Abstract
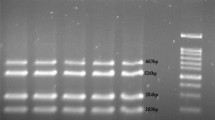
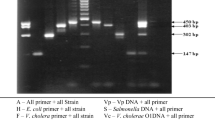
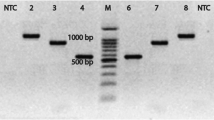
In this study, polymerase chain reaction (PCR) method and culturing strategies were used to detect carriers of Salmonella spp. in cattle and buffalo. In order to perform the assay, 100 faecal samples were obtained from both buffalo and cattle in Urmia, Iran, in July 2008. Samples were tested for Salmonella isolation by culturing techniques and biochemical testes. Faecal samples were enriched in enrichment broth and DNA was extracted and amplified by PCR using specific primers of Salmonella invasion gene (invA). PCR products were visualised using 1.2% agarose gel electrophoresis. The bacterial culturing results were negative for all samples, whilst results from the PCR method and electrophoresis confirmed the presence of Salmonella in one cattle faecal sample (1%) and three buffalo faecal samples (3%). These results indicate that the PCR method is highly sensitive and rapid for Salmonella detection in faeces compared to other routine methods.


Similar content being viewed by others
References
Chiu CH, Ou JT (1996) Rapid identification of Salmonella serovars in faeces by specific detection of virulence genes, invA and spvC, by an enrichment broth culture–multiplex PCR combination assay. J Clin Microbiol 34:2619–2622
Cocolin L, Manzano M, Cantoni C et al (1998) Use of polymerase chain reaction and restriction enzyme analysis to directly detect and identify Salmonella typhimurium in food. J Appl Microbiol 85:673–677. doi:10.1111/j.1365-2672.1998.00575.x
Cohen ND, Neibergs HL, McGruder ED et al (1993) Genus-specific detection of Salmonellae using the polymerase chain reaction (PCR). J Vet Diagn Invest 5:368–371
Cohen ND, Wallis DE, Neibergs HL et al (1995) Detection of Salmonella enteritidis equine faeces using the polymerase chain reaction and genus-specific oligonucleotide primers. J Vet Diagn Invest 7:219–222
Cohen HJ, Mechanda SM, Lin W (1996) PCR amplification of the fimA gene sequence of Salmonella typhimurium, a specific method for detection of Salmonella spp. Appl Environ Microbiol 62:4303–4308
Dombek PE, Johnson LK, Zimmerely ST et al (2000) Use of repetitive DNA sequences and the PCR to differentiate Escherichia coli isolates from human and animal sources. Appl Environ Microbiol 66:2572–2577. doi:10.1128/AEM.66.6.2572-2577.2000
Duguid JP, Campbell I (1967) Antigens of type 1 fimbriae of Salmonellae and other Enterobacteriaceae. J Med Microbiol 2:535–553
Duguid JP, Anderson S, Campbell I (1966) Fimbriae and adhesive properties of Salmonellae. J Pathol Bacteriol 92:107–138. doi:10.1002/path.1700920113
Fadl AA, Nguyen AV, Khan MI (1995) Analysis of Salmonella enteritidis isolates by arbitrarily primed PCR. J Clin Microbiol 33:987–989
Feder I, Nietfeld JC, Gallant J et al (2001) Comparison of cultivation and PCR-hybridization for detection Salmonella in porcine faecal and water samples. J Clin Microbiol 39:2477–2484. doi:10.1128/JCM.39.7.2477-2484.2001
Freschi CR, Carvalho LFOS, Oliveira CJB (2005) Comparison of DNA-extraction methods and selective enrichment broths on the detection of Salmonella typhimurium in swine faeces by polymerase chain reaction (PCR). Brasil J Microbiol 36:363–367
Hatta M, Smits HL (2007) Detection of Salmonella typhi by nested polymerase chain reaction in blood, urine and stool samples. Am J Trop Med Hyg 76:139–143
Horiuchi S, Ignagaki Y, Okamura N et al (1992) Type pili enhance the invasion of Salmonella braenderup and Salmonella typhimurium to Hela cell. Microbiol Immunol 36:352–602
Krysinski EP, Heimsch RC (1977) Use of enzyme-labeled antibodies to detect Salmonella in foods. Appl Environ Microbiol 33:947–954
Kumar S, Balakrishna K, Singh GP et al (2005) Rapid detection of Salmonella typhi in foods by combination of immunomagnetic separation and polymerase chain reaction. J Microbiol Biotechnol 21:625–628. doi:10.1007/s11274-004-3553-x
Lampel KA, Orlandi PA, Kornegay L (2000) Improved template preparation for PCR-based assay for detection of food-borne bacterial pathogens. Appl Environ Microbiol 66:4539–4542. doi:10.1128/AEM.66.10.4539-4542.2000
Malorney B, Hoorfar J, Hugas M et al (2003) Interlaboratory diagnostic accuracy of Salmonella specific PCR-based method. Int J Food Microbiol 89:241–249. doi:10.1016/S0168-1605(03)00154-5
Mattingly JA (1984) Enzyme-linked immunosorbent assay for detection of all Salmonella using a combination of myeloma protein and hybridoma antibody. J Immunol Methods 73:147–156. doi:10.1016/0022-1759(84)90040-1
Muganedi KLM, Goyvaerts EMA, Venter SN et al (2007) Optimisation of the PCR-invA primers for the detection of Salmonella in drinking and surface waters following a pre-cultivation step. Water SA 33:195–202
Myint MS, Jonson YJ, Tablante NL et al (2006) The effect of pre-enrichment protocol on the sensitivity and specificity of PCR for detection of naturally contaminated Salmonella in raw poultry compared to conventional culture. Food Microbiol 23:599–604. doi:10.1016/j.fm.2005.09.002
Rahn K, Grandis SA, Clarke RC et al (1992) Amplification of an invA gene sequence of Salmonella typhimurium by polymerase chain reaction as specific method for detection of Salmonella. Mol Cell Probes 6:271–279. doi:10.1016/0890-8508(92)90002-F
Singer RS, Cooke CL, Maddox CW et al (2006) Use of pooled samples for the detection of Salmonella in faeces by polymerase chain reaction. J Vet Diagn Invest 18:319–325
Wray C, Wray A (2000) Salmonella in domestic animals. CABI, London, pp 171–172
Wyatt GM, Langley MN, Lee HA et al (1993) Further studies on the feasibility of one-day Salmonella detection by enzyme-linked immunosorbent assay. Appl Environ Microbiol 59:1383–1390
Zahraei TS, Mahzounieh M, Saeedzadeh A (2005) Detection of invA gene in isolated Salmonella from broilers by PCR method. Int J Poult Sci 4:557–559. doi:10.3923/ijps.2005.557.559
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ahmadi, M., Dalirnaghadeh, B., Shirzad Aski, H. et al. Comparison of polymerase chain reaction (PCR) and conventional cultivation methods for detection of carriers of Salmonella spp. in cattle and buffalo. Comp Clin Pathol 19, 251–255 (2010). https://doi.org/10.1007/s00580-009-0867-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00580-009-0867-y