Abstract
The tripartite motif-containing (TRIM) protein family has steadily become a hotspot in tumor-related research. As a member of the E3 ubiquitin ligase family, TRIM is working on many crucial biological processes, including the regulation of tumor cell proliferation, metastasis, apoptosis, and autophagy. Among the diverse TRIM superfamily members, TRIM3 operates via different mechanisms in various types of tumors. This review primarily focuses on the current state of research regarding the antitumor mechanisms of TRIM3 in different cancers. A more in-depth study of TRIM3 may provide new directions for future antitumor treatments. Our review focuses on TRIM3 proteins and cancer. We searched for relevant articles on the mechanisms by which TRIM3 affects tumorigenesis and development from 1997 to 2023 and summarized the latest progress and future directions. Triad-containing motif protein 3 (TRIM3) is an important protein, which plays a key role in the process of tumorigenesis and development. The comprehensive exploration of TRIM3 is anticipated to pave the way for future advancements in antitumor therapy, which is expected to be a new hallmark for cancer detection and a novel target for drug action. TRIM3 is poised to become a significant milestone in cancer detection and a promising focal point for drug intervention. Recent years have witnessed notable progress in research aimed at unraveling the antitumor mechanism of TRIM3, with far-reaching implications for practical tumor diagnosis, treatment protocols, efficacy evaluation, economics, and pharmaceutical utilization.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The tripartite motif-containing (TRIM) protein family, a group of highly conserved E3 ubiquitin ligase proteins, plays indispensable roles in the initiation, progression, and resistance of various cancers (Hatakeyama 2011). These proteins exhibit bifunctionality, acting as either cancer promoters or tumor suppressors, the function of which is contingent on the specific cancer type (Connacher and Goldstrohm 2021). Furthermore, it turns out that the TRIM family could regulate copious biological processes. These include, but are not limited to, epithelial–mesenchymal transition (EMT), the adoption of cancer stem cell (CSC) phenotypes (Wang et al. 2022), cell cycle modulation, DNA repair, stress response, apoptosis, and autophagy (Crawford et al. 2018).
TRIM family plays a crucial role in human cancers. TRIM3 is an important member of this family, which has been under the spotlight of cancer research in recent years. Investigations have shown that the human TRIM3 gene resides in the chromosomal region 11p15.5. Intriguingly, the deletion of this region is correlated with various cancer types, hinting at a potential tumor-suppressive role of TRIM3 (Ozato et al. 2008). This review provides a comprehensive examination of the physiological function of TRIM3, focusing on its participation in antitumorigenesis and cancer development. This information should serve as a valuable foundation for future studies on TRIM3.
The structure and biological function of TRIM3
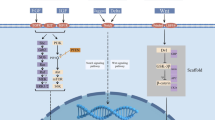
The TRIM3 protein could be portrayed through three primary domains: a RING finger domain, one or two B-BOX type zinc finger (BB1 and BB2) structures, and a coiled-coil (CC) structure (Figs. 1 and 2) (Reymond et al. 2001). The RING domain comprises a consistent arrangement of cysteine (Cys-C) and histidine (His-H) residues, which interact synergically with zinc ions in a cross-brace configuration (Fig. 3). Specifically, Cys residues at sites 1, 2, 5, and 6 bind to the first zinc ion, while His residues at sites 3, 4, 7, and 8 bind to the second zinc ion (Freemont 2000). The RING domain of TRIM3 falls within the single-subunit classification of E3 ubiquitin-protein ligases, bestowing TRIM3 with E3 ubiquitin ligase activity. The B-BOX domain, encompassing BB1 and BB2, is organized from the N-terminus to the C-terminus. Each B-BOX can adopt one of two distinct zinc-binding motifs. The consensus sequence for B-BOX type 1 (B-BOX1) is C-X(2)-C-X(7–10)-C-X(2)-C-X(4–5)-C-X(2)-C(H)-X(3–6)H-X(2–8)-H(C5(C/H)H2), whereas for B-BOX type 2 (B-BOX2), it is C-X(2)-4-C/H-X(7)10-C-X(7)-C-X(2)-C-X(3)-6-H-X(2)-8-H(C(C/H)C3H2). The existence of B-BOX2 is a distinguishing feature of RBCC (TRIM) proteins. While the specific work of the CC domain has yet to be fully clarified, it is known to be embroiled in mRNA regulation (Roshanazadeh et al. 2022). The NHL domain, positioned at the C-terminus of the TRIM3 protein, derives its name from NCL-1/HT2A/LIN-41, the original protein designation. Typically, an immunoglobulin-like domain precedes the NHL domain. Typically, an immunoglobulin-like domain precedes the NHL domain (Deyne et al. 1998).
