Abstract
Recent technological advances and efforts, including powerful metagenomic and metatranscriptomic analyses, have led to a tremendous growth in our understanding of microbial communities. Changes in microbial abundance or composition of human microbial communities impact human health or disease state. However, explorations into the mechanisms underlying host–microbe interactions in health and disease are still in their infancy. Although changes in the gut microbiota have been described in patients with kidney disease, the relationships between pathogenesis, mechanisms of disease progression, and the gut microbiome are still evolving. Here, we review changes in the host–microbiome symbiotic relationship in an attempt to explore the bidirectional relationship in which alterations in the microbiome affect kidney disease progression and how kidney disease may disrupt a balanced microbiome. We also discuss potential targeted interventions that may help re-establish this symbiosis and propose more effective ways to deploy traditional treatments in patients with kidney disease.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Background
The human body harbors a complex community of bacteria, archaea, viruses, and eukaryotic microbes that inhabit interactive interfaces exposed or connected to the external environment [1]. This collection of microorganisms consists of about 100 trillion microbial cells called the human microbiota, and the genes encoded by these microbes collectively form the human microbiome [2]. The microbiome is an integral part of the human genetic landscape; therefore, for a complete understanding of our genetics, it is essential to study the composition of our microbiome [2].
Microbes have a tremendous impact on human health and well-being, with the potential to impact our physiology both in health and in disease [3]. They protect against invading pathogens, educate our immune cells, contribute to various metabolic functions, and—through these basic functions—affect directly or indirectly most of our physiologic functions [4, 5]. The breakthroughs in high-throughput techniques used to analyze the composition of the microbiome have substantially advanced our knowledge of the microbial communities colonizing various human niches [6]. Moreover, the decrease in cost of sequencing using those high-throughput technologies has enabled large-scale studies of the human microbiome [7–9]. Large-scale studies revealed that each body site, such as the oronasopharyngeal sphere, skin, vagina, and gastrointestinal tract, contains ecological communities of microbial species that exist in a mutualistic relationship with the host, also known as symbiosis [1]. Each person’s microbiome is thought to be unique [10]. Differences in species, abundance, and diversity of microbial communities within the same individual and among different individuals and various body sites have been previously described [10, 11] and are summarized in Fig. 1 [12]. Much of this intrapersonal diversity has been attributed to differences in host genetics, geographical origin and location, age, lifestyle, eating habits, and early microbial exposure, as well as antibiotics or probiotics intake [10]. Early in life, the mode of birth delivery, gestational age, hospitalization, diet, and the nature of feeding are among the factors that contribute to microbiome diversity [13].
Diversity in the human microbiome. The human microbiome is dominated by four phyla: Actinobacteria, Bacteroidetes, Firmicutes, and Proteobacteria. In the center is a phylogenetic tree of organisms abundant in the human microbiome. Commensal microbes are indicated by circles, and potential pathogens are indicated by stars. The middle ring corresponds to body sites at which various taxa are abundant and is color-coded by site [e.g., Ruminococcus (blue) is found mostly in the gut, whereas Lactobacillus (purple) is found mostly in the vagina]. Bar heights on the outside of the circle are proportional to taxa abundance at the body site of greatest prevalence [e.g., Streptococcus mitis (yellow) dominates the inside of the cheek, whereas the gut is abundant in a variety of Bacteroides]. The intensity of external colors corresponds to species prevalence in each body site (adapted with permission from [12])
Methodologies used to study the microbiome
The study of the human microbiome has been facilitated by technological advances in performing culture-independent analyses and has yielded remarkable insights into the complex diversity of the human microbiome [14]. While characterizing the microbial phylogenetic composition from a given body site describes only the archaeal and bacterial portion of the microbiome, metagenomic approaches identify all genomes existing in an environment, including bacteria, archaea, viruses, and eukaryotic microbes. Microbial phylogenetic and taxonomic applications are used to identify the microbiota composition using sequencing of the 16S ribosomal-RNA (rRNA)-encoding gene, followed by comparison to known bacterial sequence databases [15]. This method has its limitations, as it only provides insights into the taxonomic composition of the microbial community and fails to resolve a substantial fraction of the diversity existing in a community [16]. In other words, this approach allows the rapid determination of species occurrence and abundance in an efficient way for numerous samples simultaneously; however, it neglects the fact that identical species found in two different microbiomes might vary significantly in their functional capabilities [6]. To overcome these limitations, metagenomic analysis is used as an alternative approach to study the uncultured microbiota by sequencing all microbial DNA in a complex community [6, 16]. It has the additional advantage of assessing the genetic information of the microbial population and provides insights into the biological functions encoded in the microbial genome [16]. Other methodologies have also been designed to analyze the microbial transcriptome, proteome, and metabolome; they provide additional information at successive levels of microbial physiology [17]. Figure 2 summarizes some of the methodologies used to study the microbiome [18].
Bioinformatic methods for functional metagenomics. Microbial community samples typically contain many species of bacteria and other microorganisms, here indicated by different colors. After total DNA has been extracted, the composition of the community is determined by amplifying and sequencing the 16S ribosomal RNA (rRNA) gene. Highly similar sequences are grouped into operational taxonomic Units (OTUs), which are labeled by comparison with databases of recognized organisms. OTUs can then be analyzed in terms of presence/absence, abundance, or phylogenetic diversity. In order to determine biomolecular and metabolic functions present in the community, the total metagenomic DNA may be sequenced and compared with function-oriented databases. Alternatively, sequenced community DNA can be compared with reference genomes. This allows identification of microbial sequence variants and polymorphisms and provides an alternative means of detecting the presence and abundance of specific organisms (adapted with permission from [18]). KEGG Kyoto Encyclopedia of Genes and Genomes, BLAST Basic Local Alignment Search Tool, SNPs single-nucleotide polymorphism(s)
The gut microbiome
The human gut is sterile at birth [19]. Colonization with a wide variety of microbes starts at birth originating from the mother’s vaginal and fecal microbiota, as well as from other environmental microbes encountered in the first days of life [19]. The adult human gut harbors a complex community of >100 trillion microbial cells, and >1000 different bacterial species constitute the gut microbiota [20]. It is estimated that this metabolically active endogenous “organ” is equivalent to 1–2 kg of body weight and is a reservoir of >1 g of endotoxin [21]. The gut microbiota composition varies greatly between individuals, with each individual harboring a unique collection of bacterial species, which is highly stable over time, and with the most abundant bacterial phyla found in the healthy human gut being the Gram-negative Bacteroidetes and the Gram-positive low-GC Firmicutes [22]. Many nutrients in the diet are digested by human enzymes and absorbed in the small intestine. However, the gut microbiota has a central role in the metabolism of dietary fibers, which are not degraded by human enzymes. In addition to its role in food digestion, the gut microbiota plays a role in stimulation of the immune system, maintenance of intestinal epithelium homeostasis, synthesis of vitamins (B and K), enhancement of gastrointestinal tract motility and function, nutrient absorption, inhibition of pathogens by creating colonization resistance, metabolism of plant-derived compounds/drugs, and production of short-chain fatty acids (SCFAs) and polyamines [22].
