Abstract
Purpose
Lymphocyte activation gene 3 (LAG3) is thought to contribute to T cell exhaustion within the tumor microenvironment of solid tumors. This study aimed to analyze the spatial distribution of LAG3 + cells in relation to clinicopathological and survival data in a large set of 580 primary resected and neoadjuvantly treated gastric cancers (GC).
Methods
LAG3 expression was evaluated in tumor center and invasive margin using immunohistochemistry and whole-slide digital image analysis. Cases were divided into LAG3-low and LAG3-high expression groups based on (1) median LAG3 + cell density, (2) cut-off values adapted to cancer-specific survival using Cutoff Finder application.
Results
Significant differences in spatial distribution of LAG3 + cells were observed in primarily resected GC, but not in neoadjuvantly treated GC. LAG3 + cell density showed evident prognostic value at following cut-offs: in primarily resected GC, 21.45 cells/mm2 in tumor center (17.9 vs. 10.1 months, p = 0.008) and 208.50 cells/mm2 in invasive margin (33.8 vs. 14.7 months, p = 0.006); and in neoadjuvantly treated GC, 12.62 cells/mm2 (27.3 vs. 13.2 months, p = 0.003) and 123.00 cells/mm2 (28.0 vs. 22.4 months, p = 0.136), respectively. Significant associations were found between LAG3 + cell distribution patterns and various clinicopathological factors in both cohorts. In neoadjuvantly treated GC, LAG3 + immune cell density was found to be an independent prognostic factor of survival (HR = 0.312, 95% CI 0.162–0.599, p < 0.001).
Conclusion
In this study, a higher density of LAG3 + cells was associated with favorable prognosis. Current results support the need for extended analysis of LAG3. Differences in the distribution of LAG3 + cells should be considered, as they could influence clinical outcomes and treatment responses.
Graphical abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Adenocarcinoma of the stomach and gastroesophageal junction (GC) is a heterogeneous disease that shows striking variations in epidemiology, etiology, risk factors, pathogenesis, topography, histological features, and prognosis. It remains the 5th most common and 4th deadliest cancer worldwide (Sung et al. 2021). Treatment options are still limited because of advanced-stage diagnoses that are common in the Western world (Arnold et al. 2019; National Cancer Institute).
Tumor immunotherapy has become one of the major therapeutic strategies in oncology (Galluzzi et al. 2014; van den Bulk et al. 2018; He and Xu 2020). Prominent of these therapies are immune checkpoint inhibitors (ICI) that target immune checkpoints, inhibitory or stimulatory proteins in immune cells, and/or tumor cells that modulate immune responses. Furthermore, immune checkpoints are involved in many processes of tumor cell metabolism and are related to epithelial–mesenchymal transition, metastasis, drug resistance, anti-apoptosis, and angiogenesis (Zhang and Zheng 2020). ICI block the transmission of inhibitory signals, stimulate the activation of cytotoxic T lymphocytes (CTLs), and boost the anti-tumor response of T lymphocytes (Shan et al. 2020). Current approved ICI are directed against programmed cell death 1 and its ligand (PD-1/PD-L1) and cytotoxic T-lymphocyte-associated protein 4 (CTLA4). However, many patients do not respond to ICI or the response is limited because of innate or acquired resistance (Shergold et al. 2019). Targeting other co-inhibitory receptors within the tumor microenvironment (TME), including simultaneous inhibition of multiple immune checkpoints, may provide new opportunities for immunotherapy.
Lymphocyte activation gene 3 (LAG3), also known as CD223, is a type I transmembrane protein expressed on a variety of immune cell types including activated CD4 + and CD8 + T cells, natural killer (NK) cells, NKT cells, and regulatory T cells (Triebel et al. 1990; Huang et al. 2004). Like other inhibitory receptors, it is essential to control T cell activation and to prevent autoimmunity (Andrews et al. 2017). LAG3 structurally resembles CD4 co-receptor and can interact with MHC class II molecules (Baixeras et al. 1992). Similar to CD4, the LAG3 gene is located on chromosome 12. Although both molecules are closely related (Triebel et al. 1990), they exhibit different functions due to dissimilarities in their cytoplasmic domains (Workman et al. 2002; Andrews et al. 2017). Unlike CD4 and any other known immune checkpoints, the LAG3 cytoplasmic tail contains specific KIEELE, FxxL, and EP motifs, which are thought to be essential for its inhibitory function (Anderson et al. 2016; Maeda et al. 2019).
Despite over 30 years of research involving LAG3, the exact mechanism by which LAG3 and its binding partner(s) contribute to T cell suppression is not completely understood. LAG3 is strongly upregulated upon continuous T cell stimulation due to persistent exposure to tumor antigens (Andrews et al. 2017). This inhibitory signal is thought to contribute to T cell exhaustion. Exhausted T cells lose their ability to proliferate and to perform effector functions including cytokine production and degranulation (Wherry and Kurachi 2015). Besides its well-recognized ligand MHC class II, several alternate LAG3 ligands have been reported, including liver sinusoidal endothelial cell lectin (Xu et al. 2014), galectin-3 (Kouo et al. 2015), and fibrinogen-like protein 1 (Wang et al. 2019). Interestingly, a recent study demonstrated that LAG3 could inhibit T cell receptor (TCR) signaling in the absence of binding MHC II class (Guy et al. 2022).
In vitro studies have shown that LAG3 expression level strongly correlates with its inhibitory function, and changes in the amount of LAG3 on the cell surface directly affect its inhibitory effect (Maeda et al. 2019). LAG3 is co-expressed with other inhibitory receptors like PD-1, CTLA4, T-cell immunoglobulin and mucin domains-containing protein 3 (TIM3), and T cell immuno-receptor with Ig and ITIM domains (TIGIT) (Blank et al. 2019). It has been suggested that the higher the number of co-expressed inhibitory receptors, the more severe is the T cell exhaustion (Wherry and Kurachi 2015). Dual blockade of PD-1 and LAG3 has led to decreased tumor growth and enhanced anti-tumor immunity in mouse models (Woo et al. 2012). In patients with previously untreated metastatic or un-resectable melanoma, combination of relatlimab (anti-LAG3 antibody) and nivolumab (anti-PD-1 antibody) has increased the median progression-free survival more than two times when compared to PD-1 inhibition alone (Tawbi et al. 2022). Recently, the first such combination immunotherapy has been approved by the U.S. Food and Drug Administration (Paik 2022).
