Abstract
Presence and possible functions of DNA methylation in plastid genomes of higher plants have been highly controversial. While a number of studies presented evidence for the occurrence of both cytosine and adenine methylation in plastid genomes and proposed a role of cytosine methylation in the transcriptional regulation of plastid genes, several recent studies suggested that at least cytosine methylation may be absent from higher plant plastid genomes. To test if either adenine or cytosine methylation can play a regulatory role in plastid gene expression, we have introduced cyanobacterial genes for adenine and cytosine DNA methyltransferases (methylases) into the tobacco plastid genome by chloroplast transformation. Using DNA cleavage with methylation-sensitive and methylation-dependent restriction endonucleases, we show that the plastid genomes in the transplastomic plants are efficiently methylated. All transplastomic lines are phenotypically indistinguishable from wild-type plants and, moreover, show no alterations in plastid gene expression. Our data indicate that the expression of plastid genes is not sensitive to DNA methylation and, hence, suggest that DNA methylation is unlikely to be involved in the transcriptional regulation of plastid gene expression.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
DNA methylation is ubiquitously present in both prokaryotes and eukaryotes and affects many cellular processes. Chromosomal DNA in eukaryotes is mainly methylated at cytosine residues by methyltransferases converting cytosine to 5-methylcytosine. Cytosine methylation controls chromatin structure, which in turn can affect transcriptional regulation of gene expression and epigenetic inheritance (for review, see e.g., Fuks 2005; Richards 2006).
In addition to cytosine methylation, prokaryotic genomes also exhibit adenine methylation. Many prokaryotic DNA methyltransferases are part of restriction-modification systems, which form a defense mechanism against invasion by foreign DNA, such as bacteriophage genomes. In addition, bacteria possess solitary DNA methyltransferases, such as the adenine methyltransferase Dam and the cytosine methyltransferase Dcm. For example, in Escherichia coli, almost all GATC tetranucleotide sequences in the genome contain N6-methyladenine generated by the action of Dam methylase. While Dam methylation is known to be involved in the control of a variety of cellular processes, including DNA replication, DNA repair, transcriptional regulation (e.g., by modulating interactions with DNA-binding proteins; Sternberg 1985) and conjugation (reviewed, e.g., in Lobner-Olesen et al. 2005; Wion and Casadesús 2006), the physiological functions of Dcm methylation are still unclear.
Occurrence and function of DNA methylation in plastid (chloroplast) genomes have been a matter of contention for more than 20 years. The first evidence for cytosine methylation in chloroplast DNA was obtained in the unicellular green alga Chlamydomonas reinhardtii (Sager et al. 1984). In Chlamydomonas, there are ‘male’ and ‘female’ cells which are morphologically indistinguishable and referred to as mating type + (mt+, ‘female’) and mating type – (mt–, ‘male’). In mt+ gametes, the chloroplast DNA is methylated by a cytosine DNA methyltransferase (Nishiyama et al. 2002, 2004). Successful mating triggers a zygotic maturation program, which leads to the selective degradation of the chloroplast DNA from the mt– parent (by a specific endonuclease; Nishimura et al. 2002), thus resulting in uniparental inheritance of the maternal chloroplast DNA (Nishimura et al. 1999). It was speculated that the difference in DNA methylation could be responsible for the selective degradation of the mt– chloroplast genomes and survival of the mt+ genomes. However, recent studies indicate that DNA methylation may not be required for protection of mt+ plastid genomes in the zygote and instead, may control the relative rates of plastid genome replication in mt– and mt+ cells (Umen and Goodenough 2001).
