Abstract
Single-nucleotide polymorphisms (SNPs) and small genomic regions with multiple SNPs (microhaplotypes, MHs) are rapidly emerging as novel forensic investigative tools to assist in individual identification, kinship analyses, ancestry inference, and deconvolution of DNA mixtures. Here, we analyzed information for 90 microhaplotype loci in 4009 individuals from 79 world populations in 6 major biogeographic regions. The study included multiplex microhaplotype sequencing (mMHseq) data analyzed for 524 individuals from 16 populations and genotype data for 3485 individuals from 63 populations curated from public repositories. Analyses of the 79 populations revealed excellent characteristics for this 90-plex MH panel for various forensic applications achieving an overall average effective number of allele values (Ae) of 4.55 (range 1.04–19.27) for individualization and mixture deconvolution. Population-specific random match probabilities ranged from a low of 10–115 to a maximum of 10–66. Mean informativeness (In) for ancestry inference was 0.355 (range 0.117–0.883). 65 novel SNPs were detected in 39 of the MHs using mMHseq. Of the 3018 different microhaplotype alleles identified, 1337 occurred at frequencies > 5% in at least one of the populations studied. The 90-plex MH panel enables effective differentiation of population groupings for major biogeographic regions as well as delineation of distinct subgroupings within regions. Open-source, web-based software is available to support validation of this technology for forensic case work analysis and to tailor MH analysis for specific geographical regions.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
For many years, the DNA markers for forensic practice have been short tandem repeat (STR) loci that are highly polymorphic with different numbers of repeat units at each locus (Budowle et al. 1998). Over the years, the numbers of standard STR loci have increased and the similarities of the different commercial panels and those in different countries have increased (Butler and Hill 2012; Schumm et al. 2013; Guo et al. 2014; Novroski et al. 2019). Other types of markers have been proposed starting with SNPs especially in the early 2000s (cf., Pakstis et al. 2007 for early studies). Early forensic studies of SNPs were focused on individual identification (Sanchez et al. 2006; Pakstis et al. 2007), on panels of SNPs for inferring population ancestry (e.g., Shriver et al. 2004; Tishkoff and Kidd 2004; Phillips et al. 2007), and on SNPs for phenotype (e.g., Lamason et al. 2005; Walsh et al. 2011; Walsh et al. 2013). Several commercial panels of SNPs have been introduced, some of which combine SNPs with STRs, for analysis using Massively Parallel Sequencing (MPS). MPS has also allowed the further development of a new type of genetic marker, microhaplotypes (Kidd et al. 2013,2014).
Microhaplotypes (microhaplotypes, MHs) have been defined as small genomic regions of less than ~ 300 bp with two or more polymorphisms, usually SNPs, resulting in at least three common haplotypes in the population (cf., review in Oldoni et al. 2019). They were first proposed as potentially highly informative and useful genetic markers for forensics, anthropology, and biomedical research generally. Their desirable characteristics include multiple alleles with high heterozygosity and low mutation rates. Since then, MHs have been studied by many researchers with clear demonstration of their potential for forensic, medical, and anthropologic applications (Bulbul et al. 2018; Chen et al. 2018; Kidd et al. 2018a,2021; Cheung et al. 2019; Phillips et al. 2019; Puente et al. 2020a; Puente et al. 2020b), but they have not yet been incorporated into routine forensic casework.
Although conceived of for use with MPS, the original studies which were designed to evaluate the potential for microhaplotypes (Kidd et al. 2013,2014) used TaqMan to type individual SNPs and then PHASE (Stephens et al. 2001) to determine the genotypes and haplotype frequencies. The SNPs that were chosen to study were those of at least modest frequency (5–10%) in some populations, those that were not in complete LD with another, and those for which TaqMan assays were available. Other factors could be included in selection of SNPs if different ultimate objectives were favored (Kidd and Speed 2015; Kidd et al. 2018a). While MPS was not used in these exploratory studies, it was clear that the existence of MPS was what made study of microhaplotypes relevant. Now, there have been several studies that have used MPS successfully to study panels of microhaplotypes on multiple individuals and/or populations (Turchi et al. 2019; Chen et al. 2018; Oldoni et al. 2019; Bennett et al. 2019; Gandotra et al. 2020; Puente et al. 2020a; Puente et al. 2020b; Kureshi et al. 2020; Wu et al. 2021a; Wu et al. 2021b).
We previously presented a panel of 90 microhaplotypes evaluated using data for 26 populations extracted from the 1000 Genomes (1 KG) project data (1000 Genomes Consortium Project 2015) as well as data on 155 individuals from four other populations studied using multiplex microhaplotype sequencing (mMHseq) of all 90 loci (Gandotra et al. 2020). The 90 loci had a high overall effective number of alleles (Ae) in the 30 populations studied (average Ae > 5.08). Analyses of frequency variation among populations showed that some of the loci had significant variation among populations. To be of maximal value in forensics as well as in other areas of research, a panel of loci needs a broad set of reference population frequencies. To that end, we have now assembled and analyzed sequence-based data on 4009 individuals in 79 populations for these 90 microhaps. These results also demonstrate the value of microhaplotypes for biomedical and anthropologic studies of human populations.
Materials and methods
Population samples
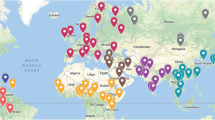
The 4009 individuals in 79 populations studied (Table 1) include 524 individuals in 16 populations that we have typed by MPS (Table 2). The DNA for the individuals sequenced was purified using phenol–chloroform from lymphoblastic cell lines that are part of the Kidd Lab collection. Greater detail of the population samples can be found in ALFRED (alfred.med.yale.edu). Comparable data for 3485 other individuals in 63 populations that were curated from public repositories: the Human Genome Diversity Project (HGDP) which includes individuals sequenced from the Kidd Lab collection of world population samples (see Table 2 and Bergstrom et al. 2020); the Genome Asia database (Genome Asia100 K Consortium 2019); and the 1000 Genomes (1000 Genomes Consortium Project et al. 2015).
The 536 sequenced individuals included 155 individuals sequenced previously (Gandotra et al. 2020) and 381 individuals that were sequenced and passed quality control steps in this study (Table 2). Twelve individuals were also successfully sequenced from other groups including 4 samples from Southern Tunisia in this study and 6 Euro-Americans and 2 Chinese from Taiwan in Gandotra et al. (2020). These were excluded from statistical analyses, because the sample sizes were too small. They will be included in future studies as more sequenced samples accumulate. The data from all samples sequenced are available on the Scharfe lab mMHseq website (see Data Availability).
Data collection
The descriptions of the 90 microhaplotype loci and the primers for MPS are described in Gandotra et al. (2020) (cf. Table S2 in that paper) as are the detailed mMHseq methods. Table 3 provides an overview of key characteristics of the 90 microhaps. The mMHseq libraries of 48 individuals and two non-template water controls were sequenced in a single Illumina MiSeq run for all 90 microhaplotypes. This number of samples per run assures that each sample receives sufficient sequence read coverage based on the assay’s empirically established performance parameters. Data analysis included sample demultiplexing, primer trimming, read alignment to the human reference genome (hg19/GRCh37), data quality control (QC), DNA variant calling (GATK UnifiedGenotyper (GATK UG), and SNP phasing for each microhaplotype using Read backed phasing tools from GATK to phase the SNP’s along the microhaplotype (McKenna et al. 2010). Following identification of variants at each of the 90 MH loci in the 536 individuals using mMHseq, base calls at the same variant sites were extracted for 3485 individuals from various whole-genome sequencing (WGS) repositories.