The effectiveness of LAG3-targeted therapies in GC has been evaluated in several clinical trials (https://clinicaltrials.gov/, accessed on 10.04.2023). However, the clinicopathological significance and prognostic role of LAG3 protein expression in GC remain unclear, showing discrepant results with regard to patient prognosis (Lee et al. 2019; Park et al. 2021; Lv et al. 2021). To our knowledge, there are no published data on LAG3 expression in GC of Western origin. Based on previous studies, we hypothesize that: (1) LAG3 expression varies among histological and molecular subtypes of GC, and (2) high density of LAG3 + cells is related to advanced clinicopathological features and poor outcome in GC. To check our hypotheses and to deepen our understanding of the role of LAG3 in GC, we investigated the clinicopathological and prognostic significance of LAG3 in a large cohort of primarily resected chemotherapy-naïve GC (PR-GC). Additionally, the need for information on the expression patterns of immune checkpoints in tumors after neoadjuvant therapy is increasing. Therefore, we evaluated LAG3 expression also in a set of neoadjuvantly treated GC (NAT-GC).
Methods
Study population
A well-characterized cohort of patients with GC was included in this study. Formalin-fixed and paraffin-embedded tissue samples from the primary tumor site were analyzed. Patients in the PR-GC sub-cohort underwent primary total gastrectomy or surgical resection between 1997 and 2009 at the University Hospital Schleswig–Holstein, Kiel, Germany. Patients in the NAT-GC sub-cohort received preoperative chemo-radiation or perioperative chemotherapy and underwent total gastrectomy or surgical resection between 1998 and 2019. The following exclusion criteria were used: (a) diagnosis other than adenocarcinoma and (b) insufficient tissue for tumor compartment analysis, that is, no residual tumor in the NAT-GC. Clinicopathological characteristics were collected from previous records, including sex, age at diagnosis, tumor localization, type by Laurén, pathologic (pTNM) or post-neoadjuvant (ypTNM) stage according to the 8th edition of the UICC guidelines (Brierley et al. 2017), status of resection lines (pR), and lymphatic or venous invasion (LVI). For survival analysis, the date of surgery and date of death or last follow-up were used. The Helicobacter pylori, microsatellite instability (MSI)-, Epstein–Barr virus (EBV)-, HER2-, MET-, and PD-1/PD-L1-status was available from previous studies of the cohort (Warneke et al. 2013; Metzger et al. 2016; Böger et al. 2016, 2017; Mathiak et al. 2017; Schoop et al. 2020).
Immunohistochemical detection of LAG3
Immunohistochemical staining was performed on whole tissue sections using the Bondmax automated system (Leica Biosystems, Wetzlar, Germany). The LAG3 monoclonal antibody (Clone D2G4O, Cell Signaling, Leiden, the Netherlands) was used at a 1:50 dilution. Slides were pre-treated with ER2 (Leica Biosystems) for 20 min, and visualization was performed using the Bond Polymer Refine Detection Kit (Leica Biosystems).
Evaluation of LAG3 immunostaining
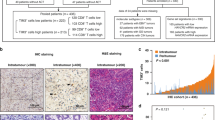
Digital images of whole tissue sections were obtained using a Leica SCN400 scanner (Leica Biosystems, Nussloch, Germany) at 40 × nominal magnification, corresponding to a resolution of 0.25 µm per pixel. To detect LAG3 + cells, image analysis was performed using Definiens Tissue Studio (version 4.4.3, Definiens, Munich, Germany). A machine learning algorithm using Definiens Composer Technology was trained with four representative images to recognize LAG3 + cells. To avoid false results (e.g., due to large areas of necrotic debris, surface ulcerations, mucin pools, folded tissue, or holes), marking of distinct tumor compartments was carried out manually using the viewer and painting program VMP (Fig. 1a–c). The invasive margin (IM) was defined as a narrow band-like area with a width of up to 1000 µm centered on the border separating cancer cells from the host tissue (Hendry et al. 2017). The tumor center (TC) was defined as the remaining tumor area comprising cancer cells and desmoplastic stroma. In NAT-GC, the tumor center could include surrounding scarred tissue that had reacted to NAT. LAG3 + cell densities were individually computed for TC and IM.
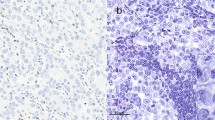
LAG3 + immune cells in gastric cancer and cancer of gastroesophageal junction: a–c, representative images from the viewer and painting program VMP, which was used to mark the tumor center (yellow) and invasive margin (orange), magenta dots c indicate LAG3 + cells identified using Definiens Tissue Studio. Anti-LAG3 immunostaining, original magnification × 12.5 (a, b) and × 50 (c); d–g, representative tissue sections with LAG3 + tumor-infiltrating immune cells in primary resected and neoadjuvantly treated tumors: d high expression in EBV-associated gastric adenocarcinoma; e low expression in unclassified gastric carcinoma with prominent tumor-associated inflammatory reaction in the stroma; f LAG3 + immune cells were present in both the stromal and intraepithelial compartments of this gastric intestinal-type adenocarcinoma; g LAG3 + immune cells in gastric cancer treated with neoadjuvant therapy. Anti-LAG3 immunostaining, original magnification × 400
Statistical analysis
Data were analyzed using SPSS 25.0.0.2 (IBM Corporation, New York, USA). A significance level of p < 0.05 was assumed. Overall survival (OS) and cancer-specific survival (CSS) were defined as the time from the date of surgery until death due to any cause and death due to GC, respectively. The median LAG3 + cell density was calculated separately for each cohort. Because of differences in median densities between both cohorts, as well as tumor compartments, it was decided to analyze them separately and use individual cut-offs. To incorporate LAG3 + immune cell density in further statistical analyses, it was dichotomized into low and high LAG3 expression groups: (1) by median number; (2) using Cutoff Finder web application (Budczies et al. 2012), in which the optimal cut-off values were generated based on CSS. Survival curves were estimated using the Kaplan–Meier method and compared using the log-rank test. Multivariate survival analysis was performed using backward stepwise (likelihood ratio) Cox regression models and included all variables with p < 0.100 in univariate survival analysis. Associations with demographic and clinicopathological variables were analyzed using cross-tabulation analysis and Kendall’s tau test for ordinal variables or Fischer’s exact test for non-ordinal variables. To account for the false discovery rate, the Benjamini–Hochberg (Simes) method was applied to the pool of all p values of this study (n = 172) (Benjamini and Hochberg 1995). All the p values are given uncorrected. Those p values which have lost significance are marked accordingly.
