Abstract
Pediatric neuroblastoma is responsible for approximately 8–10% of pediatric tumors, and it is one of the leading causes of tumor-related deaths in children. Although significant progress has been made in the characterization of neuroblastoma in recent years, the mechanisms influencing the prognosis of neuroblastoma patients remain largely unknown. Our aim was to investigate if the major neuroendocrine-associated transcriptional drivers, including ASCL1, NEUROD1, DLL3, NOTCH1, INSM1, MYCL1, POU2F3 and YAP1 are correlated with specific clinical and pathological characteristics. We selected a retrospective series of 46 primary pediatric neuroblastoma, composed of 30 treatment-naïve and 16 post-chemotherapy cases. Gene expression levels were explored by means of quantitative real-time PCR. An increased expression of NOTCH1 (p = 0.005), NEUROD1 (p = 0.0059), and YAP1 (p = 0.0008) was found in stage IV tumors, while the highest levels of MYCL1 and ASCL1 were seen in stages IVS and III, respectively (p = 0.0182 and p = 0.0134). A higher level of NOTCH1 (p = 0.0079) and YAP1 (p = 0.0026) was found in cases with differentiating morphology, while high mitosis-karyorrhexis index cases demonstrated significantly lower levels of POU2F3 (p = 0.0277). High expression of NOTCH1 (p = 0.008), NEUROD1 (p = 0.026), INSM1 (p = 0.010), and YAP1 (p = 0.005) together with stage IV (p = 0.043) was associated with shorter disease-free survival. In summary, our data indicate that the assessment of gene expression levels of neuroendocrine-lineage transcription factors might help to identify neuroblastoma patients with the risk of relapse.
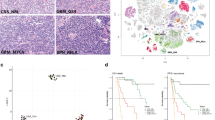
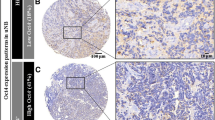
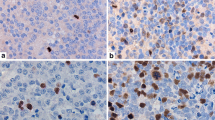
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Neuroblastic tumors represent the most common extra-cranial solid malignancy in the pediatric age. Neuroblastoma is a tumor arising from developing sympathetic nervous system and is responsible for approximately 8–10% of pediatric tumors. Despite advances in molecular profiling and therapeutic options, survival of high-risk neuroblastoma patients remains poor. Neuroblastoma development mechanisms are incompletely understood, but linked to oncogene mutations and/or amplifications [1,2,3,4]. In particular, MYCN represents one of the most important driver genes in neuroblastoma, being gene amplification strongly correlated to unfavorable outcome, although in vitro and in vivo data suggest that there is no direct correlation between a high cellular MYCN protein content and aggressive tumor cell behavior or loss of differentiation [5]. Moreover, basic helix-loop-helix (bHLH) class of transcription factors plays a pivotal role in tissue-specific differentiation and their dysregulation is associated to solid tumor development. Neuronal and neuroendocrine (NE) cell growth and differentiation, as well as their related tumors, are regulated by genes of the Notch, NEUROD and Achaete Scute families [6,7,8]. In particular, in high-grade NE small cell carcinomas, mostly of the lung, a complex genetic regulation has been recently reported with several pathways differentially activated in subgroups of such tumors. Among relevant regulators of NE growth and differentiation are Notch family genes, ASCL1, NEUROD1, HES1, INSM1, POU2F3, YAP1, MYCL1 [9,10,11,12]. In large cell NE carcinomas of the lung, two different molecular profiles were also identified, partly overlapping with those of small cell lung cancer or sharing gene alterations seen in non-small lung carcinomas [13,14,15,16]. Some of these genetic differences were linked to different outcome and response to specific therapies in both types of pulmonary high-grade NE carcinomas [10, 17]. In addition, some of these transcription factors are also express in extra-pulmonary NE carcinomas of various locations and are used as markers of NE differentiation [18,19,20].