Intestinal microbial dysbiosis and disease
Changes in composition and structure of the human microbiota, also known as dysbiosis, may predispose individuals to different disease conditions and explain why some people are more susceptible or resistant to certain diseases [23]. Alterations in the microbiota can result from exposure to various environmental factors, including diet, toxins, drugs, and pathogens [10]. Multiple studies have described relationships between gut microbial communities and disease states. Although these relationships are not cause-and-effect relationships, it is clear that the microbiome is an important contributor in many disease states, a factor that has been previously overlooked. In fact, changes in the microbiome are increasingly linked to the development of noncommunicable disease (NCD). Those include obesity [24], cancer [25, 26], diabetes [27, 28], inflammatory bowel disease (IBD) [29, 30], asthma [31], cardiovascular disease (CVD) [32, 33], kidney disease [21], and others. Therefore, understanding the interface between microbes and NCD may help uncover disease etiologies and pathogenesis. This may be achieved by identifying novel microbial causes or inflammatory intermediates that may be used as diagnostic and therapeutic targets for prediction, prevention, and treatment of common diseases. Deciphering the possible interindividual variations in microbial contents of the different body regions, and identifying changes in the human microbiota during onset or progression of various diseases is expected to leverage the application of microbiota-driven personalized medicine. While extensive research describing the role of the microbiome in obesity, IBD, cancer, and diabetes have been reported, only relatively recently have a few studies described the role of the microbiome in kidney disease [21, 34]. In this review, we address the role of dysbiosis in kidney disease, with a special focus on the role of the gut microbiome in chronic kidney disease (CKD).
Chronic kidney disease
CKD is a global health issue associated with loss of kidney function, CVD, infectious diseases, and premature death [35]. This lethal synergy between CKD and CVD and the increased awareness of the limitations of current treatment options has prompted the nephrology and research communities to explore alternative therapeutics to improve outcomes. Loss of kidney function in CKD results in major alterations in the blood concentration of numerous molecules [36]. In particular, substances that would normally be excreted or metabolized by the kidney accumulate as renal function declines, resulting in increased blood concentrations [36]. These uremic retention molecules (URM) constitute a long and ever-expanding list of substances that upon accumulation substantially contribute to the syndrome of uremia [36]. URM are classified according to their origin: endogenous (mammalian metabolism), microbial, or exogenous (such as diet) [37, 38]. Although the majority of URMs originate endogenously from mammalian metabolism—like dimethylarginines, homocysteine, and oxalate—it is increasingly recognized that intestinal microbial metabolism also contributes to the generation of numerous URMs.
CKD and the gut microbiome
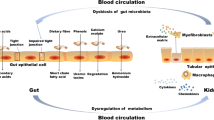
Understanding the role of human gut microbiota in the progression of CKD requires a clear comprehension of the microbiota composition, dynamics, and stability within a patient. Gut microbes produce compounds that are normally excreted by the kidneys but can be considered as potential URM [39]. The principal role of the colon is to absorb salt and water and to provide a mechanism for orderly disposal of waste products of digestion. Moreover, the colon is responsible for salvage of energy and possibly nitrogen from carbohydrates and proteins that are not digested in the upper gastrointestinal tract. This is achieved through the metabolism of anaerobic bacteria, a process known as fermentation [39]. Fermentation of the amino acids tyrosine (obtained usually from consuming turkey, chicken, beef, brown rice, nuts, fish, milk, eggs, cheese, fruit, and vegetables) and tryptophan (e.g., from beef, poultry, pork, fish, milk, yogurt, eggs, cheese, and soy products) by intestinal microbiota generates p-cresol and indole, respectively. After absorption, these compounds are further metabolized in the liver to generate p-cresyl sulfate and p-indoxyl sulfate. Indoxyl sulfate and p-cresyl sulfate circulate in equilibrium between a free solute fraction and a fraction bound to serum proteins. The best characterized binding site is albumin, for which indoxyl sulfate and p-cresyl sulfate are competitive binding inhibitors [40]. These toxins are eliminated mainly by tubular secretion in the kidneys and, therefore, are considered to be uremic toxins, with increased levels indicative of renal impairment and advancing CKD [41].
Dysbiosis in CKD patients may contribute to increased uremic toxin levels that in turn contribute to CKD progression. In a prospective, observational study of 268 patients with CKD, Wu and colleagues found the baseline concentration of indoxyl sulfate to be predictive of CKD progression [42]. Meijers and colleagues measured p-cresol levels in 499 patients with mild-to-moderate CKD and showed that p-cresol sulfate levels increased with decreasing estimated glomerular filtration rate (GFR) [43]. Likewise, an elevated p-cresol concentration was associated with increased risk of death in end-stage renal disease (ESRD) patients treated with maintenance hemodialysis [44]. Trimethylamine N-oxide (TMAO) is another uremic toxin produced by the gut microbiome, and its role in CKD has also been examined [45]. In a large cohort of CKD patients, Tang and colleagues found elevated TMAO concentrations in patients with CKD. These elevated concentrations were associated with a 70 % higher risk for all-cause mortality, even after adjusting for traditional risk factors and C-reactive protein [46].