Results
Cohort characteristics
A total of 580 patients (441 with PR-GC and 139 with NAT-GC) were included in this study. For further information on baseline characteristics, see Supplementary Table 1. The median OS for the PR-GC cohort was 14.7 months [95% confidence interval (CI) 12.6–16.7 months], and the median CSS was 16.0 months (95% CI 13.5–18.5). In NAT-GC, the median OS was 22.4 months (95% CI 17.3–27.4), and CSS was 24.6 months (95% CI 19.9–29.4).
Accumulation of LAG3 + cells in GC
LAG3 + immune cells were detected in both the stromal and intraepithelial compartments. The staining pattern varied from weak and dot-like to strong and membranous/cytoplasmic (Fig. 1d–g), which was also recently demonstrated in melanoma (Johnson et al. 2022). The distribution of LAG3 + cell densities differed significantly between the two sub-cohorts (p < 0.0001). In PR-GC, the median density of LAG3 + immune cells was 55.15 cells/mm2 (range 3.57–1687.63) in TC and 70.35 cells/mm2 (1.91–1858.35) in IM, and it differed significantly between both compartments (p < 0.001). This pattern of distribution was not observed in the NAT-GC, where the median numbers were 41.37 (3.56–543.43) in the TC and 43.28 (3.60–537.73) in the IM.
Survival analysis
To incorporate LAG3 expression in the survival analysis, median LAG3 + cell densities were used in the first step to dichotomize cohorts into low and high LAG3 expression groups. In PR-GC, no significant differences were found in OS (data not shown) and CSS between the two groups (Suppl. Table 2; Suppl. Fig. 1).
Similar results were found in the NAT-GC sub-cohort when comparing low vs. high LAG3 expression in TC (Suppl. Table 2; Suppl. Fig. 2). Interestingly, NAT-GC patients with higher LAG3 + cell density at the IM had significantly longer OS than in the LAG3 low subgroup (28.0 vs. 13.3 months, p = 0.025, log-rank test; data not shown).
LAG3 expression predicts favorable prognosis when divided by biological cut-off
Dichotomization of patient cohorts at median values may not reflect biological relevant cut-offs, as it was previously shown by our group, e.g., for neutrophil counts in GC (Clausen et al. 2020). Therefore, we explored an alternative dichotomization approach for LAG3 using Cutoff Finder (Budczies et al. 2012) and CSS as outcome measure. Cutoff Finder is a web application, which enables detection of optimal biomarker cut-offs. Interestingly, using this tool in PR-GC, the most significant cut-offs were 21.45 cells/mm2 in TC and 208.50 cells/mm2 at IM. Now, the median CSS of patients with high LAG3 expression in TC was 17.9 months (95% CI 13.7–22.1 months) in comparison to 10.1 months (95% CI 7.5–12.7) of LAG3-low patients (p = 0.008). After splitting cases with high and low LAG3 expression in the IM, the median CSS reached 33.8 months (95% CI 21.4–46.3) vs. 14.7 months (95% CI 12.1–17.2), respectively (p = 0.006) (Fig. 2; Suppl. Table 2). However, when using multivariate analysis, LAG3 expression was not found to be an independent prognostic factor (Table 1).
Kaplan–Meier curves in primary resected gastric cancer: cancer-specific survival according to LAG3 + cell density groups split by determined cut-off values (a tumor center, p = 0.008, log-rank test; b invasive margin, p = 0.006, log-rank test; small vertical lines in the graph indicate censored data)
In NAT-GC, the most significant cut-off of LAG3 + cell density in TC was 12.62 cells/mm2. The median CSS was 27.3 months (95% CI 19.6–35.1) and 13.2 months (95% CI 5.6–20.8) in patients with high vs. low LAG3 expression, respectively (p = 0.003). LAG3 expression in TC was an independent prognosticator of CSS (Table 1). There was no significant difference in median CSS among LAG3 groups of IM even at the most optimal calculated cut-off (123.00 cells/mm2): 28.0 months (95% CI non-calculable), LAG3-high, vs. 22.4 months (15.5–29.2), LAG3-low (p = 0.136) (Fig. 3; Suppl. Table 2). These data show that the biological effect of LAG3 depends on cell numbers, location, and treatment.
Kaplan–Meier curves in neoadjuvantly treated gastric cancer: cancer-specific survival according to LAG3 + cell density groups split by determined cut-off values (a tumor center, p = 0.003, log-rank test; b invasive margin, p = 0.136, log-rank test; small vertical lines in the graph indicate censored data)
Association with demographical and clinicopathological features
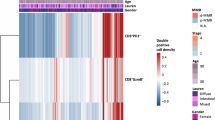
The associations between LAG3 expression (dichotomized by biological cut-offs) and clinicopathological characteristics are summarized in Tables 2 and 3. In PR-GC (Table 2), a significant association was found between LAG3 expression in IM and the Lauren phenotype (p < 0.001). Enrichment of LAG3 + cells in the IM was more likely to be observed in males (p = 0.050). Higher LAG3 + density in both compartments was associated with PD-1 expression (p = 0.004, TC, and p = 0.002, IM). LAG3 + cell density in IM showed a correlation with EBV, MSI- and PD-L1-status (p < 0.001). No significant associations were found between LAG3 expression and patient age, tumor location, pTNM categories, presence of LVI, pR status, H. pylori infection, and MET-status. For analysis based on median values, see Suppl. Table 3.
