Abstract
The role of a highly conserved YEATS protein motif is explored in the context of the Taf14 protein of Saccharomyces cerevisiae. In S. cerevisiae, Taf14 is a protein physically associated with many critical multisubunit complexes including the general transcription factors TFIID and TFIIF, the chromatin remodeling complexes SWI/SNF, Ino80 and RSC, Mediator and the histone modification enzyme NuA3. Taf14 is a member of the YEATS superfamily, conserved from bacteria to eukaryotes and thought to have a transcription stimulatory activity. However, besides its ubiquitous presence and its links with transcription, little is known about Taf14’s role in the nucleus. We use structure–function and mutational analysis to study the function of Taf14 and its well conserved N-terminal YEATS domain. We show here that the YEATS domain is not necessary for Taf14’s association with these transcription and chromatin remodeling complexes, and that its presence in these complexes is dependent only on its C-terminal domain. Our results also indicate that Taf14’s YEATS domain is not necessary for complementing the synthetic lethality between TAF14 and the general transcription factor TFIIS (encoded by DST1). Furthermore, we present evidence that the YEATS domain of Taf14 has a negative impact on cell growth: its absence enables cells to grow better than wild-type cells under stress conditions, like the microtubule destabilizing drug benomyl. Moreover, cells expressing solely the YEATS domain grow worser than cells expressing any other Taf14 construct tested, including the deletion mutant. Thus, this highly conserved domain should be considered part of a negative regulatory loop in cell growth.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
RNA polymerase II transcription is a tightly regulated process in which the chromatin near the gene to be transcribed is re-organized and remodeled, followed by the formation of the transcription pre-initiation complex and the processes of initiation, elongation and termination. Different multi-protein complexes are involved in these processes for regulation under normal or stress conditions. A similar phenomenon of complex regulation occurs during cell cycle progression where many changes take place including modulation of the cytoskeleton (Sawin and Tran 2006). Taf14 is one of the few proteins thought to play a role in all of these vital processes but its specific role is still unknown, and it is the focus of this work.
TAF14 was first identified (as ANC1) in a screen for mutations that enhance defects caused by a mutation in the actin gene (Welch et al. 1993). Cells devoid of Taf14 are viable but thermo- and osmo-sensitive (Henry et al. 1994) have defects in actin organization (Welch et al. 1993), and aberrant morphology and cytoskeletal defects (Henry et al. 1994). Additionally, Taf14 is physically associated with a variety of transcription and chromatin remodeling complexes (Fig. 1a). It is present in, at least, three different ATP-dependent chromatin remodeling complexes: Swi/Snf, which possesses DNA-stimulated ATPase activity and activates transcription by helping transcription factors access their binding sites (Cairns et al. 1996; Sudarsanam and Winston 2000; Dechassa et al. 2008); Ino80, which has helicase activity and is involved in transcription, replication and repair (Shen et al. 2003a, b; Conaway and Conaway 2009); and RSC, essential for cell cycle progression (Du et al. 1998; Eberharter et al. 1998), and known for repositioning and ejecting nucleosomes from active sites (Parnell et al. 2008). Taf14 also has been found in the histone–acetyltransferase complex, NuA3 (John et al. 2000; Kabani et al. 2005) that stimulates transcription or replication elongation through nucleosomes by providing a coupled acetyltransferase activity (John et al. 2000). All of these complexes help transcriptional activators bind to occluded chromatin sites by perturbing histone–DNA contacts or by covalently modifying histones (Clark-Adams et al. 1988; Imbalzano et al. 1994; Steger et al. 1998; Shen et al. 2000; Neely and Workman 2002; Cairns 2005). Roles for these complexes were first identified in transcriptional regulation (Kingston and Narlikar 1999; Davie and Kane 2000; Narlikar et al. 2002), but it has recently been shown that they also play diverse and complex roles in the DNA damage response (for review (Costelloe et al. 2006; Park et al. 2009)).
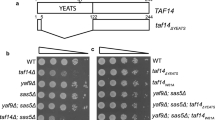
a Seven nuclear complexes that contain or bind Taf14. The sizes are not to scale. The names of the proteins tagged in this study from each of the complexes are shown. b Truncated versions of Taf14 used in this study: TAF14, taf14[YEATS] (amino acids 6–113), taf14[CT] (amino acids 113–244), and taf14[-NT] (amino acids 61–244). c Expression of the different Taf14 constructs. Equal amounts of cell extract were resolved on a 10% SDS-Laemmli protein gel and transferred to a nitrocellulose membrane for western blotting. Taf14 proteins were detected with Polyclonal Taf14 antibody. Left paneltaf14Δ strain (CMKy70) containing plasmids expressing Taf14 constructs under its natural promoter. The YEATS fragment (Y second line) is practically not detected. Empty plasmid was used as negative control, PGK protein as loading control. Right paneltaf14Δ strain (CMKy70) containing high copy plasmids expressing Tap-tagged Taf14 constructs under the PGK promoter. Empty pVV233 vector (TAP tag alone) was used as the negative control. The TAP tag is visible with the Taf14 antibody because it contains Protein A, recognized by IgG’s in every polyclonal antibody. H+ vacuolar ATPase was used as loading control
Taf14 is also a subunit of the yeast transcription factors TFIID (Poon et al. 1995) and TFIIF (Henry et al. 1994; Cairns et al. 1996) and seems to interact with TFIIB (Kimura and Ishihama 2004). TFIID recognizes the TATA box or other promoter elements and enables the binding of TFIIB and TFIIA. It is also known to modify histones (Mizzen et al. 1996; Cler et al. 2009). TFIIF assists the recruitment of RNA polymerase II, TFIIE and TFIIH into the pre-initiation complex (Buratowski et al. 1989; Flores et al. 1991, 1992; Henry et al. 1994). It is also involved in start site selection (Robert et al. 1998; Majovski et al. 2005; Khaperskyy et al. 2008) and elongation (Robert et al. 1998; Conaway et al. 2000; Zhang et al. 2005). Although a direct interaction has not always been detected (Kabani et al. 2005), Taf14 has also been suggested as a component of the transcriptional activator Mediator (Kim et al. 1994; Erlich et al. 2008), a global regulator important in recruiting of the transcription machinery and modulating RNA polymerase II activity after the alteration of the chromatin template by remodelers (Myers and Kornberg 2000; Baek et al. 2006; He et al. 2008). Supporting a role for Taf14 in initiation, in the ADH1 gene (Krogan et al. 2002) and the GAL1 gene (Kabani et al. 2005), Taf14 is recruited to the promoter region and not to the coding region, suggesting that Taf14 may be involved in the assembly of the RNA polymerase II pre-initiation complex. This recruitment is likely due to its presence in TFIID and TFIIF, but Taf14 also could have been recruited to that region through its association with the chromatin remodeling complexes described above.
In addition to the roles of Taf14 in chromatin remodeling and transcription initiation, in Saccharomyces cerevisiae there is a strong genetic interaction between Taf14 and the transcription elongation factor TFIIS (Fish et al. 2006). TFIIS stimulates RNA polymerase II during elongation to recover from an arrested state by inducing the polymerase to cleave its nascent transcript, thus realigning the new RNA 3′ end in the active site (Fish and Kane 2002; Ghavi-Helm et al. 2008). Neither TFIIS nor Taf14 is essential, but the double mutant results in synthetic lethality (Davie and Kane 2000; Fish et al. 2006). These two proteins are not functionally redundant, but have a functional interaction within a cellular process, most likely during transcriptional elongation and in concert with TFIIF (Fish et al. 2006). Taf14 (as well as proteins Gal1, Sua7, Rts1 or Tys1) has additionally been reported as a strong suppressor of the genetic interaction between DST1 (the gene encoding for TFIIS) and SOH1 (when overexpressed) (Malagon et al. 2004). SOH1 is a member of Mediator and, in Saccharomyces, is synthetically lethal with mutations in genes encoding subunits of the RNA polymerase II, TFIIB (Fan et al. 1996) and the histone H3 methyl transferase factor Set2 (Krogan et al. 2003). Taf14 is also one of the 5% of intron containing genes in yeast and, surprisingly, splicing of its intron is necessary to suppress a temperature-sensitive cell cycle arrest of cdc40Δ cells (Dahan and Kupiec 2004).