It is worth noting that the interaction between the gut microbiota and CKD is not unidirectional. CKD also affects the structure of the gut microbiota and contributes to dysbiosis. In healthy individuals, gut microbiota are classified into different enterotypes based on the abundance of specific bacterial groups, which are dominated by Bacteroides, Prevotella, or Ruminococcus [47]; these enterotypes are strongly associated with long-term diets, particularly the levels of proteins and animal fat (Bacteroides) versus carbohydrates (Prevotella) [48]. However, the intestinal microbiota in patients with CKD is altered, with lower numbers of Lactobacillaceae and Prevotellaceae families (both are considered normal colonic microbiota) and 100 times higher Enterobacteria and Enterococci species (which are normally present in lower proportions) [49].
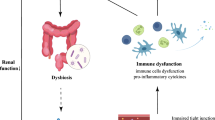
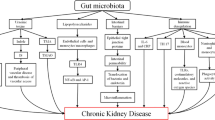
Kidney disease is associated with decreased consumption of dietary fibers [50], frequent use of antibiotics [51], slow colonic transit, metabolic acidosis, volume overload with intestinal wall congestion, intestinal wall edema, and oral iron intake [52–54]. These factors are also associated with microbial dysbiosis and higher numbers of pathogenic microbes in the gut. Many of these factors affect intestinal tight junctions and result in increased intestinal permeability and translocation of bacterial products across the intestinal barrier that will, in turn, trigger an immune response. The latter could explain the systemic inflammation that is associated with and contributes to worsening CKD and CVD [55]. Another possible mechanism of microbial dysbiosis in patients with CKD results from increased gastrointestinal urea secretion [56]. Urea is hydrolyzed by gut microbes, resulting in the formation of large quantities of ammonia, which affects the growth of commensal bacteria and causes imbalance in the gut microbiota [56]. Figure 3 summarizes mechanisms and pathways of dysbiosis in patients with CKD [57].
Dysbiosis and chronic kidney disease (CKD). CKD impairs the balance between symbionts and pathobionts in a way that favors pathobiont overgrowth. Consequences are as follows: a Impairment of the intestinal barrier by disrupting the colonic epithelial tight junction (ETJ) and decreasing epithelial survival. An increase in loss of integrity in intestinal permeability allows translocation of bacteria and lipopolysaccharide (LPS). b Dysregulation of immune response and inflammation. LPS could activate innate immune cells through toll-like receptor 4 (TLR4)-dependent and nuclear factor kappa B (NF-κB) pathways. Pathobionts stimulate dendritic cells (DCs) that activate a Th17/Th1 T-cell response and enhance production of inflammatory cytokines. c Modification of carbohydrates, protein, and bile acid (BA) fermentation. Proteins are fermented by intestinal pathobionts, which are then converted preferentially into indoxyl-sulfate (IS), p-cresyl sulfate (PCS), and trimethylamine n-oxide (TMAO). The reduction in symbionts, specifically Bifidobacterium, induces a decrease in short-chain fatty acids (SCFAs). Dysbiosis modifies BA levels and composition. INF-γ interferon γ, IL-1 interleukin-1, TNF-α tumor necrosis factor-α. (Adapted with permission from [57])
Therefore, targeting the large intestine and understanding the composition of the gut microbial communities might be a promising adjuvant approach to tackle the high morbidity and mortality in patients with CKD.
Hypertension and the gut microbiome
About 70 million Americans (29 %) have high blood pressure, which accounts for one out of three adults [58]. It is estimated that >3 % of children have hypertension [59]; this number is much higher in obese children, since the prevalence of hypertension rises progressively with increases in body mass index (BMI) percentile from ≤5th (2 %) to ≥95th (11 %) [60]. The relationship between kidney disease and hypertension is bidirectional. The microbiota of a small cohort of hypertensive patients was described as less rich and diverse than that of controls [61]. Pluznick et al. reported that major components of the olfactory signaling pathway are present in the kidney; Olfr78 is an olfactory receptor expressed in the renal juxtaglomerular apparatus, where it mediates renin secretion in response to SCFAs. Those fatty acids are end products of fermentation by the gut microbiota and are absorbed into the circulation. Treatment with antibiotics reduced the biomass of the gut microbiota and elevated blood pressure in Olfr78 knockout mice [62]. Another possible link between the gut microbiota and hypertension comes from the intestinal microbiota metabolism of choline and phosphatidylcholine, which produces trimethylamine (TMA), which is further metabolized to a proatherogenic species: TMAO. Koeth and colleagues demonstrated that metabolism by intestinal microbiota of dietary L-carnitine, a TMA abundant in red meat, produces TMAO and accelerates atherosclerosis in mice [63].