Some genes belonging to the abovementioned families have been individually investigated in neuroblastoma and found to be overexpressed and involved in response to specific therapies. In particular MYCN gene is pivotal in neuroblastoma development and progression [1,2,3,4]. Central (intracranial) and peripheral neuroectodermal tumors and cell lines are known to express NEUROD1 and ASCL1 genes [21]. Transcriptional regulators of these two genes were recently identified in neuroblastoma, including the neuronal differentiation markers of Purkinje cells PCP4/PEP19 [22], and ERK [23]. NEUROD1 seems to act mainly through ALK to favor neuroblastoma cell proliferation, directly binding to the promoter region of this gene [24]. ASCL1 is downregulated during neuroblastoma cell differentiation along with upregulation of several genes including IGF2 [25]; moreover, the negative correlation of ASCL1 expression with neuronal differentiation is independent from MYCN gene expression, suggesting that targeting ASCL1 might increase the efficacy of retinoic acid-based differentiating therapies in neuroblastoma [26]. More recently, other regulatory genes were investigated, including INSM1 that is activated by MYCN gene, expressed in a large fraction of neuroblastomas and associated to a shorter survival [27,28,29,30].
Neuroblastic tumors encompass a spectrum of lesions with different pathological features, response to therapies and outcome. We studied the role of major drivers of neuroendocrine-associated transcriptional clustering in a retrospective series of surgically resected pediatric neuroblastomas, with the aim of exploring the correlation with specific pathological and clinical characteristics including response to treatment and survival.
Materials and methods
Case series
We selected a retrospective series of 46 cases of pediatric neuroblastoma, all primitive, operated from 2007 to 2019 at “Città della Salute e della Scienza,” Turin, and treated at the Pediatric Oncology Division of the “Regina Margherita” Children’s Hospital, with sufficient residual histological material for molecular and immunohistochemical analyses. The case series was composed 30 treatment-naïve and 16 post-chemotherapy cases. The chemotherapy consisted of combination treatment, including carboplatin and etoposide.
Clinical and pathological data such as age at diagnosis, sex, tumor location, stage (according to INSS system, see Table 1), International Neuroblastoma Pathology classification (INPC) category [31], mitosis-karyorrhexis index (MKI) according to Shimada classification [32], presence of necrosis, calcifications, and follow-up data were collected from clinical charts and inserted in a dedicated database.
RNA extraction from formalin-fixed paraffin-embedded tissues and gene expression analyses
Ten µm thick sections were cut were cut from formalin fixed paraffin embedded blocks of the tumor in RNase-free conditions, following microdissection using a scalpel at a magnification of × 100 from hematoxylin–eosin (H&E) stained slides. The suitability of the material was evaluated by hematoxylin and eosin staining, and care was taken to select tumor areas. Total RNA isolation was performed by commercially available RNA extraction kits designed for paraffin material according to the manufacturer’s instructions (miRNeasy FFPE kit; Qiagen, Hilden, Germany).
RT reactions were performed using 10 ng total RNA in a volume of 15 μl with the following conditions: 16 °C for 30 min, 42 °C for 30 min, 85 °C for 5 min, and 4 °C for 5 min. Expression levels of all genes studied and internal reference were examined using a fluorescence-based real-time detection method (ABI PRISM 7900 Sequence Detection System—Taqman; Applied Biosystems, Foster City, CA,). The following TaqMan gene expression assays (Applied Biosystems) were used according to the manufacturer’s instructions: ASCL1 (HS00269932_m1), DLL3 (HS01085096_m1), INSM1 (Hs00357871_s1), MYCL1 (Hs00420495_m1), NEUROD1 (HS01922995_s1), NOTCH1 (Hs01062014_m1), POU2F3 (Hs00205009_m1), YAP1 (Hs00902712_g1), ACTB (Hs01060665_g1) assay served as housekeeping reference gene for the analyses.
Each measurement was performed in duplicate. The ΔΔCt values were calculated subtracting ΔCt values of sample and ΔCt value of Stratagene (a pool of RNA derived from normal different tissues; Stratagene, CA), and converted to ratio by the following formula: 2 − ΔΔCt.
Statistical analysis
Statistical analyses were carried out using the Stata 15.0 software (StataCorp, College Station, TX, U.S.A.). The differences in the distribution of the variables evaluated based on clinical-pathological parameters were analyzed using parametric and non-parametric tests (Student’s t test, Pearson’s chi-square test and Bonferroni’s correction, Wilcoxon’s rank test).