Taf14 contains a highly conserved YEATS domain at its N-terminus and along with Yaf9 and Sas5 comprises the YEATS domain family in yeast. The name for this domain derives from the first discovered proteins containing the domain (Yaf9, ENL, AF9, Taf14, Sas5) (Harborth et al. 2000). Its function has not yet been described, but it is thought to be implicated in regulating transcription and chromatin remodeling (Schulze et al. 2009). This domain is found in many eukaryotic species as diverse as Arabidopsis, Drosophila, Caenorhabditis elegans, yeast and human. ENL, AF-9 (Corral et al. 1996) and Gas41 (Fischer et al. 1997) are the most extensively studied human proteins in this group. These three proteins are directly implicated in leukemias and gliomas whether by fusion of their non-YEATS domain to a protein called MLL in the case of ENL and AF-9 (Adler et al. 1999) or by amplification of its expression in the case of Gas41 (Munnia et al. 2001; Park and Roeder 2006). As mentioned above, the Taf14 YEATS domain presents striking homology with two other yeast nuclear proteins, Yaf9 and Sas5. Yaf9 is the ortholog of human Gas41, and is a component of the NuA4 HAT complex and the SWR1 complex, which deposits the histone variant H2A.Z (Le Masson et al. 2003; Zhang et al. 2004). Sas5 is a component of the SAS complex involved in chromatin silencing by acetylating free histones (Sutton et al. 2003; Shia et al. 2005).
Here, we present the first evidence that the YEATS domain of Taf14 exerts a negative effect on cell growth, and deletion of the YEATS domain results in a growth advantage under certain stress conditions. We further show that the C-terminus but not the YEATS domain of Taf14 is necessary and sufficient for the rescue of the synthetic lethality with TFIIS. Moreover, the C-terminus is also sufficient for association of Taf14 with the complexes involved in transcription and chromatin remodeling described above, revealing that the YEATS domain is not required for interaction with these complexes. These results suggest that, rather than a structural role, the YEATS domain has a regulatory function, possibly coordinating the transcription and replication processes during the cell cycle.
Materials and methods
Plasmids, strains, genetic methods and media
The S. cerevisiae strains and plasmids used in this study are listed in Tables 1 and 2. Taf14 point mutations were generated based on oligonucleotide-directed mutagenesis (Kunkel et al. 1987) using the “Quick Change” mutagenesis strategy, as directed by the manufacturer (Stratagene). Escherichia coli calcium-manganese-based transformations were used with the Top10 E. coli strain. Yeast cells were transformed using lithium acetate (Gietz et al. 1995).
Media such as YPD or synthetic complete (SC) were prepared according to standard procedures (Guthrie et al. 1991). All chemicals were from Sigma–Aldrich unless otherwise noted. SE media was prepared as SC but by substituting the nitrogen source with 1 g/L of glutamic acid (monosodium salt). G418 Sulfate was added to autoclaved YPD or SE media at a concentration of 200 μg/mL. ClonNAT (WERNER BioAgents) was added to autoclaved media at 100 μg/mL. 6-Azauracil (6-AU) was added to autoclaved media at a final concentration of 60 μg/mL; NaCl, KCl, and sorbitol were included at a final concentration of 0.9, 1.2 and 1.8 M, respectively. Benomyl was added immediately after autoclaving media at a concentration of 35 μg/mL, from a stock solution at 10 mg/mL in dimethyl sulfoxide.
For the growth studies on plates, cells in exponential phase were diluted to an OD600 of 0.1 and four serial 10× dilutions were made. The same volume of cells was applied onto YPD or SC-URA plates with or without additions as indicated in the text. Cells were grown at 24, 30, or 36°C. For the growth studies in liquid media, cells in early stationary phase were diluted to an OD600 of 0.05 and grown at 30°C. Samples were taken every 1 h. The mean of the doubling times was calculated with data from exponential phase (OD600 0.18–2.0). All the RP2P were larger than 0.9.
Strain recovery and tetrad analysis
The ability of various TAF14 constructs to complement the synthetic lethality in the taf14Δdst1Δ strain was determined by tetrad analysis of heterozygous diploids. After sporulation of the TAF14/taf14Δ::kan r , dst1Δ::hisG/dst1Δ::hisG diploid (CMKy91) containing the different TAF14 constructs on a URA-marked plasmid, the tetrads were dissected onto YPD solid media (plate A) which was then incubated at 26°C for 5–7 days. The colony size at every spore position was noted. In the case of extremely small colonies, the cells were transferred to fresh YPD media (plate B) and incubated for 1 week at 26°C. Colonies on YPD were replica plated to SC-URA media with or without G418 to determine genotypes of the colonies.
Whole cell extract and protein analysis
Cell extracts for protein expression analyses were made using the post-alkaline extraction method (Kushnirov 2000).
Standard Western blotting conditions with SDS-PAGE gels were used (Sambrook and Russell 2001). Protein was transferred to nitrocellulose at 100 mV for 1 h and processed for immunoblotting. Primary antibodies were used in the following concentration: polyclonal anti-Taf14, 1:1,000; monoclonal anti-Flag, M2 (Sigma), 1:1,000; monoclonal anti-H+-vacuolar ATPase (Molecular Probes), 1:5,000; anti-PGK (molecular Probes), 1:5,000; and monoclonal anti-HA, clone HA-7 (Sigma), 1:5,000. The secondary ECLTM anti-rabbit IgG horseradish peroxidase (HRP) labeled antibody (Amersham) was used at 1:5,000, and the secondary ECLTM anti-mouse IgG horseradish peroxidase labeled antibody (Amersham) was used at 1:10,000. The detection of the immobilized proteins conjugated to HRP labeled antibodies was made by using ECL Plus western blotting detection reagents (Amersham-GE Healthcare).
Immunofluorescence microscopy
Asynchronous early log phase yeast cells (5 mL) were fixed with 5% formaldehyde (from a 37% stock solution added directly to the culture) for 1 h at 25°C on a rotator. Formaldehyde was removed by washing twice with 2 mL 0.1 M potassium phosphate buffer (pH 7.5). Cells were resuspended in 1 mL 0.1 M potassium phosphate buffer (pH 7.5) and 500 μL of cell suspension was permeabilized by the addition of 1 μL 14 M β-mercaptoethanol and 5.1 μL 5 mg/mL Zymolyase 20T (USB). After digestion at 37°C for 35 min, the cells were pelleted, and resuspended in 500 μl 0.1 M potassium phosphate buffer (pH 7.5). Ten microliters of cell suspension was then applied to polylysine coated slides (Fisher) and air dried. After rehydrating by immersion in 1× PBS for 5 min, cells were fixed for 6 min in methanol and 30 s in acetone, both at −20°C. Cells were blocked with BLOTTO [1× phosphate buffered saline (PBS), 0.05% Tween-20, 5% nonfat dry milk] for 15 min at room temperature. Affinity purified anti-Taf14 antibody (kind gift of David Drubin, UC Berkeley) was added in BLOTTO at a 1:50 dilution. Incubations continued for 12–16 h at 4°C in a humidified chamber. After washing twice with BLOTTO, 15 min each, Alexa Fluor 488® goat anti-rabbit IgG (H + L) (Molecular Probes) was added in BLOTTO at a 1:200 dilution. Incubations continued for 1 h at 25°C in the dark in a humidified chamber. Slides were washed twice for 15 min each in PBS. Slides were mounted in DABCO mounting medium (2.5% w/v DABCO, Sigma #D-2522, 50 mM Tris pH 7.5, 90% glycerol) containing 2 μg/mL DAPI. Coverslips were sealed with nail polish and stored at 4°C in the dark.