Gut microbiome and other renal conditions
Recently, multiple studies have examined the relationship between different kidney disease entities and the gut microbiome. De Angelis and colleagues examined the gut microbiome in progressor vs nonprogressor immunoglobulin A (IgA) nephropathy (IgAN) patients and compared them with healthy controls [64]. They showed that some traits of the gut microbiota and levels of urinary metabolites (free amino acids and organic volatile compounds) vary significantly between the progressor and nonprogressor groups [64]. It was hypothesized that the increased free amino acids in the serum due to IgAN pathology is possibly related to the decreased absorption of gastrointestinal proteins, which allowed for increased microbial proteolysis, altered microbiota, and contributed to elevated fecal p-cresol levels. Recent data indicate that intestinal microbiota can modify acute kidney injury (AKI) also. One possible mechanism is via the renoprotective action of SCFAs against ischaemia–reperfusion in animal models; SCFAs (which have anti-inflammatory properties) are produced by the intestinal microbiota [65]. Therapy with three main SCFAs (acetate, propionate, and butyrate) improved renal dysfunction caused by injury and was associated with reductions in the levels of reactive oxygen species, inflammation, infiltrating immune cells, and apoptotic cells in the injured kidneys; an increase in proliferation of kidney epithelial cells; and modulation of DNA methylation status. Another possible mechanism by which the microbiota affect AKI outcome is related to the hygiene hypothesis [66]; Jang and colleagues reported that germ-free animals when subjected to ischemia-induced AKI have significantly worse structural/functional renal injuries and inflammation compared with control mice. This may be due to a T-helper 1 type response similar to that seen in autoimmune disease. Furthermore, the microbiota may have a wider influence and role in autoimmune kidney disease through its immunomodulatory effects, recognized by its influence on polarization of T-cell subsets and natural killer cells [67]. Abnormalities in the immune system can also induce kidney damage, either by provoking an autoimmune phenomenon or induction of molecular mimicry. Staphylococcus aureus may be a direct pathogenetic factor in granulomatosis with polyangiitis (former Wegener’s granulomatosis) [68]. Chronic bacterial colonization or chronic infections of the upper respiratory tract have been suspected to be a trigger of IgA vasculitis and IgAN [69]. Microbial antigens may play a role in membranous nephropathy and lupus nephritis, as suggested by the abundance of Helicobacter pylori antigen deposition in renal biopsy specimens and evidence of infection in blood [70]. Undergoing kidney transplantation and the use of posttransplant medications has a major impact on the microbiota composition as well [71]. On the other hand, differences in the gut microbiota may affect medication bioavailability and dosing to achieve therapeutic levels; this may help further explain interindividual differences in tacrolimus dosing [72] and how antibiotic intake may increase the bioavailability of amlodipine and possibly change its therapeutic potency by suppressing gut microbial metabolic activities [73].
Restoring the balance: current and potential interventions
These interventions can be classified as being designed to lower uremic toxin production by restricting intake of uremic toxin precursors (lower protein intake), by restoring a more balanced gut microbiome (food supplements), and by interventions aiming to enhance the disposal of these toxins (adsorptive therapies).
Food supplements
Prebiotics
Prebiotics is a general term to refer to nondigestible (by the host) food ingredients that induce the growth or activity of microorganisms (e.g., bacteria and fungi) that contribute to the well-being of their host. Inulin, fructo-oligosaccharides, galacto-oligosaccharides, soya-oligosaccharides, xylo-oligosaccharides, and pyrodextrins are among known prebiotics. There are limited studies examining their effect in CKD patients. In a randomized, controlled, single-blind clinical trial with a crossover design, Bliss and colleagues studied the effect of gum arabic (acacia gum) fiber supplementation (50 g/day) in 16 CKD patients consuming a low-protein diet [74]. Four weeks after receiving gum arabic fiber, patients had increased fecal nitrogen excretion and lower serum urea nitrogen concentration compared with the placebo group. Only one small study of three pediatric patients with ESRD examined the efficacy of acacia gum (1 g/kg per day in divided doses) and reported improved quality of life [75]. Another randomized controlled trial examined the effect of increasing dietary fiber on plasma levels of colon-derived solutes in 56 hemodialysis patients [76]. After 6 weeks, patients on increasing dietary fiber had significantly reduced unbound, free plasma levels of indoxyl sulfate, while the reduction of p-cresol sulfate levels did not achieve significance. The authors concluded that increasing dietary fiber in hemodialysis patients might reduce plasma levels of the colon-derived solutes indoxyl sulfate and possibly p-cresol sulfate without the need to intensify dialysis treatments.
Probiotics
Probiotics are living microorganisms that, when administered in adequate amounts, confer a health benefit on the host. Studies examining the effects of probiotics in CKD patients using serum tumor necrosis factor alpha (TNF-α), interleukin (IL)-5 and -6, and endotoxin show conflicting results. In a randomized, double-blind, placebo-controlled trial examining the effect of probiotics on serum cytokine and endotoxin levels of patients on peritoneal dialysis, patients who received probiotics had lower proinflammatory cytokines and endotoxin levels while levels of serum IL-10 significantly increased [77]. In a recent study, Yacoub et al. analyzed National Health and Nutrition Survey (NHANES) data to examine the association of yogurt/probiotic with kidney parameters. Frequent yogurt and/or probiotics use was associated with decreased risk of proteinuric kidney disease [78].
Synbiotics
Synbiotics refer to nutritional supplements combining probiotics and prebiotics in a form of synergism. In a randomized, placebo-controlled trial, Guida and colleagues studied the effect of a 4-week synbiotic treatment on plasma p-cresol levels in 30 patients with stage 3–4 CKD [79]. The authors found the group on synbiotics to have lower total plasma p-cresol concentrations and suggested that because high plasma concentrations of p-cresol in early phases of CKD are predictive of progression to ESRD, synbiotics deserve attention as possible tools to delay CKD progression. Others have reported similar beneficial effects of synbiotic and low-protein treatment on CKD progression [80]. In a recent randomized trial, Rossi et al. evaluated the effect of synbiotics therapy on gut microbiota and serum concentrations of indoxyl sulfate and p-cresyl sulfate in predialysis CKD [81]. Synbiotic therapy did not significantly reduce serum indoxyl sulfate levels but reduced levels of p-cresyl sulfate and favorably altered the stool microbiome, particularly with enrichment of Bifidobacterium and depletion of Ruminococcaceae [81].
Adsorbent therapies
AST-120 is an orally ingested intestinal spherical carbon adsorbent consisting of porous carbon particles of 0.2–0.4 mm in diameter and is insoluble in water and common organic solvents. It adsorbs indole, the precursor of indoxyl sulfate derived from the metabolism of tryptophan by bacteria within the gastrointestinal tract and therefore is used to attenuate indoxyl sulfate accumulation in patients with CKD [82]. AST-120 has been available in Japan since 1991. In prospective trials and retrospective analyses, AST-120 has been shown to prolong the time to initiation of hemodialysis [83] and slow the decline in GFR and the increase in serum creatinine [84]. In an initial randomized, double-blind, placebo-controlled trial in the United States, AST-120 was associated with a significant dose-dependent reduction in serum indoxyl sulfate levels and a decrease in uremia-related malaise [85]. The Evaluating Prevention of Progression in CKD (EPPIC) trials, two double-blind, placebo-controlled trials undertaken in North America/Latin America and Europe, evaluated the efficacy of AST-120 for preventing progression of CKD in 2035 adults with moderate to severe disease. Patients we randomized to receive either placebo or AST-120 (9 g/d). The study found the time to primary end point (a composite of dialysis initiation, kidney transplantation, and serum creatinine doubling) to be similar between the AST-120 and placebo groups in both trials. The authors concluded that the benefit of adding AST-120 to standard therapy in patients with moderate to severe CKD is not supported by their study data [86].