Time to relapse (disease-free interval, DFI) was assessed from the date of diagnosis to the date of relapse or the date of the last checkup. All dead patients were considered as events. Survival analysis was determined by the Kaplan–Meier curves and Mantel log-rank test was used to compare statistical differences. Cox regression analyses were carried out on DFI to calculate HRs and 95% CIs for the different study groups. All statistical tests were two sided. p-values < 0.05 were considered significant.
Results
Clinico-pathological characteristics of the study group
The clinical and pathological features of 46 neuroblastoma cases are summarized in Supplementary Table 1. In brief, the case series was composed of 20 females (43.5%) and 26 male patients (56.5%), 27 of them aged < 18 months (58.7%) and 19 patients ≥ 18 months (41.3%). Stages I–II, III, IV, and IVS were diagnosed in 15 (32.6%), 5 (10.9%), 18 (39.1%), and 8 (17.4%) cases, respectively. The poorly differentiated subtype (according to INPC) was present in the majority of cases (26/46, 56.5%), while undifferentiated and differentiated forms were seen in 17/46 (37%) and 3/46 (6.5%) cases, respectively. Low, intermediate, and high MKI were present in 21 (45.7%), 20 (43.5%), and 5 (10.8%) cases, respectively. Both calcifications and necrosis were noted in 28/46 cases (60.9%). Relapse occurred in 19/46 (41.3%) patients and 8/46 died of disease (17.4%).
Gene expression profiles and correlations with clinical-pathological features
A strong reciprocal positive correlation was observed across the series between NOTCH1 and ASCL1 (R = 0.4561; p = 0.0014), NEUROD1 (R = 0.4982; p = 0.0004), INSM1 (R = 0.4763; p = 0.0008), and YAP1 (R = 0.8475; p < 0.0001). Moreover, ASCL1 was found significantly correlated with DLL3 (R = 0.3685; p = 0.0117), NEUROD1 (R = 0.3976; p = 0.0062) and INSM1 (R = 0.5996; p < 0.0001). Furthermore, NEUROD1 was significantly correlated with MYCL1 (R = − 0.2968; p = 0.0452) and YAP1 (R = 0.5290; p = 0.0002), while INSM1 resulted significantly correlated with YAP1 (p = 0.0257).
As to concern clinical and pathological correlations (Supplementary Table 2), NOTCH1, NEUROD1, MYCL1, and YAP1 were found significantly correlated with tumor stage. In detail, NOTCH1 (p = 0.005), NEUROD1 (p = 0.0059), and YAP1 (p = 0.0008) were more expressed in stage IV tumors, while highest levels of MYCL1 and ASCL1 were seen in stages IVS and III, respectively (p = 0.0182 and p = 0.0134). Moreover, a higher level of both NOTCH1 (p = 0.0079) and YAP1 (p = 0.0026) was found in cases with differentiating morphology according to INPC classification. Finally, cases with high MKI according to Shimada demonstrated significantly lower levels of POU2F3 (p = 0.0277).
Gene expression levels and survival
In the overall series, cases with a high gene expression level (as determined using the median value) of NOTCH1, INSM1, NEUROD1, and YAP1 demonstrated more frequently recurrence of the disease than those with low expression (Fig. 1). Considering the 30 treatment-naïve specimens only, we observed significantly higher gene expression levels of ASCL1 (p = 0.0007), INSM1 (p = 0.0016), and DLL3 (p = 0.0064) in cases that relapsed during the follow-up (Table 2).
As to concern univariate survival analyses, higher levels of INSM1 were significantly correlated with shorter disease-free interval (DFI) in the whole group (Fig. 2A, p = 0.0012), as well as when separately analyzed in the treatment-naïve (Fig. 2B, p = 0.0147) and the post-chemotherapy groups of patients (Fig. 2C, p = 0.0365). Moreover, the patients with higher levels of NOTCH1 had a shorter DFI, both in the whole cohort (Fig. 2D, p = 0.0001) and in the 30 treatment naïve patients (Fig. 2E, p = 0.0085), (but not in the 16 post-chemotherapy patients, Fig. 2F, p = 0.158). In addition, considering the whole case series, patients with high expression of YAP1 and NEUROD1 had shorter DFI, compared to those with low expression (Supplementary Fig. 1: A, p = 0.0007 and B, p = 0.0128). However, no significance was observed when patients were stratified according to the chemotherapy status (data not shown).