Inmunoprecipitation
N-terminal TAP-tagged Taf14 constructs were made with GatewayPTMP Cloning technology (Invitrogen) (Van Mullem et al. 2003). Three entry clones were generated containing the Taf14 constructs: Taf14 (1–244), taf14[CT] (113–244) and taf14[YEATS] (6–113), following the manufacturer’s instructions. From each clone, the ORF was subcloned into the high copy plasmid pVV223 creating N-terminal TAP-tagged expression clones under the PGK promoter.
Genomic N-terminal FLAG tagged Taf2, Sth1, Sas3, Ino80, Tfg1, Taf1 and genomic N-terminal HA-tagged Snf5 (strains: W303-A-Snf5, Ino80-FLAG, W303-A-Sth1, W303-A-Sas3, W303-A-Tfg1, W303-A-Taf1 and W303-A-Taf2) were transformed with the three different truncated TAP-Taf14 mutants. The strains were grown in 150 mL SE-URA/ClonNAT/G418 to log phase. Next, 150 OD600 units were harvested (~0.4 g pellet). The cell pellet was washed in 10 mL of cold PBS (130 mM NaCl, 7 mM Na2HPO4-2H2O, 3 mM NaH2PO4-2H2O adjusted to pH 7.0), pelleted again, and resuspended in 1 mL PBS. After another centrifugation, the cell pellet was resuspended in 0.6 mL TAP-Ca2+-Buffer [50 mM Tris–Cl pH 7.5, 200 mM NaCl, 1.5 mM MgAc, 2 mM CaClB2, 0.15% NP40, 1 mM DTT and one mini protease inhibitor tablet (Roche) per 5 mL], and the tube was filled with acid-washed glass beads (425–600 µm, Sigma). The cells were disrupted mechanically using a bead beater (BioSpec Products, Bartlesville, Oklahoma, USA) 5 times for 20 s with 1-min steps on ice in between. The glass beads were removed by centrifugation at 250 × g. The unbroken cells and the insoluble material were removed by centrifugation at 4°C, 20,000 × g for 20 min. The supernatant of each sample was mixed with 50 μL of TAP-Ca2+-Buffer equilibrated Calmodulin Affinity Resin (Stratagene) and incubated with rotating at 4°C for 60 min. The beads were pelleted and washed four times with 0.6 mL TAP-Ca2+-Buffer, rotating for 10 min during each wash. They were then resuspended in 100 μL 2× SDS sample buffer (50 mM Tris–Cl pH 6.8, 2% SDS, 10% glycerol, 0.025% bromophenol blue) and resolved on a 7.5% SDS-polyacrylamide gel. After transferring to a nitrocellulose membrane, the gel was immunoblotted with monoclonal antibody against FLAG or HA peptides, and a polyclonal antibody against Taf14 as described above under protein analysis.
Results
The YEATS domain inhibits growth under different stress conditions
To dissect the functional regions of Taf14 and decipher the role of its N-terminal YEATS domain, we created several deletion mutants (Fig. 1b). All the mutants were expressed on a plasmid under TAF14’s own promoter and regulatory sequences in a taf14 disruption strain. The expression of all mutant proteins was similar to wild type except for the taf14[YEATS] construct which was expressed or detected poorly (Fig. 1c, left panel). Despite this detection problem, there is no doubt of its expression as the phenotypes for this mutant are completely different than the phenotypes for the taf14 deletion mutant (see below). The cellular localization of all the constructs was also studied and while the taf14[CT] and the taf14[-NT] proteins were located in the nucleus as was the wild-type protein (Fig. 2), taf14[YEATS] was practically undetectable with the Taf14 antibody (Fig. 2). This result is consistent with its low detection by Western blots (Fig. 1c, left panel).
Fluorescence microscopy of taf14 mutants’ intracellular localization. taf14Δ cells containing different plasmids expressing Taf14 constructs under its natural promoter were fixed and immunostained with a polyclonal Taf14 antibody (left panels, green). The center panels show nuclei in DAPI stained cells (blue). The right panels show the result of merging the immunostaining and the DAPI staining. Pictures represent a single image plane following deconvolution. Cells contained the wild-type Taf14 (top panel); or the taf14[CT] construct (second panel); or the taf14[-NT] construct (third panel); or the taf14[YEATS] construct (fourth panel); or the empty vector pCP3 with the 5′ and 3′ UTR of Taf14 as a negative control to determine the background (bottom panel)
In synthetic media lacking uracil (SC-URA) at 30°C, cells expressing the Taf14 construct missing the complete YEATS domain (taf14[CT]) had a growth rate comparable to cells expressing wild-type Taf14. In contrast, cells expressing the YEATS domain alone (taf14[YEATS]) or a construct with partial deletion of the YEATS domain (taf14[-NT]) showed a diminished growth rate (Table 3). In the case of cells expressing taf14[YEATS], the doubling time was 1.6 times slower than the taf14Δ mutant (CMKy70) and 2.8 times slower than wild-type cells. In addition, the lag phase of cells expressing taf14[YEATS] was more than four times longer than that of the wild-type cells (Table 3).
The Taf14 deletion constructs were further tested for their ability to complement the temperature sensitivity of a taf14 disruption strain CMKy70 (Table 4, left panel). Taf14[CT] grew as well as wild type at 36°C (and at 24°C). In contrast, taf14[YEATS] could not complement the temperature sensitivity at 36°C. Interestingly, taf14[-NT] could partially complement the growth at 36°C as it did during growth at 30°C cited above. These phenotypes were fully recessive to wild-type Taf14 (Fig. S1).
Since a taf14 deletion strain is sensitive to different genotoxic stressors, we also tested the effect of the constructs on cell growth under different stress conditions (Table 4; Fig. 3) at both permissive (24°C) and non-permissive (36°C) temperatures. As expected from the described results above, taf14[CT] showed no growth disadvantage compared to the wild-type strain. Unexpectedly however, it displayed a growth advantage under certain conditions. This advantage was pronounced in the presence of the microtubule destabilizing drug benomyl, as well as on formamide or caffeine at 36C. A less pronounced advantage was seen in the presence of the nucleotide depleting drug 6-azauracil (6-AU). In contrast, under all tested conditions taf14[-NT] allowed very slow or no growth while taf14[YEATS] could not complement the growth defects of the taf14 disruption mutant (Table 4). We conclude that under a wide range of conditions, the YEATS domain of Taf14 limits cell growth both alone and in the context of the wild-type protein. These results may also indicate that in wild-type cells, a negative function of the YEATS domain is being regulated. This possibility is further explored in the “Discussion”.