A number of studies reported the occurrence of cytosine methylation also in plastid genomes of higher plants, including sycamore (Acer pseudoplatanus; Ngernprasirtsiri et al. 1988b; Ngernprasirtsiri et al. 1990), tomato (Solanum lycopersicum; Ngernprasirtsiri et al. 1988a; Kobayashi et al. 1990) and maize (Zea mays; Ngernprasirtsiri et al. 1989b; Gauly and Kössel 1989). Cytosine methylation was evidenced by chromatographic detection of 5-methylcytosine (Ngernprasirtsiri et al. 1988b; Ngernprasirtsiri et al. 1990; Ngernprasirtsiri et al. 1989b) and by cleavage with pairs of isoschizomeric restriction endonucleases, of which one was methylation-sensitive and the other methylation insensitive (Ngernprasirtsiri et al. 1988a, b; Gauly and Kössel 1989; Ngernprasirtsiri et al. 1989a, b). Based on detected difference in cytosine methylation between chloroplasts and amyloplasts, which correlated with differences in transcript abundance, it was suggested that DNA methylation plays a key role in the transcriptional regulation of plastid genes in different tissues and plastid types (Ngernprasirtsiri et al. 1988b; Ngernprasirtsiri et al. 1990). In tomato plastids, DNA methylation was reported to occur around lowly expressed genes during fruit development and chloroplast-to-chromoplast conversion (Ngernprasirtsiri et al. 1988a; Kobayashi et al. 1990). In maize, differences in plastid DNA methylation in mesophyll versus bundle sheath cells were reported and interpreted as a regulatory mechanism involved in the differential expression of photosynthesis genes in C4 plants (Ngernprasirtsiri et al. 1989b). In contrast, other investigations in tomato chloroplasts and chromoplasts (Marano and Carrillo 1991) and barley plastids during leaf development (Tomas et al. 1992) failed to detect cytosine methylation in plastid DNA and have cast considerable doubt on the occurrence and physiological significance of cytosine methylation in plastids (Fojtová et al. 2001).
In addition to 5-methylcytosine, the occurrence of substantial amounts of N6-methyladenine in plastid (amyloplast) DNA has also been reported (Ngernprasirtsiri et al. 1988b), but was not further investigated. In order to clarify the role of both adenine and cytosine DNA methylation in plastids and its possible functional significance for the transcriptional regulation of plastid gene expression, we have introduced cyanobacterial adenine and cytosine methyltransferase genes into the plastid genome of tobacco plants. We report here that neither adenine nor cytosine methylation of the plastid DNA leads to detectable effects on transcript abundance of any gene in the plastid genome suggesting that DNA methylation, if it occurs in plastids, is unlikely to play a role in transcriptional regulation.
Materials and methods
Plant material, growth conditions and phenotypical assays
Sterile tobacco (Nicotiana tabacum cv. Petit Havana) plants were grown on agar-solidified MS medium supplemented with 30 g/l sucrose (Murashige and Skoog 1962). Homoplasmic transplastomic lines were rooted and propagated on the same medium. Rooted homoplasmic plants were transferred to soil and grown to maturity under standard greenhouse conditions. Transgene inheritance was assayed by germination of seeds on spectinomycin-containing MS medium (500 mg/l spectinomycin). Growth tests under different light conditions were performed by raising wild-type and transplastomic plants from seeds in soil at 26°C under the following light intensities: 50, 200, 600 and 1,100 μmol m−2 s−1.
Construction of chloroplast transformation vectors
The adenine (dam; slr1803) and cytosine DNA methyltransferase (M.Ssp6803I; slr0214; Scharnagl et al. 1998) genes from Synechocystis sp. strain PCC 6803 (Kaneko et al. 1996) were cloned as NcoI/XbaI fragments into plastid transformation vector pRB96 (Wurbs et al. 2007), a derivative of vector pRB94 (Ruf et al. 2001). To this end, the coding regions were amplified from Synechocystis total DNA introducing a 5′ NcoI site and a 3′ XbaI site with the PCR primers. The following synthetic oligonucleotides were used as primers (introduced restriction sites: underlined; start and stop codons: bold):
slr1803
- P5′dam:
-
5′ AAAACC ATG GTGAAAATTAAACAAGCTATTGC 3′
- P3′dam:
-
5′ AAAAT CTA GAGGCGATGGTAATTCACAACC 3′
slr0214
- P5′dcm:
-
5′AAAACC ATG GCCAGACCCATTGCC 3′
- P3′dcm:
-
5′ AAAAT CTA GAGGGAATGGGATTTGGCACATA 3′
Correctness of cloning and PCR amplification was verified by DNA sequencing of both genes.
Plastid transformation
Young leaves from sterile tobacco plants were bombarded with plasmid DNA-coated 0.6 µm gold particles (BioRad, Munich, Germany) using a biolistic gun (PDS1000He; BioRad). Primary spectinomycin-resistant lines were selected on plant regeneration medium containing 500 mg/l spectinomycin (Svab et al. 1990; Svab and Maliga 1993). Double resistance tests on medium containing both spectinomycin and streptomycin (500 mg/l each; Bock 2001) eliminated spontaneous spectinomycin-resistant mutants and identified true plastid transformants. For each transgene, several independent transplastomic lines were subjected to three additional rounds of regeneration on spectinomycin-containing regeneration medium to obtain homoplasmic tissue.