Data analyses
Effective number of alleles (Ae) is a measure that standardizes the information among diverse populations for their different frequencies among the multiple alleles (Kimura and Crow 1964; Kidd and Speed 2015). Ae for a locus is calculated as the inverse of homozygosity, Ae = 1/sum(pi2). As such, it is the number of equally frequent alleles that would yield the same heterozygosity as the observed set of alleles with diverse frequencies. This measure is good for evaluating multiallelic loci (such as microhaplotypes) for individualization and mixture analysis. Informativeness (In) for measuring allele frequency differences among populations was calculated according to Rosenberg et al. (2003). This measure is appropriate for evaluating loci for their ability to infer population ancestry of an individual and relationships among populations.
For the extracted data that were not phased in the respective repositories, the haplotypes were inferred using PHASE version 2.1.1 (Stephens et al. 2001; Stephens and Scheet 2005). For all of the QC passed samples, the phasing was obtained directly from the reads for each of the MH loci.
Structure, PCA, and population trees
To help visualize clustering of individuals in populations, we used version 2.3.4 of the STRUCTURE software (Pritchard et al. 2000). The program was run under the standard admixture model assuming correlated allele frequencies. The input data consisted of the microhaplotype genotype profiles for all individuals in the 79 populations. The program was run 20 times at each K level from K = 2 to K = 16 with 10,000 burn-in and 10,000 Markov Chain Monte Carlo (MCMC) iterations. The result with the highest likelihood of the 20 runs was selected to illustrate the results for a given K value.
To help visualize clustering of populations, we used Principal Component Analyses (PCA). We used the XLSTAT 2019 software (http://www.xlstat.com/en/about-us/company.html) on the matrix of haplotype allele frequencies for all 90 microhaplotype loci in the populations relevant to each analysis.
We also conducted tree analyses for the 79 populations using pairwise Tau genetic distances (Kidd and Cavalli-Sforza 1974) and methods and logic described in Kidd and Sgaramella-Zonta (1971) and Cherni et al. (2016). Analyses started with the Neighbor Joining tree (Saitou and Nei 1987), which gives an approximate Least Squares fit, and then explored similar tree structures by an exact Least Squares fit to the defining set of linear equations. The Neighbor Joining (NJ) program employed is part of the PHYLIP software package (Felsenstein 1989,2009). The Drawtree utility (version 3.69) in the PHYLIP package was used to plot the postscript images of the best population trees.
Results
mMHseq data analysis and quality control
Assay performance was assessed using our algorithms for monitoring sequence read coverage on three levels: samples, amplicons (loci), and sequence bases (Fig. S1, Table S2). Any sample that failed this QC was removed from further analysis. The first QC metric (sample coverage), defined as the number of reads per sample, was used for detecting samples that failed in the multiplex PCR. An average read depth across 384 samples was 705,536 reads per sample. Eight out of 384 samples had lower read depth coverage of less than 150,000 reads and were flagged for further analysis of amplicon and base coverage (Table S2 and Fig. S1). The second QC metric (amplicon coverage) was used to identify samples with partially failed amplification, such as individual amplicons that may have been insufficiently covered despite an overall normal read count for that sample. For each sample, we obtained the number of amplicons that had > 0.2-fold the mean amplicon coverage and used a threshold of 2 standard deviations below the mean to flag samples for review. This metric identified 4 samples with poor amplicon uniformity (Table S2 and Fig. S1). The third QC metric (base pairs) assessed base coverage for each sample, reasoning that if base coverage was sufficiently high, even samples with lower amplicon uniformity could be analyzed further. Five samples had a lower base coverage (< 75% of bases with 100 × reads per nucleotide per amplicon). Three samples failed QC at all three levels and were removed from the analysis, while the other samples flagged in one of the three QC steps yielded interpretable results in sequence analysis. Thus, final analyses are based on data for 381 individuals (Tables 2 and S2). Additionally, we investigated the data for MH genotypes that could have been due to allele dropouts. We found 4 MH alleles that were present only as homozygous MH genotypes in a single individual (but in different sequenced individuals for each allele type) and the inferred two alleles were the only occurrences of those alleles in the whole dataset; so, these genotypes were removed from the analyses.
We estimate that each genotype call was based, on average, by 7067 reads. That number is the average of the sequencing reads per locus (amplicon) in the last five sequencing runs, each of which involved sequencing of 48 individuals. Thus, sequencing of a total of 240 individuals contributed to this number. These are the right-most 62% of the reads in supplemental Fig. S1. Some variation in read numbers occurred among the five runs considered, but the variation in reads per locus was consistent; the distribution of the number of loci by the number of reads is given in Fig. S2. We note that except for 13 loci, there were more than 500 reads per allele per locus per individual. Only one locus, mh01KK-001, averaged fewer than 100 reads per allele with 75.3 reads per allele. In general, coverage per locus exceeds the clinical exome sequencing standard of 80×. It is unclear whether the differences in reads per locus per individual are inherent to the locus or are inherent just to the sequence or concentration of the specific primer pair used for the sequencing. A future effort will be made to better balance across loci to assure a higher minimum number of reads for all loci.
In summary, the mMHseq 90-plex data for the sequenced individuals from 16 populations are available at the Scharfe lab mMHseq website and have also been deposited in the Zenodo archive (see Data Availability). Our previous study (Gandotra et al. 2020) identified 717 SNPs in the 90 MHs for 30 populations, while this study of 79 populations recorded 905 SNPs in the 90 MHs (Table S1), which included 65 novel SNPs in 39 of the 90 MHs.
Characteristics of MH markers
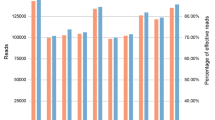
As noted earlier, two statistics characterize the information in the markers with respect to variation within populations (Ae) and variation among populations (In). Figure 1 is a scatterplot of all 90 MHs according to In by average Ae for the total of 79 populations. Some of the markers rank very high by both criteria. The six MHs that are highest for Ae are shaded and included in Table 4. The clinal decrease in the average Ae across loci for populations that are farther from Africa is evident in Fig. 2. The markers have high heterozygosity with mean values of Ae ranging from 3.0 to more than 6.0 (Fig. 2) depending on the population. Among the 7110 individual population values (79 × 90) for Ae, it is noteworthy that 81.7% are ≥ 3.0 and 96.8% are ≥ 2.0. Supplemental Fig. S3 plots the average Ae value for each of the 90 microhaplotypes. The most common genotype frequency in each population is also plotted in Fig. 3. Note that the specific genotype will likely be different in each population, the point being that no genotype is common anywhere when all 90 loci are considered.