Growth rates of the different taf14 truncated mutants. Growth comparison of CMKy70 containing Taf14, taf14[CT] or taf14[-NT] on a plasmid. Tenfold serial dilutions from exponential phase cells were plated onto SC-URA with or without the indicated drugs (concentrations as in Table 4). Cells on SC-URA and SC-URA with formamide were photographed after 3 days growth; SC-URA with benomyl, after 5 days; SC-URA with 6-AU, after 6 days
Amino acids W81 and W84 form part of a putative regulator interaction site on the YEATS domain surface
Given the possible regulatory role of the YEATS domain in the cell cycle, we decided to perform a mutagenic scan of the domain to localize sites within the YEATS domain that were important for its function. We used the taf14[-NT] construct since its sensitized phenotype allowed for the observation of mutations that resulted in either increased or decreased growth.
Knowing that charged amino acids are usually necessary for protein–protein interactions, ten different single, double, or triple alanine substitutions of charged amino acid residues were made in the taf14[-NT] construct (Fig. 4a; Table 1). In addition, we created five different alanine substitutions of highly conserved residues hoping to target an essential docking site (Fig. 4a; Table 1). We compared the growth of cells transformed with each of the mutants to that of the strain expressing taf14[-NT] on solid media. Fourteen of the mutants grew similarly to cells harbouring taf14[-NT] (data not shown). However, the double mutant W81A/F84A improved growth (Fig. 4b). In fact, the growth of this mutant became comparable to that of cells expressing the full length protein at 24, 30 and 37°C. W81A/F84A also showed slightly better growth than unmutagenized taf14[-NT] on both sorbitol and benomyl (data not shown). There was no significant difference between the protein levels of any of these taf14[-NT] mutants (Fig. S2), arguing against the fact that altered expression levels were responsible for the observed phenotypes. Furthermore, the conserved residues W81 and F84 are predicted to lie on the surface of Taf14 (confirmed by the prediction programs: 3D-PSSM and JNet Secondary Structure Prediction, and Wang et al. 2009) suggesting that they are not serving a structural role, but instead are involved in binding interactions between Taf14 and other protein regulators.
a Multiple sequence alignment of the carboxy terminal half of the YEATS domain of Taf14 (61–113 aa) compared with other YEATS family proteins. Conserved residues in bold type. The point mutations used in this study are marked below Taf14; the grey boxes correspond to aminoacids 81 and 84. b Quantitative growth comparison of cells with Taf14 mutant constructs. CMY70 (taf14Δ) strains containing either pJS02 (TAF14), pCP5 (taf14[CT]), pCP6 (taf14[-NT]), pJS06 (taf14[-NT W81A-F84A ]), pCP4 (taf14[YEATS]) or pJS06-A (taf14[YEATS W81A-F84A ]) were grown to exponential phase (OD600 of 0.8–2.0) and 10× serial dilution were plated onto SC-URA. Cells were grown at 24°C and after 3 days the plate was photographed
Interestingly, the same double mutant slightly decreased growth when incorporated into the YEATS construct itself (Table 3) and had no impact on growth in either liquid or on solid media when inserted into the wild-type Taf14 protein. These results suggest that the putative YEATS domain interaction site extends beyond these two residues, perhaps along the fragment of the YEATS domain missing in the taf14[-NT] mutant.
Taf14 residues in the C-terminal domain are important for the synthetic lethal phenotype between taf14Δ and dst1Δ
Although TAF14 is not essential, its deletion shows a synthetic lethality with deletion of DST1, the Taf14 protein is therefore required for the survival of cells lacking the transcript elongation factor TFIIS (encoded by DST1) (Fish et al. 2006). To determine if the YEATS domain of Taf14 is required for this genetic interaction, we analyzed the different mutants of TAF14 for their ability to complement the synthetic lethality between taf14Δ and dst1Δ. Plasmids containing the taf14 deletion mutants (Fig. 1b) were transformed into the heterozygous TAF14/taf14Δ::kan r, dst1Δ::hisG/dst1Δ::hisG diploid strain and transformants were sporulated at 30°C (Fig. 5). The taf14[YEATS] construct did not complement the synthetic lethality of the double deletion mutant (Fig. 5b), and taf14[-NT] grew slowly, taking 5–7 days for the germinating spores to make visible colonies (Fig. 5c). However, taf14[CT] complemented to equal levels as the wild-type protein (Fig. 5d).
Efficiency of different Taf14 constructs in complementing the synthetic lethality between dst1Δ and taf14Δ. Heterozygous diploids were transformed with plasmids containing the indicated Taf14 constructs. Tetrad analysis was carried out with each strain. Grey boxes represent the taf14Δ/dst1Δ double mutant. The taf14∆::kan r spores were identified by their G418r phenotype. The presence of the URA3 marked plasmid was determined by testing growth on SC-URA media. White boxes represent the dst1Δ/TAF14 mutant. Boxes with X did not have spores. a Tetrad analysis following sporulation of CMKy91 containing pJS02 (TAF14). The tetrads were dissected onto YPD medium, and the plate was photographed after 2 days at 24°C. b Similar to a with plasmid pCP4 (taf14[YEATS]). Plates were photographed after 3 days at 24°C. c Similar to a with plasmid pCP6 (taf14[-NT]). Plates were photographed after 7 days at 24°C. d Similar to a with plasmid pCP5 (taf14[CT]). Plates were photographed after 3 days at 24°C. Even after 9 days, no growth was detectable in the blank spots
We also studied the ability of taf14[CT] or taf14[-NT] mutants to complement the temperature sensitivity in the taf14Δdst1Δ background (conferred by the absence of Taf14). As in the taf14Δ background (Fig. 3, left bottom panel), the taf14Δdst1Δ strain expressing the taf14[CT] construct grew as well as the strain expressing the wild-type construct at 36°C. But the cells containing the taf14[-NT] construct grew poorly although this protein was able to complement the temperature sensitivity.
We next tested the ability of point mutants in taf14[-NT] (Table 1) to complement the synthetic lethality between taf14Δ and dst1Δ (Fig. 6). The double mutant W81A/F84A conferred a growth advantage in the taf14Δdst1Δ cells, as it did in the taf14 null background (Fig. 4b). Surprisingly, in contrast to our result in the taf14 null background, three additional mutations outside the YEATS domain conferred a growth advantage in the taf14Δdst1Δ background: K124A, E129A/E130A, and K133A, suggesting that these three C-terminal residues are particularly relevant for the TAF14 and DST1 genetic interaction. Thus, we concluded that genetic interaction between TAF14 and DST1 is not dependent on the YEATS domain of Taf14 either.
Complementation of the taf14Δ dst1Δ synthetic lethality by the taf14[-NT*] point mutants. CMKy91 (taf14Δ dst1Δ) cells containing either pJS02 (TAF14), pCP5 (taf14[CT]), pCP6 (taf14[-NT]) or one of the plasmids with point mutations in taf14[-NT*] were plated in serial dilutions onto SC-URA and YPD plates. Cells were grown at 24°C for 3 days. Mutants that could rescue the taf14[-NT] slow growth phenotype are marked with arrows. The slight growth advantage seen for the H103A/D104A mutant in this figure was not reproducible
Taf14 interaction with proteins related to transcription and chromatin
To determine the region of Taf14 protein required for the physical interaction of Taf14 with RSC, Swi/Snf, Ino80, NuA3, TFIIF and TFIID, we performed small-scale interaction assays.