Isolation of nucleic acids and DNA gel blot analyses
Total cellular DNA was isolated from fresh leaf tissue by a cetyltrimethylammoniumbromide (CTAB)-based method (Doyle and Doyle 1990). After RNase A digestion, phenol/chloroform extraction and ethanol precipitation, DNA samples were digested with restriction enzymes, separated by gel electrophoresis on 0.8% agarose gels, and transferred onto Hybond nylon membranes (GE Healthcare, Buckinghamshire, UK) by capillary blotting. Hybridizations were performed at 65°C in Rapid-Hyb buffer (GE Healthcare) following the manufacturer’s protocol. Total cellular RNA was extracted using the peqGOLD TriFast reagent (Peqlab GmbH, Erlangen, Germany). RNA samples for cDNA synthesis were further purified by treatment with RNase-free DNase I (Roche, Mannheim, Germany).
Microarray analysis
Plastid transcript abundance was analyzed using an oligonucleotide microarray covering all genes and reading frames in Solanaceous plastid genomes (Kahlau and Bock 2008). 100 pg HPLC-purified oligonucleotide was printed per spot onto SuperEpoxy 2 DNA Substrates (ArrayIt, Sunnyvale, USA) using a Piezorray spotter (Perkin-Elmer, Waltham, USA). Labeled cDNA probes for microarray analyses were generated by reverse transcription of 10 μg total RNA spiked with 2 μl of reference RNAs (Lucidea Universal ScoreCard, GE Healthcare) using the SuperScript® Indirect cDNA Labeling Kit (Invitrogen, Carlsbad, USA) and random hexanucleotide primers. The fluorescent dye Cy3 (GE Healthcare) was incorporated into the cDNA according to the manufacturer’s instructions (Invitrogen). Sample preparation and hybridization experiments were performed as described previously (Kahlau and Bock 2008). Following hybridization and washing, the slides were dried by centrifugation and scanned using a FLA-8000 scanner (Fujifilm). Hybridization signal intensities were determined using the GeneSpotter software (MicroDiscovery GmbH, Berlin Germany) and quantitated as described (Kahlau and Bock 2008).
Results
Introduction of cyanobacterial genes for DNA methyltransferases into the tobacco plastid genome
As chloroplasts are of cyanobacterial origin, we selected two DNA methyltransferase genes from the fully sequenced genome of the cyanobacterium Synechocystis sp. strain PCC6803 (Kaneko et al. 1996) for expression in plastids. The Synechocystis genome contains an open reading frame (slr1803) whose conceptual translation displays strong sequence similarity to the E. coli Dam methylase. The E. coli Dam methylase methylates adenine in the sequence 5′ GATC 3′ (Lobner-Olesen et al. 2005). The Synechocystis genome also contains at least one functional cytosine methyltransferase (slr0214). The gene product has been functionally characterized and shown to methylate cytosines in the recognition sequence of the restriction endonuclease PvuI (Scharnagl et al. 1998). This cytosine methylase has been termed M.Ssp6803I (Scharnagl et al. 1998) in accordance with the standard nomenclature for DNA methyltransferases, but for simplicity, it will be referred to here as Dcm for deoxycytosine deaminase, although it recognizes a different sequence than the classical Dcm methylase from E. coli.
Using the Synechocystis slr0214 and dam (slr1803) genes, we constructed two vectors for plastid transformation, pDcm and pDam (Fig. 1). The methylase transgenes are driven by the plastid atpI promoter, which contains multiple binding sites for both RNA polymerases present in higher plant plastids and is active in both photosynthetic and non-photosynthetic tissues (Miyagi et al. 1998). The coding regions of the two transgenes were integrated into plasmid pRB96 (Wurbs et al. 2007) as NcoI/XbaI-digested PCR products with the NcoI restriction site containing the start codon and the XbaI site containing the stop codon.