Kaplan–Meier estimates of DFI according to the INSM1 gene expression level in the whole series (A p = 0.0012) and in treatment naïve (B p = 0.0147) and post-chemotherapy cases (C p = 0.0365). Kaplan–Meier estimates of DFI according to the NOTCH1 gene expression level in the whole series (D p = 0.0001) and in treatment naïve (E p = 0.0085) and post-chemotherapy cases (F p = 0.1580)
Furthermore, Cox regression analyses to estimate HRs and 95% CIs for DFI in the different study groups shown in Table 3 demonstrated in the whole series that stage IV (HR 8.57; CI 1.1–68.6 p = 0.043) and high expression of NOTCH1 (HR 15.6; CI 2.05–118.7, p = 0.008), NEUROD1 (HR 3.17;1.14–8.78, p = 0.026), INSM1 (HR 5.24, CI 1.49–18.5, p = 0.010), and YAP1 (HR 8.57, CI 1.93–37.9, p = 0.005) were associated with adverse prognosis. Moreover, poorly differentiated forms (HR 0.24; 0.06–0.98, p = 0.048) of disease were significantly associated to better DFI survival compared to undifferentiating lesions.
Gene combination analyses
Considering the different combinations of INSM1, NOTCH1, NEUROD1, and YAP1 gene expression levels according to the recurrent disease status (Supplementary Table 3), patients with low expression of all four genes did not experience recurrent disease while patients with high levels of at least one of the aforementioned genes demonstrated significantly more frequent recurrences (p = 0.003).
Moreover, we observed a statistically significant increase in the odds of the patients with the high levels of ≥ 3 above indicated genes when considering the whole case series (OR = 8.75, 95% CI = 2.19–34.81, p = 0.002). However, when stratified according to the treatment status, no significance was observed the treatment naïve group (OR = 4.75, 95% CI = 0.63–35.48, p = 0.129) and post-chemotherapy group of patients (OR = 2.75, 95% CI = 0.16–46.79, p = 0.484) analyzed separately (Supplementary Table 4).
Discussion
We demonstrated in a series of 46 pediatric neuroblastomas that neuroendocrine-lineage transcriptional genes are expressed in neuroblastoma and that their profiles of expression may identify subgroups of patients with increased risk of recurrence and/or shorter survival. The study design focused on gene expression analysis and not immunohistochemical determination of the corresponding protein expression due to the following considerations. First, reliable and robust antibodies are not available for all molecules. Second, tissue material available was limited in some cases preventing the possibility to perform both RNA extraction and subsequent sectioning for a multi-target immunohistochemical procedure. Moreover, even in the more extensively investigated small cell lung cancer, model gene expression data have been used for molecular classification purposes [11, 12], whereas no study clearly demonstrated a linear correlation between protein and mRNA expression for these targets. The strength of the transcriptional data observed in this study is proven by the very high positive reciprocal correlation among most of the markers investigated, being NOTCH1 the one showing the correlation with the highest number of genes. Interestingly, at variance with other models in which these transcription factors are active, such as small cell lung carcinoma, the expression of the genes here investigated did not segregate neuroblastoma cases into different families characterized by alterative transcriptional profiles, since no inverse correlation among any of the genes tested was identified.
Our data on the association of gene expression profiles with aggressive clinical outcome and survival represent the first global evidence that the expression of neuroendocrine differentiation transcriptional drivers may be used to further characterize neuroblastoma patients, and several preclinical evidence exist supporting their role as potentially relevant clinical prognostic biomarkers.
Among all genes, four of them—NOTCH1, INSM1, YAP1, and NEUROD1—emerge as the most significant, being associated with both shorter disease-free interval and high tumor stage and/or risk of recurrence.