In NAT-GC (Table 3), LAG3 expression (dichotomized by adapted cut-off value) in TC was associated with Lauren phenotype (p < 0.001), presence of distant metastases (p = 0.015) and enrichment of PD-L1 + immune cells (p = 0.001). LAG3 + cell density in IM inversely correlated with ypT (p = 0.013) and UICC stage (p = 0.009). No associations were observed between LAG3 + cell infiltration and patient age, sex, tumor location, ypN, presence of LVI, and HER2 status. LAG3 expression in both compartments was associated with PD-L1 expression in tumor cells (p = 0.040, TC, and p = 0.001, IM). No significant associations were found between LAG3 expression and EBV, MSI, MET, and PD1 status, probably because of the limited number of cases. For analysis based on median values, see Suppl. Table 4.
Discussion
Recently, LAG3 has gained increasing attention in immuno-oncology, and its putative role as a biomarker has been studied in various solid tumors. In this retrospective study of GC, we demonstrated the distribution patterns of LAG3 + cells within tumor tissues and their association with clinicopathological data and survival. The main findings of the current study are as follows: (1) LAG3 + immune cell density differs significantly between PR-GC and NAT-GC; (2) LAG3 + cell density shows significant differences in spatial distribution patterns between tumor compartments in PR-GC, which is not seen in NAT-GC; (3) CSS is significantly longer for patients with LAG3 expression above the calculated cut-offs in both cohorts; (4) LAG3 + immune cell density in TC is an independent prognostic factor of CSS in NAT-GC. Moreover, LAG3 + immune cell density is associated with various parameters, including sex, tumor location, Lauren phenotype, and HER2-, EBV-, MSI, and PD-1/PD-L1 status in PR-GC. In NAT-GC, an association with ypT and ypN categories, as well as PD-L1 expression in tumor cells and immune cells is observed.
High LAG3 + expression has been associated with poor prognosis in non-small cell lung cancer (He et al. 2017; Shepherd et al. 2022), renal cell cancer (Giraldo et al. 2015), hepatocellular cancer (Guo et al. 2020), and pancreatic cancer (Seifert et al. 2021) but with favorable prognosis in breast cancer (Burugu et al. 2017), colon cancer (Rhyner Agocs et al. 2021), and esophageal cancer (Zhang et al. 2018; Gebauer et al. 2020). In a large cohort of colorectal cancers, the prognostic effect differed based on the spatial location of LAG3 + tumor-infiltrating lymphocytes (TILs), and showed poor CSS in cases with high intra-tumoral LAG3 + TILs and improved CSS when LAG3 was identified in stromal immune cells (Al-Badran et al. 2021). Another interesting finding was an association of LAG3 with better outcome in early-stage tumors (Saleh et al. 2019). What is more, although LAG3 expression has been associated with more aggressive tumor features in breast cancer, patients with high LAG3 + intraepithelial TILs showed improved survival (Burugu et al. 2017). Interestingly, low LAG3 + expression became an independent predictor of favorable prognosis in breast cancers that were treated with neoadjuvant chemotherapy (Asano et al. 2022). To sum up, previous results suggest that the prognostic effect of LAG3 may depend on the tumor type, spatial location of TILs, clinical stage, and therapeutic approach (primary surgery vs. NAT).
LAG3 expression has also been observed in GC. A prospective study of solid tumors performed by Lee et al. (2019) included 53 metastatic GCs cases. They analyzed TMAs (two tissue cores from each GC) and regarded positive cases as any presence of LAG3 + immune cells regardless of their number (Lee et al. 2019). LAG3 expression was found in 24.7% of all GCs and was associated only with EBV status. No significant differences in sex, age, primary tumor site, Lauren classification, HER2 status, or OS were found between patients with or without LAG3 expression.
Lv et al. (2021) investigated the clinical and molecular correlations of LAG3 + cell infiltration in a large set of 464 GCs. Using TMAs (one core from each GC), they evaluated the mean density of LAG3 + immune cells and dichotomized all cases into LAG3 + low and LAG3 + high groups based on the median value of 59 cells/HPF. In their study, LAG3 + cell infiltration was associated with male sex and immuno-evasive contexture, and was found to be an independent adverse prognostic factor for both OS and disease-free survival (Lv et al. 2021). LAG3 showed higher expression in EBV + subtype and defective MLH1 subtype, as well as predicted poor survival in these subtypes (Lv et al. 2021).
Park et al. (2021) performed conventional and multiplex IHC on GC TMAs (n = 385, two tissue cores from each case) for immune cell markers and immune checkpoint receptors, including LAG3, as well as survival analysis. Positive LAG3 expression was defined as immunostaining in ≥ 5% of the immune cells, and it was found in 45.5%, 29.6%, and 50.1% of cases in the TC, IM, and TC or IM, respectively. LAG3 was expressed more commonly by CD3+/CD8+ T cells in the tumor area than in the stromal compartment (Park et al. 2021). It was associated with male sex, distal location, intestinal and mixed subtypes by Lauren, and better prognosis in multivariate survival analysis (Park et al. 2021).
These varying findings regarding the role of LAG3 in malignant tumors could be explained by the use of different assessment and scoring methods. Currently, there is no standardized assay for LAG3 expression in GC tissue. Several approaches, including absolute number per HPF (Lv et al. 2021) or per mm2 (Shepherd et al. 2022), and cut-offs, such as any positivity (Burugu et al. 2017; Lee et al. 2019; Rhyner Agocs et al. 2021), median number (Lv et al. 2021), 1% of all nucleated cells (Johnson et al. 2022), or 5% of all TILs (Park et al. 2021), have been used in GC and other solid malignancies. Furthermore, the use of the TMA technique might influence the results because of the limited amount of tissue, uneven distribution of infiltrating immune cells, and heterogeneous expression of immune checkpoint molecules. It has been demonstrated on TMAs as substitutes for core biopsies that at least five biopsies are needed to reflect the objective status of PD-L1 expression as in whole sections of GC (Ye et al. 2020).
