Abstract
Pancreatic adenocarcinoma is a lethal cancer with poor response to chemotherapy and immune checkpoint inhibitors. Recent studies suggest that epigenetic alterations contribute to its aggressive biology and the tumor microenvironment which render it unresponsive to immune checkpoint blockade. Here, we review our current understandings of epigenetic dysregulation in pancreatic adenocarcinoma, its effect on the tumor immune microenvironment, and the potential for epigenetic therapy to be combined with immune checkpoint inhibitors.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
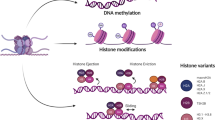
Pancreatic adenocarcinoma (PDAC) is highly fatal malignancy with a 5-year survival rate of less than 10% and is the 4th leading cause of cancer-related death in USA [1, 2]. While surgical resection is the only cure for PDAC, 80% of patients are diagnosed with advanced-stage disease and are not eligible for surgery [3]. Recent genomic analysis facilitated the sub-classification of pancreatic adenocarcinoma based on molecular signatures; however, the use of most of molecular targeted therapies has failed to show clinical benefit to date [4,5,6,7]. Epigenetic regulation of gene expression includes the post-translational modification of histone proteins and methylation of DNA. These modifications are controlled by enzymes and adaptor proteins that are referred to as epigenetic ‘writers’, ‘erasers’, and ‘readers’ [8]. Dysregulation of these proteins contributes to carcinogenesis and pathobiology, including pancreatic cancer [9, 10]. These epigenetic regulators are getting attention as therapeutic targets for cancer therapy.
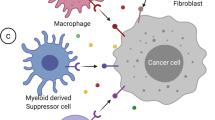
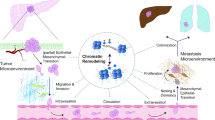
Immune checkpoint inhibitors targeting cytotoxic T-lymphocyte-associated protein 4 (CTLA4) and programmed cell death 1/programmed death ligand 1 (PD1/PDL1) have been used with great success in several cancers and are now widely accepted as one of the standard therapies in cancer treatment [11]. Cancers with high tumor mutation burden and mismatch repair deficiency are associated with favorable response to immune checkpoint inhibitors, but their rates are less than 1% in pancreatic adenocarcinoma [12,13,14]. Moreover, the tumor microenvironment in pancreatic adenocarcinoma is characterized by abundant fibrosis and cancer-associated fibroblast, which contribute to the immune evasion and immunotherapy failure [15,16,17]. However, the precise mechanism of resistance is not fully understood in pancreatic cancer.
Several epigenetic modifiers affect the tumor immune microenvironment and are being investigated as anticancer treatment in combination with immune checkpoint inhibitors [18]. In this review article, we summarize the epigenetic drug therapies that affect tumor immunity against pancreatic cancer.
DNA methylation
DNA methylation regulates gene expression and chromatin structure, which is mainly controlled by DNA methylation writers (DNA methyltransferase; DNMT family) and erasers (ten–eleven translocation methylcytosine dioxygenase; TET family). The genes encoding DNMT and TET family members are frequently mutated in cancers including pancreatic cancer [19]. DNMT3a and DNMT3b promote de novo DNA methylation during embryogenesis and germ cell development, whereas DNMT1 is required for the maintenance of DNA methylation during cell replication [20, 21]. Aberrant focal DNA hypomethylation by loss of function mutation in DNMT3a promotes hematologic malignancies and is associated with poor prognosis [22, 23]. In addition, DNMT family members are more highly expressed in cancer cells than that in normal cells [24, 25] and the resulting aberrant DNA hypermethylation contributes to pancreatic cancer development [26, 27].
DNA methyltransferase (DNMT) inhibitors are the most commonly used epigenetic drug for cancer treatment. These inhibitors are nucleoside cytidine analogues [5-Azacitidine (5-Aza), 5-aza-2′-deoxycytidine (decitabine), and SGI-110 (guadecitabine)] that are incorporated into DNA and bind DNMTs, resulting in reactivation of the genes silenced by aberrant methylation by DNMTs [28]. In pancreatic cancer cell lines with low DNMT1 expression, the DNMT inhibitor decitabine depletes DNMT1 and exerts anti-tumor effects [25]. While guadecitabine sensitizes pancreatic cancer cells to chemotherapeutic agents [29], there is only one phase I study evaluating the anti-tumor effect of DNMT inhibitor on pancreatic cancer [30].
The ten–eleven translocation (TET) family of α-ketoglutarate-dependent dioxygenases indirectly promotes DNA demethylation by hydroxylation of 5-methylcytocine to 5-Hydroxymethylcytocine [31]. Characterization of pancreatic pre-cancerous lesions from pancreatic intraepithelial neoplasia, intraductal papillary mucinous neoplasms, intraductal event during pancreatic oncocytic papillary neoplasms, and mucinous cystic neoplasms by immunohistochemistry revealed that downregulation of 5-hydroxymethylcytocine and TET1 is an early carcinogenesis [32]. Moreover, CRISPR-Cas9-induced knockout of TET1 showed that TET1 suppress pancreatic cancer progression via inhibition of the Wnt pathway [33]. Taken together, these studies demonstrate that increased DNA methylation, either by TET inhibition or DNMT activation, has an important role for pancreatic cancer development.