The association of NOTCH1 overexpression with adverse clinical outcome in our series is expanding previous data obtained by means of the immunohistochemical analysis of Notch1 protein in a large series of neuroblastomas [33]. At variance with such previous series, we did not compare the expression of NOTCH1 (and of all other genes) with MYCN amplification status, and this limitation should be considered in future studies. Interestingly, in a translational view, the demonstration of NOTCH1 expression in neuroblastoma and its positive correlation with aggressiveness is paralleled by in vitro data on the efficacy of Notch1 inhibition in neuroblastoma cells. In a study on a variety of neuroblastoma cell lines, gamma-secretase inhibition (in particular GSI-I) was shown to impair cell proliferation and to promote apoptosis in vitro and in vivo through targeting Notch signaling [34]. Moreover, the Notch1 inhibitor NSI-1 has recently shown to suppress the viability of SH-SY5Y neuroblastoma cells characterized by a constitutive Notch1 activation [35]. The impact of NOTCH1 overexpression in the adverse clinical behavior of neuroblastoma patients in our series is to be further validated in a biological perspective. However, among the possible mechanisms, Notch1 has been shown to actively maintain a stem cell phenotype in neuroblastoma cells that confer highly tumorigenic properties [36].
INSM1 transcription factor has emerged in vitro as a neuroblastoma biomarker that plays critical role in facilitating tumor cell growth and transformation [30]. Its protein nuclear expression has been documented in 84% of neuroblastomas, with a suggested association with clinical outcome, being the three INSM1-negative neuroblastoma patients in the published study all alive with a median survival of 15 years as opposed to a median of 5 years in 9 out of 13 INSM1-positive neuroblastoma patients [28]. No definitive data are present in the literature on its possible role as prognostic biomarker in neuroblastoma. However, in other tumor models—such as pulmonary high-grade neuroendocrine carcinoma-positive INSM1 protein expression has been associated with a dismal prognosis [37], in line with the data on INSM1 gene expression levels from the present study.
Among other markers associated with neuroblastoma aggressiveness in our series, NEUROD1 was already shown to promote cell growth in vitro [24] and tumor formation in vivo [38] in neuroblastoma cells. YAP1 expression has been reported to significantly increase cell proliferation and growth through inhibition of 27Kip1 activity in neuroblastoma cell lines SH-SY5Y and SK-N-SH [39]. Moreover, YAP1 overexpression has been associated to the increased resistance to platinum-based [40] and MEK-inhibiting [41] therapeutic strategies.
In terms of correlation with differentiation, NOTCH1 and YAP1 expression was significantly higher in differentiating compared to poorly differentiated tumors according to the INPC classification. This observation cannot find a clear explanation in the current literature. A previous in vitro study is partly in contrast with our findings and shows that Notch1 inhibition prevents neurite formation in neuroblastoma cells [42]. Moreover, these data are apparently in contrast with the adverse impact on clinical outcome for both markers in our same series and might more probably represent a bias related to the limited number of cases in the differentiation group.
The potential clinical impact of our findings is twofold. First, the integration of gene expression profiles of the biomarkers investigated in the present study might assist to better predict the clinical behavior of neuroblastoma patients, and to improve a personalized approach in tailoring adjuvant chemotherapeutic regimens and/or predict response to treatment. Second, the pathways regulated by the genes herein investigated are potential targets for selective therapies. In particular, DLL3 has been tested as a target for rovalpituzumab tesirine therapy in neuroblastoma models with promising results [43].
In conclusion, we identify a strong prognostic impact of neuroendocrine-lineage transcriptional profiles in neuroblastoma, and suggest that the evaluation of NOTCH1, INSM1, YAP1, and NEUROD1 might help to further characterize the risk of relapse in neuroblastoma patients.