Fecal microbiota transplantation (FMT)
FMT is becoming increasingly accepted as an effective and safe intervention in patients with recurrent Clostridium difficile infection, aiming at the restoration of a disrupted microbiome [87]. While no large studies examining the effect of fecal transplantation on restoring the gut microbiome in patients with CKD are available, a case showing successful eradication of a pathogenic organism in a patient with ESRD has been reported [88]. Whether healthy microbial transfer “in a pill” will be part of our future management of patients with CKD and dysbiosis remains to be seen. Until then, further studies are needed to describe what a healthy microbiome is in a patient with CKD and what hard clinical outcomes can be achieved from manipulating the microbiome.
Conclusion
The relationship between the human microbiome and kidney disease is bidirectional. Recent studies have described how kidney disease contributes to dysbiosis and how dysbiosis contributes to progression of kidney disease. Clinicians must be aware of the potential, unintended effects of treatments that may alter the gut microbiome, exercise self-discipline, and weigh risks and benefits when prescribing prophylactic antibiotics to patients with recurrent urinary tract infections, vesicourethral reflux, and other infections. There is a pressing need for more studies that characterize the microbiome profile in children with CKD and explore the relationship between different pediatric kidney disease parameters and the microbiome of the growing child. This is not only needed to establish relationships by association but to examine the interaction between certain early-life microbial and antibiotic exposure on the pathogenesis of kidney disease. Multiple promising interventions have been described to restore a more balanced microbiome and possibly slow the progression of CKD; such interventions need to be further examined in large controlled trials before they can become part of our mainstream management.
References
Hamady M, Knight R (2009) Microbial community profiling for human microbiome projects: Tools, techniques, and challenges. Genome Res 19:1141–1152
Solt I, Kim MJ, Offer C (2011) The human microbiome. Harefuah 150:484–488
Pflughoeft KJ, Versalovic J (2012) Human microbiome in health and disease. Annu Rev Pathol 7:99–122
Hooper LV, Midtvedt T, Gordon JI (2002) How host-microbial interactions shape the nutrient environment of the mammalian intestine. Annu Rev Nutr 22:283–307
Backhed F, Ley RE, Sonnenburg JL, Peterson DA, Gordon JI (2005) Host-bacterial mutualism in the human intestine. Science 307:1915–1920
Kuczynski J, Lauber CL, Walters WA, Parfrey LW, Clemente JC, Gevers D, Knight R (2012) Experimental and analytical tools for studying the human microbiome. Nat Rev Genet 13:47–58
Conlan S, Kong HH, Segre JA (2012) Species-level analysis of DNA sequence data from the NIH Human Microbiome Project. PLoS ONE 7, e47075
NIH HMP Working Group, Peterson J, Garges S, Giovanni M, McInnes P, Wang L, Schloss JA, Bonazzi V, McEwen JE, Wetterstrand KA, Deal C, Baker CC, Di Francesco V, Howcroft TK, Karp RW, Lunsford RD, Wellington CR, Belachew T, Wright M, Giblin C, David H, Mills M, Salomon R, Mullins C, Akolkar B, Begg L, Davis C, Grandison L, Humble M, Khalsa J, Little AR, Peavy H, Pontzer C, Portnoy M, Sayre MH, Starke-Reed P, Zakhari S, Read J, Watson B, Guyer M (2009) The NIH Human Microbiome Project. Genome Res 19:2317–2323
Blaser MJ (2010) Harnessing the power of the human microbiome. Proc Natl Acad Sci U S A 107:6125–6126
Ursell LK, Clemente JC, Rideout JR, Gevers D, Caporaso JG, Knight R (2012) The interpersonal and intrapersonal diversity of human-associated microbiota in key body sites. J Allergy Clin Immunol 129:1204–1208
Cho I, Blaser MJ (2012) The human microbiome: at the interface of health and disease. Nat Rev Genet 13:260–270
Morgan XC, Segata N, Huttenhower C (2013) Biodiversity and functional genomics in the human microbiome. Trends Genet 29:51–58
Johnson CL, Versalovic J (2012) The human microbiome and its potential importance to pediatrics. Pediatrics 129:950–960
Sankar SA, Lagier JC, Pontarotti P, Raoult D, Fournier PE (2015) The human gut microbiome, a taxonomic conundrum. Syst Appl Microbiol 38:276–286
Ram JL, Karim AS, Sendler ED, Kato I (2011) Strategy for microbiome analysis using 16S rRNA gene sequence analysis on the Illumina sequencing platform. Syst Biol Reprod Med 57:162–170
Logares R, Sunagawa S, Salazar G, Cornejo-Castillo FM, Ferrera I, Sarmento H, Hingamp P, Ogata H, de Vargas C, Lima-Mendez G, Raes J, Poulain J, Jaillon O, Wincker P, Kandels-Lewis S, Karsenti E, Bork P, Acinas SG (2014) Metagenomic 16S rDNA Illumina tags are a powerful alternative to amplicon sequencing to explore diversity and structure of microbial communities. Environ Microbiol 16:2659–2671
Boyang Ji JN (2015) New insight into the gut microbiome through metagenomics. Adv Genom Genet 5:77–91