Isocitrate dehydrogenase (IDH) enzymes IDH1 and IDH2 are frequently mutated in several cancers and a gain-of-function activity promotes conversion of α-ketoglutarate (α-KG) to the oncometabolite 2-Hydroxyglutarate (2-HG) [34, 35]. The accumulation of 2-HG inhibits α-ketoglutarate-dependent dioxygenases including TET family. Subsequently, IDH-mutated cancers have global DNA hypermethylation leading to an epigenetic gene silencing [36, 37]. A recent study in glioma revealed that IDH mutations are associated with decreased PD-L1 expression via hypermethylation of the STAT1 promoter and impaired IFN-γ signaling [38]. Employing a mouse model of IDH-mutant glioma, the authors demonstrated that selective inhibition of mutant IDH suppresses 2-HG production, restoring T-cell immunity and enhancing the anti-tumor effect of immune checkpoint inhibitor. Mutation and amplification of IDH are not common in pancreatic cancer, but the relationship between IDH inhibition and immune checkpoint blockade remains to be clarified.
There is little study to evaluate the combination therapy of immune checkpoint inhibitors and DNA demethylating agents for pancreatic cancer. However, recent studies revealed that DNA demethylation increase tumor antigen and reactivate endogenous retroviral sequences [39, 40]. In terms of relationship to immune checkpoint inhibitor, the DNMT inhibitor 5-azacytidine inhibits immune evasion in cancer through up-regulation of tumor antigen presentation and T-cell chemokine expression [41, 42]. Interestingly, decitabine improve the effectiveness of immune checkpoint inhibitor via modulation of stroma-rich tumor microenvironment by inducing STAT1 and IFN-γ signaling [43, 44]. These findings facilitate the ongoing study of DNMT inhibitor combined with immune checkpoint inhibitors for pancreatic cancer.
Histone acetylation
Gene expression regulated by histone acetylation is controlled by writers; histone acetyltransferase (HATs), and erasers; histone deacetylases (HDACs) [45]. Histone acetylation is associated with open chromatin structure and promotes gene expression. HDAC inhibitors exert anti-tumor effect by re-expression of silenced genes by promoting histone acetylation [46].
In humans, there are 18 HDACs grouped into four classes (class I, II, III, and IV) based on their homology to yeast HDAC. There are several HDAC inhibitors approved by FDA [47]. In preclinical models, class I HDAC and BET-inhibitors (discussed below) showed synergistic anti-tumor effects against pancreatic cancer through the dysregulation of FOSL-1 [48]. A class II HDAC inhibitor combined with the proteasome inhibitor carfilzomib induces cell cycle arrest in pancreatic cancer through FOXO3a activation [49]. Moreover, there is growing evidence that modulating histone acetylation can impact the immune microenvironment. HDAC3 regulates tumor immunity by increasing STAT3 signaling and PDL1 expression [50]. Trichostatin-A, a class I and II inhibitor, suppresses M2 macrophage polarization and MDSCs infiltration in syngeneic cancer mouse models [51]. Entinostat, class I HDAC inhibitor, modulates immune-resistant pancreatic cancer microenvironment by suppressing the myeloid-derived suppressor cells (MDSCs) [52].
Efficacy of HDAC inhibitors was well established in hematologic malignancies, but most clinical trials failed to demonstrate clinical benefit as a single agent in solid cancers, including pancreatic cancer [53]. For solid tumors, the anticancer effect of HDAC inhibition is dependent on the tumor immune microenvironment [54]. In melanoma, HDAC inhibition upregulates PDL1 expression and enhances the anti-tumor effect of immune checkpoint inhibitor [55]. While there have been several phase I studies evaluating the combination therapy with HDAC inhibitor [30, 56,57,58,59,60,61,62,63,64], phase II trials have not yet revealed clinical benefit. In a phase II trial, an oral HDAC inhibitor, CI994 (tacedinaline), combined with gemcitabine did not improve overall survival in pancreatic cancer as compared to gemcitabine alone [65]. Another phase II study demonstrated that combination therapy of panobinostat and bortezomib provided a modest median progression free survival of 2.1 months in patients with pancreatic cancer and was not recommended [66].
HATs have become one of the top targets of cancer immunotherapy. HATs are composed of CREB-binding protein (CBP) and its homologue E1A-associated protein p300 (P300) and augment cancer immunotherapy through induction of MHC-I antigen presentation [67]. On the other hand, P300 acetyltransferase regulates the expression of PDL1 on plasma membrane by acetylating the cytoplasmic domain and prevents the evasion of immune surveillance via nuclear translocation [68]. HAT1 is overexpressed in PDAC patients and associated with poor prognosis. Moreover, HAT1 inhibition improved the immune checkpoint blockade treatment by decreasing the PDL1 expression in PDAC xenograft model via cell-intrinsic signals, including the regulation of autophagy and the mTOR pathway [69]. The effect of HAT1 on tumor immunotherapy remains unrevealed.
Bromodomain and extraterminal (BET) family members are ‘readers’ of acetylated lysine residue on histones. In pancreatic cancer cell lines, BET proteins promote cancer progression, at least in part, by interaction with GLI transcription factors as well as impacting the tumor microenvironment by regulating the expression of SHH and IL6 [70, 71]. Moreover, BET proteins play a key role in mediating the PDAC-cancer-associated fibroblast (CAF) cross-talk that is required for the formation of PDAC matrisome [72]. BET bromodomain inhibitor JQ1 combined with HDAC inhibitor SAHA demonstrated synergistic antiproliferative effect by de-repression of p57 [73]. BET proteins induce PDL1 expression in several cancers by binding to its promoter and enhancer [74]. JQ1 combined with PD-1 blockade enhances the anti-tumor response in Kras-mutant lung cancer by reducing the T-reg and increasing T-cell infiltration with T-helper type 1 cytokine [75]. In pancreatic cancer, a dual inhibitor of BET proteins and HATs suppress KRAS/MAPK signaling and augment PDL1 blockade through recruitment of cytotoxic T-cell [76]. Several BET bromodomain inhibitors are vigorously investigated in clinical trials of anticancer treatment and only one phase I study is evaluating the combination therapy of BET inhibitor and immune checkpoint inhibitor [77, 78]. However, these studies highlight that epigenetic silencing by BET protein inhibitors promotes anti-tumor immunity and potentiates anti-tumor effect of immune checkpoint inhibitor in pancreatic cancer.