TRIM3 protein structural domain pattern diagram. TRIM3 protein structural domain pattern diagram. A TRIM3 protein pattern diagram; B Filamin structural domain longitudinal schematic; C Filamin structural domain transverse schematic; D NHL structural domain longitudinal schematic; E NHL structural domain transverse schematic. Photograph credit: α Folded Protein Structure Database (ebi.ac.uk)
TRIM3, one kind of E3 ubiquitin ligases, is widely distributed within several subcellular organelles including the cell nucleus, cytosol, and endoplasmic reticulum and is expressed in a variety of cell types. It possesses both transcriptional repression activity and E3 ligase activity (Shen et al. 1997, 1998). Importantly, TRIM3 exerts substantial influence over multiple essential physiological processes within an organism, including cell proliferation, DNA damage repair, intracellular signaling, and immune response (Lau et al. 2010). By enhancing the activity of caspase 3, it induces apoptosis in normal cell processes. It also intervenes in the NF-κB signaling pathway by ubiquitinating and degrading NF-κB inhibitor IκBα, thereby exercising critical regulatory functions in innate immunity (Weatheritt et al. 2014). Furthermore, TRIM3’s binding affinity with p21 WAF1/CIP1 impedes its nuclear accumulation, effectively inhibiting tumor growth by impeding the promotion of cyclin D1-CDK4 in the cell cycle (Wulczyn et al. 2007). Recent studies have further illuminated TRIM3’s involvement in key signaling pathways during cancer progression. For example, it modulates the Notch signaling pathway, the P38 signaling pathway, and mechanisms related to cell cycle, tumor cell stemness, and EMT (Zuo et al. 2023; Dai et al. 2021).
The role of TRIM3 in antitumorigenesis and tumor development.
The antitumor effect of TRIM3 on glioblastoma
Glioblastoma multiforme (GBM) is an aggressive primary brain tumor and ranks among the deadliest central nervous system (CNS) tumors in adults. Despite significant advancements in clinical treatments, such as surgical interventions, temozolomide drug therapy, and gamma-ray therapy, the long-term survival rate for patients is only around 5% after five years (Stupp et al. 2005; Ostrom et al. 2018; Wen and Kesari 2008). Research suggests that the loss of heterozygosity on chromosome segment 11p15.5 in malignant gliomas may indicate TRIM3 as a potential candidate tumor suppressor gene for brain tumors (Tian et al. 2022).
Inhibition on GBM cells in vitro and in vivo
The growth of GBM cells in vitro and in vivo can be influenced by various factors, including environmental and nutritional factors and the regulation of certain tumor suppressor genes. Among these, TRIM3, a prototypical E3 ubiquitin ligase, assumes a vital role in glioblastoma. In experiments utilizing orthotopic tumor-bearing nude mice, it was viewed that mice engrafted with U87-TRIM3 cells exhibited a longer median survival time, indicating that TRIM3 expression exerts an inhibitory effect on glioblastoma in vivo. Moreover, GBM cells overexpressing TRIM3 demonstrated a marked reduction in nonadherent colony formation in soft agar during a 30-day period, providing strong evidence that TRIM3 expression stifles the proliferation and colony formation of GBM cells in vitro (Chen et al. 2014).
Inhibiting the expression and activity of c-myc and affecting cell transformation
The c-myc gene is an important constituent of the myc gene family which has a prominent function in cell apoptosis and has been implicated in the development of various tumors (Dang 1999). Analysis of GBM data from TCGA revealed a pattern of c-myc overexpression and TRIM3 underexpression in nearly all GBM samples. Fruit fly studies have illustrated that the Brat protein partially restrains c-myc, facilitating asymmetric cell division and neural differentiation (Lee et al. 2006). In cultured glioblastoma cell lines, TRIM3 has been noticed to inhibit the levels and transcriptional activity of c-myc. Evidence points to a marked negative correlation between TRIM3 expression and oncogene expression within the same sample. They demonstrated TRIM3's inhibitory effect on c-myc transcriptional activity and expression levels in glioblastoma by employing the c-myc-driven luciferase reporter gene in U87 MG and LN 229 GBM cell lines (Chen et al. 2014).
Inhibiting the cyclin-CDK complex by binding to P21 and affecting the proliferation
P21 is a pivotal player within the cyclin-dependent kinase (CDK) inhibitor family and is intimately associated with tumor suppressor functions. It orchestrates cell cycle progression, DNA replication, and repair by reducing the activity of CDK complexes, thereby establishing a tight nexus between tumor suppression and cell cycle control processes (Abbas and Dutta 2009). Recent studies have shown that in the RCASPDGF-HA/nestin-TvA model, p21 acts as an assembly factor for cyclin D-CDK4, and TRIM3 can bind to p21 to inhibit cell proliferation. The underlying mechanism may involve TRIM3 and the cell cycle protein-CDK complex competing interaction with p21 (Mukherjee et al. 2016).