Morgan XC, Huttenhower C (2012) Chapter 12: Human microbiome analysis. PLoS Comput Biol 8, e1002808
Dominguez-Bello MG, Costello EK, Contreras M, Magris M, Hidalgo G, Fierer N, Knight R (2010) Delivery mode shapes the acquisition and structure of the initial microbiota across multiple body habitats in newborns. Proc Natl Acad Sci U S A 107:11971–11975
Ley RE, Peterson DA, Gordon JI (2006) Ecological and evolutionary forces shaping microbial diversity in the human intestine. Cell 124:837–848
Ramezani A, Raj DS (2014) The gut microbiome, kidney disease, and targeted interventions. J Am Soc Nephrol 25:657–670
Dominguez-Bello MG, Blaser MJ, Ley RE, Knight R (2011) Development of the Human Gastrointestinal Microbiota and Insights From High-Throughput Sequencing. Gastroenterology 140:1713–1719
Carding S, Verbeke K, Vipond DT, Corfe BM, Owen LJ (2015) Dysbiosis of the gut microbiota in disease. Microb Ecol Health Dis 26:26191
Bradlow HL (2014) Obesity and the gut microbiome: pathophysiological aspects. Horm Mol Biol Clin Investig 17:53–61
Rogers CJ, Prabhu KS, Vijay-Kumar M (2014) The microbiome and obesity-an established risk for certain types of cancer. Cancer J 20:176–180
Francescone R, Hou V, Grivennikov SI (2014) Microbiome, inflammation, and cancer. Cancer J 20:181–189
Dunne JL, Triplett EW, Gevers D, Xavier R, Insel R, Danska J, Atkinson MA (2014) The intestinal microbiome in type 1 diabetes. Clin Exp Immunol 177:30–37
Gross M (2013) Does the gut microbiome hold clues to obesity and diabetes? Curr Biol 23:R359–362
Major G, Spiller R (2014) Irritable bowel syndrome, inflammatory bowel disease and the microbiome. Curr Opin Endocrinol Diabetes Obes 21:15–21
Kostic AD, Xavier RJ, Gevers D (2014) The microbiome in inflammatory bowel disease: current status and the future ahead. Gastroenterology 146:1489–1499
Fujimura KE, Lynch SV (2015) Microbiota in allergy and asthma and the emerging relationship with the gut microbiome. Cell Host Microbe 17:592–602
Dinakaran V, Rathinavel A, Pushpanathan M, Sivakumar R, Gunasekaran P, Rajendhran J (2014) Elevated levels of circulating DNA in cardiovascular disease patients: metagenomic profiling of microbiome in the circulation. PLoS ONE 9, e105221
Rajendhran J, Shankar M, Dinakaran V, Rathinavel A, Gunasekaran P (2013) Contrasting circulating microbiome in cardiovascular disease patients and healthy individuals. Int J Cardiol 168:5118–5120
Anders HJ, Andersen K, Stecher B (2013) The intestinal microbiota, a leaky gut, and abnormal immunity in kidney disease. Kidney Int 83:1010–1016
Levey AS, Eckardt KU, Tsukamoto Y, Levin A, Coresh J, Rossert J, De Zeeuw D, Hostetter TH, Lameire N, Eknoyan G (2005) Definition and classification of chronic kidney disease: a position statement from Kidney Disease: Improving Global Outcomes (KDIGO). Kidney Int 67:2089–2100
Meyer TW, Hostetter TH (2007) Uremia. N Engl J Med 357:1316–1325
Jankowski J, van der Giet M, Jankowski V, Schmidt S, Hemeier M, Mahn B, Giebing G, Tolle M, Luftmann H, Schluter H, Zidek W, Tepel M (2003) Increased plasma phenylacetic acid in patients with end-stage renal failure inhibits iNOS expression. J Clin Invest 112:256–264
Wikoff WR, Anfora AT, Liu J, Schultz PG, Lesley SA, Peters EC, Siuzdak G (2009) Metabolomics analysis reveals large effects of gut microflora on mammalian blood metabolites. Proc Natl Acad Sci U S A 106:3698–3703
Evenepoel P, Meijers BK, Bammens BR, Verbeke K (2009) Uremic toxins originating from colonic microbial metabolism. Kidney Int Suppl 114:S12–19
Fukagawa M, Watanabe Y (2011) Role of uremic toxins and oxidative stress in chronic kidney disease. Ther Apher Dial 15:119
Poesen R, Meijers B, Evenepoel P (2013) The Colon: An Overlooked Site for Therapeutics in Dialysis Patients. Semin Dial 26:323–332
Wu IW, Hsu KH, Lee CC, Sun CY, Hsu HJ, Tsai CJ, Tzen CY, Wang YC, Lin CY, Wu MS (2011) p-Cresyl sulphate and indoxyl sulphate predict progression of chronic kidney disease. Nephrol Dial Transpl 26:938–947
Meijers BK, Claes K, Bammens B, de Loor H, Viaene L, Verbeke K, Kuypers D, Vanrenterghem Y, Evenepoel P (2010) p-Cresol and cardiovascular risk in mild-to-moderate kidney disease. Clin J Am Soc Nephrol 5:1182–1189
Bammens B, Evenepoel P, Keuleers H, Verbeke K, Vanrenterghem Y (2006) Free serum concentrations of the protein-bound retention solute p-cresol predict mortality in hemodialysis patients. Kidney Int 69:1081–1087
Missailidis C, Hallqvist J, Qureshi AR, Barany P, Heimburger O, Lindholm B, Stenvinkel P, Bergman P (2016) Serum Trimethylamine-N-Oxide Is Strongly Related to Renal Function and Predicts Outcome in Chronic Kidney Disease. PLoS ONE 11, e0141738
Tang WHW, Wang ZN, Kennedy DJ, Wu YP, Buffa JA, Agatisa-Boyle B, Li XMS, Levison BS, Hazen SL (2015) Gut Microbiota-Dependent Trimethylamine N-Oxide (TMAO) Pathway Contributes to Both Development of Renal Insufficiency and Mortality Risk in Chronic Kidney Disease. Circ Res 116:448–455