Scatterplot of 90 microhaplotypes by In and average Ae for 79 populations (79p). 6 MHs with highest Ae values in all 6 biogeograhic regions (cf Table 4) are shown as patterned circles
The high Ae for many loci individually and on average across all populations indicates considerable variation within populations. A forensic measure, Random Match Probability (RMP), at a single locus is the sum over all the possible genotypes in the population of the squares of the genotype frequencies. In other words, it is the expected frequency (probability) for the population of, having randomly selected one individual, another unrelated individual will have that same specific genotype. For multi-locus genotypes, RMP becomes the product of the individual locus probabilities. It is often used in criminal cases to note how unlikely it is that someone else has the same genotype as a defendant. The RMP values are quite small for these 90 MH loci. However, RMP is population-specific and has a dramatic difference of 50 orders of magnitude depending on the population (Fig. 3). The range goes from the very small RMP values for Africans up to the much larger, but still highly probative, values for the Pacific and Native American populations: 10–115 up to 10–66. Globally, the probability of two unrelated individuals having the same genotype for these markers is vanishingly small. Note, this RMP is not the same as the probability that a random person will have the same genotype as a specific evidentiary genotype profile.
Informativeness (In) across the 79 population samples likewise shows considerable variation by locus (Fig. 4). The specific loci with the highest In values are clearly distinct in Fig. 1 as are those loci with the lowest In values.
Rosenberg informativeness (In) across 79 populations for each of 90 microhaplotypes. The 6 dark triangles correspond to the 6 MH with the highest Ae values in Fig. 1
Inference of population relationships
Structure
STRUCTURE analyses were run on all 79 population samples from K = 2 through K = 16. The first K value at which all major biogeographic regions are distinct is K = 6 (Fig. 5, Fig. S4). Those six clusters are the ones that correspond to “continental” clusters when representatives of all “continents” are present: Sub-Sahara Africa; North Africa, Southwest Asia, and Europe; South Central Asia; East Asia; the Pacific; and the Americas. These six are the commonly seen clusters from many studies based on SNPs (Soundararajan et al. 2016; Li et al. 2016; Cherni et al. 2016; Santos et al. 2016; Fondevila et al. 2017; Pakstis et al. 2017; Pakstis et al. 2019;Xavier et al. 2020), on studies of microhaplotypes (Kidd et al. 2017,2018a; Bulbul et al. 2018; Gandotra et al. 2020; Puente et al. 2020b; Staadig and Tillmar 2021), and on studies combining single SNPs and MHs (Phillips et al. 2019; Kidd et al. 2021). K = 6 provides a convenient basis for summarizing aspects of the data such as the MHs with the highest regional Ae values. K = 6 is also the point at which the likelihood increases with increasing K values begin to be progressively smaller until the curve is nearly flat at K = 14 to K = 16 (Fig. S5). K = 7 shows that these loci can begin to distinguish among the sub-Saharan Africans. Yet, when all 79 populations were analyzed up to K = 16, the African clustering looks identical to the K = 7 pattern (Fig. S6). In contrast, the East Asia pattern became much more complex at K = 16. This panel of 90 loci is capable of more refined STRUCTURE clustering when subsets are analyzed separately. When the 21 African and Southwest Asia populations were analyzed as a group, K = 6 showed five clusters within sub-Saharan Africa (Fig. 6) distinct from the Southwest Asians. When the 21 Siberian, East Asian, and Pacific populations were analyzed as a group, K = 7 showed the clearest set of clusters (Fig. 7).
PCA
The African populations are a distinct group and their distinctiveness is the primary driver of PC1 when all 79 populations are analyzed (Suppl. Fig. S7). All other populations are primarily distributed according to PC2. To separate those non-African populations better, a separate analysis was done omitting all of the sub-Saharan populations (Fig. 8). This analysis clusters the European and SW Asia populations close together at one end of PC#1 followed by the South Central Asian populations with an internal differentiation along a West-to-East axis. The Native Americans form a distinct cluster as do the East Asians. The Oceania populations form a loose cluster next to the tight East Asian cluster. The two North Asian populations (BUR and YAK) are very close together but far from the Western Siberian Khanty (KTY) which is not part of any cluster. Similarly, the Hazara (HZR) is a distinct population.
Tree analysis
The tree analysis of Tau genetic distances on all 79 populations involved evaluations of a total of 294 different additive tree structures of which 31 had no internal negative segments. The best of these 31 is shown in Suppl. Fig. S8. There are two small negative segments connecting the two mostly West African populations (ACB and ASW) to the African branch of the tree. This is an indication that these do not conform to the underlying assumption of an additive tree for which only random genetic drift has caused divergence of populations. Indeed, these two populations are admixed and do not meet the assumptions, but were included as part of the 1 KG set of populations.
In general, many of the clusters of populations are similar to those seen in the STRUCTURE and PCA analyses. The South Asians are divided into four different clusters in the tree. One is close to the European and SW Asia cluster; the others are more differentiated.
Discussion
The utility/value of a locus in forensics can relate to at least four different questions: individualization, ancestry inference, kinship analysis, and mixture resolution. Individualization is often noted as the random match probability (RMP) reflecting the low likelihood that a match between evidence and an accused individual would have occurred by chance alone. Ancestry inference can be pursued as the identification of the population for which the probability of the observed genotype is highest (Kidd et al. 2018b; Rajeevan et al. 2020). The value of a panel of loci in anthropology is related to what the genetic data can tell about population relationships and histories (Kidd et al. 2021). Kinship analysis compares DNA sequence or dense markers among individuals to determine the likely degrees of relationship. Paternity testing is one form of kinship analysis. Mixture deconvolution is a developing field with probabilistic genotyping available for STR analysis but not yet for microhaplotypes. As discussed in the following sections, microhaplotypes are useful in all of these areas.
Individualization
SNPs are overwhelmingly di-allelic and hence provide less information per locus than the polymorphic STRs when comparing a forensic sample with a reference sample. High levels of individualization measured by random match probability (RMP) are a consequence of the high Ae values of the loci. Figure 3 plots the RMP by population based on all 90 microhaplotypes. Although the scales are very different, Figs. 2 and 3 show otherwise similar variation among populations, because both are based on the heterozygosities of the 90 loci in the 79 populations. Both show high Ae values in African and significantly lower values in the Pacific and Native American populations. The range of population-specific RMP values is close to 50 orders of magnitude from a minimum of 10–115 to a maximum of 10–66. Even at the maximum value, the RMP based on all 90 loci is highly probative.
There is a significant range in the average Ae values (3.00–6.25) across all 79 populations among the 90 microhaplotypes (Fig. 2). While some of the loci are at the low end of the distribution overall, a relevant question is whether or not some of the better markers exist in different regions of the world. The STRUCTURE software can show reliable clusters of populations at higher K values (Fig. S6), but K = 6 provides a convenient basis for summarizing aspects of the data such as the MHs with the highest regional Ae values. Table S3 summarizes the top 20 MHs ranked by Ae value for each of the six biogeographic regions defined in Fig. 5. The averages of the average Ae values for the 20 highest loci are lower for the non-African regions with the smallest for the Pacific populations, but the decrease is not great compared to the overall decrease seen in Fig. 2. Overall, there are 38 different loci in this tabulation. Many of the loci have a high Ae in more than one broad region of the world. Only 6 of these 38 loci occur in all six biogeographic regions (cf. Figure 5) and are listed in Table 4. These are the highlighted loci in Fig. 1. The averages for those loci that rank among the top 20 are above 4.0 (See Suppl. Table S3). Many markers have good Ae values for random match probabilities and for mixture deconvolution for nearly all populations.
