Abstract
Purpose
It was reported that individual heterogeneity among malignancies (IHAM) might correlate well to the prognosis of lung cancer; however, seldom radiomic study is on this field. Standard deviation (SD) in statistics could scale average amount of variability of a variable; therefore, we used SD of CT feature (FeatureSD) among primary tumor and malignant lymph nodes (LNs) in an individual to represent IHAM, and its prognostic ability was explored.
Methods
The enrolled patients who had accepted PET/CT scans were selected from our previous study (ClinicalTrials.gov, NCT03648151). The patients had primary tumor and at least one LN, and standardized uptake value of LN higher than 2.0 and 2.5 were enrolled as the cohort 1 (n = 94) and 2 (n = 88), respectively. FeatureSD from the combined or thin-section CT were calculated among primary tumor and malignant LNs in each patient, and were separately selected by the survival XGBoost method. Finally, their prognostic ability was compared to the significant patient characteristics identified by the Cox regression.
Results
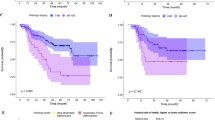
In the univariate and multi-variate Cox analysis, surgery, target therapy, and TNM stage were significantly against OS in the both cohorts. In the survival XGBoost analysis of the thin-section CT dataset, none FeatureSD could be repeatably ranked on the top list of the both cohorts. For the combined CT dataset, only one FeatureSD ranked in the top three of both cohorts, but the three significant factors in the Cox regression were not even on the list. Both in the cohort 1 and 2, C-index of the model composed of the three factors could be improved by integrating the continuous FeatureSD; furthermore, that of each factor was obviously lower than FeatureSD.
Conclusion
Standard deviation of CT features among malignant foci within an individual was a powerful prognostic factor in vivo for lung cancer patients.






Similar content being viewed by others
Data availability
The datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request.
References
Bashir U, Siddique MM, McLean E, Goh V, Cook GJ (2016) Imaging heterogeneity in lung cancer: techniques, applications, and challenges. AJR Am J Roentgenol 207(3):534–543. https://doi.org/10.2214/AJR.15.15864
Carkaci S, Adrada BE, Rohren E, Wei W, Quraishi MA, Mawlawi O, Yang W (2012) Semiquantitative analysis of maximum standardized uptake values of regional lymph nodes in inflammatory breast cancer: is there a reliable threshold for differentiating benign from malignant? Acad Radiol 19(5):535–541. https://doi.org/10.1016/j.acra.2012.01.001
Galvan A, Ioannidis JP, Dragani TA (2010) Beyond genome-wide association studies: genetic heterogeneity and individual predisposition to cancer. Trends Genet 26(3):132–141. https://doi.org/10.1016/j.tig.2009.12.008
Hata A, Katakami N, Yoshioka H, Kaji R, Masago K, Fujita S, Yatabe Y (2015) Spatiotemporal T790M heterogeneity in individual patients with EGFR-mutant non-small-cell lung cancer after acquired resistance to EGFR-TKI. J Thorac Oncol 10(11):1553–1559. https://doi.org/10.1097/JTO.0000000000000647
Liegmann AS, Heselmeyer-Haddad K, Lischka A, Hirsch D, Chen WD, Torres I, Ried T (2021) Single cell genetic profiling of tumors of breast cancer patients aged 50 years and older reveals enormous intratumor heterogeneity independent of individual prognosis. Cancers (basel). https://doi.org/10.3390/cancers13133366
Lim SB, Yeo T, Lee WD, Bhagat AAS, Tan SJ, Tan DSW, Lim CT (2019) Addressing cellular heterogeneity in tumor and circulation for refined prognostication. Proc Natl Acad Sci 116(36):17957–17962. https://doi.org/10.1073/pnas.1907904116
Liu Y, Kim J, Balagurunathan Y, Li Q, Garcia AL, Stringfield O, Gillies RJ (2016) Radiomic features are associated with EGFR mutation status in lung adenocarcinomas. Clin Lung Cancer 17(5):441-448.e446. https://doi.org/10.1016/j.cllc.2016.02.001
Liu J, Dang H, Wang XW (2018) The significance of intertumor and intratumor heterogeneity in liver cancer. Exp Mol Med 50(1):e416–e416. https://doi.org/10.1038/emm.2017.165
Liu WX, Shi M, Su H, Wang Y, He X, Xu LM, Li YX (2019) Effect of age as a continuous variable on survival outcomes and treatment selection in patients with extranodal nasal-type NK/T-cell lymphoma from the China Lymphoma Collaborative Group (CLCG). Aging (albany NY) 11(19):8463–8473. https://doi.org/10.18632/aging.102331
Makohon-Moore AP, Zhang M, Reiter JG, Bozic I, Allen B, Kundu D, Iacobuzio-Donahue CA (2017) Limited heterogeneity of known driver gene mutations among the metastases of individual patients with pancreatic cancer. Nat Genet 49(3):358–366. https://doi.org/10.1038/ng.3764
Pizon M, Zimon D, Carl S, Pachmann U, Pachmann K, Camara O (2013) Heterogeneity of circulating epithelial tumour cells from individual patients with respect to expression profiles and clonal growth (sphere formation) in breast cancer. Ecancermedicalscience 7:343. https://doi.org/10.3332/ecancer.2013.343
Si H, Hao X, Zhang L, Xu X, Cao J, Wu P, Li S (2021) Total lesion glycolysis estimated by a radiomics model from CT image alone. Front Oncol. https://doi.org/10.3389/fonc.2021.664346
Sollini M, Cozzi L, Antunovic L, Chiti A, Kirienko M (2017) PET Radiomics in NSCLC: state of the art and a proposal for harmonization of methodology. Sci Rep 7(1):358. https://doi.org/10.1038/s41598-017-00426-y
VanderWeele DJ, Finney R, Katayama K, Gillard M, Paner G, Imoto S, Nakagawa H (2019) Genomic heterogeneity within individual prostate cancer foci impacts predictive biomarkers of targeted therapy. Eur Urol Focus 5(3):416–424. https://doi.org/10.1016/j.euf.2018.01.006
Yang F, Dong J, Wang X, Fu X, Zhang T (2017) Non-small cell lung cancer: spectral computed tomography quantitative parameters for preoperative diagnosis of metastatic lymph nodes. Eur J Radiol 89:129–135. https://doi.org/10.1016/j.ejrad.2017.01.026
Yip SS, Aerts HJ (2016) Applications and limitations of radiomics. Phys Med Biol 61(13):R150-166. https://doi.org/10.1088/0031-9155/61/13/R150
Zhang L, Fried DV, Fave XJ, Hunter LA, Yang J, Court LE (2015) IBEX: an open infrastructure software platform to facilitate collaborative work in radiomics. Med Phys 42(3):1341–1353. https://doi.org/10.1118/1.4908210
Funding
The work was funded by grants from Science and Technology project of Health Commission of Anhui Province (No. AHWJ2021b148), Collaborative Innovation Center for Molecular Imaging and Precise D&T Center (Grant No. MP201604) and Shenzhen Science and Technology Innovation Commission (JCYJ20200109120205924).
Author information
Authors and Affiliations
Contributions
SH, LS, XS and CJ contributed to the study conception and design. Material preparation, data collection and analysis were performed by XS, WR, LL and. The first draft of the manuscript was written by HX and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Hongwei, S., Xinzhong, H., Huiqin, X. et al. Standard deviation of CT radiomic features among malignancies in each individual: prognostic ability in lung cancer patients. J Cancer Res Clin Oncol 149, 7165–7173 (2023). https://doi.org/10.1007/s00432-023-04649-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00432-023-04649-7