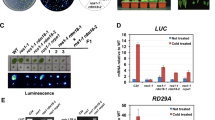
Introduction of cyanobacterial genes for DNA methylation enzymes into the tobacco plastid genome. a Map of the plastid genome (ptDNA) region to which the transgenes were targeted. Genes above the line are transcribed from the left to the right, genes below the line are transcribed in the opposite direction. Restrictions sites used for transgene insertion (SpeI) and RFLP analyses (BamHI) are indicated. b Map of the plastid transformation vectors pDam and pDcm constructed in this study. The vectors are based on the previously constructed plasmids pRB94 (Ruf et al. 2001) and pRB96 (Wurbs et al. 2007). The chimeric selectable marker gene aadA is driven by the ribosomal RNA operon promoter (Prrn; Svab and Maliga 1993). The DNA methyltransferase genes (dam and dcm) are transcribed from the atpI promoter (PatpI; Miyagi et al. 1998; Wurbs et al. 2007). The 3′ UTR of the aadA cassette is from the psbA gene (TpsbA; Svab and Maliga, 1993), the 3′ UTR of the methylase transgenes from the rps16 gene (Wurbs et al. 2007). c Southern blot analysis to confirm transgene integration by homologous recombination. DNA samples from the wild-type (Nt-Wt), three Nt-dam lines and three Nt-dcm plants (representing two independently generated transplastomic lines, Nt-dcm3 and Nt-dcm9) were digested with BamHI and hybridized to a psaB-derived probe. The size difference between the hybridizing bands in the wild-type and the transplastomic lines corresponds to the combined size of the two transgene cassettes (aadA + methylase gene). The faint wild-type-like band represents promiscuous chloroplast DNA in the nuclear genome as demonstrated earlier (Wurbs et al. 2007). M: DNA size marker (band sizes given in kb). d Seed assays to confirm homoplasmy of the transplastomic lines. Exemplarily, seed assays for one Nt-dam and one Nt-dcm plant are shown. As a control, the wild-type on medium with (+Spec) and without spectinomycin (–Spec) is also shown. Resistance to spectinomycin and lack of phenotypic segregation in the T1 generation demonstrate homoplasmy of the Nt-dam and Nt-dcm lines
The constructs were introduced into tobacco chloroplasts by biolistic plastid transformation (Svab and Maliga 1993). Several transplastomic lines were obtained for each construct after selection for spectinomycin resistance conferred by the chimeric aadA gene present in the transformation vectors (Fig. 1b). The lines will be subsequently referred to as Nt-dam and Nt-dcm, respectively (followed by the number of the transplastomic line and an uppercase letter denoting the individual plant analyzed; Figs. 1, 2). Successful chloroplast transformation was verified by tests for double resistance on medium containing two aminoglycoside antibiotics, spectinomycin and streptomycin (Svab and Maliga 1993; Bock 2001). Transgene integration was preliminarily confirmed by PCR assays using transgene-specific primers (data not shown). Plastid transformants were subsequently purified to homoplasmy by conducting additional regeneration cycles under antibiotic selection.
Analysis of DNA methylation in the plastid genome of Nt-dam and Nt-dcm plants. The same plant lines as in Fig. 1c were analyzed. a Test for adenine methylation with two isoschizomeric restriction enzymes differing in their sensitivity to adenine methylation in their recognition sequence (GATC). Whereas MboI cleaves the unmethylated sequence GATC but not the methylated sequence GmetATC, DpnI cleavage is dependent on Dam methylation and the unmethylated sequence is not cut (http://rebase.neb.com/rebase/rebase.html). The blot was hybridized to a 7.2 kb SalI/StuI fragment derived from the trnR(UCU)-rps2 region of the tobacco plastid genome (Shinozaki et al. 1986). Complete digestion of the plastid DNA with MboI in the wild-type (Nt-Wt) and all Nt-dcm lines indicates absence of Dam methylation. Strongly inhibited MboI cleavage in all Nt-dam lines suggests intense Dam methylation. Efficient cleavage of the plastid genome in all Nt-dam lines with the methylation-dependent restriction enzyme DpnI confirms efficient adenine methylation of GATC sequences in the Nt-dam transplastomic plants, but not the wild-type and the Nt-dcm plants. Minor differences between transplastomic lines obtained from the same construct are due to unequal loading. b Test for cytosine methylation by DNA digestion with the methylation-sensitive restriction endonuclease PvuI (Scharnagl et al. 1998). PvuI does not cut if the cytosines in its recognition sequence are methylated. Digestion of the plastid DNA with PvuI and hybridization to a 19.5 kb SalI fragment covering the ycf1-ndhB region of the tobacco plastid genome (Shinozaki et al. 1986) detects the expected four fragments of 24.5, 13, 5.5 and 2 kb in the wild-type (Nt-Wt). Note that the 5.5 kb fragment produces only a weak hybridization signal which is due to its limited coverage by the probe. Lack of any detectable DNA cleavage in the Nt-dcm plants suggests that the PvuI recognition sequences are efficiently methylated. M: DNA size marker (band sizes given in kb)
Homoplasmy of the transplastomic lines (i.e., absence of residual copies of the wild-type chloroplast genome) was first assessed by restriction fragment length polymorphism (RFLP) analysis (Fig. 1c). In addition to a strong band for the transplastomic fragment, RFLP analysis of all transplastomic lines also yielded a faint hybridization signal that corresponds in size to the restriction fragment from the wild-type plastid genome (Fig. 1c). Persistence of a wild type-like hybridization signal even after multiple rounds of selection and regeneration is often seen in plastid transformation experiments and usually is not caused by true heteroplasmy of the transplastomic lines, but rather by the presence of promiscuous plastid DNA in the nuclear genome (Ayliffe and Timmis 1992; Hager et al. 1999; Ruf et al. 2000). As previous plastid transformation experiments using the same vector backbone (pRB96; Wurbs et al. 2007) had revealed promiscuous DNA as the source of the wild type-like hybridization signal seen in Southern blot analyses, we strongly suspected our Nt-dam and Nt-dcm lines to be homoplasmic. To ultimately confirm homoplasmy of all transplastomic lines, seed assays were performed which provide a highly sensitive test to assess homoplasmy (Bock 2001; Maliga 2004). As expected, lack of segregation of the spectinomycin resistance in the T1 generation demonstrated homoplasmy (Fig. 1d) of the Nt-dam and Nt-dcm lines.