The large number of MH alleles varying in the six biogeographic regions are illustrated in Fig. 9. There are 3018 total different MH alleles in the dataset analyzed with 1337 occurring at common frequencies ≥ 5% in specific populations, while a total of 1810 MH alleles occur at frequencies > 2%. The remaining 1208 alleles mostly occur at very low (usually rare) frequencies; for example, 910 of the 1208 very-low-frequency mh-alleles are only counted to occur once or twice in the whole dataset. Supplemental Table S3 lists the 20 highest ranking MHs by average Ae in each of six world regions. The average MH allele frequencies in each of six major geographic regions are shown as bar plots for the microhaps, mh01KK-212 (Fig. 10) and mh05KK-170 (Fig. 11), with the highest In values (0.88 and 0.81) in 79 populations and the highest average Ae (9.708 and 9.750) in the 79 populations.
Average allele frequency bar plot for mh01KK-212 for each of 6 major biogeographic regions. This microhaplotype has the largest value for Rosenberg’s In in 79 populations (0.88; Fig. 4) and the second higheset average Ae for 79 populations (9.708; Suppl. Fig. S3). The 34 alleles with frequencies ≥ 5% in specific populations are plotted separately with different colors/patterns; the 58 alleles with frequencies < 5% are pooled (bars shown with black diagonal lines and green background)
Average allele frequency bar plot for mh05KK-170 for each of 6 major biogeographic regions. This microhaplotype has the second largest value for Rosenberg’s In in 79 populations (0.81; Fig. 4) and the highest average Ae for 79 populations (9.750; Suppl. Fig. S3). The 33 alleles with frequencies ≥ 5% in specific populations are plotted separately with different colors/patterns; the 24 alleles with frequencies < 5% are pooled (bars shown with black diagonal lines and yellow background)
Ancestry inference: population relationships
High In markers require a reference database to determine allele frequencies for calculating RMP values and for use in forensic attempts to identify the population ancestry of the donor of a DNA profile. This study provides reference data on 79 population samples. Several of those populations are smaller than ideal for forensic reference, but as seen in Fig. 5, the clusters at K = 6 and K = 7 define Mendelian populations of considerable size in some cases. It is clear that an amalgam of European population samples in one STRUCTURE cluster is as valid a reference population as a forensic reference population such as “U.S. White”.
The PCA and STRUCTURE results presented show that the extensive genetic variation in the 79 populations analyzed with the 90 MH panel can both differentiate clear population groupings for major geographical areas of the world as well as delineate distinct subgroupings of populations, especially when analyses are restricted to particular biogeographic regions.
There were no real surprises in the population relationships seen in STRUCTURE analyses and PCA. Indeed, as noted earlier, several other sets of markers on similar collections of populations have shown similar relationships (e.g., Bulbul et al. 2018) to those seen in Figs. 5 and 8. What these analyses do demonstrate is that this set of markers is highly informative for population similarities and differences at K values > 6. The new marker data do provide new information on some of the populations as discussed and also presented separately for African and East Asian populations below.
Comments of overall analyses of 79 populations using these 90 microhaplotypes
The six main clusters of populations seen in Fig. 5 and Fig. S4 remained distinct at higher K values. Figure S5 shows that likelihoods increased through K = 14 but at progressively lower increases as K increases until likelihoods remain almost constant after K = 14. What happened is that the six major regions have been subdivided at the higher K values and the “intermediate” populations (i.e., the magnified blocks in Fig. 5) with small sample sizes have differing patterns at the higher K values. In supplemental material, we present analyses at K = 16 (Fig. S6) which is a higher K value than the likelihood increases warrant, but illustrates the general pattern for subdivisions of the six major regions. For Africa, the change from K = 7 (Fig. 5) occurred at K = 13 in the 79-population analysis when the Biaka Pygmies became distinct from the East Africans. That pattern persisted through K = 16 (Fig. S6) but with the Ethiopian Jews showing differing patterns at higher K levels. The North African and Southwest Asian populations became a separate cluster from the Europeans at K = 9 and the cluster persisted through higher K values. The South-Central Asia cluster separates off the Pakistani populations with a distinct admixture component at K = 13 and that distinction remains through K = 16. Three of the South-Central Asia populations show inconsistent patterns of clustering after K = 13. In contrast to the small refinements of the African and European patterns, the East Asian patterns became more subdivided with increasing K value, as discussed below. The Oceania populations show several different patterns at the different K values.
Comments on African ancestry inference of these 90 microhaplotypes
Based on the overall analyses, we chose 21 populations for a more detailed analysis: the African and Southwest Asian samples. STRUCTURE analyses stabilized at K = 5 and K = 6 (Fig. 6). The Mozabites clustered with the SW Asian populations as a distinct group. The Ethiopian Jews were intermediate between the SW Asian and Sandawe from Tanzania. Other East African populations form a distinct cluster and the Central African Biaka population was distinct. The West African populations show some indication of two distinct groups with the Gambians and Mandenka distinct from both Yoruba samples and the Esan. This pattern of subdivision of the African cluster does not occur in the larger analyses of all 79 populations (Fig. S6). PCA of all 79 populations (Fig. S7) showed a distinct African cluster but no clear separation of Eastern vs. Western African populations. The Ethiopian Jews were distinct. PCA of the 21 populations showed that these populations generally are distributed along PC#1 (24.5%) as West Africa, East Africa, Ethiopian Jews, the Mozabites, and the SW Asian populations. PC#2 (9.1%) essentially separated the Biaka from all others (Suppl. Fig. S9a). PC#3 (8.2%) more clearly separated the East Africans and Ethiopian Jews from all the others (Suppl. Fig. S9b). PCA provided barely any evidence of clustering among the West African populations with only the Mandeka slightly different from the others. The two samples of admixed African-European origin cluster with the African populations by PCA but closer to the East Africans.
Comments on East Asia and the Pacific
The most striking result for the 79 population analysis is that at K = 11, the three samples of Han Chinese all show an “admixture” pattern with many individuals showing mixed membership in the Northeast Asia (Koreans and Japanese) cluster and the Southeast Asia (Dai, Vietnamese, and Cambodians) cluster. That pattern persisted through K = 16. If it has any meaning, it is probably that the Han Chinese are intermediate in a North-to-South cline in far East Asia and not that they are individually admixed of those flanking populations. At K = 9, the Atayal became distinct. At K = 10–16, the Khanty became distinct and usually (for K = 10 to 14) group with the Buryat and Yakut; in both cases, they remained distinct through K = 16 (Fig. S6). Oceania showed inconsistent clustering among the populations except for the consistent clustering of the two Melanesian populations together.