Histone methylation
The effect of post-translational histone methylation on gene expression depends on the methylation status of specific histone tail residues [79]. For example, histone H3 trimethylated at lysine 4 (H3K4me3) promotes gene expression, while H3K27me3 suppresses it [80]. Histone methylation is regulated by histone methyltransferases (HMTs, writers) and demethylases (HDMs, erasers). Histone methylation is dysregulated in many cancers [81]. Low levels of H3K4me2, H3K18ac, and H3K27me3 are associated with poor survival in pancreatic cancer [82, 83]. Mutation in histone proteins can also have an oncogenic effect [84]. One of the mutations, H2BG53D (termed oncohistone), is associated with the progression of pancreatic cancer [85, 86]. However, the effect of oncohistone mutation on pancreatic cancer progression is not well elucidated.
HMTs are being investigated as a therapeutic target for several cancers. Enhancer of zeste homologue 2 (EZH2) is one of the most widely investigated HMTs and is responsible for the formation of H3K27me3. EZH2 promotes the progression of several cancer [87] and promotes cell migration and invasion of pancreatic cancer cell lines [88]. EZH2 is one of four core subunits of Polycomb repressive complex 2 (PRC2), which maintains bivalency at the promoters of MHC-1 antigen processing pathway gene to silence MHC-1 expression and promote evasion of T-cell-mediated immunity. Pharmacological inhibition of EZH2 reverses PRC2-mediated MHC-1 silencing by suppressing the repressive H3K27me3, leading to re-establishment of effective T-cell-mediated anti-tumor immunity [89]. In head and neck cancer cell lines, EZH2-inhibitors (GSK126 and EPZ6438) enhance antigen presentation and overcome immune checkpoint resistance [90]. EZH2 is also associated with silencing of TH-1 type chemokines, including CXCL9 and CXCL10 [91]. GSK126 combined with a DNMT inhibitor (GSK-J4) epigenetically reprogrammed the tumor immune microenvironment by restoring the expression of these chemokines. There is only one phase I study evaluating EZH2-inhibitors combined with immune checkpoint blockade for a solid tumor including melanoma, lung cancer, and renal cell carcinoma; however, targeting the epigenetic silencing of antigen presentation is an attractive mechanism to induce tumor immunity in pancreatic cancer.
SET domain bifurcated 1 (SETDB1) catalyzes the di- and trimethylation of H3K9 (H3K9me3) to switch from euchromatic to heterochromatic states. Aberrant SETDB1 expression is observed in various cancers and is associated with carcinogenesis [92]. In pancreatic cancer, SETDB1 inhibits apoptosis by regulating p53 expression [93]. A recent CRISPR-Cas9 chromatin regulator screen revealed that SETDB1 plays an important role for suppressing tumor-intrinsic immunogenicity and its amplification is associated with immune evasion and resistance to immune checkpoint inhibitors [94]. CRISPR-induced knockdown of SETDB1 in lung and melanoma cell lines induces transposable elements-encoded viral antigens and triggers cytotoxic T-cell response. While there is no SETDB1-specific methyltransferase inhibitor available, these preclinical studies suggest that it might be a therapeutic target of cancer immunotherapy for pancreatic cancer.
Disruptor of telomeric silencing 1-like (DOT1L) is a histone H3 lysine 79 methyltransferase. DOT1L is well investigated in prostate cancer and MLL-fusion leukemia [95, 96]. Decreased DOT1L along with low levels of H3K79me3 is associated with epithelial–mesenchymal transition (EMT) and PDL1 expression, but the effect of DOT1L on tumor immunity is still under investigation [97].
There are additional studies suggesting the promise of other histone methyltransferases as candidates for drug development to enhance tumor immunity. Nuclear receptor-binding SET domain protein 1 (NSD1) is a histone H3 lysine 36 methyltransferase. NSD1 mutations are observed in head and neck cancer and are associated with increased PDL1 expression [98, 99]. Knockdown of NSD1 in head and neck cancer cells resulted in decreased expression of CCL5 and suppressed T-cell infiltration into the tumor microenvironment, suggesting that NSD1 inactivation in cancer has implications for cancer immunotherapy [100]. In pancreatic cancer, high NSD1 expression is associated with poor prognosis [101]. Histone-lysine N-methyltransferase (KMT2) family catalyze the formation of trimethylation of H3K4 (H3K4me3). KMT2s are frequently mutated especially in hematologic malignancies and are associated with favorable response to cancer immunotherapy [102, 103]. KMT2A is highly expressed in pancreatic cancer cells followed by H3K4me3-mediated PD-L1 expression and immune evasion [104, 105]. There is no FDA approved drug to inhibit NSD1or KMT2 family members, but inhibition of these enzymes combined with immune checkpoint blockade may be promising strategies.
Unlike the HMTs described above, protein arginine methyltransferases (PRMTs) catalyze the methylation of arginine residues of histones. PRMTs are classified into three enzymes (type I, II and III). PRMT5 (type II PRMT) promotes EMT and PRMT1 (type I PRMT) is associated with poor prognosis in pancreatic cancer [106, 107]. Recent work revealed that inhibition of PRMTs promotes anti-tumor immunity by modulating RNA splicing [108]. Type 1 and 2 PRMT inhibition promotes T-cell-mediated anti-tumor immunity [109, 110]. In a xenograft model of mouse pancreatic cancer, a type I PRMT inhibitor combined with PD-L1 blockade demonstrated anti-tumor effect through increasing infiltration of CD8 + T cells [111].