References
Jung M, Russell AJ, Liu B, George J, Liu PY, Liu T, DeFazio A, Bowtell DD, Oberthuer A, London WB, Fletcher JI, Haber M, Norris MD, Henderson MJ (2017) A Myc activity signature predicts poor clinical outcomes in Myc-associated cancers. Cancer Res 77:971–981. https://doi.org/10.1158/0008-5472.CAN-15-2906
Matthay KK, Maris JM, Schleiermacher G, Nakagawara A, Mackall CL, Diller L, Weiss WA (2016) Neuroblastoma. Nat Rev Dis Primers 2:16078. https://doi.org/10.1038/nrdp.2016.78
Zafar A, Wang W, Liu G, Wang X, Xian W, McKeon F, Foster J, Zhou J, Zhang R (2021) Molecular targeting therapies for neuroblastoma: progress and challenges. Med Res Rev 41:961–1021. https://doi.org/10.1002/med.21750
Liang WH, Federico SM, London WB, Naranjo A, Irwin MS, Volchenboum SL, Cohn SL (2020) Tailoring therapy for children with neuroblastoma on the basis of risk group classification: past, present, and future. JCO Clin Cancer Inform 4:895–905. https://doi.org/10.1200/CCI.20.00074
Edsjö A, Nilsson H, Vandesompele J, Karlsson J, Pattyn F, Culp LA, Speleman F, Påhlman S (2004) Neuroblastoma cells with overexpressed MYCN retain their capacity to undergo neuronal differentiation. Lab Invest 84:406–417. https://doi.org/10.1038/labinvest.3700061
Borromeo MD, Savage TK, Kollipara RK, He M, Augustyn A, Osborne JK, Girard L, Minna JD, Gazdar AF, Cobb MH, Johnson JE (2016) ASCL1 and NEUROD1 reveal heterogeneity in pulmonary neuroendocrine tumors and regulate distinct genetic programs. Cell Rep 16:1259–1272. https://doi.org/10.1016/j.celrep.2016.06.081
Crabtree JS, Singleton CS, Miele L (2016) Notch signaling in neuroendocrine tumors. Front Oncol 6:94. https://doi.org/10.3389/fonc.2016.00094
Kudoh S, Tenjin Y, Kameyama H, Ichimura T, Yamada T, Matsuo A, Kudo N, Sato Y, Ito T (2020) Significance of achaete-scute complex homologue 1 (ASCL1) in pulmonary neuroendocrine carcinomas; RNA sequence analyses using small cell lung cancer cells and Ascl1-induced pulmonary neuroendocrine carcinoma cells. Histochem Cell Biol 153:443–456. https://doi.org/10.1007/s00418-020-01863-z
Augustyn A, Borromeo M, Wang T, Fujimoto J, Shao C, Dospoy PD, Lee V, Tan C, Sullivan JP, Larsen JE, Girard L, Behrens C, Wistuba II, Xie Y, Cobb MH, Gazdar AF, Johnson JE, Minna JD (2014) ASCL1 is a lineage oncogene providing therapeutic targets for high-grade neuroendocrine lung cancers. Proc Natl Acad Sci USA 111:14788–14793. https://doi.org/10.1073/pnas.1410419111
Baine MK, Hsieh MS, Lai WV, Egger JV, Jungbluth AA, Daneshbod Y, Beras A, Spencer R, Lopardo J, Bodd F, Montecalvo J, Sauter JL, Chang JC, Buonocore DJ, Travis WD, Sen T, Poirier JT, Rudin CM, Rekhtman N (2020) SCLC subtypes defined by ASCL1, NEUROD1, POU2F3, and YAP1: a comprehensive immunohistochemical and histopathologic characterization. J Thorac Oncol 15:1823–1835. https://doi.org/10.1016/j.jtho.2020.09.009
Rudin CM, Poirier JT, Byers LA, Dive C, Dowlati A, George J, Heymach JV, Johnson JE, Lehman JM, MacPherson D, Massion PP, Minna JD, Oliver TG, Quaranta V, Sage J, Thomas RK, Vakoc CR, Gazdar AF (2019) Molecular subtypes of small cell lung cancer: a synthesis of human and mouse model data. Nat Rev Cancer 19:289–297. https://doi.org/10.1038/s41568-019-0133-9