Similar population groupings are seen in the PCA results (Fig. S10). The Khanty from northwest Siberia is a clearly distinct population in this analysis. Note that in the full global context, it was intermediate between the Europeans and East Asians. We chose 21 population samples from Western Siberia to the Pacific omitting the South Central Asian samples that were a clearly distinct cluster in Fig. 5. STRUCTURE analysis of these 21 populations showed clear clusters at K = 7 (Fig. 7). The Buryat and Yakut samples cluster together both in the STRUCTURE analysis of the 21 samples and in the PCA of all 79 populations (Fig. 5). The Koreans and the three samples of Japanese ancestry form a clean cluster in STRUCTURE at all K levels, but are close to the Chinese in the PCA analyses. The three Chinese samples appear admixed between the Japanese and the three South East Asia populations that form a clean cluster. The STRUCTURE data constitute evidence for a North-to-South cline of genetic differentiation in Far East Asia. The Atayal sample defined its own isolated cluster in STRUCTURE at K = 9, 10, and 16 but group with the South East Asian populations from K = 11 to 15. The various Oceania populations form a noisy cluster with evidence of admixture except for the two Melanesian samples from Papua New Guinea that are distinct at all K values in analyses of both the full (79) and restricted (21) sets of populations.
A general comment
Overall, these 90 microhaplotype markers are a powerful set for population relationships, but it was impossible from these analyses to determine when a subset of populations would provide an answer not inferable from the full set of populations. The Africans, in the separate 21 population analysis, clearly show clustering at K = 5 that is not seen in any of the results for all 79 populations. In contrast, the East Asians by themselves cluster in ways that are similar (but never identical) to the clustering of all populations at K levels up to K = 16. We do not fully understand the cause in this case of the different patterns. We know that different markers are most relevant to different regions; the magnitude of the allele frequency differences is undoubtedly relevant. How well this regional inconsistency in finer clustering generalizes to other datasets is unknown at present.
Kinship
Any multiallelic genetic system is useful for kinship analysis. Indeed, even a di-allelic locus provides evidence of relationship by allele sharing. In this respect, the high Ae values of this set of MHs should be especially informative, because the probability of allele sharing identical by state can be much less than sharing identical by descent for close relatives. However, no direct test has been done. Recent papers by Puente et al. (2020a), Staadig and Tillmar (2021), and Wu et al. (2021b) have assessed microhaplotypes in kinship analyses to varying degrees. Based on (Wu et al. 2021b) with 54 high In MHs that were problematic at relationships beyond second degree, we cannot expect the 90 MHs in our study to be good at distant relationships. How good the 90 will be is for future research.
Mixture deconvolution
Three questions arise when considering the existence of mixtures in a forensic sample. First, is there a mixture? The essential proof that a mixture exists is the presence of at least three alleles at several of the loci. Note that this criterion cannot be met by a di-allelic SNP. The only way a di-allelic SNP can contribute to the inference of a mixture is if a quantitative method is used and the two alleles differ in their values, e.g., sequence read number, more than heterozygote read imbalance would explain. Second, how many contributors are there to this mixture? At any one locus, the minimum number of contributors is the number of alleles seen divided by 2 and, if a fraction, rounded to the next whole number: five alleles seen implicates 3 contributors; six alleles also implicates 3 contributors. The loci with the largest numbers of alleles seen provide an overall minimum estimate of contributors that applies to all loci. Note that sensitivity issues and diminishing concentrations with larger numbers of contributors prevents any realistic estimate of the maximum number of contributors. However, the global sum of all the alleles seen at all the loci can implicate more contributors than the maximum seen at individual loci (see Fig. 2 in Bennett et al. (2019) for an illustration). Also, quantitative variation in allele “intensity” may also provide hints at larger numbers of contributors, but some model of the relationships of numbers of copies of alleles to their intensity is required.
Finally, can the individual multi-locus genotypes of the contributors be determined? It may be possible to readily infer the contributing genotypes at a single locus using allele “intensity” (e.g., read count in MPS) as seen at locus mh05KK-170 in Bennett et al. (2019). However, the permutations of the individual locus results overwhelm such single locus approaches. This becomes an issue for probabilistic genotyping of microhaplotypes analogous to the use of STRMix (Buckleton et al. 2019) for probabilistic genotyping of forensic STR data. In the forensic case, the question is usually whether a known sample can be part of a mixture. This is a different question than attempting to fully deconvolute a mixture. This is an area that needs development for microhaplotypes because of the many variables that are involved. Elements of such deconvolution methods include the number of contributors, the relative amounts of each contributor, and the allele frequencies in the relevant population(s). The 90 MHs provide a set of highly heterozygous loci that can help with some of these issues and have the advantage of low mutation rates and the absence of stutter.
Optimizing the panel
This panel of 90 MH loci was designed to have high Ae and high In. This has resulted in loci with, on average, greater extent to encompass more SNPs. Eliminating the loci with the lowest Ae and/or In values globally should improve the efficiency of the panel. However, a careful analysis should be undertaken to assure that the lowest In marker for all populations is not providing significant differentiation of some population(s). We generated exploratory STRUCTURE runs from K = 2 to K = 8 for 79 populations after excluding 19 MH with In ≤ 0.25. The cluster patterns of the highest likelihood runs for the 71 MHs were all very similar to those obtained with all 90 MH. The most noticeable difference occurred at K = 7 where the Biaka from central Africa clustered with the West African groups instead of the East African cluster. Some of the excluded MH markers undoubtedly have value in differentiating among the sub-Saharan groups. Given the high level of informativeness of the panel for obtaining results at 90 loci, efficiency is not an issue. Rather, any pruning would allow space for adding additional marker loci with higher values, including some of the best of the loci identified by others, e.g., (Wu et al. 2021a), have identified many MHs with global average Ae values > 5.0. Those are issues for future research.
General utility of microhaplotypes
While the loci studied here are human specific and will not be relevant to other species, the general molecular approach and methods (Gandotra et al. 2020) are applicable tools in population genetic studies of other organisms. The fields of ecology and conservation are increasingly using molecular techniques and some researchers are already using microhaplotypes (Meek and Larson 2019). Microhaplotypes have been shown to be much more informative per locus than SNPs in studying the familial relationships among Kelp Rockfish (Baetscher et al. 2018). Microhaplotypes have also been used to study porpoises (Morin et al. 2021) and salmon (Larson et al. 2016; McKinney et al. 2017). Tessema et al. (2020) identified 93 microhaplotypes in Plasmodium falciparum. Those P. falciparum microhaplotypes had a median Ae of 3.33 and provided good discrimination between related and unrelated polyclonal infections.
Impact on forensic practice
In spite of their technical advantages over the forensic STR markers, SNPs have not been incorporated in routine forensic practice. Part of the reason has been the need for separate methodologies to type STR loci and SNPs. With the advent of MPS, it is now possible to use one technology and multiplex the standard STR markers with a reasonable panel of SNP-based markers in the same sequencing run. We show in this study that microhaplotypes with high Ae, rivaling the Ae values for STR markers, can be found and are far superior to individual SNPs. We believe that such microhaplotypes will supplant individual SNPs in future applications. As more laboratories acquire sequencing technology, it may be possible for microhaplotypes to become a tool in forensic practice while maintaining the standard STR markers and the national databases of convicted felons. However, the costs of new equipment and training of personnel and the absence of an agreed upon panel of highly informative microhaplotypes remain major obstacles.