Arumugam M, Raes J, Pelletier E, Le Paslier D, Yamada T, Mende DR, Fernandes GR, Tap J, Bruls T, Batto JM, Bertalan M, Borruel N, Casellas F, Fernandez L, Gautier L, Hansen T, Hattori M, Hayashi T, Kleerebezem M, Kurokawa K, Leclerc M, Levenez F, Manichanh C, Nielsen HB, Nielsen T, Pons N, Poulain J, Qin J, Sicheritz-Ponten T, Tims S, Torrents D, Ugarte E, Zoetendal EG, Wang J, Guarner F, Pedersen O, de Vos WM, Brunak S, Dore J, Meta HITC, Antolin M, Artiguenave F, Blottiere HM, Almeida M, Brechot C, Cara C, Chervaux C, Cultrone A, Delorme C, Denariaz G, Dervyn R, Foerstner KU, Friss C, van de Guchte M, Guedon E, Haimet F, Huber W, van Hylckama-Vlieg J, Jamet A, Juste C, Kaci G, Knol J, Lakhdari O, Layec S, Le Roux K, Maguin E, Merieux A, Melo Minardi R, M’Rini C, Muller J, Oozeer R, Parkhill J, Renault P, Rescigno M, Sanchez N, Sunagawa S, Torrejon A, Turner K, Vandemeulebrouck G, Varela E, Winogradsky Y, Zeller G, Weissenbach J, Ehrlich SD, Bork P (2011) Enterotypes of the human gut microbiome. Nature 473:174–180
Wu GD, Chen J, Hoffmann C, Bittinger K, Chen YY, Keilbaugh SA, Bewtra M, Knights D, Walters WA, Knight R, Sinha R, Gilroy E, Gupta K, Baldassano R, Nessel L, Li H, Bushman FD, Lewis JD (2011) Linking long-term dietary patterns with gut microbial enterotypes. Science 334:105–108
Vaziri ND, Wong J, Pahl M, Piceno YM, Yuan J, DeSantis TZ, Ni Z, Nguyen TH, Andersen GL (2013) Chronic kidney disease alters intestinal microbial flora. Kidney Int 83:308–315
Kalantar-Zadeh K, Kopple JD, Deepak S, Block D, Block G (2002) Food intake characteristics of hemodialysis patients as obtained by food frequency questionnaire. J Ren Nutr 12:17–31
Jakobsson HE, Jernberg C, Andersson AF, Sjolund-Karlsson M, Jansson JK, Engstrand L (2010) Short-Term Antibiotic Treatment Has Differing Long-Term Impacts on the Human Throat and Gut Microbiome. Plos One 5, e9836
Fouque D, Cruz Casal M, Lindley E, Rogers S, Pancirova J, Kernc J, Copley JB (2014) Dietary trends and management of hyperphosphatemia among patients with chronic kidney disease: an international survey of renal care professionals. J Ren Nutr 24:110–115
Nakao T (2012) Chronic kidney disease (CKD)--recent progress. Topics: VII. Management of chronic kidney disease (CKD) and treatment; 1. Dietary treatment and life-style management in CKD patients. Nihon Naika Gakkai Zasshi 101:1340–1346
Dietary management of chronic kidney disease patients: protein-restricted diets supplemented with keto/amino acids. Abstracts from the International Advisory Board Meetings 2003/2004 (2005). Am J Nephrol 25 Suppl 1:1–28
Wing MR, Patel SS, Ramezani A, Raj DS (2016) Gut microbiome in chronic kidney disease. Exp Physiol 101:471–477
Sud K, Sakhuja V (1997) The gastrointestinal tract in uremia. J Assoc Physicians India 45:833–834
Koppe L, Mafra D, Fouque D (2015) Probiotics and chronic kidney disease. Kidney Int 88:958–966
Gu QP, Burt VL, Dillon CF, Yoon S (2012) Trends in Antihypertensive Medication Use and Blood Pressure Control Among United States Adults With Hypertension The National Health and Nutrition Examination Survey, 2001 to 2010. Circulation 126:2105–2114
Falkner B (2010) Hypertension in children and adolescents: epidemiology and natural history. Pediatr Nephrol 25:1219–1224
Sorof JM, Lai DJ, Turner J, Poffenbarger T, Portman RJ (2004) Overweight, ethnicity, and the prevalence of hypertension in school-aged children. Pediatrics 113:475–482
Yang T, Santisteban MM, Rodriguez V, Li E, Ahmari N, Carvajal JM, Zadeh M, Gong MH, Qi YF, Zubcevic J, Sahay B, Pepine CJ, Raizada MK, Mohamadzadeh M (2015) Gut Dysbiosis Is Linked to Hypertension. Hypertension 65:1331–1340
Pluznick JL, Protzko RJ, Gevorgyan H, Peterlin Z, Sipos A, Han J, Brunet I, Wan LX, Rey F, Wang T, Firestein SJ, Yanagisawa M, Gordon JI, Eichmann A, Peti-Peterdi J, Caplan MJ (2013) Olfactory receptor responding to gut microbiota-derived signals plays a role in renin secretion and blood pressure regulation. Proc Natl Acad Sci U S A 110:4410–4415
Koeth RA, Wang ZE, Levison BS, Buffa JA, Org E, Sheehy BT, Britt EB, Fu XM, Wu YP, Li L, Smith JD, DiDonato JA, Chen J, Li HZ, Wu GD, Lewis JD, Warrier M, Brown JM, Krauss RM, Tang WHW, Bushman FD, Lusis AJ, Hazen SL (2013) Intestinal microbiota metabolism of L-carnitine, a nutrient in red meat, promotes atherosclerosis. Nat Med 19:576–585
De Angelis M, Montemurno E, Piccolo M, Vannini L, Lauriero G, Maranzano V, Gozzi G, Serrazanetti D, Dalfino G, Gobbetti M, Gesualdo L (2014) Microbiota and metabolome associated with immunoglobulin A nephropathy (IgAN). PLoS ONE 9, e99006
Andrade-Oliveira V, Amano MT, Correa-Costa M, Castoldi A, Felizardo RJ, de Almeida DC, Bassi EJ, Moraes-Vieira PM, Hiyane MI, Rodas AC, Peron JP, Aguiar CF, Reis MA, Ribeiro WR, Valduga CJ, Curi R, Vinolo MA, Ferreira CM, Camara NO (2015) Gut Bacteria Products Prevent AKI Induced by Ischemia-Reperfusion. J Am Soc Nephrol 26:1877–1888
Jang HR, Gandolfo MT, Ko GJ, Satpute S, Racusen L, Rabb H (2009) Early exposure to germs modifies kidney damage and inflammation after experimental ischemia-reperfusion injury. Am J Physiol Renal Physiol 297:F1457–1465