In previous studies, LAG3 expression was correlated with EBV + and MSI-H GC molecular subtypes. Our results confirm this association in a Western GC cohort, however only when using the median cut-off values (an approach used by Lv et al.). Both subtypes are known to contain a high number of TILs. DNA mismatch repair-deficient tumors harbor large amounts of somatic mutations and tumor-specific neo-antigens that trigger neo-antigen-specific T cells and make them sensitive to immune checkpoint blockade (Le et al. 2017). Similarly, EBV evokes an active immune response that leads to enrichment in activated CD8 + T cells (Landais et al. 2005; van Beek et al. 2006). In a study of Hodgkin’s lymphoma, EBV infection increased gene expression of LAG3 and immunosuppressive cytokines associated with type-1 T regulatory cells (Tr1) (Morales et al. 2014).
Although expression of inhibitory receptors is a hallmark of T cell exhaustion, they are transiently expressed already on activated effector T cells (Wherry and Kurachi 2015). According to a study by Bae et al. (2014), the majority of LAG3 is localized in lysosomes in resting cells and translocates to the cell surface upon stimulation. This could explain the enrichment of LAG3 + cells in early-invasive tumors (pT1/2) despite LAG3 classical role in immunosuppressive TME. We assume that, in such cases, LAG3 expression could rather be a sign of an activated immune response. This feature makes the LAG3 protein an attractive marker for the assessment of potential anti-tumor response and, thus, patient stratification for immunotherapy.
Another interesting finding is higher LAG3 expression in males, which was also demonstrated by both larger Asian studies (Park et al. 2021; Lv et al. 2021). There is increasing awareness about sexual dimorphism in the immune response in solid cancers and its effect on patient outcomes. In a meta-analysis of 17 clinical trials, males appeared to benefit more frequently from ICI therapy than females (Parmar et al. 2022). Higher tumor mutational burden/antigenicity and T cell-dominating inflammation are thought to be key features of such differences in males (Conforti et al. 2019). Previously, tumor-associated neutrophils at the invasive front were found to be an independent predictor of CSS in females of the same PR-GC cohort (Clausen et al. 2020). Current findings expand the sex-based differences in TME of PR-GC, now also including LAG3.
Furthermore, NAT also affects TME. Chemotherapeutics and anticancer agents increase tumor antigenicity and response of CTLs or inhibit immunosuppressive pathways, thus favoring anti-tumor response (Galluzzi et al. 2015). Previous GC studies of paired pre-NAT biopsies and post-NAT resection specimens demonstrated that chemotherapy increases CD8 + T cell density (Yu et al. 2019; Wei et al. 2021; Christina Svensson et al. 2021). It also induces macrophage markers CD68 and CD163 (Wei et al. 2021) and immune checkpoint molecules PD1, PD-L1, and TIM3 (Yu et al. 2019), but decreases CD20 + B cell density (Christina Svensson et al. 2021). However, there are no data regarding LAG3 in GC. Previous studies on esophageal and rectal cancers revealed upregulation of immune checkpoint molecules, including LAG3, after chemo-radiation (Kelly et al. 2018; Peng et al. 2021). Unfortunately, we were unable to characterize the direct effect of NAT on the TME. However, in our study, the median density of LAG3 + cells was lower in NAT-GC than that in PR-GC. There was no significant difference in LAG3 + cell distribution between TC and IM and among molecular subtypes (i.e., EBV and MSI) in NAT-GC, as observed in PR-GC. Collectively, these findings support the notion that NAT also affects the expression of LAG3 in GC.
Our study has several limitations. This retrospective study was restricted to immunohistochemical assessment of LAG3. No further subtyping of LAG3 + immune cells was performed. In the NAT-GC sub-cohort, different treatment schemes and doses were used (according to the changing treatment protocols). No matched pre- and post-treatment samples were compared. Digital analysis is performed only on sections where marking of tumor compartments was possible. Some samples lacked IM values because the stained sections included neoplastic tissue only (marked as TC). However, this is a large and well-characterized set of both PR-GC and NAT-GC. To our knowledge, this is the first study to investigate LAG3 expression in GC of Caucasian origin. The assessment was performed on whole tissue sections to reduce the under- or over-representation of LAG3 + cells due to their heterogeneous distribution within the TME. The same marking techniques and software settings were used for the entire cohort. To increase the reproducibility of the results, a dichotomous scoring method was used.
In conclusion, despite the postulated immunosuppressive role of LAG3 within the TME of solid tumors, the current findings demonstrate differences in the TME among the main histological and molecular subtypes of GC. A high density of LAG3 + cells predicts favorable prognosis in NAT-GC. LAG3 may have different, stage-based functional roles within the TME. Increased numbers of LAG3 + cells within GC tissue could be a sign of crosstalk between cancer and immune cells rather than a sign of exhausted, dysfunctional T cells. The precise mechanisms by which LAG3 regulates T cell function require further investigation. The current findings also raise an important issue in GC immune checkpoint studies based on TMA analysis using single-tissue cores. In the present study, LAG3 + immune cells were detectable in all cases and demonstrated spatial heterogeneity. Considering the popularity of LAG3 as a potential biomarker in cancer studies, a robust LAG3 assay should be developed. Cut-offs may vary between TC and IM, as well as between primarily resected and neo-adjuvantly treated GCs, which merits specific consideration. Matters are far more complicated than anticipated.