Future studies
Refining and optimizing the microhaplotype markers that have already been identified for more localized geographic regions will likely be productive. Identifying additional useful microhaplotypes would be helpful. Some may emerge as more diverse human populations are studied routinely. While we have studied 79 populations from major geographical regions of the world, there is still a need to obtain better coverage of the diversity of human populations, especially in Africa, North Asia, Southeast Asia, and the Americas. Recent reviews and population genetic studies (Ramsay et al. 2021), for example, continue to indicate that the diversity of African populations is greater than what has been routinely studied. Indigenous populations of the Americas (Moreno-Estrada et al. 2014; Homburger et al. 2015; Barbieri et al. 2019) also need better coverage.
Conclusions
Our results document this panel of microhaplotype markers as the best one so far with highest overall values of Ae and In in the largest number of populations studied. The combination of multiplex mMHseq) and the expanded set of populations studied from around the world revealed a highly informative set of markers that has characteristics that can serve a range of forensic, medical, and anthropological applications. Additional useful microhaplotypes will likely emerge from other and future studies (e.g., Wu et al. 2021a). New analyses can focus on tailoring the best subsets and supersets of MH markers for use in specific geographical regions as well as for major world regions. As more extensive sampling and analyses of world populations occur, it can be expected that the ability to distinguish more refined population relationships in multiple world regions will increase, especially in Africa.
Data availability
Genotype profiles on 90 MHs for 556 individuals in 16 Kidd lab population samples (including the 524 sequenced Kidd lab individuals and 32 individuals from HGDP studies of the same population samples) have been deposited in the Zenodo archive and can be freely accessed at https://doi.org/10.5281/zenodo.5095364. Data for the additional individuals included in the analyses were taken from public datasets as indicated in the text. The mMHseq 90-plex data for 524 sequenced individuals from 16 Kidd lab population samples are available at the Scharfe lab website, https://mmhseq.shinyapps.io/mMHseq.
References
1000 Genomes Consortium Project, Auton A, Brooks LD, Durbin RM, Garrison EP, Kang HM, Korbel JO, Marchini JL, McCarthy S, McVean GA, Abecasis GR (2015) A global reference for human genetic variation. Nature 526(7571):68–74
Baetscher DS, Clemento AJ, Ng TC, Anderson EC, Garza JC (2018) Microhaplotypes provide increased power from short-read DNA sequences for relationship inference. Mol Ecol Resour 18:296–305. https://doi.org/10.1111/1755-0998.12737
Barbieri C, Barquera R, Arias L, Sandoval JR et al (2019) The current genomic landscape of western South America: Andes, Amazonia, and Pacific coast. Mol Biol Evol 6:2698–2713. https://doi.org/10.1093/molbev/msz174
Bennett L, Oldoni F, Long K, Cisana S, Madella K, Wootton S, Chang J, Hasegawa R, Lagace R, Kidd KK, Podini D (2019) Mixture deconvolution by massively parallel sequencing of microhaplotypes. Int J Legal Med 133:719–729
Bergstrom A, McCarthy SA, Hui R, Almarri MA, Ayub Q, Danecek P, Chen Y, Felkel S, Hallast P et al (2020) Insights into human genetic variation and population history from 929 diverse genomes. Science 367:5012. https://doi.org/10.1126/science.aay5012
Buckleton JS, Bright JA, Gittelson S, Moretti TR, Onorato AJ, Bieber FR, Budowle B, Taylor DA (2019) The probabilistic genotyping software STRmix: utility and evidence for its validity. J Forensic Sci 64:393–405. https://doi.org/10.1111/1556-4029.13898
Budowle B, Moretti TR, Niezgoda SJ, Brown BL (1998) CODIS and PCR-based short tandem repeat loci: law enforcement tools. In: Second European symposium on human identification, Promega Corporation, Madison
Bulbul O, Pakstis AJ, Soundararajan U, Gurkan C, Brissenden JE, Roscoe JM, Evsanaa B, Togtokh A, Paschou P, Grigorenko EL, Gurwitz D, Wootton S, Lagace R, Chang J, Speed WC, Kidd KK (2018) Ancestry inference of 96 population samples using microhaplotypes. Int J Legal Med 132:703–711
Butler JM, Hill CR (2012) Biology and genetics of new autosomal STR loci useful for forensic DNA analysis. Forensic Sci Rev 24(1):15–26
Chen P, Yin C, Li Z, Pu Y, Yu Y, Zhao P, Chen D, Liang W, Zhang L, Chen F (2018) Evaluation of the microhaplotypes panel for DNA mixture analyses. Forensic Sci Int Genet 35:149–155. https://doi.org/10.1016/j.fsigen.2018.05.003
Cherni L, Pakstis AJ, Boussetta S, Elkamel S, Frigi S, Khodjet-El-Khil H, Barton A, Haigh E, Speed WC, BenAmmarElgaaied A, Kidd JR, Kidd KK (2016) Genetic variation in Tunisia in the context of human diversity worldwide. Am J Phys Anthropol 161:62–71
Cheung EYY, Phillips C, Eduardoff M, Lareu MV, McNevin D (2019) Performance of ancestry-informative SNP and microhaplotype markers. Forensic Sci Int Genet 43:102141. https://doi.org/10.1016/j.fsigen.2019.102141
de la Puente M, Phillips C, Xavier C, Amigo J, Carracedo A, Parson W, Lareu MV (2020a) Building a custom large-scale panel of novel microhaplotypes for forensic identification using MiSeq and Ion S5 massively parallel sequencing systems. Forensic Sci Int Genet 45:102213. https://doi.org/10.1016/j.fsigen.2019.102213
de la Puente M, Ruiz-Ramirez J, Ambroa-Conde A, Xavier C, Amigo J, Casares de Cal MA, Gomez-Tato A, Carracedo A, Parson W, Phillips C, Lareu MV (2020b) Broadening the applicability of a custom multi-platform panel of microhaplotypes: bio-geographical ancestry inference and expanded reference data. Front Genet 11:581041. https://doi.org/10.3389/fgene.2020.581041
Felsenstein J (1989) PHYLIP-phylogeny inference package (Version 3.2). Cladistics 5:164–166
Felsenstein J (2009) PHYLIP (Phylogeny Inference Package) version 3.7a. Distributed by the author. Department of Genome Sciences, University of Washington, Seattle. https://evolution.genetics.washington.edu/phylip.html
Fondevila M, Børsting C, Phillips C, de la Puente M, Carracedo A, Morling N, Lareu MV, Consortium EN (2017) Forensic SNP genotyping with SNaPshot: technical considerations for the development and optimization of multiplexed SNP assays. Forensic Sci Rev 29:57–76
Gandotra N, Speed WC, Qin W, Tang Y, Pakstis AJ, Kidd KK, Scharfe C (2020) Validation of novel forensic DNA markers using multiplex microhaplotype sequencing. Forensic Sci Int Genet. https://doi.org/10.1016/j.fsigen.2020.102275