Proteins from each complex were FLAG- or HA-tagged in their chromosomal loci. Tagged proteins included: (1) Snf5, thought to coordinate the assembly and nucleosome-remodeling activities of Swi/Snf (Geng et al. 2001); (2) Sth1, the DNA-dependent ATPase of RSC (Du et al. 1998); (3) Sas3, the histone acetyl transferase catalytic subunit of NuA3 (John et al. 2000; Howe et al. 2002); (4) Ino80, the DNA-dependent ATPase of the Ino80 complex (Shen et al. 2003a, b); (5) Tfg1, the largest subunit of TFIIF that functions in transcription initiation and elongation (Fish et al. 2006; Khaperskyy et al. 2008); (6) Taf1, the largest subunit of TFIID, with histone acetyltransferase activity (Mizzen et al. 1996) and involved in promoter binding and G1/S progression (Tansey and Herr 1997); (7) Taf2, also a TFIID subunit, stabilizes TFIID binding to the core promoter (Walker et al. 1997; Martinez et al. 1998). The addition of the tag to these proteins did not affect the growth of these strains, suggesting that the tag did not impair function (data not shown).
Full length Taf14, taf14[CT] and taf14[YEATS] were TAP-tagged and expressed from plasmids under the PGK promoter (Fig. 1c, right panel). Each strain carrying one of the seven FLAG- or HA-tagged proteins was transformed with each of the three different Taf14-encoding plasmids. The incorporation of the Taf14 constructs into the different nuclear complexes was tested by immunoprecipitation experiments using the TAP tag on Taf14, followed by Western blotting with FLAG or HA antibodies to detect the protein in the complexes (Fig. 7).
Interaction assay between Taf14 and different nuclear complexes. The same amount of soluble protein from extracts of taf14Δ/FLAG-Tfg1, taf14Δ/FLAG-Sas3, taf14Δ/FLAG-Taf1, taf14Δ/FLAG-Taf2, taf14Δ/HA-Snf5, and taf14Δ/FLAG-Sth1 strains containing plasmids expressing either TAP-tagged wild-type Taf14 (pJS40), TAP-taf14[CT] (pJS41), or the TAP tag alone (pVV223) were processed as described in “Materials and methods”. The Western-blotted membranes were incubated with anti-FLAG M2 or anti-HA to visualize the protein representing each complex. The protein A present in the TAP tag allowed for the visualization of all versions of Taf14 (black arrows). Red stars mark the FLAG and HA-tagged proteins. (CT) represents the TAP-taf14[CT]. (FL) represents the TAP-Taf14. + indicates a cell extract of that strain without expression of Taf14 and was used as a positive control. − represents the negative control expressing the TAP tag alone
Both full length Taf14 and mutant taf14[CT] were associated with Swi/Snf, RSC, NuA3, TFIIF and TFIID (Fig. 7). We could draw no conclusions about the association with Ino80 as it could not be clearly detected in any of the five pull down experiments performed. Additionally, in the case of Swi/Snf, the Taf14[CT] was not always well incorporated and sometimes taf14[CT] appeared to immunoprecipitate more Taf2 than the full length protein itself. These results prevented us from making quantitative conclusions about the associations. Taf14[YEATS] was convincingly (three out of three experiments performed, data not shown) associated with Taf1, one of the TFIID components tested. However, in general the TAP-taf14[YEATS] peptide did not immunoprecipitate successfully and was not readily detected in association with the tested proteins even when the polyclonal anti-Taf14 antibody was used to maximize detection of the YEATS domain. Nevertheless, it is clear that the YEATS domain is not essential for Taf14’s association with Swi/Snf, RSC, NuA3, TFIIF or TFIID.
Discussion
The work presented here explores the cellular function of Taf14 and studies the role of its conserved YEATS domain by structure–function and mutational analysis of various regions of Taf14. This protein is a member of at least six nuclear proteins complexes that are involved in transcription and chromatin modification (RSC, SWI/SNF, Ino80, NuA3, TFIIF, TFIID and possibly Mediator) (Henry et al. 1994; Poon et al. 1995; Cairns et al. 1996; Du et al. 1998; Shen et al. 2003a, b; Kabani et al. 2005). Its deletion renders cells sensitive to high temperature and osmolarity, causes deficiency in both constitutive and induced transcription and leads to altered actin cytoskeleton (Welch et al. 1993; Henry et al. 1994; Kimura and Ishihama 2004). The associations of Taf14 with different complexes and the resulting phenotypes of the disruption strain indicate that this protein might facilitate many nuclear processes in the cell, or that it links different processes to a general common mechanism. Exploring the cellular function of Taf14 and determining the role of its conserved YEATS domain was the focus of this work. We show that the YEATS domain has a negative role in cell growth, and it is not needed to bind the nuclear complexes that contain Taf14 nor to complement the Taf14 synthetic lethality with TFIIS when both proteins are absent from the cell.
In S. cerevisiae, there are three proteins that contain the YEATS domain (Taf14, Yaf9, Sas5), and cells require at least one of these proteins to survive (Zhang et al. 2004). For this reason, when Taf14 deletion mutations in the YEATS domain were constructed, the initial expectation was that the YEATS domain would stimulate cell growth. However, contrary to expectations, we found that in all the cases tested, the C-terminal domain of Taf14 (taf14[CT]) was sufficient to complement all of the phenotypes of a taf14 deletion strain. Furthermore, the YEATS domain deletion provides a growth advantage compared to wild-type cells under a variety of conditions (Table 3; Fig. 3). In addition, a deletion of just the first 61 amino acids in the Taf14 YEATS domain (the taf14[-NT] construct) creates a strain with a growth disadvantage compared to a full deletion of the YEATS domain (aa 6–113, the taf14[CT] construct). These results, coupled with the extremely slow growth of cells expressing the YEATS domain alone, strongly suggest that the YEATS domain of Taf14 has an important negative regulatory function. We show that the YEATS domain is not needed for the association of Taf14 with any of the tested nuclear complexes (Fig. 7). This is consistent with the observation that only the coiled-coil C-terminal domain of Yaf9 is required for its interaction with Esa1, the histone acetyltransferase catalytic subunit of, NuA4, and not its YEATS domain (Zhang et al. 2004). In the same way, the non-YEATS domain (also a coiled-coil region) of GAS41 is required for interaction with the human TIP60 and SRCAP complexes (Park and Roeder 2006). Together, these observations suggest that the hypothesized negative effect of the YEATS domain would not be transmitted via the complexes themselves. While it is still formally possible that there may be interactions between Taf14 and other subunits of these complexes that are affected by deletion of the YEATS domain, we prefer a model where the YEATS domain functions outside of the complexes themselves, possibly by binding and modifying growth regulators through a specific binding site on its surface.
Compatible with the possibility that the YEATS domain contains such a regulatory motif, site-specific mutations (W81A, F84A) in the remaining YEATS residues of taf14[-NT] conferred a growth advantage to cells (Fig. 4b). These residues are predicted to reside in one of the loops of a β-sandwich, likely on the protein surface. Thus, this region might be critical for interaction with a protein or proteins in the cellular pathway in which the YEATS domain of Taf14 plays a role. The growth rescue of this sensitized mutant via these point mutations argues against the possible misfolding of the construct. This increased growth is not observed when the mutations are incorporated into the full length TAF14, but the reason for this result could be that we cannot detect increased growth over that of an already healthy strain. Since the YEATS domain is not required for interaction with at least five of the six complexes in which Taf14 is a subunit, these residues (W81 and F84) are unlikely to be necessary for interaction with the complexes themselves. It seems more likely that these residues are highlighting a regulatory region interacting with other proteins important for Taf14 function. Further, it seems plausible that the YEATS domain is carrying out a function separate from that of the C-terminal domain of Taf14.