Macpherson AJ, Harris NL (2004) Interactions between commensal intestinal bacteria and the immune system. Nat Rev Immunol 4:478–485
Peschel A, Basu N, Benharkou A, Brandes R, Brown M, Dieckmann R, Rees AJ, Kain R (2014) Autoantibodies to hLAMP-2 in ANCA-negative pauci-immune focal necrotizing GN. J Am Soc Nephrol 25:455–463
Popa ER, Stegeman CA, Bos NA, Kallenberg CG, Tervaert JW (2003) Staphylococcal superantigens and T cell expansions in Wegener’s granulomatosis. Clin Exp Immunol 132:496–504
Moriyama T, Kaneko T, Fujii M, Tsubakihara Y, Kawano S, Imai E (2006) High prevalence of Helicobacter pylori infection in Japanese patients with membranous nephropathy. Aliment Pharmacol Ther Symp Ser 2:189–193
Fricke WF, Maddox C, Song Y, Bromberg JS (2014) Human microbiota characterization in the course of renal transplantation. Am J Transplant 14:416–427
Lee JR, Muthukumar T, Dadhania D, Taur Y, Jenq RR, Toussaint NC, Ling L, Pamer E, Suthanthiran M (2015) Gut microbiota and tacrolimus dosing in kidney transplantation. PLoS ONE 10, e0122399
Yoo HH, Kim IS, Yoo DH, Kim DH (2016) Effects of orally administered antibiotics on the bioavailability of amlodipine: gut microbiota-mediated drug interaction. J Hypertens 34:156–162
Bliss DZ, Stein TP, Schleifer CR, Settle RG (1996) Supplementation with gum arabic fiber increases fecal nitrogen excretion and lowers serum urea nitrogen concentration in chronic renal failure patients consuming a low-protein diet. Am J Clin Nutr 63:392–398
Al-Mosawi AJ (2004) Acacia gum supplementation of a low-protein diet in children with end-stage renal disease. Pediatr Nephrol 19:1156–1159
Sirich TL, Plummer NS, Gardner CD, Hostetter TH, Meyer TW (2014) Effect of increasing dietary fiber on plasma levels of colon-derived solutes in hemodialysis patients. Clin J Am Soc Nephrol 9:1603–1610
Wang IK, Wu YY, Yang YF, Ting IW, Lin CC, Yen TH, Chen JH, Wang CH, Huang CC, Lin HC (2015) The effect of probiotics on serum levels of cytokine and endotoxin in peritoneal dialysis patients: a randomised, double-blind, placebo-controlled trial. Benefic Microbes 6:423–430
Yacoub R, Kaji D, Patel SN, Simoes PK, Busayavalasa D, Nadkarni GN, He JC, Coca SG, Uribarri J (2016) Association between probiotic and yogurt consumption and kidney disease: insights from NHANES. Nutr J 15:10
Guida B, Germano R, Trio R, Russo D, Memoli B, Grumetto L, Barbato F, Cataldi M (2014) Effect of short-term synbiotic treatment on plasma p-cresol levels in patients with chronic renal failure: a randomized clinical trial. Nutr Metab Cardiovasc Dis 24:1043–1049
Pavan M (2014) Influence of prebiotic and probiotic supplementation on the progression of chronic kidney disease. Minerva Urol Nefrol. ISSN 0393–2249
Rossi M, Johnson DW, Morrison M, Pascoe EM, Coombes JS, Forbes JM, Szeto CC, McWhinney BC, Ungerer JP, Campbell KL (2016) Synbiotics Easing Renal Failure by Improving Gut Microbiology (SYNERGY): A Randomized Trial. Clin J Am Soc Nephrol 11:223–231
Niwa T, Emoto Y, Maeda K, Uehara Y, Yamada N, Shibata M (1991) Oral sorbent suppresses accumulation of albumin-bound indoxyl sulphate in serum of haemodialysis patients. Nephrol Dial Transplant 6:105–109
Ueda H, Shibahara N, Takagi S, Inoue T, Katsuoka Y (2007) AST-120, an oral adsorbent, delays the initiation of dialysis in patients with chronic kidney diseases. Ther Apher Dial 11:189–195
Akizawa T, Asano Y, Morita S, Wakita T, Onishi Y, Fukuhara S, Gejyo F, Matsuo S, Yorioka N, Kurokawa K, CAP-KD Study Group (2009) Effect of a carbonaceous oral adsorbent on the progression of CKD: a multicenter, randomized, controlled trial. Am J Kidney Dis 54:459–467
Schulman G, Agarwal R, Acharya M, Berl T, Blumenthal S, Kopyt N (2006) A multicenter, randomized, double-blind, placebo-controlled, dose-ranging study of AST-120 (Kremezin) in patients with moderate to severe CKD. Am J Kidney Dis 47:565–577
Schulman G, Berl T, Beck GJ, Remuzzi G, Ritz E, Arita K, Kato A, Shimizu M (2015) Randomized Placebo-Controlled EPPIC Trials of AST-120 in CKD. J Am Soc Nephrol 26:1732–1746
Rohlke F, Stollman N (2012) Fecal microbiota transplantation in relapsing Clostridium difficile infection. Ther Adv Gastroenter 5:403–420
Singh R, van Nood E, Nieuwdorp M, van Dam B, ten Berge IJM, Geerlings SE, Bemelman FJ (2014) Donor feces infusion for eradication of Extended Spectrum beta-Lactamase producing Escherichia coli in a patient with end stage renal disease. Clin Microbiol Infect 20:O977–O978
Acknowledgments
The authors thank Dr. Bernice Lo for her assistance in the proofreading of the review.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare no conflict of interest.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Al Khodor, S., Shatat, I.F. Gut microbiome and kidney disease: a bidirectional relationship. Pediatr Nephrol 32, 921–931 (2017). https://doi.org/10.1007/s00467-016-3392-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00467-016-3392-7