Genome Asia100 K Consortium (2019) The GenomeAsia 100K Project enables genetic discoveries across Asia. Nature 576:106–111. https://doi.org/10.1038/s41586-019-1793-z
Guo F, Shen H, Tian H, Jin P, Jiang X (2014) Development of a 24-locus multiplex system to incorporate the core loci in the Combined DNA Index System (CODIS) and the European Standard Set (ESS). Forensic Sci Int Genet 8:44–54. https://doi.org/10.1016/j.fsigen.2013.07.007
Homburger JR, Morenao-Estrada A, Gignoux CR et al (2015) Genomic insights into the ancestry and demographic history of South America. PLoS Genet 11(12):e1005602. https://doi.org/10.1371/journal.pgen.1005602
Kidd KK, Cavalli-Sforza LL (1974) The role of genetic drift in the differentiation of Icelandic and Norwegian cattle. Evolution 28(3):381–395
Kidd KK, Sgaramella-Zonta LA (1971) Phylogenetic analysis: concepts and methods. Am J Hum Genet 23(3):235–252
Kidd KK, Speed WC (2015) Criteria for selecting microhaplotypes: mixtures and deconvolution. Invest Genet 6:1
Kidd KK, Pakstis AJ, Speed WC, Lagace R, Chang J, Wootton S, Ihuegbu N (2013) Microhaplotype loci are a powerful new type of forensic marker. Forensic Sci Int Genet Suppl Series 4:e123–e124
Kidd KK, Pakstis AJ, Speed WC, Lagace R, Chang J, Wootton S, Haigh E, Kidd JR (2014) Current sequencing technology makes microhaplotypes a powerful new type of genetic marker for forensics. Forensic Sci Int Genet 12:215–224
Kidd KK, Speed WC, Pakstis AJ, Podini DS, Lagace R, Chang J, Wootton S, Haigh E, Soundararajan U (2017) Evaluating 130 microhaplotypes across a global set of 83 populations. Forensic Sci Int Genet 29:29–37
Kidd KK, Pakstis AJ, Speed WC, Lagace R, Wootton S, Chang J (2018a) Selecting microhaplotypes optimized for different purposes. Electrophoresis 39:2815–2823
Kidd KK, Soundararajan U, Rajeevan H, Pakstis AJ, Moore KN, Ropero-Miller JD (2018b) The redesigned forensic Research/Reference on genetics-knowledge base, FROG-Kb. Forensic Sci Int Genet 33:33–37
Kidd KK, Bulbul O, Gurkan C, Dogan M, Dogan S, Neophytou PI, Cherni L, Gurwitz D, Speed WC, Murtha M, Kidd JR, Pakstis AJ (2021) Genetic relationships of Southwest Asian and Mediterranean populations. Forensic Sci Int Genet. https://doi.org/10.1016/j.fsigen.2021.102528
Kimura M, Crow JF (1964) The number of alleles that can be maintained in a finite population. Genetics 49:725–738
Kureshi A, Li J, Wen D, Sun S, Yang Z, Zha L (2020) Construction and forensic application of 20 highly polymorphic microhaplotypes. R Soc Open Sci 7(5):191937. https://doi.org/10.1098/rsos.191937
Lamason RL, Mohideen MPK, Mest JR, Wong AC, Norton HL, Aros MC, Jurynec MJ et al (2005) SLC24A5, a putative cation exchanger, affects pigmentation in zebrafish and humans. Science 310(5755):1782–1786. https://doi.org/10.1126/science.1116238
Larson WA, Limborg MT, McKinney GJ, Schindler DE, Seeb JE, Seeb LW (2016) Genomic islands of divergence linked to ecotypic variation in sockeye salmon. Mol Ecol 26:554–570. https://doi.org/10.1111/mec.13933
Li C-X, Pakstis AJ, Jiang L, Wei Y-L, Sun Q-F, Wu H, Bulbul O, Wang P, Kang L-L, Kidd JR, Kidd KK (2016) A panel of 74 AISNPs: improved ancestry inference within Eastern Asia. Forensic Sci Int Genet 23:101–110
McKenna A, Hanna M, Banks E, Sivachenko A, Cibulskis K, Kernytsky A, Garimella K, Altshuler D, Gabriel S, Daly M, DePristo MA (2010) The genome analysis toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res 20:1297–1303
McKinney GJ, Seeb JE, Seeb LW (2017) Managing mixed-stock fisheries: genotyping multi-SNP haplotypes increases powerfor genetic stock identification. Can J Fish Aquat Sci 74:429–434
Meek MH, Larson WA (2019) The future is now: amplicon sequencing and sequence capture usher in the conservation genomics era. Mol Ecol Resour 19:795–803. https://doi.org/10.1111/1755-0998.12998
Moreno-Estrada A, Gignoux CR, Fernández-López JC, Zakharia F, Sikora M, Contreras AV et al (2014) The genetics of Mexico recapitulates Native American substructure and affects biomedical traits. Science 344:1280–1285. https://doi.org/10.1126/science.1251688
Morin PA, Forester BR, Forney KA, Crossman CA, Hancock-Hanser BL, Robertson KM, Barrett-Lennard LG, Baird RW, Calambokidis J, Gearin P, Hanson MB, Schumacher C, Harkins T, Fontaine MC, Taylor BL, Parsons KM (2021) Population structure in a continuously distributed coastal marine species, the harbor porpoise, based on microhaplotypes derived from poor-quality samples. Mol Ecol 30:1457–1476. https://doi.org/10.1111/mec.15827
Novroski NMM, Wendt FR, Woerner AE, Bus MM, Coble M, Budowle B (2019) Expanding beyond the current core STR loci: an exploration of 73 STR markers with increased diversity for enhanced DNA mixture deconvolution. Forensic Sci Int Genet 38:121–129. https://doi.org/10.1016/j.fsigen.2018.10.013
Oldoni F, Kidd KK, Podini D (2019) Microhaplotypes in forensic genetics. Forensic Sci Int Genet 38:54–69. https://doi.org/10.1016/j.fsigen.2018.09.009
Pakstis AJ, Speed WC, Kidd JR, Kidd KK (2007) Candidate SNPs for a universal individual identification panel. Hum Genet 121:305–317
Pakstis AJ, Kang L, Liu L, Zhang Z, Jin T, Grigorenko EL, Wendt FR, Budowle B, Hadi S, AlQahtani MS, Morling N, Mogensen HS, Themudo GE, Soundararajan U, Rajeevan H, Kidd JR, Kidd KK (2017) Increasing the reference populations for the 55 AISNP panel: the need and benefits. Int J Legal Med 131:913–917
Pakstis AJ, Gurkan C, Dogan M, Balkaya HE, Dogan S, Neophytou PI, Cherni L, Boussetta S, Khodjet-El-Khil H, Ben Ammar ElGaaied A, Salvo NM, Janssen K, Olsen GH, Hadi S, Almohammed EK, Pereira V, Truelsen DM, Bulbul O, Soundararajan U, Rajeevan H, Kidd JR, Kidd KK (2019) Genetic relationships of European, Mediterranean, and SW Asian populations using a panel of 55 AISNPs. Eur J Hum Genet 27:1885–1893. https://doi.org/10.1038/s41431-019-0466-6
Phillips C, Salas A, Sánchez JJ, Fondevila M, Gómez-Tato A, Alvarez-Dios J, Calaza M, de Cal MC, Ballard D, Lareu MV, Carracedo A, SNPforID Consortium (2007) Inferring ancestral origin using a single multiplex assay of ancestry-informative marker SNPs. Forensic Sci Int Genet 1:273–280. https://doi.org/10.1016/j.fsigen.2007.06.008
Phillips C, McNevin D, Kidd KK, Lagace R, Wootton S, de la Puente M, Freire-Aradas A, Mosquera-Miguel A, Eduardoff M, Gross TE, Dagostino L, Power D, Olsen S, Hashiyada D, Oz C, Parson W, Schneider PM, Lareu MV, Daniel R (2019) MAPlex—a massively parallel sequencing ancestry analysis multiplex for Asia-Pacific populations. Forensic Sci Int Genet 42:213–226