Another feature of TAF14 is that its disruption makes the gene encoding TFIIS (DST1) essential (Fish and Kane 2002). TFIIS is a transcript elongation factor that allows RNA polymerase II to recover from the backtracked state and read through blocks to elongation such as the nucleosome (Kireeva et al. 2005). The synthetic lethality between TAF14 and DST1 appears to be mediated through TFIIF, both an initiation and an elongation factor in transcription (Fish et al. 2006). Both taf14[CT] and taf14[-NTW81A-F84A] complement the synthetic lethality in a taf14Δdst1Δ strain as effectively as wild-type Taf14 (Figs. 5, 6). In addition, three other mutants (K124A, E129A/E130A, or K133A) in the taf14[-NT] construct complement as well as wild-type Taf14 (Fig. 6). Interestingly, none of these residues are located in the YEATS domain, and these mutants do not rescue the slow growth phenotype of the taf14[-NT] mutant in the haploid taf14Δ strain. Thus, since these mutant residues do rescue the taf14Δdst1Δ synthetic lethality, they are residues that are needed specifically for the TFIIS function. This result reinforces the likelihood that the YEATS domain and the C-terminal domain are involved in distinct functions.
Disruption of TAF14 also renders genes encoding subunits of SWI/SNF essential (Davie and Kane 2000), and mutant alleles in the two larger subunits of TFIIF are also synthetically lethal with taf14Δ (unpublished results). It is possible that more of this type of genetic interaction might be found if direct experiments were done with genes in the other Taf14 bearing-complexes as well. Alternatively, the defects seen in taf14 null mutants might be explained by disassembled complexes (one or more) or by assembled but functionally inactive complexes. For example, in a snf5Δ mutant, Swi/Snf is not well assembled leading to an inactive complex (Geng et al. 2001). In contrast, taf14Δ mutants still contain a stable Swi/Snf complex with ATPase activity (Cairns et al. 1996) and yeast TFIIF without Taf14 is still active in vitro (Ziegler et al. 2003). In any case, Taf14’s presence in the six nuclear complexes suggests that it might perform a broad function designed to enlist these complexes in particular processes or locations, perhaps at particular times. Clearly Taf14 is not needed in all circumstances as the complexes can function in its absence at the permissive temperature.
All these observations, together with several others found in the literature (see below), lead to plausible and testable hypotheses. First, it has been reported that the taf14Δ cells have elongated buds and actin hyperpolarization (Henry et al. 1994), phenotypes often associated with cell cycle regulating proteins (Welch and Drubin 1994). Second, cells lacking Taf14 spend much longer in G1 than do wild-type cells (Erlich et al. 2008). Third, Taf14 is essential for the G2/M transition in cdc40Δ cells (Dahan and Kupiec 2004), important for cell cycle progression. Fourth, Taf14 is related to two YEATS domain containing mammalian proteins Gas41 and AF-9, both implicated in spindle function. Fifth, Taf14 is a subunit of RSC, a chromatin remodeling complex also essential for cell cycle progression (Du et al. 1998; Yamada et al. 2008). Sixth, the YEATS domain containing protein Yaf9 is important for the deposition of H2A.Z (Le Masson et al. 2003), a histone variant required for proper timing of transcription of cell cycle regulatory proteins. Along this line of thought, the results reported here also suggest a role for Taf14 and its YEATS domain during the cell cycle. While the doubling times of cells containing taf14[-NT] and YEATS constructs are longer than for cells containing the normal Taf14 or the taf14[CT] constructs, the lag time prior to starting cell division is profoundly increased, especially for cells containing the YEATS construct (Table 3). This observation suggests a problem in reentering the cell cycle after stationary phase, perhaps from G0 to late G1 where cells commit to DNA replication. Additionally, the taf14[CT] mutant grows better than wild type in the presence of benomyl, a microtubule destabilizing drug (Fig. 3), as if the absence of the YEATS domain actually stabilized nuclear microtubules.
In summary, expression of only the YEATS domain makes the cells sick: a partial YEATS domain in Taf14 imparts slowed growth; a full deletion of the YEATS domain of Taf14 generates healthy cells. Additional experiments will be necessary to find the YEATS-binding regulator, to test, in a quantitative way, what fraction of each nuclear complex has Taf14 and the mutant versions bound, and to determine the importance of the YEATS domain for Taf14’s function in the nucleus.
References
Adler HT, Chinery R, Wu DY, Kussick SJ, Payne JM, Fornace AJ, Tkachuk DC (1999) Leukemic HRX fusion proteins inhibit GADD34-induced apoptosis and associate with the GADD34 and hSNF5/INI1 proteins. Mol Cell Biol 19:7050–7060
Baek HJ, Kang YK, Roeder RG (2006) Human mediator enhances basal transcription by facilitating recruitment of transcription factor IIB during preinitiation complex assembly. J Biol Chem 281:15172–15181
Buratowski S, Hahn S, Guarente L, Sharp PA (1989) Five intermediate complexes in transcription initiation by RNA polymerase II. Cell 56:549–561
Cairns BR (2005) Chromatin remodeling complexes: strength in diversity, precision through specialization. Curr Opin Genet Dev 15:185–190
Cairns BR, Henry NL, Kornberg RD (1996) TFG/TAF30/ANC1, a component of the yeast SWI/SNF complex that is similar to the leukemogenic proteins ENL and AF-9. Mol Cell Biol 16:3308–3316
Clark-Adams CD, Norris D, Osley MA, Fassler JS, Winston F (1988) Changes in histone gene dosage alter transcription in yeast. Genes Dev 2:150–159
Cler E, Papai G, Schultz P, Davidson I (2009) Recent advances in understanding the structure and function of general transcription factor TFIID. Cell Mol Life Sci 66:2123–2134
Conaway R, Conaway J (2009) The INO80 chromatin remodeling complex in transcription, replication and repair. Trends Biochem Sci 34:71–77
Conaway JW, Shilatifard A, Dvir A, Conaway RC (2000) Control of elongation by RNA polymerase II. Trends Biochem Sci 25:375–380
Corral J, Lavenir I, Impey H, Warren AJ, Forster A, Larson TA, Bell S, McKenzie AN, King G, Rabbitts TH (1996) An Mll-AF9 fusion gene made by homologous recombination causes acute leukemia in chimeric mice: a method to create fusion oncogenes. Cell 85:853–861
Costelloe T, Fitzgerald J, Murphy NJ, Flaus A, Lowndes NF (2006) Chromatin modulation and the DNA damage response. Exp Cell Res 312:2677–2686
Dahan O, Kupiec M (2004) The Saccharomyces cerevisiae gene CDC40/PRP17 controls cell cycle progression through splicing of the ANC1 gene. Nucleic Acids Res 32:2529–2540