Gay CM, Stewart CA, Park EM, Diao L, Groves SM, Heeke S, Nabet BY, Fujimoto J, Solis LM, Lu W, Xi Y, Cardnell RJ, Wang Q, Fabbri G, Cargill KR, Vokes NI, Ramkumar K, Zhang B, Della Corte CM, Robson P, Swisher SG, Roth JA, Glisson BS, Shames DS, Wistuba II, Wang J, Quaranta V, Minna J, Heymach JV, Byers LA (2021) Patterns of transcription factor programs and immune pathway activation define four major subtypes of SCLC with distinct therapeutic vulnerabilities. Cancer Cell 39:346-360.e7. https://doi.org/10.1016/j.ccell.2020.12.014
George J, Walter V, Peifer M, Alexandrov LB, Seidel D, Leenders F, Maas L, Müller C, Dahmen I, Delhomme TM, Ardin M, Leblay N, Byrnes G, Sun R, De Reynies A, McLeer-Florin A, Bosco G, Malchers F, Menon R, Altmüller J, Becker C, Nürnberg P, Achter V, Lang U, Schneider PM, Bogus M, Soloway MG, Wilkerson MD, Cun Y, McKay JD, Moro-Sibilot D, Brambilla CG, Lantuejoul S, Lemaitre N, Soltermann A, Weder W, Tischler V, Brustugun OT, Lund-Iversen M, Helland Å, Solberg S, Ansén S, Wright G, Solomon B, Roz L, Pastorino U, Petersen I, Clement JH, Sänger J, Wolf J, Vingron M, Zander T, Perner S, Travis WD, Haas SA, Olivier M, Foll M, Büttner R, Hayes DN, Brambilla E, Fernandez-Cuesta L, Thomas RK (2018) Integrative genomic profiling of large-cell neuroendocrine carcinomas reveals distinct subtypes of high-grade neuroendocrine lung tumors. Nat Commun 9:1048. https://doi.org/10.1038/s41467-018-03099-x
Rekhtman N, Pietanza MC, Hellmann MD, Naidoo J, Arora A, Won H, Halpenny DF, Wang H, Tian SK, Litvak AM, Paik PK, Drilon AE, Socci N, Poirier JT, Shen R, Berger MF, Moreira AL, Travis WD, Rudin CM, Ladanyi M (2016) Next-generation sequencing of pulmonary large cell neuroendocrine carcinoma reveals small cell carcinoma-like and non-small cell carcinoma-like subsets. Clin Cancer Res 22:3618–3629. https://doi.org/10.1158/1078-0432.CCR-15-2946
Alcala N, Leblay N, Gabriel AAG, Mangiante L, Hervas D, Giffon T, Sertier AS, Ferrari A, Derks J, Ghantous A, Delhomme TM, Chabrier A, Cuenin C, Abedi-Ardekani B, Boland A, Olaso R, Meyer V, Altmuller J, Le Calvez-Kelm F, Durand G, Voegele C, Boyault S, Moonen L, Lemaitre N, Lorimier P, Toffart AC, Soltermann A, Clement JH, Saenger J, Field JK, Brevet M, Blanc-Fournier C, Galateau-Salle F, Le Stang N, Russell PA, Wright G, Sozzi G, Pastorino U, Lacomme S, Vignaud JM, Hofman V, Hofman P, Brustugun OT, Lund-Iversen M, Thomas de Montpreville V, Muscarella LA, Graziano P, Popper H, Stojsic J, Deleuze JF, Herceg Z, Viari A, Nuernberg P, Pelosi G, Dingemans AMC, Milione M, Roz L, Brcic L, Volante M, Papotti MG, Caux C, Sandoval J, Hernandez-Vargas H, Brambilla E, Speel EJM, Girard N, Lantuejoul S, McKay JD, Foll M, Fernandez-Cuesta L (2019) Integrative and comparative genomic analyses identify clinically relevant pulmonary carcinoid groups and unveil the supra-carcinoids. Nat Commun 10:3407. https://doi.org/10.1038/s41467-019-11276-9
Simbolo M, Barbi S, Fassan M, Mafficini A, Ali G, Vicentini C, Sperandio N, Corbo V, Rusev B, Mastracci L, Grillo F, Pilotto S, Pelosi G, Pelliccioni S, Lawlor RT, Tortora G, Fontanini G, Volante M, Scarpa A, Bria E (2019) Gene expression profiling of lung atypical carcinoids and large cell neuroendocrine carcinomas identifies three transcriptomic subtypes with specific genomic alterations. J Thorac Oncol 14:1651–1661. https://doi.org/10.1016/j.jtho.2019.05.003