Regulating NOTCH signaling to balance the stem cell proliferation and differentiation
The receptors encoded by NOTCH genes are members of a highly conserved class of cell surface receptors that regulate the development of various biological cells, ranging from sea urchins to humans. NOTCH signaling influences multiple processes in normal cell morphogenesis, such as the differentiation of pluripotent progenitor cells, cell apoptosis, proliferation, and the formation of cell boundaries. The myriad phenotypic alterations resulting from mutations in NOTCH gene loci underscore the diversity of NOTCH signaling effects (Bray 2006). NOTCH has a strong regulatory effect on GBM stem cell behavior (Battiste et al. 2007). As an RNA-binding protein, Musashi has the function of triggering NOTCH signaling by hindering Numb translation and averting the degradation of NOTCH-1. Typically, Numb affixes to the intracellular domain of NOTCH-1 (NICD) and lowers the expression of downstream targets, e.g., Hes-1. In NHNP-controlled neural spheres (NHNP-GFP) studied by Subhas Mukherjee et al., high basal TRIM3 expression was found to be associated with low Musashi expression, high Numb expression, and low Notch activity. In NHNP neural spheres (NHNP-sh-TRIM3), inhibiting TRIM3 caused an upregulation of Musashi, downregulation of Numb, and elevated expression of the NOTCH-1 target Hes-1. Mirroring these results in NHNP, the repair of TRIM3 in GBM neural spheres resulted in a decrease in Musashi and Hes-1 levels, thus establishing an inverse relationship between TRIM 3 and Musashi expression (Ji et al. 2021). Therefore, TRIM3 is crucial in regulating the Musashi–Numb–Notch signaling.
The antitumor effect of TRIM3 on liver cancer
Liver cancer is a prevalent malignancy with escalating incidence and mortality rate globally. Addressing those challenges and limitations in liver cancer treatment is essential to enhance patient survival. Emerging research has demonstrated that TRIM3 expression is diminished in liver cancer and exhibits pronounced anticancer effects (Huang et al. 2017). Therefore, exploring TRIM3 as a potential therapeutic target could pave the way for novel strategies to combat liver cancer and improve patient outcomes.
Inhibition of hepatoma cell proliferation
Overexpression of TRIM3 can curtail the proliferation capacity of liver cancer cells by regulating related signaling pathways, such as PI3K/Akt, MAPK, and Wnt/β-catenin, which affect the cell cycle and proliferation-related gene expression (Mukherjee et al. 2016). Additionally, the enhanced expression of TRIM3 can suppress their migration and invasive ability by blocking EMT and expression of metastasis-related genes, such as E-cadherin, N-cadherin, and vimentin (Lu et al. 2022). In addition, TRIM3 overexpression reduces the number of tumor stem cells and hinders tumor angiogenesis, effectively restraining tumor growth and metastasis in liver cancer.
Inducing Go/G phase arrest and affecting cell cycle progression
TRIM3 mitigates tumor growth in liver cancer by making cell cycle arrest in tumor cells. Besides liver cancer cells with low TRIM3 expression, those with high TRIM3 expression exhibit a noteworthy boost in the ratio of cells in the Go/G1 phase and a significant decrease in the percentage of cells in the G2/M phase. Conversely, when the expression of TRIM3 is reduced, the proportion of liver cancer cells in the S and G2/M phases increases. These finding clearly indicates that TRIM3 can induce Go-/G1-phase arrest in liver cancer cells, thereby influencing the cell cycle progression and inhibiting tumor cell proliferation. Hence, TRIM3 has a certain inhibitory job in the occurrence and development of liver cancer (Huang et al. 2017).
Inhibitory effect of TRIM3 on colorectal cancer
Colorectal cancer, a malignancy deriving from the epithelial cells of the colon or rectum, is a common digestive system tumor predominantly affecting individuals aged 50 and above, with incidence rates increasing with age (Kahi et al. 2016). Recent studies showed that the expression level of TRIM3 in colorectal cancer is much less than that in normal colorectal tissue (Hiltunen et al. 2022). A particular study identified a significant downregulation of TRIM3 in colorectal cancer tissues, and the expression level of TRIM3 was closely associated with the clinical and pathological hallmarks of colorectal cancer. Furthermore, the expression level of TRIM3 has a direct link with the prognosis of colorectal cancer, as sufferers with lesser TRIM3 expression tend to experience poorer outcomes (Betschinger and Knoblich 2004).
Inhibiting the proliferation
The overexpression of TRIM3 significantly curbs the proliferation and growth, while simultaneously resulting in apoptosis of colorectal cancer cells (Han et al. 2023). The Wnt/β-catenin signaling pathway has a critical influence on the occurrence and growth of colorectal cancer. Studies have shown that TRIM3 can block the activation of the Wnt/ beta-catenin signaling pathway, thereby hindering cell proliferation and invasion. By reducing the stability of β-catenin, TRIM3 effectively obstructs the activation of the Wnt/β-catenin signaling pathway (Hu and Gan 2017). Furthermore, TRIM3 exerts its inhibitory action by suppressing the expression of downstream target genes in the Wnt/β-catenin signaling pathway, such as c-Myc and CyclinD1, thus curtailing the proliferation and invasion of colorectal cancer cells (Song et al. 2019). Consequently, TRIM3 also limits the proliferation and self-renewing capabilities of colorectal cancer stem cells, thus inhibiting the onset and development of colorectal cancer.
