Abstract
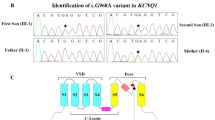
Next-generation sequencing platforms are being increasingly applied in clinical genetic settings for evaluation of families with suspected heritable disease. These platforms potentially improve the diagnostic yield beyond that of disease-specific targeted gene panels, but also increase the number of rare or novel genetic variants that may confound precise diagnostics. Here, we describe a functional testing approach used to interpret the results of whole exome sequencing (WES) in a family presenting with syncope and sudden death. One individual had a prolonged QT interval on electrocardiogram (ECG) and carried a diagnosis of long QT syndrome (LQTS), but a second individual did not meet criteria for LQTS. Filtering WES results for uncommon variants with arrhythmia association identified four for further analyses. In silico analyses indicated that two of these variants, KCNH2 p.(Cys555Arg) and KCNQ1 p.(Arg293Cys), were likely to be causal in this family’s LQTS. We subsequently performed functional characterization of these variants in a heterologous expression system. The expression of KCNQ1-Arg293Cys did not show a deleterious phenotype but KCNH2-Cys555Arg demonstrated a loss-of-function phenotype that was partially dominant. Our stepwise approach identified a precise genetic etiology in this family, which resulted in the establishment of a LQTS diagnosis in the second individual as well as an additional asymptomatic family member, enabling personalized clinical management. Given its ability to aid in the diagnosis, the application of functional characterization should be considered as a value adjunct to in silico analyses of WES.






Similar content being viewed by others
Availability of data and materials
Data is available upon reasonable request to the authors.
References
Adler A, Novelli V, Amin AS, Abiusi E, Care M, Nannenberg EA, Feilotter H, Amenta S, Mazza D, Bikker H, Sturm AC, Garcia J, Ackerman MJ, Hershberger RE, Perez MV, Zareba W, Ware JS, Wilde AAM, Gollob MH (2020) An international, multicentered, evidence-based reappraisal of genes reported to cause congenital long QT syndrome. Circulation 141(6):418–428
Adzhubei IA, Schmidt S, Peshkin L, Ramensky VE, Gerasimova A, Bork P, Kondrashov AS, Sunyaev SR (2010) A method and server for predicting damaging missense mutations. Nat Methods 7(4):248–249
Alders M, Bikker H, Christiaans I (2018) In: Long QT Syndrome. GeneReviews((R)). Adam MP, Ardinger HH, Pagon RA et al. Seattle (WA). https://www.ncbi.nlm.nih.gov/books/NBK1129/
Al-Hassnan ZN, Al-Fayyadh M, Al-Ghamdi B, Shafquat A, Mallawi Y, Al-Hadeq F, Tulbah S, Shinwari ZMA, Almesned A, Alakhfash A, Al Fadly F, Hersi AS, Alhayani A, Al-Hashem A, Arafah D, Dzimiri N, Meyer B, Rababh M, Al-Manea W (2017) Clinical profile and mutation spectrum of long QT syndrome in Saudi Arabia: the impact of consanguinity. Heart Rhythm 14(8):1191–1199
Anderson CL, Kuzmicki CE, Childs RR, Hintz CJ, Delisle BP, January CT (2014) Large-scale mutational analysis of Kv11.1 reveals molecular insights into type 2 long QT syndrome. Nat Commun 5:5535
Ashkenazy H, Abadi S, Martz E, Chay O, Mayrose I, Pupko T, Ben-Tal N (2016) ConSurf 2016: an improved methodology to estimate and visualize evolutionary conservation in macromolecules. Nucleic Acids Res 44(W1):W344-350
Bartos DC, Duchatelet S, Burgess DE, Klug D, Denjoy I, Peat R, Lupoglazoff JM, Fressart V, Berthet M, Ackerman MJ, January CT, Guicheney P, Delisle BP (2011) R231C mutation in KCNQ1 causes long QT syndrome type 1 and familial atrial fibrillation. Heart Rhythm 8(1):48–55