Availability of data and material
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
Abbreviations
- CI:
-
Confidence interval
- CSS:
-
Cancer-specific survival
- CTLA4:
-
Cytotoxic T-lymphocyte-associated protein 4
- CTLs:
-
Cytotoxic T lymphocytes
- EBV:
-
Epstein–Barr virus
- GC:
-
Adenocarcinoma of the stomach and gastroesophageal junction; gastric cancer
- ICI:
-
Immune checkpoint inhibitors
- IM:
-
Invasive margin
- LAG3:
-
Lymphocyte activation gene 3
- LVI:
-
Lymphovascular invasion
- MSI:
-
Microsatellite instability
- NAT-GC:
-
Neoadjuvantly treated gastric cancer
- OS:
-
Overall survival
- PD-1:
-
Programmed cell death 1
- PD-L1:
-
Programmed cell death 1 ligand
- pR:
-
Status of resection lines
- PR-GC:
-
Primarily resected gastric cancer
- TC:
-
Tumor center
- TILs:
-
Tumor-infiltrating lymphocytes
- TMAs:
-
Tissue microarrays
- TME:
-
Tumor microenvironment
References
Al-Badran SS, Grant L, Campo MV et al (2021) Relationship between immune checkpoint proteins, tumour microenvironment characteristics, and prognosis in primary operable colorectal cancer. J Pathol Clin Res 7:121–134. https://doi.org/10.1002/cjp2.193
Anderson AC, Joller N, Kuchroo VK (2016) Lag-3, Tim-3, and TIGIT: co-inhibitory receptors with specialized functions in immune regulation. Immunity 44:989–1004. https://doi.org/10.1016/j.immuni.2016.05.001
Andrews LP, Marciscano AE, Drake CG, Vignali DAA (2017) LAG3 (CD223) as a cancer immunotherapy target. Immunol Rev 276:80–96. https://doi.org/10.1111/imr.12519
Arnold M, Rutherford M, Lam F, et al (2019) ICBP SURVMARK-2 online tool: International Cancer Survival Benchmarking. http://gco.iarc.fr/survival/survmark. Accessed 1 Mar 2023
Asano Y, Kashiwagi S, Takada K et al (2022) Clinical significance of expression of immunoadjuvant molecules (LAG-3, TIM-3, OX-40) in neoadjuvant chemotherapy for breast cancer. Anticancer Res 42:125–136. https://doi.org/10.21873/anticanres.15466
Bae J, Lee SJ, Park C-G et al (2014) Trafficking of LAG-3 to the surface on activated T cells via its cytoplasmic domain and protein kinase C signaling. J Immunol 193:3101–3112. https://doi.org/10.4049/jimmunol.1401025
Baixeras E, Huard B, Miossec C et al (1992) Characterization of the lymphocyte activation gene 3-encoded protein. A new ligand for human leukocyte antigen class II antigens. J Exp Med 176:327–337. https://doi.org/10.1084/jem.176.2.327
Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc Ser B (methodol) 57:289–300
Blank CU, Haining WN, Held W et al (2019) Defining “T cell exhaustion.” Nat Rev Immunol 19:665–674. https://doi.org/10.1038/s41577-019-0221-9
Böger C, Behrens H-M, Mathiak M et al (2016) PD-L1 is an independent prognostic predictor in gastric cancer of Western patients. Oncotarget 7:24269–24283. https://doi.org/10.18632/oncotarget.8169
Böger C, Krüger S, Behrens HM et al (2017) Epstein-Barr virus-associated gastric cancer reveals intratumoral heterogeneity of PIK3CA mutations. Ann Oncol 28:1005–1014. https://doi.org/10.1093/annonc/mdx047
Brierley JD, Gospodarowicz MK, Wittekind C (2017) TNM classification of malignant tumours. Wiley, New York
Budczies J, Klauschen F, Sinn BV et al (2012) Cutoff Finder: a comprehensive and straightforward Web application enabling rapid biomarker cutoff optimization. PLoS One 7:e51862. https://doi.org/10.1371/journal.pone.0051862
Burugu S, Gao D, Leung S et al (2017) LAG-3+ tumor infiltrating lymphocytes in breast cancer: clinical correlates and association with PD-1/PD-L1+ tumors. Ann Oncol 28:2977–2984. https://doi.org/10.1093/annonc/mdx557
Christina Svensson M, Lindén A, Nygaard J et al (2021) T cells, B cells, and PD-L1 expression in esophageal and gastric adenocarcinoma before and after neoadjuvant chemotherapy: relationship with histopathological response and survival. Oncoimmunology 10:1921443. https://doi.org/10.1080/2162402X.2021.1921443
Clausen F, Behrens H-M, Krüger S, Röcken C (2020) Sexual dimorphism in gastric cancer: tumor-associated neutrophils predict patient outcome only for women. J Cancer Res Clin Oncol 146:53–66. https://doi.org/10.1007/s00432-019-03082-z
Conforti F, Pala L, Bagnardi V et al (2019) Sex-based differences of the tumor mutational burden and T-cell inflammation of the tumor microenvironment. Ann Oncol 30:653–655. https://doi.org/10.1093/annonc/mdz034
Galluzzi L, Vacchelli E, Bravo-San Pedro J-M et al (2014) Classification of current anticancer immunotherapies. Oncotarget 5:12472–12508
Galluzzi L, Buqué A, Kepp O et al (2015) Immunological effects of conventional chemotherapy and targeted anticancer agents. Cancer Cell 28:690–714. https://doi.org/10.1016/j.ccell.2015.10.012
Gebauer F, Krämer M, Bruns C et al (2020) Lymphocyte activation gene-3 (LAG3) mRNA and protein expression on tumour infiltrating lymphocytes (TILs) in oesophageal adenocarcinoma. J Cancer Res Clin Oncol 146:2319–2327. https://doi.org/10.1007/s00432-020-03295-7
Giraldo NA, Becht E, Pagès F et al (2015) Orchestration and prognostic significance of immune checkpoints in the microenvironment of primary and metastatic renal cell cancer. Clin Cancer Res 21:3031–3040. https://doi.org/10.1158/1078-0432.CCR-14-2926
Guo M, Yuan F, Qi F et al (2020) Expression and clinical significance of LAG-3, FGL1, PD-L1 and CD8(+)T cells in hepatocellular carcinoma using multiplex quantitative analysis. J Transl Med 18:306. https://doi.org/10.1186/s12967-020-02469-8