Histone demethylases are also being investigated as epigenetic regulators of pancreatic cancer. Lysine-specific histone demethylase (KDM) family members remove methylated lysine residue of histone tails. KDM1A maintains pancreatic cancer progression through HIF1α-dependent glycolysis [112]. In a subcutaneous synergistic mouse tumor model, KDM1A is a potent inhibitor of anti-tumor immunity, and its ablation activates the type I interferon along with CD8 + T-cell infiltration [33, 113]. High expression of KDM5A is associated with improved response to immune checkpoint blockade by enhancing TLR signaling in several mouse models of cancer, including breast cancer, melanoma and fibrosarcoma [114]. KDM5C mutations predict the favorable outcome of immune checkpoint inhibitor by pan-cancer data base analysis [115]. KDM4A inhibition activates anti-tumor immunity and enhances anti-PD1 immunotherapy in squamous cell carcinoma [116]. Interestingly, KDM2A expression in cancer-associated fibroblast promotes carcinogenesis and inhibition of KDM2A suppress the tumor growth via inhibiting PDL1 expression in breast cancer fibroblast cell lines [117]. Recent large data base analysis revealed that up-regulation of KDM5A/B/C expression was associated with infiltrating immune cells in pancreatic cancer [118]. Since the activity of KDM family members regulate tumor immunity, therapeutically targeting these proteins in combination with immune checkpoint inhibitors against pancreatic adenocarcinoma needs investigation.
In summary, several epigenetic drugs can augment the tumor immunity in pancreatic adenocarcinoma. There is still a long way to go before these therapies become a standard treatment in the daily practice. Considering the unique tumor microenvironment of PDAC characterized by abundant non-cancer components that create additional barriers to immune checkpoint inhibitors, therapies that target the tumor microenvironment might enhance the effect of epigenetic drugs on cancer immunotherapy. Table 1 summarizes the current clinical trial combining epigenetic therapy with immune checkpoint inhibitors for pancreatic cancer. The improvement in specificity of current epigenetic drugs will provide further possibilities for combination therapies. Collectively, epigenetic therapy combined with immune checkpoint blockade turns immune cold into hot tumor microenvironment and is promising strategy for the treatment of pancreatic adenocarcinoma.
Change history
17 September 2022
A Correction to this paper has been published: https://doi.org/10.1007/s00535-022-01922-3
Abbreviations
- PD1:
-
Programmed cell death 1
- PDL1:
-
Programmed death ligand 1
- CTLA4:
-
Cytotoxic T-lymphocyte-associated protein 4
- DNMT:
-
DNA methyltransferase
- HDAC:
-
Histone deacetylase
- HMT:
-
Histone methyltransferase
- HAT:
-
Histone acetyltransferase
- TET:
-
Ten–eleven translocation
References
Siegel RL, Miller KD, Fuchs HE, et al. Cancer statistics, 2021. CA Cancer J Clin. 2021;71:7–33.
Cancer Facts & Figures 2020. Atlanta: American Cancer Society; 2021.Avalable at https://cancerstatisticscenter.cancer.org/#!/cancer-site/Pancreas. Accessed 31 Mar 2022
Ferrone CR, Pieretti-Vanmarcke R, Bloom JP, et al. Pancreatic ductal adenocarcinoma: long-term survival does not equal cure. Surgery. 2012;152:S43–9.
Collisson EA, Sadanandam A, Olson P, et al. Subtypes of pancreatic ductal adenocarcinoma and their differing responses to therapy. Nat Med. 2011;17:500–3.
Moffitt RA, Marayati R, Flate EL, et al. Virtual microdissection identifies distinct tumor- and stroma-specific subtypes of pancreatic ductal adenocarcinoma. Nat Genet. 2015;47:1168–78.
Bailey P, Chang DK, Nones K, et al. Genomic analyses identify molecular subtypes of pancreatic cancer. Nature. 2016;531:47–52.
Ciliberto D, Staropoli N, Chiellino S, et al. Systematic review and meta-analysis on targeted therapy in advanced pancreatic cancer. Pancreatology. 2016;16:249–58.
Egger G, Liang G, Aparicio A, et al. Epigenetics in human disease and prospects for epigenetic therapy. Nature. 2004;429:457–63.
Jones PA, Baylin SB. The fundamental role of epigenetic events in cancer. Nat Rev Genet. 2002;3:415–28.
Lomberk G, Blum Y, Nicolle R, et al. Distinct epigenetic landscapes underlie the pathobiology of pancreatic cancer subtypes. Nat Commun. 2018;9:1978.
Ribas A, Wolchok JD. Cancer immunotherapy using checkpoint blockade. Science. 2018;359:1350–5.
Le DT, Durham JN, Smith KN, et al. Mismatch repair deficiency predicts response of solid tumors to PD-1 blockade. Science. 2017;357:409–13.
Yarchoan M, Hopkins A, Jaffee EM. Tumor Mutational Burden and Response Rate to PD-1 Inhibition. N Engl J Med. 2017;377:2500–1.
Hu ZI, Shia J, Stadler ZK, et al. Evaluating Mismatch Repair Deficiency in Pancreatic Adenocarcinoma: Challenges and Recommendations. Clin Cancer Res. 2018;24:1326–36.
Chakravarthy A, Khan L, Bensler NP, et al. TGF-beta-associated extracellular matrix genes link cancer-associated fibroblasts to immune evasion and immunotherapy failure. Nat Commun. 2018;9:4692.