Bartos DC, Anderson JB, Bastiaenen R, Johnson JN, Gollob MH, Tester DJ, Burgess DE, Homfray T, Behr ER, Ackerman MJ, Guicheney P, Delisle BP (2013) A KCNQ1 mutation causes a high penetrance for familial atrial fibrillation. J Cardiovasc Electrophysiol 24(5):562–569
Bdier AY, Al-Ghamdi S, Verma PK, Dagriri K, Alshehri B, Jiman OA, Ahmed SE, Wilde AAM, Bhuiyan ZA, Al-Aama JY (2017) Autosomal recessive long QT syndrome, type 1 in eight families from Saudi Arabia. Mol Genet Genomic Med 5(5):592–601
Bertalovitz AC, Badhey MLO, McDonald TV (2018) Synonymous nucleotide modification of the KCNH2 gene affects both mRNA characteristics and translation of the encoded hERG ion channel. J Biol Chem 293(31):12120–12136
Cerrone M, Remme CA, Tadros R, Bezzina CR, Delmar M (2019) Beyond the one gene-one disease paradigm: complex genetics and pleiotropy in inheritable cardiac disorders. Circulation 140(7):595–610
Chen J, Liu Z, Creagh J, Zheng R, McDonald TV (2020) Physical and functional interaction sites in cytoplasmic domains of KCNQ1 and KCNE1 channel subunits. Am J Physiol Heart Circ Physiol 318(2):H212–H222
Choi Y, Chan AP (2015) PROVEAN web server: a tool to predict the functional effect of amino acid substitutions and indels. Bioinformatics 31(16):2745–2747
Clemens DJ, Tester DJ, Giudicessi JR, Bos JM, Rohatgi RK, Abrams DJ, Balaji S, Crotti L, Faure J, Napolitano C, Priori SG, Probst V, Rooryck-Thambo C, Roux-Buisson N, Sacher F, Schwartz PJ, Silka MJ, Walsh MA, Ackerman MJ (2019) International triadin knockout syndrome registry. Circ Genom Precis Med 12(2):e002419
Crotti L, Lundquist AL, Insolia R, Pedrazzini M, Ferrandi C, De Ferrari GM, Vicentini A, Yang P, Roden DM, George AL Jr, Schwartz PJ (2005) KCNH2-K897T is a genetic modifier of latent congenital long-QT syndrome. Circulation 112(9):1251–1258
Crotti L, Spazzolini C, Tester DJ, Ghidoni A, Baruteau AE, Beckmann BM, Behr ER, Bennett JS, Bezzina CR, Bhuiyan ZA, Celiker A, Cerrone M, Dagradi F, De Ferrari GM, Etheridge SP, Fatah M, Garcia-Pavia P, Al-Ghamdi S, Hamilton RM, Al-Hassnan ZN, Horie M, Jimenez-Jaimez J, Kanter RJ, Kaski JP, Kotta MC, Lahrouchi N, Makita N, Norrish G, Odland HH, Ohno S, Papagiannis J, Parati G, Sekarski N, Tveten K, Vatta M, Webster G, Wilde AAM, Wojciak J, George AL, Ackerman MJ, Schwartz PJ (2019) Calmodulin mutations and life-threatening cardiac arrhythmias: insights from the International Calmodulinopathy Registry. Eur Heart J 40(35):2964–2975
Delisle BP, Underkofler HA, Moungey BM, Slind JK, Kilby JA, Best JM, Foell JD, Balijepalli RC, Kamp TJ, January CT (2009) Small GTPase determinants for the Golgi processing and plasmalemmal expression of human ether-a-go-go related (hERG) K+ channels. J Biol Chem 284(5):2844–2853
Giudicessi JR, Wilde AAM, Ackerman MJ (2018) The genetic architecture of long QT syndrome: a critical reappraisal. Trends Cardiovasc Med 28(7):453–464
Goldenberg I, Horr S, Moss AJ, Lopes CM, Barsheshet A, McNitt S, Zareba W, Andrews ML, Robinson JL, Locati EH, Ackerman MJ, Benhorin J, Kaufman ES, Napolitano C, Platonov PG, Priori SG, Qi M, Schwartz PJ, Shimizu W, Towbin JA, Vincent GM, Wilde AA, Zhang L (2011) Risk for life-threatening cardiac events in patients with genotype-confirmed long-QT syndrome and normal-range corrected QT intervals. J Am Coll Cardiol 57(1):51–59
Karczewski KJ, Francioli LC, Tiao G, Cummings BB, Alfoldi J, Wang Q, Collins RL, Laricchia KM, Ganna A, Birnbaum DP, Gauthier LD, Brand H, Solomonson M, Watts NA, Rhodes D, Singer-Berk M, England EM, Seaby EG, Kosmicki JA, Walters RK, Tashman K, Farjoun Y, Banks E, Poterba T, Wang A, Seed C, Whiffin N, Chong JX, Samocha KE, Pierce-Hoffman E, Zappala Z, O’Donnell-Luria AH, Minikel EV, Weisburd B, Lek M, Ware JS, Vittal C, Armean IM, Bergelson L, Cibulskis K, Connolly KM, Covarrubias M, Donnelly S, Ferriera S, Gabriel S, Gentry J, Gupta N, Jeandet T, Kaplan D, Llanwarne C, Munshi R, Novod S, Petrillo N, Roazen D, Ruano-Rubio V, Saltzman A, Schleicher M, Soto J, Tibbetts K, Tolonen C, Wade G, Talkowski ME, Neale BM, Daly MJ, MacArthur DG, Genome Aggregation Database (2020) The mutational constraint spectrum quantified from variation in 141,456 humans. Nature 581(7809):434–443