Inhibiting the invasion and metastasis
TRIM3 demonstrates significant potential in inhibiting the invasion and metastasis of colorectal cancer cells. A study has revealed that overexpression of TRIM3 leads to a substantial diminution in the invasion and metastasis capabilities of colorectal cancer cells, along with a decreased migration ability. Additionally, TRIM3 effectively curbs the colorectal EMT process, hindering their metastatic and invasive properties (Song et al. 2018). Furthermore, TRIM3 induces apoptosis in colorectal cancer cells and regulates their metabolism, effectively impeding the occurrence and progression of colorectal cancer. The NF-κB signaling pathway also holds importance in the development and advancement of colorectal cancer (Boulay et al. 2009). Research has demonstrated that TRIM3 acts as a hindrance to the NF-κB signaling pathway, creating a decreased proliferation and invasion of colorectal cancer cells. By hindering the nuclear transport and DNA-binding capacity of NF-κB, TRIM3 effectively deactivates this signaling pathway and downregulates the downstream target genes, such as IL-6 and TNF-α, thus curbing the inflammatory response and invasion abilities of colorectal cancer cells (Zhu et al. 2023). As a result, TRIM3 emerges as a promising new target for the treatment of colorectal cancer, given its potential to inhibit invasion and metastasis, induce apoptosis, and regulate essential signaling pathways involved in colorectal cancer progression.
Regulation of cellular metabolism
TRIM3 can regulate the metabolism of colorectal cancer cells. One study found that overexpression of TRIM3 can inhibit metabolic pathways such as glycolysis and oxidative phosphorylation in colorectal cancer cells, causing the quelling of their growth and invasion (Xie et al. 2020). TRIM3 can also modulate the mitochondrial function of colorectal cancer cells, further contributing to the impediment of their growth and invasion.
Regulation of PI3K/Akt signaling pathway
The PI3K/Akt signaling pathway also possesses an extremely important mission in the happening and development of colorectal cancer. Research has shown that TRIM3 can hinder the energizing of the PI3K/Akt signaling, effectively restraining the proliferation and invasion of colorectal cancer cells (Eberhardt et al. 2020). By reducing the phosphorylation level of Akt, TRIM3 effectively hampers the activation of the PI3K/Akt signaling pathway. In addition, TRIM3 can also suppress the expression of downstream target genes in the PI3K/Akt signaling pathway, such as Bcl-2 and Mcl-1, thus promoting apoptosis in colorectal cancer cells (Wang et al. 2020).
Tumor-suppressive effect of TRIM3 in cervical cancer
Inhibiting the proliferation
TRIM3 overexpression has a substantial inhibitory effect on the proliferation capacity of cervical cancer cells. Research has found that TRIM3 regulates multiple signaling pathways and related molecules to suppress cell proliferation (Jiang et al. 2022). For instance, TRIM3 downregulates the activity of the PI3K/Akt and MAPK signaling pathways, causing the inhibition of cell proliferation and proliferation-related gene expression. Additionally, TRIM3 overexpression also inhibits the activity of the Wnt/β-catenin signaling pathway, thereby influencing cell proliferation and transcriptional regulation (Chen et al. 2017).
Inhibiting the migration and invasion
TRIM3 overexpression significantly hinders the migration and invasion capabilities of cervical cancer cells. This inhibition is accomplished through the regulation of transcription factors and mediation of signaling molecules crucial for the EMT process, which is essential for cell migration and invasion. Notably, TRIM3 downregulates the expression of EMT transcription factors, such as Snail, Slug, and TWIST, thus curbing the EMT process in cervical cancer cells and weakening their migration and invasion potentials (Diao et al. 2020). Furthermore, TRIM3 overexpression exhibits tumor-inhibiting effects in cervical cancer. Research reveals that TRIM3 reduces the population of cervical cancer tumor stem cells and actively participates in regulating their apoptosis and differentiation by engaging various signaling pathways (Diao et al. 2020). Moreover, TRIM3 overexpression effectively suppresses angiogenesis, resulting in decreased tumor blood supply and growth. These findings underscore the potential of TRIM3 as a valuable target for therapeutic interventions in cervical cancer treatment.