Analysis of DNA methylation of the chloroplast genome
Having generated homoplasmic transplastomic lines containing cyanobacterial DNA methyltransferase genes, we next wanted to test whether the introduced methyltransferase genes are expressed and indeed methylate the chloroplast DNA. To this end, we employed restriction endonucleases that distinguish methylated from unmethylated DNA. Two types of such restriction enzymes exist: (1) enzymes that cannot cut if their recognition sequence is methylated (methylation-sensitive enzymes) and (2) enzyme that can only cleave if their recognition sequence is methylated (methylation-dependent enzymes). While many methylation-sensitive restriction enzymes are known and commercially available, only very few methylation-dependent enzymes have been identified so far (http://rebase.neb.com/rebase/rebase.html). However, for the sequence methylated by the bacterial Dam methylase (GATC), with the enzyme DpnI such a methylation-dependent restriction enzyme is available.
To test for adenine methylation in the Nt-dam transplastomic lines, we employed the two isoschizomeric restriction enzymes MboI (methylation-sensitive enzyme) and DpnI (methylation-dependent enzyme). Interestingly, while the plastid DNA was completely digested with MboI in the wild-type and the Nt-dcm transplastomic plants, only very little digestion occurred in the Nt-dam lines (Fig. 2a), suggesting that the chloroplast genome was efficiently Dam methylated in the Nt-dam lines and therefore, digestion with MboI was largely inhibited. Conversely, digestion with the methylation-dependent restriction enzyme DpnI revealed efficient cleavage of the plastid genome in Nt-dam plants, but lack of cleavage in all Nt-dcm lines and the wild-type (Fig. 2a). This finding ultimately confirmed that the plastid DNA in the Nt-dam lines undergoes efficient adenine methylation.
To test whether cytosine methylation occurs in the Nt-dcm transplastomic lines, we used the methylation-sensitive restriction enzyme PvuI (Scharnagl et al. 1998). As PvuI is a six-base pair cutter and therefore does not cut as frequently in the chloroplast genome as MboI and DpnI, we used a large restriction fragment of the plastid genome as hybridization probe, which detects altogether four PvuI fragments covering more than 40 kb of the plastid genome (Fig. 2b). While the expected four fragments were generated in both the wild-type and the Nt-dam transplastomic lines, the plastid DNA from the Nt-dcm lines was completely resistant to cleavage with PvuI (Fig. 2b), indicating that the PvuI recognition sequences in the plastid genome of Nt-dcm lines are modified by cytosine methylation.
Insensitivity of plastid gene expression to DNA methylation
Having established that the two cyanobacterial DNA methyltransferases, when expressed as plastid transgenes, are enzymatically active and efficiently methylate the chloroplast genome, we next wanted to know, whether adenine or cytosine methylation of chloroplast DNA has any effect on plant growth and/or plastid gene expression.
When grown under a variety of different light and temperature regimes, all transplastomic Nt-dam and Nt-dcm lines were phenotypically indistinguishable from the wild-type control (data not shown) indicating that expression of the methyltransferase transgenes is phenotypically neutral and does not appreciably affect plant growth and development.
We also wanted to test the possibility that the methylation of the chloroplast DNA leads to alterations in plastid gene expression that do not translate into mutant phenotypes. If it has an effect on gene expression, DNA methylation should act at the level of transcription rather than on any of the later (DNA-independent) steps in gene expression. We, therefore, performed a set of microarray hybridization experiments (Kahlau and Bock 2008), in which we comparatively analyzed the transcript accumulation for all plastid genome-encoded genes and conserved open reading frames (Kahlau et al. 2006). These analyses revealed no detectable difference in mRNA accumulation for any plastid gene or open reading frame between wild-type plants, Nt-dam plants and Nt-dcm plants (Fig. 3 and data not shown). This suggests that plastid transcription is largely insensitive to adenine and cytosine methylation of the plastid DNA.