Davie JK, Kane CM (2000) Genetic interactions between TFIIS and the Swi-Snf chromatin-remodeling complex. Mol Cell Biol 20:5960–5973
Dechassa ML, Zhang B, Horowitz-Scherer R, Persinger J, Woodcock CL, Peterson CL, Bartholomew B (2008) Architecture of the SWI/SNF-nucleosome complex. Mol Cell Biol 28:6010–6021
Du J, Nasir I, Benton BK, Kladde MP, Laurent BC (1998) Sth1p, a Saccharomyces cerevisiae Snf2p/Swi2p homolog, is an essential ATPase in RSC and differs from Snf/Swi in its interactions with histones and chromatin-associated proteins. Genetics 150:987–981005
Eberharter A, John S, Grant PA, Utley RT, Workman JL (1998) Identification and analysis of yeast nucleosomal histone acetyltransferase complexes. Methods 15:315–321
Erlich R, Fry R, Begley T, Daee D, Lahue R, Samson L (2008) Anc1, a protein associated with multiple transcription complexes, is involved in postreplication repair pathway in S. cerevisiae. PLoS ONE 3:e3717
Fan HY, Cheng KK, Klein HL (1996) Mutations in the RNA polymerase II transcription machinery suppress the hyperrecombination mutant hpr1 delta of Saccharomyces cerevisiae. Genetics 142:749–759
Fischer U, Heckel D, Michel A, Janka M, Hulsebos T, Meese E (1997) Cloning of a novel transcription factor-like gene amplified in human glioma including astrocytoma grade I. Hum Mol Genet 6:1817–1822
Fish R, Kane C (2002) Promoting elongation with transcript cleavage stimulatory factors. Biochim Biophys Acta 1577:287–307
Fish RN, Ammerman ML, Davie JK, Lu BF, Pham C, Howe L, Ponticelli AS, Kane CM (2006) Genetic interactions between TFIIF and TFIIS. Genetics 173:1871–1884
Flores O, Lu H, Killeen M, Greenblatt J, Burton ZF, Reinberg D (1991) The small subunit of transcription factor IIF recruits RNA polymerase II into the preinitiation complex. Proc Natl Acad Sci USA 88:9999–10003
Flores O, Lu H, Reinberg D (1992) Factors involved in specific transcription by mammalian RNA polymerase II. Identification and characterization of factor IIH. J Biol Chem 267:2786–2793
Geng F, Cao Y, Laurent BC (2001) Essential roles of Snf5p in Snf-Swi chromatin remodeling in vivo. Mol Cell Biol 21:4311–4320
Ghavi-Helm Y, Michaut M, Acker J, Aude JC, Thuriaux P, Werner M, Soutourina J (2008) Genome-wide location analysis reveals a role of TFIIS in RNA polymerase III transcription. Genes Dev 22:1934–1947
Gietz RD, Schiestl RH, Willems AR, Woods RA (1995) Studies on the transformation of intact yeast cells by the LiAc/SS-DNA/PEG procedure. Yeast 11:355–360
Guthrie T, Tsuji J, Wells J (1991) A synthetic pseudosubstrate peptide of protein kinase C inhibits the phorbol-12, 13-dibutyrate effect on permeabilized coronary artery smooth muscle. Mol Pharmacol 39:621–624
Harborth J, Weber K, Osborn M (2000) GAS41, a highly conserved protein in eukaryotic nuclei, binds to NuMA. J Biol Chem 275:31979–31985
He Q, Battistella L, Morse R (2008) Mediator requirement downstream of chromatin remodeling during transcriptional activation of CHA1 in yeast. J Biol Chem 283:5276–5286
Henry NL, Campbell AM, Feaver WJ, Poon D, Weil PA, Kornberg RD (1994) TFIIF-TAF-RNA polymerase II connection. Genes Dev 8:2868–2878
Howe L, Kusch T, Muster N, Chaterji R, Yates JR, Workman JL (2002) Yng1p modulates the activity of Sas3p as a component of the yeast NuA3 histone acetyltransferase complex. Mol Cell Biol 22:5047–5053
Imbalzano AN, Kwon H, Green MR, Kingston RE (1994) Facilitated binding of TATA-binding protein to nucleosomal DNA. Nature 370:481–485
John S, Howe L, Tafrov ST, Grant PA, Sternglanz R, Workman JL (2000) The something about silencing protein, Sas3, is the catalytic subunit of NuA3, a yTAF(II)30-containing HAT complex that interacts with the Spt16 subunit of the yeast CP (Cdc68/Pob3)-FACT complex. Genes Dev 14:1196–1208
Kabani M, Michot K, Boschiero C, Werner M (2005) Anc1 interacts with the catalytic subunits of the general transcription factors TFIID and TFIIF, the chromatin remodeling complexes RSC and INO80, and the histone acetyltransferase complex NuA3. Biochem Biophys Res Commun 332:398–403
Khaperskyy D, Ammerman M, Majovski R, Ponticelli A (2008) Functions of Saccharomyces cerevisiae TFIIF during transcription start site utilization. Mol Cell Biol 28:3757–3766
Kim YJ, Bjorklund S, Li Y, Sayre MH, Kornberg RD (1994) A multiprotein mediator of transcriptional activation and its interaction with the C-terminal repeat domain of RNA polymerase II. Cell 77:599–608
Kimura M, Ishihama A (2004) Tfg3, a subunit of the general transcription factor TFIIF in Schizosaccharomyces pombe, functions under stress conditions. Nucleic Acids Res 32:6706–6715
Kingston RE, Narlikar GJ (1999) ATP-dependent remodeling and acetylation as regulators of chromatin fluidity. Genes Dev 13:2339–2352
Kireeva M, Hancock B, Cremona G, Walter W, Studitsky V, Kashlev M (2005) Nature of the nucleosomal barrier to RNA polymerase II. Mol Cell 18:97–108
Krogan NJ, Kim M, Tong A, Golshani A, Cagney G, Canadien V, Richards DP, Beattie BK, Emili A, Boone C, Shilatifard A, Buratowski S, Greenblatt J (2002) RNA polymerase II elongation factors of Saccharomyces cerevisiae: a targeted proteomics approach. Mol Cell Biol 22:6979–6992
Krogan NJ, Kim M, Tong A, Golshani A, Cagney G, Canadien V, Richards DP, Beattie BK, Emili A, Boone C, Shilatifard A, Buratowski S, Greenblatt J (2003) Methylation of histone H3 by Set2 in Saccharomyces cerevisiae is linked to transcriptional elongation by RNA polymerase II. Mol Cell Biol 23:4207–4218
Kunkel TA, Roberts JD, Zakour RA (1987) Rapid and efficient site-specific mutagenesis without phenotypic selection. Methods Enzymol 154:367–382
Kushnirov VV (2000) Rapid and reliable protein extraction from yeast. Yeast 16:857–860
Le Masson I, Yu DY, Jensen K, Chevalier A, Courbeyrette R, Boulard Y, Smith MM, Mann C (2003) Yaf9, a novel NuA4 histone acetyltransferase subunit, is required for the cellular response to spindle stress in yeast. Mol Cell Biol 23:6086–6102
Majovski RC, Khaperskyy DA, Ghazy MA, Ponticelli AS (2005) A functional role for the switch 2 region of yeast RNA polymerase II in transcription start site utilization and abortive initiation. J Biol Chem 280:34917–34923
Malagon F, Tong AH, Shafer BK, Strathern JN (2004) Genetic interactions of DST1 in Saccharomyces cerevisiae suggest a role of TFIIS in the initiation–elongation transition. Genetics 166:1215–1227
Martinez E, Ge H, Tao Y, Yuan C, Palhan V, Roeder R (1998) Novel cofactors and TFIIA mediate functional core promoter selectivity by the human TAFII150-containing TFIID complex. Mol Cell Biol 18:6571–6583
Mizzen CA, Yang XJ, Kokubo T, Brownell JE, Bannister AJ, Owen-Hughes T, Workman J, Wang L, Berger SL, Kouzarides T, Nakatani Y, Allis CD (1996) The TAF(II)250 subunit of TFIID has histone acetyltransferase activity. Cell 87:1261–1270
