diversity. Proc Roy Soc B 274:2409–2412. doi:10.1098/rspb.2007.0674
Young A, Boyle T, Brown T (1996) The population genetic consequences of habitat fragmentation for plants. Trends Ecol Evol 11:413–418. doi:10.1016/0169-5347(96)10045-8
Acknowledgements
The authors thank the Deutsche Forschungsgemeinschaft (German Research Foundation, DFG) for funding this research (GE 1242/12-1). We also very much acknowledge field support and organizational help during our collection trip by I. Giladi, and Y. Ziv from Ben-Gurion University of the Negev, F. Jeltsch and K. Bergholz from the Plant Ecology and Nature Conservation department of the University of Potsdam. We thank A. Fichtner for graphical design (Systematic Botany, JLU Giessen).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Handling editor: Christoph Oberprieler.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Information on Electronic Supplementary Material
Information on Electronic Supplementary Material
Online Resource 1. Geropogon hybridus data matrix: first column number of populations; second ID of individuals; followed columns the three primer combinations (AGC_CCA, AGC_CTC, ACA_CGA).
Online Resource 2. Summary of the Geropogon hybridus data matrix. Primer combination and detected loci per individual.
Online Resource 3. Principal Component Analysis (PCA) of genetic distances between individuals of Geropogon hybridus.
Online Resource 4. Optimal ∆K detection by STRUCTURE HARVESTER.
Online Resource 5. Pairwise ΦST-Matrix for the surveyed populations of Geropogon hybridus.
Online Resource 6. Results of hierarchical analysis of molecular variance (AMOVA) performed by grouping the 12 Geropogon hybridus populations in two groups N vs. C + S and N + C vs. S.
Online Resource 7. List of putative adaptive AFLP loci in the 12 surveyed populations of Geropogon hybridus as assessed by differentiation-based genome scan approaches (DFDIST/FDIST & BAYESCAN) and SAM analyses for correlation with mean annual precipitation.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Müller, C.M., Schulz, B., Lauterbach, D. et al. Geropogon hybridus (L.) Sch.Bip. (Asteraceae) exhibits micro-geographic genetic divergence at ecological range limits along a steep precipitation gradient. Plant Syst Evol 303, 91–104 (2017). https://doi.org/10.1007/s00606-016-1354-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00606-016-1354-y