Function of TRIM3 in esophageal squamous cell carcinoma
In the current research, it has been demonstrated that TRIM3 coacts with the E4 ligase-dependent proteasomal turnover of importin α3 and α-actinin-4 (ACTN4), which successively modulates the nuclear factor κB (NF-κB) at a stable level(Zhu et al. 2019 Apr). Heterozygous deletion-mediated downregulation of TRIM3 disrupts the NF-κB-IκB-α negative-feedback loop, thereby promoting symmetric dimethylation of NF-κB/p65 at Arg65 and Arg65. This enhances p65 DNA-binding affinity and transcriptional activity, consequently facilitating lymphatic metastasis of esophageal squamous cell carcinoma (ESCC) cells and triggering the activation of NF-κB signaling(Zhao et al. 2022 Sep).
Role of TRIM3 in lung cancer
Lung cancer stands as one of the most prevalent and deadliest malignancies worldwide. Recent investigations have highlighted the crucial role of TRIM3 in suppressing lung cancer (Zhan et al. 2015). A study demonstrated significantly lower expression levels of TRIM3 in lung cancer tissue compared to normal lung tissue, with its expression level closely correlating with the clinical and pathological characteristics of lung cancer. Further experiments evidenced TRIM3’s ability to inhibit lung cancer cell proliferation and invasion and induce apoptosis (Altinoz et al. 2016). Additionally, TRIM3 can also regulate the metabolism of lung cancer cells, curtailing metabolic pathways such as glycolysis and oxidative phosphorylation, thus hindering the growth and invasion of lung cancer cells. The anticancer effect of TRIM3 is mainly achieved by regulating the NF-κB signaling pathway (Choi et al. 2014). TRIM3 can diminish the activation of NF-κB, thereby suppressing the proliferation and invasion of lung cancer cells.
Summary of TRIM3 antitumor mechanisms in different tumors
Abnormal expression of TRIM3 is closely linked to the occurrence and progression of various tumors, including glioma, liver cancer, and colorectal cancer (Table 1). When TRIM3 is overexpressed in these tumors, it exhibits a promising anticancer effect, and it is mainly not only through the regulation of the NF-κB signaling pathway, Notch signaling pathway, and P38 signaling pathway, but also through the regulation of cell cycle, regulation of tumor cell dryness, EMT, and other mechanisms to play a tumor inhibitory role, although the specific biological behaviors and underlying mechanisms may vary among different tumor types. At present, there are few studies on this gene protein, and the antitumor mechanism of TRIM3 is still not completely clear. Its molecular mechanism and signaling pathway will be further explored in future. Therefore, further comprehensive and in-depth multi-omics studies are warranted to elucidate the specific mechanisms through which TRIM3 operates in these varied tumors.
Conclusion
Targeted TRIM3 therapy has certain possibilities and has become one of the research hotspots in the field of cancer therapy. Future, by targeting overexpressed TRIM3 in individual tumors, a personalized treatment regimen can be achieved to improve cure and survival rates. In addition, compared with traditional chemoradiotherapy and other therapies, TRIM3-targeted therapies may have better specificity and reduce damage to normal cells and side effects. However, the current targeted therapy for TRIM3 is still in the early stages, and there are still some challenges and limitations: Similar to other targeted therapies, TRIM3-targeted therapy may also face resistance problems.
In summary, TRIM3 is an ubiquitin-related protein, which is related to cell proliferation, invasion, metastasis, cell cycle, apoptosis, etc., during the occurrence and development of tumors. However, due to the heterogeneity and complexity of tumors, the functional mechanism of TRIM3 in tumors is not fully understood, and its specific signaling pathway and molecular mechanism need to be further clarified. This makes it a challenge for us to use TRIM3 molecules to intervene in tumor development. In future, we will further explore the functional mechanism of TRIM3 molecules in different types of tumors and explore the expression patterns and functional changes of TRIM3 in different tumor subtypes, which may provide valuable insights for molecular diagnosis and gene therapy of cancer patients at the gene level in future and have important clinical significance for diagnosis and treatment.
In my opinion, first of all, TRIM3 is important in tumor diagnosis. Examining the expression levels of TRIM3 in different types of tumors can assist physicians in determining the type and grade of the tumor and predicting the prognosis of the patient. This provides an important reference for clinicians and helps to develop more individualized and precise treatment plans. The study of TRIM3 antitumor mechanism also has a positive impact on the development of treatment guidelines. The in-depth exploration of the relationship between TRIM3 and tumor development can provide a basis for the formulation of more scientific and rational treatment guidelines. For example, in some specific types of tumors, the expression of TRIM3 is closely related to the sensitivity of a certain treatment, so TRIM3 can be used as a predictive indicator to choose the best treatment for patients. The study of TRIM3 antitumor mechanism is also important for evaluating the effectiveness of the treatment methods. By observing the changes of TRIM3 in the course of treatment, it is possible to understand the effect of treatment in time and adjust the treatment program. This helps to improve the accuracy and effectiveness of tumor therapy and reduce unnecessary therapeutic risks. In addition, from the economic point of view, the study of TRIM3 antitumor mechanism also possesses certain value. By gaining a deeper understanding of the relationship between TRIM3 and tumor development, drug use strategies applicable to patients with different types of tumors can be more precisely determined. This can help avoid unnecessary drug waste and side effects and improve the efficiency of medical resource utilization.