Ford K, Hanley CJ, Mellone M, et al. NOX4 Inhibition potentiates immunotherapy by overcoming cancer-associated fibroblast-mediated cd8 t-cell exclusion from tumors. Cancer Res. 2020;80:1846–60.
Kieffer Y, Hocine HR, Gentric G, et al. single-cell analysis reveals fibroblast clusters linked to immunotherapy resistance in cancer. Cancer Discov. 2020;10:1330–51.
Topper MJ, Vaz M, Marrone KA, et al. The emerging role of epigenetic therapeutics in immuno-oncology. Nat Rev Clin Oncol. 2020;17:75–90.
Baylin SB, Jones PA. Epigenetic determinants of cancer. Cold Spring Harb Perspect Biol. 2016;8: a019505.
Okano M, Bell DW, Haber DA, et al. DNA methyltransferases dnmt3a and dnmt3b are essential for de novo methylation and mammalian development. Cell. 1999;99:247–57.
Robert MF, Morin S, Beaulieu N, et al. DNMT1 is required to maintain CpG methylation and aberrant gene silencing in human cancer cells. Nat Genet. 2003;33:61–5.
Hou HA, Kuo YY, Liu CY, et al. DNMT3A mutations in acute myeloid leukemia: stability during disease evolution and clinical implications. Blood. 2012;119:559–68.
Russler-Germain DA, Spencer DH, Young MA, et al. The R882H DNMT3A mutation associated with Aml dominantly inhibits wild-type Dnmt3a by blocking its ability to form active tetramers. Cancer Cell. 2014;25:442–54.
Robertson KD, Uzvolgyi E, Liang G, et al. The human DNA methyltransferases (DNMTs) 1, 3a and 3b: coordinate mRNA expression in normal tissues and overexpression in tumors. Nucleic Acids Res. 1999;27:2291–8.
Li A, Omura N, Hong SM, et al. Pancreatic cancer DNMT1 expression and sensitivity to DNMT1 inhibitors. Cancer Biol Ther. 2010;9:321–9.
Sato N, Maitra A, Fukushima N, et al. Frequent hypomethylation of multiple genes overexpressed in pancreatic ductal adenocarcinoma. Cancer Res. 2003;63:4158–66.
Rhee I, Bachman KE, Park BH, et al. DNMT1 and DNMT3b cooperate to silence genes in human cancer cells. Nature. 2002;416:552–6.
Da Costa EM, McInnes G, Beaudry A, et al. DNA Methylation-Targeted Drugs Cancer J. 2017;23:270–6.
Thakar M, Hu Y, Morreale M, et al. A novel epigenetic modulating agent sensitizes pancreatic cells to a chemotherapy agent. PLoS ONE. 2018;13: e0199130.
Von Hoff DD, Rasco DW, Heath EI, et al. Phase I study of CC-486 Alone and in Combination with Carboplatin or nab-Paclitaxel in Patients with Relapsed or Refractory Solid Tumors. Clin Cancer Res. 2018;24:4072–80.
Tahiliani M, Koh KP, Shen Y, et al. Conversion of 5-methylcytosine to 5-hydroxymethylcytosine in mammalian DNA by MLL partner TET1. Science. 2009;324:930–5.
Fujikura K, Alruwaii ZI, Haffner MC, et al. Downregulation of 5-hydroxymethylcytosine is an early event in pancreatic tumorigenesis. J Pathol. 2021;254:279–88.
Wu J, Li H, Shi M, et al. TET1-mediated DNA hydroxymethylation activates inhibitors of the Wnt/beta-catenin signaling pathway to suppress EMT in pancreatic tumor cells. J Exp Clin Cancer Res. 2019;38:348.
Xu W, Yang H, Liu Y, et al. Oncometabolite 2-hydroxyglutarate is a competitive inhibitor of alpha-ketoglutarate-dependent dioxygenases. Cancer Cell. 2011;19:17–30.
Borger DR, Tanabe KK, Fan KC, et al. Frequent mutation of isocitrate dehydrogenase (IDH)1 and IDH2 in cholangiocarcinoma identified through broad-based tumor genotyping. Oncologist. 2012;17:72–9.
Nunez FJ, Mendez FM, Kadiyala P, et al. IDH1-R132H acts as a tumor suppressor in glioma via epigenetic up-regulation of the DNA damage response. Sci Transl Med. 2019. https://doi.org/10.1126/scitranslmed.aaq1427.
Turcan S, Rohle D, Goenka A, et al. IDH1 mutation is sufficient to establish the glioma hypermethylator phenotype. Nature. 2012;483:479–83.
Kadiyala P, Carney SV, Gauss JC, et al. Inhibition of 2-hydroxyglutarate elicits metabolic reprogramming and mutant IDH1 glioma immunity in mice. J Clin Invest. 2021;131:e139542. https://doi.org/10.1172/JCI139542.
Roulois D, Loo Yau H, Singhania R, et al. DNA-Demethylating Agents Target Colorectal Cancer Cells by Inducing Viral Mimicry by Endogenous Transcripts. Cell. 2015;162:961–73.
Chiappinelli KB, Strissel PL, Desrichard A, et al. Inhibiting DNA Methylation Causes an Interferon Response in Cancer via dsRNA Including Endogenous Retroviruses. Cell. 2015;162:974–86.
Ebelt ND, Zuniga E, Johnson BL, et al. 5-Azacytidine Potentiates Anti-tumor Immunity in a Model of Pancreatic Ductal Adenocarcinoma. Front Immunol. 2020;11:538.
Li H, Chiappinelli KB, Guzzetta AA, et al. Immune regulation by low doses of the DNA methyltransferase inhibitor 5-azacitidine in common human epithelial cancers. Oncotarget. 2014;5:587–98.