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155(2):945–959
Rajeevan H, Soundararajan U, Pakstis AJ, Kidd KK (2020) FrogAncestryCalc: a standalone batch likelihood computation tool for ancestry inference panels catalogued in FROG-kb. Forensic Sci Int Genet. https://doi.org/10.1016/j.fsigen.2020.102237
Ramsay M, Schlebush C, Davies K (2021) Evolutionary genomics in Africa. Hum Mol Genet. https://doi.org/10.1093/hmg/ddab030
Rosenberg NA, Li LM, Ward R, Pritchard JK (2003) Informativeness of genetic markers for inference of ancestry. Am J Hum Genet 73(6):1402–1422
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4(4):406–425. https://doi.org/10.1093/oxfordjournals.molbev.a040454
Sanchez JJ, Phillips C, Børsting C, Balogh K, Bogus M, Fondevila M, Harrison CD, Musgrave-Brown E, Salas A, Syndercombe-Court D, Schneider PM, Carracedo A, Morling N (2006) A multiplex assay with 52 single nucleotide polymorphisms for human identification. Electrophoresis 27:1713–1724. https://doi.org/10.1002/elps.200500671
Santos C, Phillips C, Fondevila M, Daniel R, van Oorschot RAH, Burchard EG, Schanfield MS, Souto L, Uacyisrael J, Via M, Carracedo Á, Lareu MV (2016) Pacifiplex: an ancestry-informative SNP panel centred on Australia and the Pacific region. Forensic Sci Int Genet 20:71–80. https://doi.org/10.1016/j.fsigen.2015.10.003
Schumm JW, Gutierrez-Mateo C, Tan E, Selden R (2013) A 27-locus STR assay to meet all United States and European law enforcement agency standards. J Forensic Sci 58:1584–1592. https://doi.org/10.1111/1556-4029.12214
Shriver MD, Kennedy GC, Parra EJ, Lawson HA, Sonpar V, Huang J, Akey JM, Jones KW (2004) The genomic distribution of population substructure in four populations using 8,525 autosomal SNPs. Hum Genomics 1(4):274–286. https://doi.org/10.1186/1479-7364-1-4-274
Soundararajan U, Yun L, Shi M, Kidd KK (2016) Minimal SNP overlap among multiple panels of ancestry informative markers argues for more international collaboration. Forensic Sci Int: Genet 23:25–32
Staadig A, Tillmar A (2021) Evaluation of microhaplotypes in forensic kinship analysis from a Swedish population perspective. Int J Legal Med 135:1151–1160. https://doi.org/10.1007/s00414-021-02509-y
Stephens M, Scheet P (2005) Accounting for decay of linkage disequilibrium in haplotype inference and missing-data imputation. Am J Hum Genet 76:449–462
Stephens M, Smith NJ, Donnelly P (2001) A new statistical method for haplotype reconstruction from population data. Am J Hum Genet 68(4):978–989
Tessema SK, Hathaway NJ, Teyssier NB, Murphy M, Chen A, Aydemir O, Duarte EM, Simone W, Colborn J, Saute F, Crawford E, Aide P, Bailey JA, Greenhouse B (2020) Sensitive, highly multiplexed sequencing of microhaplotypes from the Plasmodium falciparum heterozygome. J Infect Dis. https://doi.org/10.1093/infdis/jiaa527
Tishkoff SA, Kidd KK (2004) Implications of biogeography of human populations for ‘race’ and medicine. Nature Genet 36(11 Suppl):S21–S27. https://doi.org/10.1038/ng1438
Turchi C, Melchionda F, Pesaresi M, Tagliabracci A (2019) Evaluation of a microhaplotypes panel for forensic genetics using massive parallel sequencing technology. Forensic Sci Int Genet 41:120–127. https://doi.org/10.1016/j.fsigen.2019.04.009
Walsh S et al (2011) IrisPlex: a sensitive DNA tool for accurate prediction of blue and brown eye colour in the absence of ancestry information. Forensic Sci Int Genet 5:170–180
Walsh S et al (2013) The HIrisPlex system for simultaneous prediction of hair and eye colour from DNA. Forensic Sci Int Genet 7:98–115
Wu R, Li H, Li R, Peng D, Wang N, Shen X, Sun H (2021a) Identification and sequencing of 59 highly polymorphic microhaplotypes for analysis of DNA mixtures. Int J Legal Med. https://doi.org/10.1007/s00414-020-02483-x
Wu R, Chen H, Li R, Zang Y, Shen X, Hao B, Wang Q, Sun H (2021b) Pairwise kinship testing with microhaplotypes: can advancements be made in kinship inference with these markers? Forensic Sci Int 325:110875. https://doi.org/10.1016/j.forsciint.2021.110875
Xavier C, de la Puente M, Mosquera-Miguel A, Freire-Aradas A, Kalamara V, Vidaki A, Gross TE, Revoir A, Pośpiech E, Kartasińska E, Spólnicka M, Branicki W, Ames CE, Schneider PM, Hohoff C, Kayser M, Phillips C, Parson W, VISAGE Consortium (2020) Development and validation of the VISAGE ampliSeq basic tool to predict appearance and ancestry from DNA. Forensic Sci Int Genet. https://doi.org/10.1016/j.fsigen.2020.102336
Acknowledgements
The authors thank Dr. Francoise R. Friedlaender for her expert help in formatting and labeling the STRUCTURE bar plots. Special thanks go to the many hundreds of individuals who volunteered to give blood or saliva samples for studies of gene frequency variation and to the many colleagues who helped collect the samples. In addition, some of the cell lines were obtained from the National Laboratory for the Genetics of Israeli Populations at Tel Aviv University.
Funding
This work was funded in part by National Institute of Justice grant 2018-75-CX-0041 awarded to KKK by the National Institute of Justice, Office of Justice Programs of the United States Department of Justice and by National Institutes of Health grant R01 HD102537 to CS. Points of view in this presentation are those of the authors and do not necessarily represent the official position or policies of the U.S. Department of Justice.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interests
None.
Informed consent
All samples collected by the authors were collected with individual informed consent and for use in population studies such as this. All samples are anonymous. Only anonymous, pre-existing DNA samples were used in this study; no human subjects were involved. The many samples were collected under Yale protocol (HIC#8711001387) also reviewed and approved by the National Institute of General Medical Sciences (NIGMS) in the U.S. and by the Center for the Study of Human Polymorphisms (CEPH) in Paris. One-third of the samples in the CEPH-HGDP collection came from our collection as well.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Pakstis, A.J., Gandotra, N., Speed, W.C. et al. The population genetics characteristics of a 90 locus panel of microhaplotypes. Hum Genet 140, 1753–1773 (2021). https://doi.org/10.1007/s00439-021-02382-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00439-021-02382-0