Data availability
The authors confirm that all data generated or analyzed during this study are included in this article.
References
Abbas T, Dutta A (2009) p21 in cancer: intricate networks and multiple activities. Nat Rev Cancer 9(6):400–414. https://doi.org/10.1038/nrc2657
Altinoz MA, Elmaci I, Ince B et al (2016) Hemoglobins, Hemorphins, and 11p15.5 Chromosomal Region in Cancer Biology and İmmunity with Special Emphasis for Brain Tumors. J Neurol Surg A Cent Eur Neurosurg. https://doi.org/10.1055/s-0035-1566120
Battiste J, Helms AW, Kim EJ et al (2007) Ascl1 defines sequentially generated lineage-restricted neuronal and oligodendrocyte precursor cells in the spinal cord. Development 134(2):285–293. https://doi.org/10.1242/dev.02727
Betschinger J, Knoblich JA (2004) Dare to be different: asymmetric cell division in Drosophila. C Elegans and Vertebrates Curr Biol 14(18):R674–R685. https://doi.org/10.1016/j.cub.2004.08.017
Boulay JL, Stiefel U, Taylor E et al (2009) Loss of heterozygosity of TRIM3 in malignant gliomas. BMC Cancer 27(9):71. https://doi.org/10.1186/1471-2407-9-71
Bray S (2006) Notch signalling: a simple pathway becomes complex. Nat Rev Mol Cell Biol 7(9):678–689. https://doi.org/10.1038/nrm2009
Chen G, Kong J, Tucker-Burden C et al (2014) Human Brat ortholog TRIM3 is a tumor suppressor that regulates asymmetric cell division in glioblastoma. Cancer Res 74(16):4536–4548. https://doi.org/10.1158/0008-5472.CAN-13-3703
Chen Y, Sun J, Ma J (2017) Proliferation and invasion of ovarian cancer cells are suppressed by knockdown of TRIM11. Oncol Lett 14(2):2125–2130. https://doi.org/10.3892/ol.2017.6432
Choi UY, Hur JY, Lee MS et al (2014) Tripartite motif-containing protein 30 modulates TCR-activated proliferation and effector functions in CD4+ T cells. PLoS ONE. https://doi.org/10.1371/journal.pone.0095805
Connacher RP, Goldstrohm AC (2021) Molecular and biological functions of TRIM-NHL RNA-binding proteins. Wiley Interdiscip Rev RNA. https://doi.org/10.1002/wrna.1620
Crawford LJ, Johnston CK, Irvine AE (2018) TRIM proteins in blood cancers. J Cell Commun Signal 12(1):21–29. https://doi.org/10.1007/s12079-017-0423-5
Dai W, Wang J, Wang Z et al (2021) Comprehensive Analysis of the Prognostic Values of the TRIM Family in Hepatocellular Carcinoma. Front Oncol. https://doi.org/10.3389/fonc.2021.767644
Dang CV (1999) c-Myc target genes involved in cell growth, apoptosis, and metabolism. Mol Cell Biol 19(1):1–11. https://doi.org/10.1128/MCB.19.1.1
De Deyne PG, O’Neill A, Resneck WG et al (1998) The vitronectin receptor associates with clathrin-coated membrane domains via the cytoplasmic domain of its beta5 subunit. J Cell Sci 111(Pt 18):2729–2740. https://doi.org/10.1242/jcs.111.18.2729
Diao W, Zhu C, Guo Q et al (2020) Tripartite motif-containing 14 regulates cell proliferation and apoptosis in cervical cancer via the Akt signaling pathway. Mol Med Rep 22(6):5145–5154. https://doi.org/10.3892/mmr.2020.11634
Eberhardt W, Haeussler K, Nasrullah U, Pfeilschifter J (2020) Multifaceted Roles of TRIM Proteins in Colorectal Carcinoma. Int J Mol Sci 21(20):7532. https://doi.org/10.3390/ijms21207532
Freemont PS (2000) RING for destruction? Curr Biol 10(2):R84–R87. https://doi.org/10.1016/s0960-9822(00)00287-6
Han Y, Lu S, Song C et al (2023) Dual roles of TRIM3 in colorectal cancer by retaining p53 in the cytoplasm to decrease its nuclear expression. Cell Death Discov 9(1):85. https://doi.org/10.1038/s41420-023-01386-1
Hatakeyama S (2011) TRIM proteins and cancer. Nat Rev Cancer 11(11):792–804. https://doi.org/10.1038/nrc3139
Hiltunen AE, Vuolteenaho R, Ronkainen VP et al (2022) Nhlrc2 is crucial during mouse gastrulation. Genesis. https://doi.org/10.1002/dvg.23470
Hu CE, Gan J (2017) TRIM37 promotes epithelial-mesenchymal transition in colorectal cancer. Mol Med Rep 15(3):1057–1062. https://doi.org/10.3892/mmr.2017.6125
Huang XQ, Zhang XF, Xia JH et al (2017) Tripartite motif-containing 3 (TRIM3) inhibits tumor growth and metastasis of liver cancer. Chin J Cancer 36(1):77. https://doi.org/10.1186/s40880-017-0240-5