Gonda TA, Fang J, Salas M, et al. A DNA Hypomethylating Drug Alters the Tumor Microenvironment and Improves the Effectiveness of Immune Checkpoint Inhibitors in a Mouse Model of Pancreatic Cancer. Cancer Res. 2020;80:4754–67.
Shakya R, Gonda T, Quante M, et al. Hypomethylating therapy in an aggressive stroma-rich model of pancreatic carcinoma. Cancer Res. 2013;73:885–96.
West AC, Johnstone RW. New and emerging HDAC inhibitors for cancer treatment. J Clin Invest. 2014;124:30–9.
Bolden JE, Peart MJ, Johnstone RW. Anticancer activities of histone deacetylase inhibitors. Nat Rev Drug Discov. 2006;5:769–84.
Bondarev AD, Attwood MM, Jonsson J, et al. Recent developments of HDAC inhibitors: Emerging indications and novel molecules. Br J Clin Pharmacol. 2021;87:4577–97.
Zhang X, Zegar T, Weiser T, et al. Characterization of a dual BET/HDAC inhibitor for treatment of pancreatic ductal adenocarcinoma. Int J Cancer. 2020;147:2847–61.
Usami M, Kikuchi S, Takada K, et al. FOXO3a Activation by HDAC Class IIa Inhibition Induces Cell Cycle Arrest in Pancreatic Cancer Cells. Pancreas. 2020;49:135–42.
Hu G, He N, Cai C, et al. HDAC3 modulates cancer immunity via increasing PD-L1 expression in pancreatic cancer. Pancreatology. 2019;19:383–9.
Kim YD, Park SM, Ha HC, et al. HDAC Inhibitor, CG-745, Enhances the Anti-Cancer Effect of Anti-PD-1 Immune Checkpoint Inhibitor by Modulation of the Immune Microenvironment. J Cancer. 2020;11:4059–72.
Christmas BJ, Rafie CI, Hopkins AC, et al. Entinostat Converts Immune-Resistant Breast and Pancreatic Cancers into Checkpoint-Responsive Tumors by Reprogramming Tumor-Infiltrating MDSCs. Cancer Immunol Res. 2018;6:1561–77.
Qiu T, Zhou L, Zhu W, et al. Effects of treatment with histone deacetylase inhibitors in solid tumors: a review based on 30 clinical trials. Future Oncol. 2013;9:255–69.
West AC, Mattarollo SR, Shortt J, et al. An intact immune system is required for the anticancer activities of histone deacetylase inhibitors. Cancer Res. 2013;73:7265–76.
Woods DM, Sodre AL, Villagra A, et al. HDAC Inhibition Upregulates PD-1 Ligands in Melanoma and Augments Immunotherapy with PD-1 Blockade. Cancer Immunol Res. 2015;3:1375–85.
Luu T, Frankel P, Beumer JH, et al. Phase I trial of belinostat in combination with 13-cis-retinoic acid in advanced solid tumor malignancies: a California Cancer Consortium NCI/CTEP sponsored trial. Cancer Chemother Pharmacol. 2019;84:1201–8.
Ikeda M, Ohno I, Ueno H, et al. Phase I study of resminostat, an HDAC inhibitor, combined with S-1 in patients with pre-treated biliary tract or pancreatic cancer. Invest New Drugs. 2019;37:109–17.
Chan E, Chiorean EG, O’Dwyer PJ, et al. Phase I/II study of mocetinostat in combination with gemcitabine for patients with advanced pancreatic cancer and other advanced solid tumors. Cancer Chemother Pharmacol. 2018;81:355–64.
Chan E, Arlinghaus LR, Cardin DB, et al. Phase I trial of vorinostat added to chemoradiation with capecitabine in pancreatic cancer. Radiother Oncol. 2016;119:312–8.
Iwahashi S, Utsunomiya T, Imura S, et al. Effects of valproic acid in combination with S-1 on advanced pancreatobiliary tract cancers: clinical study phases I/II. Anticancer Res. 2014;34:5187–91.
Deming DA, Ninan J, Bailey HH, et al. A Phase I study of intermittently dosed vorinostat in combination with bortezomib in patients with advanced solid tumors. Invest New Drugs. 2014;32:323–9.
Pili R, Salumbides B, Zhao M, et al. Phase I study of the histone deacetylase inhibitor entinostat in combination with 13-cis retinoic acid in patients with solid tumours. Br J Cancer. 2012;106:77–84.
Millward M, Price T, Townsend A, et al. Phase 1 clinical trial of the novel proteasome inhibitor marizomib with the histone deacetylase inhibitor vorinostat in patients with melanoma, pancreatic and lung cancer based on in vitro assessments of the combination. Invest New Drugs. 2012;30:2303–17.
Lassen U, Molife LR, Sorensen M, et al. A phase I study of the safety and pharmacokinetics of the histone deacetylase inhibitor belinostat administered in combination with carboplatin and/or paclitaxel in patients with solid tumours. Br J Cancer. 2010;103:12–7.
Richards DA, Boehm KA, Waterhouse DM, et al. Gemcitabine plus CI-994 offers no advantage over gemcitabine alone in the treatment of patients with advanced pancreatic cancer: results of a phase II randomized, double-blind, placebo-controlled, multicenter study. Ann Oncol. 2006;17:1096–102.
Wang H, Cao Q, Dudek AZ. Phase II study of panobinostat and bortezomib in patients with pancreatic cancer progressing on gemcitabine-based therapy. Anticancer Res. 2012;32:1027–31.
Zhou Y, Bastian IN, Long MD, et al. Activation of NF-kappaB and p300/CBP potentiates cancer chemoimmunotherapy through induction of MHC-I antigen presentation. Proc Natl Acad Sci U S A. 2021;118: e2025840118.