Munnia A, Schutz N, Romeike BF, Maldener E, Glass B, Maas R, Nastainczyk W, Feiden W, Fischer U, Meese E (2001) Expression, cellular distribution and protein binding of the glioma amplified sequence (GAS41), a highly conserved putative transcription factor. Oncogene 20:4853–4863
Myers LC, Kornberg RD (2000) Mediator of transcriptional regulation. Annu Rev Biochem 69:729–749
Narlikar GJ, Fan H-Y, Kingston RE (2002) Cooperation between complexes that regulate chromatin structure and transcription. Cell 108:475–487
Neely KE, Workman JL (2002) Histone acetylation and chromatin remodeling: which comes first? Mol Genet Metab 76:1–5
Park JH, Roeder RG (2006) GAS41 is required for repression of the p53 tumor suppressor pathway during normal cellular proliferation. Mol Cell Biol 26:4006–4016
Park J, Park E, Hur S, Kim S, Kwon J (2009) Mammalian SWI/SNF chromatin remodeling complexes are required to prevent apoptosis after DNA damage. DNA Repair (Amst) 8:29–39
Parnell T, Huff J, Cairns B (2008) RSC regulates nucleosome positioning at Pol II genes and density at Pol III genes. EMBO J 27:100–110
Poon D, Bai Y, Campbell AM, Bjorklund S, Kim YJ, Zhou S, Kornberg RD, Weil PA (1995) Identification and characterization of a TFIID-like multiprotein complex from Saccharomyces cerevisiae. Proc Natl Acad Sci USA 92:8224–8228
Puig O, Caspary F, Rigaut G, Rutz B, Bouveret E, Bragado-Nilsson E, Wilm M, Seraphin B (2001) The tandem affinity purification (TAP) method: a general procedure of protein complex purification. Methods 24:218–229
Robert F, Douziech M, Forget D, Egly JM, Greenblatt J, Burton ZF, Coulombe B (1998) Wrapping of promoter DNA around the RNA polymerase II initiation complex induced by TFIIF. Mol Cell 2:341–351
Sambrook J, Russell D (2001) Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory Press, Cold Spring Harbor
Sawin K, Tran P (2006) Cytoplasmic microtubule organization in fission yeast. Yeast 23:1001–1014
Schulze J, Wang A, Kobor M (2009) YEATS domain proteins: a diverse family with many links to chromatin modification and transcription. Biochem Cell Biol 87:65–75
Shen X, Mizuguchi G, Hamiche A, Wu C (2000) A chromatin remodelling complex involved in transcription and DNA processing. Nature 406:541–544
Shen X, Ranallo R, Choi E, Wu C (2003a) Involvement of actin-related proteins in ATP-dependent chromatin remodeling. Mol Cell 12:147–155
Shen X, Xiao H, Ranallo R, Wu W-H, Wu C (2003b) Modulation of ATP-dependent chromatin-remodeling complexes by inositol polyphosphates. Science 299:112–114
Shia W-J, Osada S, Florens L, Swanson SK, Washburn MP, Workman JL (2005) Characterization of the yeast trimeric-SAS acetyltransferase complex. J Biol Chem 280:11987–11994
Sikorski RS, Hieter P (1989) A system of shuttle vectors and yeast host strains designed for efficient manipulation of DNA in Saccharomyces cerevisiae. Genetics 122:19–27
Steger DJ, Utley RT, Grant PA, John S, Eberharter A, Cote J, Owen-Hughes T, Ikeda K, Workman JL (1998) Regulation of transcription by multisubunit complexes that alter nucleosome structure. Cold Spring Harb Symp Quant Biol 63:483–491
Sudarsanam P, Winston F (2000) The Swi/Snf family nucleosome-remodeling complexes and transcriptional control. Trends Genet 16:345–351
Sutton A, Shia WJ, Band D, Kaufman PD, Osada S, Workman JL, Sternglanz R (2003) Sas4 and Sas5 are required for the histone acetyltransferase activity of Sas2 in the SAS complex. J Biol Chem 278:16887–16892
Tansey WP, Herr W (1997) TAFs: guilt by association? Cell 88:729–732
Van Mullem V, Wery M, De Bolle X, Vandenhaute J (2003) Construction of a set of Saccharomyces cerevisiae vectors designed for recombinational cloning. Yeast 20:739–746
Walker SS, Shen WC, Reese JC, Apone LM, Green MR (1997) Yeast TAF(II)145 required for transcription of G1/S cyclin genes and regulated by the cellular growth state. Cell 90:607–614
Wang AY, Schulze JM, Skordalakes E, Gin JW, Berger JM, Rine J, Kobor MS (2009) Asf1-like structure of the conserved Yaf9 YEATS domain and role in H2A.Z deposition and acetylation. PNAS 106(51):21573–21578
Welch MD, Drubin DG (1994) A nuclear protein with sequence similarity to proteins implicated in human acute leukemias is important for cellular morphogenesis and actin cytoskeletal function in Saccharomyces cerevisiae. Mol Biol Cell 5:617–632
Welch MD, Vinh DB, Okamura HH, Drubin DG (1993) Screens for extragenic mutations that fail to complement act1 alleles identify genes that are important for actin function in Saccharomyces cerevisiae. Genetics 135:265–274
Yamada K, Hirota K, Mizuno K, Shibata T, Ohta K (2008) Essential roles of Snf21, a Swi2/Snf2 family chromatin remodeler, in fission yeast mitosis. Genes Genet Syst 83:361–372
Zhang H, Richardson DO, Roberts DN, Utley R, Erdjument-Bromage H, Tempst P, Cote J, Cairns BR (2004) The Yaf9 component of the SWR1 and NuA4 complexes is required for proper gene expression, histone H4 acetylation, and Htz1 replacement near telomeres. Mol Cell Biol 24:9424–9436
Zhang C, Zobeck KL, Burton ZF (2005) Human RNA polymerase II elongation in slow motion: role of the TFIIF RAP74 alpha1 helix in nucleoside triphosphate-driven translocation. Mol Cell Biol 25:3583–3595
Ziegler LM, Khaperskyy DA, Ammerman ML, Ponticelli AS (2003) Yeast RNA polymerase II lacking the Rpb9 subunit is impaired for interaction with transcription factor IIF. J Biol Chem 278:48950–48956
Acknowledgments
This work was supported by NSF Grant MCB:0316224 grant to C.M.K., Studienstiftung des deutschen Volkes (German National Academic Foundation) to J.M.S., and a Postdoctoral fellowship from the Basque Government to A.R.M.
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by M. Collart.
Electronic supplementary material
Below is the link to the electronic supplementary material.
438_2010_523_MOESM1_ESM.tif
Supplementary Fig. S1 Effect of taf14 mutants on a wild type strain. Same amounts of CH1305 containing TAP-TAF14 (pJS40), TAP-taf14[CT] (pJS41) and TAP-taf14[YEATS] (pJS42) on plasmids were grown onto SC-URA plates. The empty vector (pVV223) was used as a negative control (TIFF 1022 kb)
438_2010_523_MOESM2_ESM.tif
Supplementary 2 Fig. S2 Expression level of the different Taf14 mutant constructs in taf14Δ cells. Expression of different Taf14[-NT] mutants. Equal amounts of cell extract were resolved on a 10% SDS-Laemmli protein gel and transferred to a nitrocellulose membrane for western blotting. Proteins were detected with Polyclonal Taf14 antibody. W81A/F84A mutant, in red, is the only one showing an improved growth (Fig. 4) (TIFF 1796 kb)
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Schulze, J.M., Kane, C.M. & Ruiz-Manzano, A. The YEATS domain of Taf14 in Saccharomyces cerevisiae has a negative impact on cell growth. Mol Genet Genomics 283, 365–380 (2010). https://doi.org/10.1007/s00438-010-0523-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-010-0523-x