Ji J, Ding K, Luo T et al (2021) TRIM22 activates NF-κB signaling in glioblastoma by accelerating the degradation of IκBα. Cell Death Differ 28(1):367–381. https://doi.org/10.1038/s41418-020-00606-w
Jiang Y, Xu W, Tao J (2022) Circular RNA circVPRBP serves as a microRNA-106b-5p sponge to regulate proliferation and metastasis of cervical cancer cells via tripartite motif-containing protein 3. Anticancer Drugs 33(9):850–860. https://doi.org/10.1097/CAD.0000000000001335
Kahi CJ, Boland CR, Dominitz JA et al (2016) Colonoscopy surveillance after colorectal cancer resection: recommendations of the US Multi-Society Task Force on Colorectal Cancer. Gastroenterology 150(3):758-768.e11. https://doi.org/10.1053/j.gastro.2016.01.001
Lau SK, Li KS, Huang Y et al (2010) Ecoepidemiology and complete genome comparison of different strains of severe acute respiratory syndrome-related Rhinolophus bat coronavirus in China reveal bats as a reservoir for acute, self-limiting infection that allows recombination events. J Virol 84(6):2808–2819. https://doi.org/10.1128/JVI.02219-09
Lee CY, Wilkinson BD, Siegrist SE et al (2006) Brat is a Miranda cargo protein that promotes neuronal differentiation and inhibits neuroblast self-renewal. Dev Cell 10(4):441–449. https://doi.org/10.1016/j.devcel.2006.01.017
Lu K, Pan Y, Huang Z et al (2022) TRIM proteins in hepatocellular carcinoma. J Biomed Sci 29(1):69. https://doi.org/10.1186/s12929-022-00854-7
Mukherjee S, Tucker-Burden C, Zhang C et al (2016) Drosophila Brat and Human Ortholog TRIM3 Maintain Stem Cell Equilibrium and Suppress Brain Tumorigenesis by Attenuating Notch Nuclear Transport. Cancer Res 76(8):2443–2452. https://doi.org/10.1158/0008-5472.CAN-15-2299
Ostrom QT, Gittleman H, Truitt G et al (2018) CBTRUS statistical report: primary brain and central nervous system tumors diagnosed in the United States in 2011–2015. Neuro Oncol. https://doi.org/10.1093/neuonc/noy131
Ozato K, Shin DM, Chang TH et al (2008) TRIM family proteins and their emerging roles in innate immunity. Nat Rev Immunol 8(11):849–860. https://doi.org/10.1038/nri2413
Reymond A, Meroni G, Fantozzi A et al (2001) The tripartite motif family identifies cell compartments. EMBO J 20(9):2140–2151. https://doi.org/10.1093/emboj/20.9.2140
Roshanazadeh MR, Adelipour M, Sanaei A et al (2022) TRIM3 and TRIM16 as potential tumor suppressors in breast cancer patients. BMC Res Notes 15(1):312. https://doi.org/10.1186/s13104-022-06193-y
Shen CP, Jan LY, Jan YN (1997) Miranda is required for the asymmetric localization of Prospero during mitosis in Drosophila. Cell 90(3):449–458. https://doi.org/10.1016/s0092-8674(00)80505-x
Shen CP, Knoblich JA, Chan YM et al (1998) Miranda as a multidomain adapter linking apically localized Inscuteable and basally localized Staufen and Prospero during asymmetric cell division in Drosophila. Genes Dev 12(12):1837–1846. https://doi.org/10.1101/gad.12.12.1837
Song Y, Guo Q, Gao S et al (2018) Tripartite motif-containing protein 3 plays a role of tumor inhibitor in cervical cancer. Biochem Biophys Res Commun 498(3):686–692. https://doi.org/10.1016/j.bbrc.2018.03.046
Song Y, Guo Q, Gao S et al (2019) miR-454-3p promotes proliferation and induces apoptosis in human cervical cancer cells by targeting TRIM3. Biochem Biophys Res Commun 516(3):872–879. https://doi.org/10.1016/j.bbrc.2019.06.126
Stupp R, Mason WP, van den Bent MJ et al (2005) Radiotherapy plus concomitant and adjuvant temozolomide for glioblastoma. N Engl J Med 352(10):987–996. https://doi.org/10.1056/NEJMoa043330
Tian Y, Liu H, Zhang C et al (2022) Comprehensive Analyses of Ferroptosis-Related Alterations and Their Prognostic Significance in Glioblastoma. Front Mol Biosci. https://doi.org/10.3389/fmolb.2022.904098
Wang X, Zhang Y, Pei X et al (2020) TRIM3 inhibits P53 signaling in breast cancer cells. Cancer Cell Int 20(1):559. https://doi.org/10.1186/s12935-020-01630-z
Wang X, Cheng H, Zhao J et al (2022) Long noncoding RNA DLGAP1-AS2 promotes tumorigenesis and metastasis by regulating the Trim21/ELOA/LHPP axis in colorectal cancer. Mol Cancer 21(1):210. https://doi.org/10.1186/s12943-022-01675-w