Gao Y, Nihira NT, Bu X, et al. Acetylation-dependent regulation of PD-L1 nuclear translocation dictates the efficacy of anti-PD-1 immunotherapy. Nat Cell Biol. 2020;22:1064–75.
Fan P, Zhao J, Meng Z, et al. Overexpressed histone acetyltransferase 1 regulates cancer immunity by increasing programmed death-ligand 1 expression in pancreatic cancer. J Exp Clin Cancer Res. 2019;38:47.
Huang Y, Nahar S, Nakagawa A, et al. Regulation of GLI Underlies a Role for BET Bromodomains in Pancreatic Cancer Growth and the Tumor Microenvironment. Clin Cancer Res. 2016;22:4259–70.
Kumar K, Raza SS, Knab LM, et al. GLI2-dependent c-MYC upregulation mediates resistance of pancreatic cancer cells to the BET bromodomain inhibitor JQ1. Sci Rep. 2015;5:9489.
Ebine K, Kumar K, Pham TN, et al. Interplay between interferon regulatory factor 1 and BRD4 in the regulation of PD-L1 in pancreatic stellate cells. Sci Rep. 2018;8:13225.
Mazur PK, Herner A, Mello SS, et al. Combined inhibition of BET family proteins and histone deacetylases as a potential epigenetics-based therapy for pancreatic ductal adenocarcinoma. Nat Med. 2015;21:1163–71.
Hogg SJ, Vervoort SJ, Deswal S, et al. BET-Bromodomain Inhibitors Engage the Host Immune System and Regulate Expression of the Immune Checkpoint Ligand PD-L1. Cell Rep. 2017;18:2162–74.
Adeegbe DO, Liu S, Hattersley MM, et al. BET Bromodomain Inhibition Cooperates with PD-1 Blockade to Facilitate Antitumor Response in Kras-Mutant Non-Small Cell Lung Cancer. Cancer Immunol Res. 2018;6:1234–45.
Principe DR, Xiong R, Li Y, et al. XP-524 is a dual-BET/EP300 inhibitor that represses oncogenic KRAS and potentiates immune checkpoint inhibition in pancreatic cancer. Proc Natl Acad Sci U S A. 2022;119: e2116764119.
Aggarwal RR, Schweizer MT, Nanus DM, et al. A Phase Ib/IIa Study of the Pan-BET Inhibitor ZEN-3694 in Combination with Enzalutamide in Patients with Metastatic Castration-resistant Prostate Cancer. Clin Cancer Res. 2020;26:5338–47.
Falchook G, Rosen S, LoRusso P, et al. Development of 2 Bromodomain and Extraterminal Inhibitors With Distinct Pharmacokinetic and Pharmacodynamic Profiles for the Treatment of Advanced Malignancies. Clin Cancer Res. 2020;26:1247–57.
Michalak EM, Burr ML, Bannister AJ, et al. The roles of DNA, RNA and histone methylation in ageing and cancer. Nat Rev Mol Cell Biol. 2019;20:573–89.
Santos-Rosa H, Schneider R, Bannister AJ, et al. Active genes are tri-methylated at K4 of histone H3. Nature. 2002;419:407–11.
Greer EL, Shi Y. Histone methylation: a dynamic mark in health, disease and inheritance. Nat Rev Genet. 2012;13:343–57.
Wei Y, Xia W, Zhang Z, et al. Loss of trimethylation at lysine 27 of histone H3 is a predictor of poor outcome in breast, ovarian, and pancreatic cancers. Mol Carcinog. 2008;47:701–6.
Manuyakorn A, Paulus R, Farrell J, et al. Cellular histone modification patterns predict prognosis and treatment response in resectable pancreatic adenocarcinoma: results from RTOG 9704. J Clin Oncol. 2010;28:1358–65.
Qiu L, Hu X, Jing Q, et al. Mechanism of cancer: Oncohistones in action. J Genet Genom. 2018;45:227–36.
Wan YCE, Leung TCS, Ding D, et al. Cancer-associated histone mutation H2BG53D disrupts DNA-histone octamer interaction and promotes oncogenic phenotypes. Signal Transduct Target Ther. 2020;5:27.
Wan YCE, Liu J, Zhu L, et al. The H2BG53D oncohistone directly upregulates ANXA3 transcription and enhances cell migration in pancreatic ductal adenocarcinoma. Signal Transduct Targ Ther. 2020;5:106.
Kim KH, Roberts CW. Targeting EZH2 in cancer. Nat Med. 2016;22:128–34.
Han T, Jiao F, Hu H, et al. EZH2 promotes cell migration and invasion but not alters cell proliferation by suppressing E-cadherin, partly through association with MALAT-1 in pancreatic cancer. Oncotarget. 2016;7:11194–207.
Burr ML, Sparbier CE, Chan KL, et al. An Evolutionarily Conserved Function of Polycomb Silences the MHC Class I Antigen Presentation Pathway and Enables Immune Evasion in Cancer. Cancer Cell. 2019;36(385–401): e8.
Zhou L, Mudianto T, Ma X, et al. Targeting EZH2 Enhances Antigen Presentation, Antitumor Immunity, and Circumvents Anti-PD-1 Resistance in Head and Neck Cancer. Clin Cancer Res. 2020;26:290–300.
Peng D, Kryczek I, Nagarsheth N, et al. Epigenetic silencing of TH1-type chemokines shapes tumour immunity and immunotherapy. Nature. 2015;527:249–53.
Strepkos D, Markouli M, Klonou A, et al. Histone Methyltransferase SETDB1: A Common Denominator of Tumorigenesis with Therapeutic Potential. Cancer Res. 2021;81:525–34.