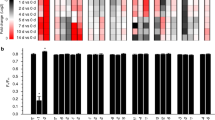
Analysis of expression of the plastid genome in Nt-dam and Nt-dcm transplastomic lines using the plastome microarray for Solanaceae (Kahlau and Bock 2008). a Spotting scheme of oligonucleotide probes on the microarray. The array contains all genes and conserved open reading frames (ycfs) present in Solanaceous plastid genomes and also a number of non-conserved open reading frames (orfs; for details see Kahlau and Bock 2008). Poorly conserved genes are covered by more than one oligonucleotide probe (indicated by the suffix ‘-S’ for Solanum-specific probe). NTC: negative control to assess non-specific background hybridization; Calib, Ratio, Utility: calibration and reference DNAs (artificial sequences corresponding to the Lucidea Universal ScoreCard; GE Healthcare; Kahlau and Bock, 2008). b Representative examples of microarray hybridization experiments for an Nt-dam and an Nt-dcm transplastomic line. No significant differences in expression of any plastid genome-encoded gene or open reading frame could be detected. Moreover, expression of the plastid genome in both Nt-dam and Nt-dcm was identical to the wild-type (not shown), indicating that neither adenine nor cytosine methylation affect plastid gene expression
Discussion
In this work, we have expressed two cyanobacterial DNA methyltransferase genes from the tobacco plastid genome. Both the adenine methyltransferase and the cytosine methyltransferase efficiently methylated their target sequences in the plastid genome. The observed insensitivity of plastid gene expression to both adenine and cytosine methylation of the plastid DNA makes it unlikely that DNA methylation is employed as a regulatory mechanism controlling expression of the plastid genome in response to environmental stimuli or developmental cues, as proposed earlier (Ngernprasirtsiri et al. 1988a, b, 1989b, 1990; Kobayashi et al. 1990).
Our digestions with methylation-sensitive restriction enzymes also revealed no evidence of adenine or cytosine methylation in the wild-type, at least not in the genomic regions analyzed here (Fig. 2). This is consistent with the absence of genes for putative plastid-targeted DNA methyltransferases from the fully sequenced genome of Arabidopsis thaliana. However, at present, it cannot be entirely excluded that some low-level methylation occurs in specific regions of the plastid genome and/or in specific tissues, developmental stages or under certain environmental conditions. However, the sole fact that methylation of the plastid DNA does not affect transcription of any gene in the plastid genome suggests that DNA methylation, even if it occurs under certain conditions, has no significant impact on plastid gene expression in higher plants. This is consistent with a number of other reports that failed to detect cytosine methylation in plastid DNA (Marano and Carrillo 1991; Tomas et al. 1992; Fojtová et al. 2001). The absence of a biological function for cytosine methylation in plastids is also supported by a recent study in which expression of the mFokI methyltransferase in tobacco plastids and targeting of the E. coli Dcm methylase to plastids appeared to be phenotypically neutral (Jaffé et al. 2008).
Clearly, cytosine methylation does occur in the plastid DNA of the unicellular green alga C. reinhardtii (Umen and Goodenough 2001; Nishiyama et al. 2002). The chloroplast DNA in mt+ gametes is methylated by a cytosine DNA methyltransferase, whereas the chloroplast genome in mt– gametes remains unmethylated (Nishiyama et al. 2002, 2004). It was therefore speculated that the difference in DNA methylation could be causally responsible for the uniparentally maternal inheritance of the plastid DNA and the selective degradation of the chloroplast genomes in mt– chloroplasts. However, not all experimental findings are compatible with this idea (Umen and Goodenough 2001) and the precise physiological role of chloroplast DNA methylation in chloroplast inheritance in Chlamydomonas remains to be determined. Although the mechanisms of plastid inheritance are fundamentally different in C. reinhardtii and higher plants (Bock 2007), we have nonetheless tested, whether or not methylation of the plastid genome in Nt-dam and Nt-dcm plants influences plastid genome transmission. No change in plastid transmission was detectable in reciprocal crosses (data not shown) and the plastid genomes of Nt-dam and Nt-dcm plants were inherited uniparentally maternally.