Guy C, Mitrea DM, Chou P-C et al (2022) LAG3 associates with TCR-CD3 complexes and suppresses signaling by driving co-receptor-Lck dissociation. Nat Immunol 23:757–767. https://doi.org/10.1038/s41590-022-01176-4
He X, Xu C (2020) Immune checkpoint signaling and cancer immunotherapy. Cell Res 30:660–669. https://doi.org/10.1038/s41422-020-0343-4
He Y, Yu H, Rozeboom L et al (2017) LAG-3 protein expression in non-small cell lung cancer and its relationship with PD-1/PD-L1 and tumor-infiltrating lymphocytes. J Thorac Oncol 12:814–823. https://doi.org/10.1016/j.jtho.2017.01.019
Hendry S, Salgado R, Gevaert T et al (2017) Assessing tumor-infiltrating lymphocytes in solid tumors: a practical review for pathologists and proposal for a standardized method from the international immunooncology biomarkers working group: part 1: assessing the host immune response, TILs in invasive breast carcinoma and ductal carcinoma in situ, metastatic tumor deposits and areas for further research. Adv Anat Pathol 24:235–251. https://doi.org/10.1097/PAP.0000000000000162
Huang C-T, Workman CJ, Flies D et al (2004) Role of LAG-3 in regulatory T cells. Immunity 21:503–513. https://doi.org/10.1016/j.immuni.2004.08.010
Johnson L, McCune B, Locke D et al (2022) Development of a LAG-3 immunohistochemistry assay for melanoma. J Clin Pathol. https://doi.org/10.1136/jclinpath-2022-208254
Kelly RJ, Zaidi AH, Smith MA et al (2018) The dynamic and transient immune microenvironment in locally advanced esophageal adenocarcinoma post chemoradiation. Ann Surg 268:992–999. https://doi.org/10.1097/SLA.0000000000002410
Kouo T, Huang L, Pucsek AB et al (2015) Galectin-3 shapes antitumor immune responses by suppressing CD8+ T cells via LAG-3 and inhibiting expansion of plasmacytoid dendritic cells. Cancer Immunol Res 3:412–423. https://doi.org/10.1158/2326-6066.CIR-14-0150
Landais E, Saulquin X, Houssaint E (2005) The human T cell immune response to Epstein–Barr virus. Int J Dev Biol 49:285–292. https://doi.org/10.1387/ijdb.041947el
Le DT, Durham JN, Smith KN et al (2017) Mismatch repair deficiency predicts response of solid tumors to PD-1 blockade. Science 357:409–413. https://doi.org/10.1126/science.aan6733
Lee SJ, Byeon S-J, Lee J et al (2019) LAG3 in solid tumors as a potential novel immunotherapy target. J Immunother 42:279–283. https://doi.org/10.1097/CJI.0000000000000283
Lv K, Li R, Cao Y et al (2021) Lymphocyte-activation gene 3 expression associates with poor prognosis and immunoevasive contexture in Epstein–Barr virus-positive and MLH1-defective gastric cancer patients. Int J Cancer 148:759–768. https://doi.org/10.1002/ijc.33358
Maeda TK, Sugiura D, Okazaki I-M et al (2019) Atypical motifs in the cytoplasmic region of the inhibitory immune co-receptor LAG-3 inhibit T cell activation. J Biol Chem 294:6017–6026. https://doi.org/10.1074/jbc.RA119.007455
Mathiak M, Warneke VS, Behrens H-M et al (2017) Clinicopathologic characteristics of microsatellite instable gastric carcinomas revisited: urgent need for standardization. Appl Immunohistochem Mol Morphol 25:12–24. https://doi.org/10.1097/PAI.0000000000000264
Metzger M-L, Behrens H-M, Böger C et al (2016) MET in gastric cancer—discarding a 10% cutoff rule. Histopathology 68:241–253. https://doi.org/10.1111/his.12745
Morales O, Mrizak D, François V et al (2014) Epstein–Barr virus infection induces an increase of T regulatory type 1 cells in Hodgkin lymphoma patients. Br J Haematol 166:875–890. https://doi.org/10.1111/bjh.12980
National Cancer Institute SEER*Explorer: an interactive website for SEER cancer statistics. https://seer.cancer.gov/explorer/. Accessed 1 Mar 2023
Paik J (2022) Nivolumab plus relatlimab: first approval. Drugs 82:925–931
Park Y, Seo AN, Koh J et al (2021) Expression of the immune checkpoint receptors PD-1, LAG3, and TIM3 in the immune context of stage II and III gastric cancer by using single and chromogenic multiplex immunohistochemistry. Oncoimmunology 10:1954761. https://doi.org/10.1080/2162402X.2021.1954761
Parmar K, Subramanyam S, Attwood K et al (2022) Anti PD-1/anti PDL-1 inhibitors in advanced gastroesophageal cancers: a systematic review and meta-analysis of phase 2/3 randomized controlled trials. Pharmaceutics. https://doi.org/10.3390/pharmaceutics14091953
Peng Q-Q, Li J-L, Xin P-L et al (2021) Assessment of the expression and response of PD-1, LAG-3, and TIM-3 after neoadjuvant radiotherapy in rectal cancer. Neoplasma 68:742–750. https://doi.org/10.4149/neo_2021_201210N1341
Rhyner Agocs G, Assarzadegan N, Kirsch R et al (2021) LAG-3 expression predicts outcome in stage II colon cancer. J Pers Med. https://doi.org/10.3390/jpm11080749
Saleh RR, Peinado P, Fuentes-Antrás J et al (2019) Prognostic value of lymphocyte-activation gene 3 (LAG3) in cancer: a meta-analysis. Front Oncol 9:1040. https://doi.org/10.3389/fonc.2019.01040
Schoop H, Bregenzer A, Halske C et al (2020) Therapy resistance in neoadjuvantly treated gastric cancer and cancer of the gastroesophageal junction is associated with an increased expression of immune checkpoint inhibitors-comparison against a therapy naïve cohort. Transl Oncol 13:165–176. https://doi.org/10.1016/j.tranon.2019.11.004