Ogawa S, Fukuda A, Matsumoto Y, et al. SETDB1 Inhibits p53-Mediated Apoptosis and Is Required for Formation of Pancreatic Ductal Adenocarcinomas in Mice. Gastroenterology. 2020;159(682–96): e13.
Griffin GK, Wu J, Iracheta-Vellve A, et al. Epigenetic silencing by SETDB1 suppresses tumour intrinsic immunogenicity. Nature. 2021;595:309–14.
Daigle SR, Olhava EJ, Therkelsen CA, et al. Potent inhibition of DOT1L as treatment of MLL-fusion leukemia. Blood. 2013;122:1017–25.
Vatapalli R, Sagar V, Rodriguez Y, et al. Histone methyltransferase DOT1L coordinates AR and MYC stability in prostate cancer. Nat Commun. 2020;11:4153.
Evanno E, Godet J, Piccirilli N, et al. Tri-methylation of H3K79 is decreased in TGF-beta1-induced epithelial-to-mesenchymal transition in lung cancer. Clin Epigenet. 2017;9:80.
Cancer Genome Atlas N. Comprehensive genomic characterization of head and neck squamous cell carcinomas. Nature. 2015;517:576.
Song D, Lyu H, Feng Q, et al. Subtyping of head and neck squamous cell cancers based on immune signatures. Int Immunopharmacol. 2021;99: 108007.
Brennan K, Shin JH, Tay JK, et al. NSD1 inactivation defines an immune cold, DNA hypomethylated subtype in squamous cell carcinoma. Sci Rep. 2017;7:17064.
Ettel M, Zhao L, Schechter S, et al. Expression and prognostic value of NSD1 and SETD2 in pancreatic ductal adenocarcinoma and its precursor lesions. Pathology. 2019;51:392–8.
Shi Y, Lei Y, Liu L, et al. Integration of comprehensive genomic profiling, tumor mutational burden, and PD-L1 expression to identify novel biomarkers of immunotherapy in non-small cell lung cancer. Cancer Med. 2021;10:2216–31.
Zhang P, Huang Y. Genomic alterations in KMT2 family predict outcome of immune checkpoint therapy in multiple cancers. J Hematol Oncol. 2021;14:39.
Lu C, Liu Z, Klement JD, et al. WDR5-H3K4me3 epigenetic axis regulates OPN expression to compensate PD-L1 function to promote pancreatic cancer immune escape. J Immunother Cancer. 2021;9: e002624.
Lu C, Paschall AV, Shi H, et al. The MLL1-H3K4me3 Axis-Mediated PD-L1 Expression and Pancreatic Cancer Immune Evasion. J Natl Cancer. 2017;109(6):djw283.
Ge L, Wang H, Xu X, et al. PRMT5 promotes epithelial-mesenchymal transition via EGFR-beta-catenin axis in pancreatic cancer cells. J Cell Mol Med. 2020;24:1969–79.
Song C, Chen T, He L, et al. PRMT1 promotes pancreatic cancer growth and predicts poor prognosis. Cell Oncol (Dordr). 2020;43:51–62.
Lu SX, De Neef E, Thomas JD, et al. Pharmacologic modulation of RNA splicing enhances anti-tumor immunity. Cell. 2021;184(4032–47): e31.
Fedoriw A, Shi L, O’Brien S, et al. Inhibiting Type I Arginine Methyltransferase Activity Promotes T Cell-Mediated Antitumor Immune Responses. Cancer Immunol Res. 2022;10:420–36.
Kim H, Kim H, Feng Y, et al. PRMT5 control of cGAS/STING and NLRC5 pathways defines melanoma response to antitumor immunity. Sci Transl Med. 2020;12(551):eaaz5683. https://doi.org/10.1126/scitranslmed.aaz5683.
Zheng NN, Zhou M, Sun F, et al. Combining protein arginine methyltransferase inhibitor and anti-programmed death-ligand-1 inhibits pancreatic cancer progression. World J Gastroenterol. 2020;26:3737–49.
Qin Y, Zhu W, Xu W, et al. LSD1 sustains pancreatic cancer growth via maintaining HIF1alpha-dependent glycolytic process. Cancer Lett. 2014;347:225–32.
Sheng W, LaFleur MW, Nguyen TH, et al. LSD1 Ablation Stimulates Anti-tumor Immunity and Enables Checkpoint Blockade. Cell. 2018;174(549–63): e19.
Wang L, Gao Y, Zhang G, et al. Enhancing KDM5A and TLR activity improves the response to immune checkpoint blockade. Sci Transl Med. 2020;12(560):eaax2282.
Chen XJ, Ren AQ, Zheng L, et al. Predictive Value of KDM5C Alterations for Immune Checkpoint Inhibitors Treatment Outcomes in Patients With Cancer. Front Immunol. 2021;12: 664847.
Zhang W, Liu W, Jia L, et al. Targeting KDM4A epigenetically activates tumor-cell-intrinsic immunity by inducing DNA replication stress. Mol Cell. 2021;81(2148–65): e9.
Chen JY, Li CF, Lai YS, et al. Lysine demethylase 2A expression in cancer-associated fibroblasts promotes breast tumour growth. Br J Cancer. 2021;124:484–93.
Duan Y, Du Y, Gu Z, et al. Expression, Prognostic Value, and Functional Mechanism of the KDM5 Family in Pancreatic Cancer. Front Cell Dev Biol. 2022;10: 887385.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no competing interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kawakubo, K., Castillo, C.Fd. & Liss, A.S. Epigenetic regulation of pancreatic adenocarcinoma in the era of cancer immunotherapy. J Gastroenterol 57, 819–826 (2022). https://doi.org/10.1007/s00535-022-01915-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00535-022-01915-2