In summary, the plastid genome of higher plants displays a remarkable insensitivity to both adenine and cytosine methylation. This makes it highly unlikely that adenine and/or cytosine methylation are employed for the regulation of plastid gene expression in higher plants.
References
Ayliffe MA, Timmis JN (1992) Tobacco nuclear DNA contains long tracts of homology to chloroplast DNA. Theor Appl Genet 85:229–238
Bock R (2001) Transgenic chloroplasts in basic research and plant biotechnology. J Mol Biol 312:425–438
Bock R (2007) Structure, function, and inheritance of plastid genomes. Top Curr Genet 19:29–63
Doyle JJ, Doyle JL (1990) Isolation of plant DNA from fresh tissue. Focus 12:13–15
Fojtová M, Kovarik A, Matyásek R (2001) Cytosine methylation of plastid genome in higher plants. Fact or artefact? Plant Sci 160:585–593
Fuks F (2005) DNA methylation and histone modifications: teaming up to silence genes. Curr Op Genet Dev 15:490–495
Gauly A, Kössel H (1989) Evidence for tissue-specific cytosine methylation of plastid DNA from Zea mays. Curr Genet 15:371–376
Hager M, Biehler K, Illerhaus J, Ruf S, Bock R (1999) Targeted inactivation of the smallest plastid genome-encoded open reading frame reveals a novel and essential subunit of the cytochrome b6f complex. EMBO J 18:5834–5842
Jaffé B, Kovács K, Andras C, Bódi Z, Liu Z, Fray RG (2008) Methylation of chloroplast DNA does not affect viability and maternal inheritance in tobacco and may provide a strategy towards transgene containment. Plant Cell Rep 27:1377–1384
Kahlau S, Bock R (2008) Plastid transcriptomics and translatomics of tomato fruit development and chloroplast-to-chromoplast differentiation: chromoplast gene expression largely serves the production of a single protein. Plant Cell 20:856–874
Kahlau S, Aspinall S, Gray JC, Bock R (2006) Sequence of the tomato chloroplast DNA and evolutionary comparison of solanaceous plastid genomes. J Mol Evol 63:194–207
Kaneko T, Sato S, Kotani H, Tanaka A, Asamizu E, Nakamura Y, Miyajima N, Hirosawa M, Sugiura M, Sasamoto S, Kimura T, Hosouchi T, Matsuno A, Muraki A, Nakazaki N, Naruo K, Okumura S, Shimpo S, Takeuchi C, Wada T, Watanabe A, Yamada M, Yasuda M, Tabata S (1996) Sequence analysis of the genome of the unicellular cyanobacterium Synechocystis sp. strain PCC6803. II. Sequence determination of the entire genome and assignment of potential protein-coding regions. DNA Res 3:109–136
Kobayashi H, Ngernprasirtsiri J, Akazawa T (1990) Transcriptional regulation and DNA methylation in plastids during transitional conversion of chloroplasts to chromoplasts. EMBO J 9:307–313
Lobner-Olesen A, Skovgaard O, Marinus MG (2005) Dam methylation: coordinating cellular processes. Curr Op Microbiol 8:154–160
Maliga P (2004) Plastid transformation in higher plants. Annu Rev Plant Biol 55:289–313
Marano MR, Carrillo N (1991) Chromoplast formation during tomato fruit ripening. No evidence for plastid methylation. Plant Mol Biol 16:11–19
Miyagi T, Kapoor S, Sugita M, Sugiura M (1998) Transcript analysis of the tobacco plastid operon rps2/atpI/H/F/A reveals the existence of a non-consensus type II (NCII) promoter upstream of the atpI coding sequence. Mol Gen Genet 257:299–307
Murashige T, Skoog F (1962) A revised medium for rapid growth and bio assays with tobacco tissue culture. Physiol Plant 15:473–497
Ngernprasirtsiri J, Kobayashi H, Akazawa T (1988a) DNA methylation occurred around lowly expressed genes of plastid DNA during tomato fruit development. Plant Physiol 88:16–20
Ngernprasirtsiri J, Kobayashi H, Akazawa T (1988b) DNA methylation as a mechanism of transcriptional regulation in nonphotosynthetic plastids in plant cells. Proc Natl Acad Sci USA 85:4750–4754
Ngernprasirtsiri J, Kobayashi H, Akazawa T (1989a) Transcriptional regulation and DNA methylation of nuclear genes for photosynthesis in nongreen plant cells. Proc Natl Acad Sci USA 86:7919–7923