Seifert L, Plesca I, Müller L et al (2021) LAG-3-expressing tumor-infiltrating T cells are associated with reduced disease-free survival in pancreatic cancer. Cancers (basel). https://doi.org/10.3390/cancers13061297
Shan C, Li X, Zhang J (2020) Progress of immune checkpoint LAG-3 in immunotherapy. Oncol Lett 20:207. https://doi.org/10.3892/ol.2020.12070
Shepherd DJ, Tabb ES, Kunitoki K et al (2022) Lymphocyte-activation gene 3 in non-small-cell lung carcinomas: correlations with clinicopathologic features and prognostic significance. Mod Pathol 35:615–624. https://doi.org/10.1038/s41379-021-00974-9
Shergold AL, Millar R, Nibbs RJB (2019) Understanding and overcoming the resistance of cancer to PD-1/PD-L1 blockade. Pharmacol Res 145:104258. https://doi.org/10.1016/j.phrs.2019.104258
Sung H, Ferlay J, Siegel RL et al (2021) Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 71:209–249. https://doi.org/10.3322/caac.21660
Tawbi HA, Schadendorf D, Lipson EJ et al (2022) Relatlimab and nivolumab versus nivolumab in untreated advanced melanoma. N Engl J Med 386:24–34. https://doi.org/10.1056/NEJMoa2109970
Triebel F, Jitsukawa S, Baixeras E et al (1990) LAG-3, a novel lymphocyte activation gene closely related to CD4. J Exp Med 171:1393–1405. https://doi.org/10.1084/jem.171.5.1393
van Beek J, zur Hausen A, Snel SN et al (2006) Morphological evidence of an activated cytotoxic T-cell infiltrate in EBV-positive gastric carcinoma preventing lymph node metastases. Am J Surg Pathol 30:59–65. https://doi.org/10.1097/01.pas.0000176428.06629.1e
van den Bulk J, Verdegaal EM, de Miranda NF (2018) Cancer immunotherapy: broadening the scope of targetable tumours. Open Biol. https://doi.org/10.1098/rsob.180037
Wang J, Sanmamed MF, Datar I et al (2019) Fibrinogen-like protein 1 is a major immune inhibitory ligand of LAG-3. Cell 176:334-347.e12. https://doi.org/10.1016/j.cell.2018.11.010
Warneke VS, Behrens H-M, Böger C et al (2013) Her2/neu testing in gastric cancer: evaluating the risk of sampling errors. Ann Oncol 24:725–733. https://doi.org/10.1093/annonc/mds528
Wei Q, Xu Q, Yuan X et al (2021) Immunological impact of chemotherapy on the tumor microenvironment in gastric cancer. J Surg Oncol 123:1708–1715. https://doi.org/10.1002/jso.26449
Wherry EJ, Kurachi M (2015) Molecular and cellular insights into T cell exhaustion. Nat Rev Immunol 15:486–499. https://doi.org/10.1038/nri3862
Woo S-R, Turnis ME, Goldberg MV et al (2012) Immune inhibitory molecules LAG-3 and PD-1 synergistically regulate T-cell function to promote tumoral immune escape. Cancer Res 72:917–927. https://doi.org/10.1158/0008-5472.CAN-11-1620
Workman CJ, Dugger KJ, Vignali DAA (2002) Cutting edge: molecular analysis of the negative regulatory function of lymphocyte activation gene-3. J Immunol 169:5392–5395. https://doi.org/10.4049/jimmunol.169.10.5392
Xu F, Liu J, Liu D et al (2014) LSECtin expressed on melanoma cells promotes tumor progression by inhibiting antitumor T-cell responses. Cancer Res 74:3418–3428. https://doi.org/10.1158/0008-5472.CAN-13-2690
Ye M, Huang D, Zhang Q et al (2020) Heterogeneous programmed death-ligand 1 expression in gastric cancer: comparison of tissue microarrays and whole sections. Cancer Cell Int 20:186. https://doi.org/10.1186/s12935-020-01273-0
Yu Y, Ma X, Zhang Y et al (2019) Changes in expression of multiple checkpoint molecules and infiltration of tumor immune cells after neoadjuvant chemotherapy in gastric cancer. J Cancer 10:2754–2763. https://doi.org/10.7150/jca.31755
Zhang Y, Zheng J (2020) Functions of immune checkpoint molecules beyond immune evasion. Adv Exp Med Biol 1248:201–226. https://doi.org/10.1007/978-981-15-3266-5_9
Zhang Y, Liu Y-D, Luo Y-L et al (2018) Prognostic value of lymphocyte activation gene-3 (LAG-3) expression in esophageal squamous cell carcinoma. J Cancer 9:4287–4293. https://doi.org/10.7150/jca.26949
Funding
Open Access funding enabled and organized by Projekt DEAL. The authors declare that no funds, grants, or other support was received during the preparation of this manuscript.
Author information
Authors and Affiliations
Contributions
Conceptualization, CR and DU; methodology, DU, HMB, and SK; software, HMB; validation, DU and HMB; investigation, DU and HMB; resources, CR, SH, SK, TB, and UE; data curation, HMB; writing—original draft, DU; writing—review and editing, CR, DU, HMB, SH, SK, TB, and UE; visualization, DU and HMB; supervision, CR; project administration, CR.
Corresponding authors
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose.
Ethics approval
All procedures followed were in accordance with the ethical standards of the responsible committee on human experimentation (institutional and national) and the Helsinki Declaration of 1964 and later versions. Ethics approval was obtained from the local ethical review board of the University Hospital Schleswig–Holstein, Kiel, Germany (D 453/10, D 525/15).
Consent to participate
Informed consent was obtained from all individual participants included in the study.
Consent to publish
Not required.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ulase, D., Behrens, HM., Krüger, S. et al. LAG3 in gastric cancer: it’s complicated. J Cancer Res Clin Oncol 149, 10797–10811 (2023). https://doi.org/10.1007/s00432-023-04954-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00432-023-04954-1