Kim S, Jhong JH, Lee J, Koo JY (2017) Meta-analytic support vector machine for integrating multiple omics data. BioData Min 10:2
Lin Y, Williams N, Wang D, Coetzee W, Zhou B, Eng LS, Um SY, Bao R, Devinsky O, McDonald TV, Sampson BA, Tang Y (2017) Applying high-resolution variant classification to cardiac arrhythmogenic gene testing in a demographically diverse cohort of sudden unexplained deaths. Circ Cardiovasc Genet. https://doi.org/10.1161/CIRCGENETICS.117.001839
Liu J, Liu D, Li M, Wu K, Liu N, Zhao C, Shi X, Liu Q (2020) Identification of a nonsense mutation in TNNI3K associated with cardiac conduction disease. J Clin Lab Anal 34(9):e23418. https://doi.org/10.1002/jcla.23418
Lundby A, Ravn LS, Svendsen JH, Olesen SP, Schmitt N (2007) KCNQ1 mutation Q147R is associated with atrial fibrillation and prolonged QT interval. Heart Rhythm 4(12):1532–1541
McKenna A, Hanna M, Banks E, Sivachenko A, Cibulskis K, Kernytsky A, Garimella K, Altshuler D, Gabriel S, Daly M, DePristo MA (2010) The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res 20(9):1297–1303
Mitcheson JS, Chen J, Lin M, Culberson C, Sanguinetti MC (2000) A structural basis for drug-induced long QT syndrome. Proc Natl Acad Sci USA 97(22):12329–12333
Nickless A, Bailis JM, You Z (2017) Control of gene expression through the nonsense-mediated RNA decay pathway. Cell Biosci 7:26
Priori SG, Napolitano C, Schwartz PJ (1999) Low penetrance in the long-QT syndrome: clinical impact. Circulation 99(4):529–533
Priori SG, Schwartz PJ, Napolitano C, Bloise R, Ronchetti E, Grillo M, Vicentini A, Spazzolini C, Nastoli J, Bottelli G, Folli R, Cappelletti D (2003) Risk stratification in the long-QT syndrome. N Engl J Med 348(19):1866–1874
Priori SG, Wilde AA, Horie M, Cho Y, Behr ER, Berul C, Blom N, Brugada J, Chiang CE, Huikuri H, Kannankeril P, Krahn A, Leenhardt A, Moss A, Schwartz PJ, Shimizu W, Tomaselli G, Tracy C (2013) HRS/EHRA/APHRS expert consensus statement on the diagnosis and management of patients with inherited primary arrhythmia syndromes: document endorsed by HRS, EHRA, and APHRS in May 2013 and by ACCF, AHA, PACES, and AEPC in June 2013. Heart Rhythm 10(12):1932–1963
Rentzsch P, Witten D, Cooper GM, Shendure J, Kircher M (2019) CADD: predicting the deleteriousness of variants throughout the human genome. Nucleic Acids Res 47(D1):D886–D894
Schwartz PJ, Ackerman MJ (2013) The long QT syndrome: a transatlantic clinical approach to diagnosis and therapy. Eur Heart J 34(40):3109–3116
Schwartz PJ, Crotti L (2011) QTc behavior during exercise and genetic testing for the long-QT syndrome. Circulation 124(20):2181–2184
Schwartz PJ, Priori SG, Spazzolini C, Moss AJ, Vincent GM, Napolitano C, Denjoy I, Guicheney P, Breithardt G, Keating MT, Towbin JA, Beggs AH, Brink P, Wilde AA, Toivonen L, Zareba W, Robinson JL, Timothy KW, Corfield V, Wattanasirichaigoon D, Corbett C, Haverkamp W, Schulze-Bahr E, Lehmann MH, Schwartz K, Coumel P, Bloise R (2001) Genotype-phenotype correlation in the long-QT syndrome: gene-specific triggers for life-threatening arrhythmias. Circulation 103(1):89–95
Schwartz PJ, Stramba-Badiale M, Crotti L, Pedrazzini M, Besana A, Bosi G, Gabbarini F, Goulene K, Insolia R, Mannarino S, Mosca F, Nespoli L, Rimini A, Rosati E, Salice P, Spazzolini C (2009) Prevalence of the congenital long-QT syndrome. Circulation 120(18):1761–1767
Schwartz PJ, Crotti L, George AL Jr (2018) Modifier genes for sudden cardiac death. Eur Heart J 39(44):3925–3931