Ngernprasirtsiri J, Chollet R, Kobayashi H, Sugiyama T, Akazawa T (1989b) DNA methylation and the differential expression of C4 photosynthesis genes in mesophyll and bundle sheath cells of greening maize leaves. J Biol Chem 264:8241–8248
Ngernprasirtsiri J, Kobayashi H, Akazawa T (1990) DNA methylation is a determinative element of photosynthesis gene expression in amyloplasts from liquid-cultured cells of sycamore. Cell Struct Function 15:285–293
Nishimura Y, Misumi O, Matsunaga S, Higashiyama T, Yokota A, Kuroiwa T (1999) The active digestion of uniparental chloroplast DNA in a single zygote of Chlamydomonas reinhardtii is revealed by using the optical tweezer. Proc Natl Acad Sci USA 96:12577–12582
Nishimura Y, Misumi O, Kato K, Inada N, Higashiyma T, Momoyama Y, Kuroiwa T (2002) An mt+ gamete-specific nuclease that targets mt− chloroplasts during sexual reproduction in C. reinhardtii. Genes Dev 16:1116–1128
Nishiyama R, Ito M, Yamaguchi Y, Koizumi N, Sano H (2002) A chloroplast-resident DNA methyltransferase is responsible for hypermethylation of chloroplast genes in Chlamydomonas maternal gametes. Proc Natl Acad Sci USA 99:5925–5930
Nishiyama R, Wada Y, Mibu M, Yamaguchi Y, Shimogawara K, Sano H (2004) Role of a nonselective de novo DNA methyltransferase in maternal inheritance of chloroplast genes in the green alga, Chlamydomonas reinhardtii. Genetics 168:809–816
Richards EJ (2006) Inherited epigenetic variation—revisiting soft inheritance. Nature Rev Genet 7:395–401
Ruf S, Biehler K, Bock R (2000) A small chloroplast-encoded protein as a novel architectural component of the light-harvesting antenna. J Cell Biol 149:369–377
Ruf S, Hermann M, Berger IJ, Carrer H, Bock R (2001) Stable genetic transformation of tomato plastids and expression of a foreign protein in fruit. Nature Biotechnol 19:870–875
Sager R, Sano H, Grabowy CT (1984) Control of maternal inheritance by DNA methylation in Chlamydomonas. Curr Topics Microbiol Immunol 108:157–173
Scharnagl M, Richter S, Hagemann M (1998) The cyanobacterium Synechocystis sp. strain PCC 6803 expresses a DNA methyltransferase specific for the recognition sequence of the restriction endonuclease PvuI. J Bacteriol 180:4116–4122
Shinozaki K, Ohme M, Tanaka M, Wakasugi T, Hayashida N, Matsubayashi T, Zaita N, Chunwongse J, Obokata J, Yamaguchi-Shinozaki K, Ohto C, Torazawa K, Meng BY, Sugita M, Deno H, Kamogashira T, Yamada K, Kusuda J, Takaiwa F, Kato A, Tohdoh N, Shimada H, Sugiura M (1986) The complete nucleotide sequence of the tobacco chloroplast genome: its gene organization and expression. EMBO J 5:2043–2049
Sternberg N (1985) Evidence that adenine methylation influences DNA-protein interactions in Escherichia coli. J Bacteriol 164:490–493
Svab Z, Maliga P (1993) High-frequency plastid transformation in tobacco by selection for a chimeric aadA gene. Proc Natl Acad Sci USA 90:913–917
Svab Z, Hajdukiewicz P, Maliga P (1990) Stable transformation of plastids in higher plants. Proc Natl Acad Sci USA 87:8526–8530
Tomas R, Vera A, Martin M, Sabater B (1992) Changes in protein synthesis without evidence of DNA methylation in barley chloroplasts during leaf growth and development. Plant Sci 85:71–77
Umen JG, Goodenough UW (2001) Chloroplast DNA methylation and inheritance in Chlamydomonas. Genes Dev 15:2585–2597
Wion D, Casadesús J (2006) N6-methyl-adenine: an epigenetic signal for DNA-protein interactions. Nature Rev Microbiol 4:183–192
Wurbs D, Ruf S, Bock R (2007) Contained metabolic engineering in tomatoes by expression of carotenoid biosynthesis genes from the plastid genome. Plant J 49:276–288
Acknowledgments
This research was supported in part by a grant from the Bundesministerium für Bildung und Forschung (BMBF).
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by R. Hagemann.
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Ahlert, D., Stegemann, S., Kahlau, S. et al. Insensitivity of chloroplast gene expression to DNA methylation. Mol Genet Genomics 282, 17–24 (2009). https://doi.org/10.1007/s00438-009-0440-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-009-0440-z