Weatheritt RJ, Gibson TJ, Babu MM (2014) Asymmetric mRNA localization contributes to fidelity and sensitivity of spatially localized systems. Nat Struct Mol Biol 21(9):833–839. https://doi.org/10.1038/nsmb.2876
Wen PY, Kesari S (2008) Malignant gliomas in adults. N Engl J Med 359(5):492–507. https://doi.org/10.1056/NEJMra0708126.Erratum.In:NEnglJMed.2008Aug21;359(8):877
Wulczyn FG, Smirnova L, Rybak A et al (2007) Post-transcriptional regulation of the let-7 microRNA during neural cell specification. FASEB J 21(2):415–426. https://doi.org/10.1096/fj.06-6130com
Xie W, Zhang Y, Wang B et al (2020) Tripartite motif containing 24 regulates cell proliferation in colorectal cancer through YAP signaling. Cancer Med 9(17):6367–6376. https://doi.org/10.1002/cam4.3310. (Epub 2020 Jul 17)
Zhan W, Han T, Zhang C et al (2015) TRIM59 Promotes the Proliferation and Migration of Non-Small Cell Lung Cancer Cells by Upregulating Cell Cycle Related Proteins. PLoS ONE 10(11):e0142596. https://doi.org/10.1371/journal.pone.0142596
Zhao B, Qiao G, Li J et al (2022) TRIM36 suppresses cell growth and promotes apoptosis in human esophageal squamous cell carcinoma cells by inhibiting Wnt/β-catenin signaling pathway. Hum Cell 35(5):1487–1498. https://doi.org/10.1007/s13577-022-00737-x
Zhu J, Wu G, Ke Z et al (2019) Targeting TRIM3 deletion-induced tumor-associated lymphangiogenesis prohibits lymphatic metastasis in esophageal squamous cell carcinoma. Oncogene 38(15):2736–2749. https://doi.org/10.1038/s41388-018-0621-5
Zhu S, Mao J, Zhang X et al (2023) CAF-derived exosomal lncRNA FAL1 promotes chemoresistance to oxaliplatin by regulating autophagy in colorectal cancer. Dig Liver Dis S1590–8658(23):00705–00713. https://doi.org/10.1016/j.dld.2023.06.010
Zuo Q, Xu Q, Li Z et al (2023) TRIM3 inhibits colorectal cancer cell migration and lipid droplet formation by promoting FABP4 degradation. Histol Histopathol 11:18627. https://doi.org/10.14670/HH-18-627
Acknowledgements
The authors would like to thank the STRING, α-Folded Protein Structure database, etc., for the availability of the data.
Funding
This study was funded by the National Natural Science Foundation of China (No:82360493) to Xingwang Zhou; Guizhou Provincial Science and Technology Projects Qiankehe Foundation-ZK [2023] General 360 to Niya Long; Qiankehe Foundation-ZK [2023] General 362 to Liangzhao Chu; National Natural Science Foundation of China (NSFC), Affiliated Hospital of Guizhou Medical University (gyfynsfc-2022–25) to Niya Long; Science and Technology Fund project of Guizhou Provincial Health Commission (gzwkj-2022–09) to Liangzhao Chu; Science and Technology Fund project of Guizhou Provincial Health Commission (gzwkj-2023–035) to Niya Long; the PhD scientific research launch fund project of the Affiliated Hospital of Guizhou Medical University (gyfybsky-2022–02) to Niya Long; and Guizhou Science and Technology Plan Project (No. Qiankehe 2016 Support [2905]) to Jian Liu.
Author information
Authors and Affiliations
Contributions
LC conceived the data; XZ, YL, and WT curated the data; YL and WT analyzed the data; WT investigated the manuscript; YL designed the methodology; YL and WT developed the software; YL and WT supervised the manuscript; XZ, YL, and NL validated the article; LC visualized the manuscript; and YL and WT wrote the original draft.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
Approval for this study was issued by the Institutional Ethics Committee of the Faculty of Affiliated Hospital of Guizhou Medical University.
Consent for publication
All authors have endorsed the manuscript and agreed to submit it to the journal.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Teng, W., Ling, Y., Liu, Z. et al. Advances in the antitumor mechanisms of tripartite motif-containing protein 3. J Cancer Res Clin Oncol 150, 105 (2024). https://doi.org/10.1007/s00432-024-05632-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00432-024-05632-6