Schwartz PJ, Ackerman MJ, Antzelevitch C, Bezzina CR, Borggrefe M, Cuneo BF, Wilde AAM (2020) Inherited cardiac arrhythmias. Nat Rev Dis Primers 6(1):58
Schwarz JM, Cooper DN, Schuelke M, Seelow D (2014) MutationTaster2: mutation prediction for the deep-sequencing age. Nat Methods 11(4):361–362
Sega AG, Mis EK, Lindstrom K, Mercimek-Andrews S, Ji W, Cho MT, Juusola J, Konstantino M, Jeffries L, Khokha MK, Lakhani SA (2019) De novo pathogenic variants in neuronal differentiation factor 2 (NEUROD2) cause a form of early infantile epileptic encephalopathy. J Med Genet 56(2):113–122
Sim NL, Kumar P, Hu J, Henikoff S, Schneider G, Ng PC (2012) SIFT web server: predicting effects of amino acid substitutions on proteins. Nucleic Acids Res 40(Web Server issue):W452–W457
Smith JL, Anderson CL, Burgess DE, Elayi CS, January CT, Delisle BP (2016) Molecular pathogenesis of long QT syndrome type 2. J Arrhythm 32(5):373–380
Stead LF, Wood IC, Westhead DR (2011) KvSNP: accurately predicting the effect of genetic variants in voltage-gated potassium channels. Bioinformatics 27(16):2181–2186
Theis JL, Zimmermann MT, Larsen BT, Rybakova IN, Long PA, Evans JM, Middha S, de Andrade M, Moss RL, Wieben ED, Michels VV, Olson TM (2014) TNNI3K mutation in familial syndrome of conduction system disease, atrial tachyarrhythmia and dilated cardiomyopathy. Hum Mol Genet 23(21):5793–5804
Wang K, Li M, Hakonarson H (2010) ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res 38(16):e164
Whiffin N, Minikel E, Walsh R, O’Donnell-Luria AH, Karczewski K, Ing AY, Barton PJR, Funke B, Cook SA, MacArthur D, Ware JS (2017) Using high-resolution variant frequencies to empower clinical genome interpretation. Genet Med 19(10):1151–1158
Zareba W, Moss AJ, Schwartz PJ, Vincent GM, Robinson JL, Priori SG, Benhorin J, Locati EH, Towbin JA, Keating MT, Lehmann MH, Hall WJ (1998) Influence of the genotype on the clinical course of the long-QT syndrome. International Long-QT Syndrome Registry Research Group. N Engl J Med 339(14):960–965
Zhou Z, Gong Q, Ye B, Fan Z, Makielski JC, Robertson GA, January CT (1998) Properties of HERG channels stably expressed in HEK 293 cells studied at physiological temperature. Biophys J 74(1):230–241
Acknowledgements
We would like to thank the Yale New Haven Hospital and Sara and Jeffery Buell for their generous support for the Pediatric Genomics Discovery Program, and the Yale Center for Genome Analysis for performing whole exome sequencing. We would also like to acknowledge the proband and her family for their willingness to share their stories with the medical community.
Funding
TVM was supported by National Institutes of Health/National Heart Lung Blood Institute HL120782. MKK was supported by National Institutes of Health/National Heart Lung Blood Institute HL149746.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
SAL is part owner of Qiyas Higher Health and Victory Genomics, startup companies unrelated to this work. MKK is part owner of Victory Genomics. No other authors have any disclosures to report.
Ethical statement
This study was approved by the Institutional Review Board of Yale University School of Medicine. Informed consent was obtained for all individual participants included in the study. All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
Additional information
Communicated by Stefan Hohmann.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Najari Beidokhti, M., Bertalovitz, A.C., Ji, W. et al. Functional testing for variant prioritization in a family with long QT syndrome. Mol Genet Genomics 296, 823–836 (2021). https://doi.org/10.1007/s00438-021-01780-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-021-01780-3