Derks JL, Dingemans AC, van Suylen RJ, den Bakker MA, Damhuis RAM, van den Broek EC, Speel EJ, Thunnissen E (2019) Is the sum of positive neuroendocrine immunohistochemical stains useful for diagnosis of large cell neuroendocrine carcinoma (LCNEC) on biopsy specimens? Histopathology 74:555–566. https://doi.org/10.1111/his.13800
La Rosa S, Marando A, Gatti G, Rapa I, Volante M, Papotti M, Sessa F, Capella C (2013) Achaete-scute homolog 1 as a marker of poorly differentiated neuroendocrine carcinomas of different sites: a validation study using immunohistochemistry and quantitative real-time polymerase chain reaction on 335 cases. Hum Pathol 44:1391–1399. https://doi.org/10.1016/j.humpath.2012.11.013
Shida T, Furuya M, Kishimoto T, Nikaido T, Tanizawa T, Koda K, Oda K, Takano S, Kimura F, Shimizu H, Yoshidome H, Ohtsuka M, Nakatani Y, Miyazaki M (2008) The expression of NeuroD and mASH1 in the gastroenteropancreatic neuroendocrine tumors. Mod Pathol 21:1363–1370. https://doi.org/10.1038/modpathol.2008.121
Juhlin CC (2021) Second-generation neuroendocrine immunohistochemical markers: reflections from clinical implementation. Biology (Basel) 10:874. https://doi.org/10.3390/biology10090874
Rostomily RC, Bermingham-McDonogh O, Berger MS, Tapscott SJ, Reh TA, Olson JM (1997) Expression of neurogenic basic helix-loop-helix genes in primitive neuroectodermal tumors. Cancer Res 57:3526–3531
Kitazono I, Hamada T, Yoshimura T, Kirishima M, Yokoyama S, Akahane T, Tanimoto A (2020) PCP4/PEP19 downregulates neurite outgrowth via transcriptional regulation of Ascl1 and NeuroD1 expression in human neuroblastoma M17 cells. Lab Invest 100:1551–1563. https://doi.org/10.1038/s41374-020-0462-z
Lee TY, Cho IS, Bashyal N, Naya FJ, Tsai MJ, Yoon JS, Choi JM, Park CH, Kim SS, Suh-Kim H (2020) ERK regulates NeuroD1-mediated neurite outgrowth via proteasomal degradation. Exp Neurobiol 29:189–206. https://doi.org/10.5607/en20021
Lu F, Kishida S, Mu P, Huang P, Cao D, Tsubota S, Kadomatsu K (2015) NeuroD1 promotes neuroblastoma cell growth by inducing the expression of ALK. Cancer Sci 106:390–396. https://doi.org/10.1111/cas.12628
Li J, Neumann I, Volkmer I, Staege MS (2011) Down-regulation of achaete-scute complex homolog 1 (ASCL1) in neuroblastoma cells induces up-regulation of insulin-like growth factor 2 (IGF2). Mol Biol Rep 38:1515–1521. https://doi.org/10.1007/s11033-010-0259-z
Kasim M, Heß V, Scholz H, Persson PB, Fähling M (2016) Achaete-Scute homolog 1 expression controls cellular differentiation of neuroblastoma. Front Mol Neurosci 9:156. https://doi.org/10.3389/fnmol.2016.00156
Chen C, Breslin MB, Guidry JJ, Lan MS (2019) 5’-Iodotubercidin represses insulinoma-associated-1 expression, decreases cAMP levels, and suppresses human neuroblastoma cell growth. J Biol Chem 294:5456–5465. https://doi.org/10.1074/jbc.RA118.006761
Wang H, Krishnan C, Charville GW (2019) INSM1 expression in peripheral neuroblastic tumors and other embryonal neoplasms. Pediatr Dev Pathol 22:440–448. https://doi.org/10.1177/1093526619843725
Chen C, Lan MS (2020) A promoter-driven assay for INSM1-associated signaling pathway in neuroblastoma. Cell Signal 76:109785. https://doi.org/10.1016/j.cellsig.2020.109785
Chen C, Breslin MB, Lan MS (2015) INSM1 increases N-myc stability and oncogenesis via a positive-feedback loop in neuroblastoma. Oncotarget 6:36700–12. https://doi.org/10.18632/oncotarget.5485
Shimada H, Delellis RA, Tissier F (2017) Neuroblastic tumors of the adrenal gland. In: Lloyd RV, Osamura RY, Klöppel G, Rosai J (eds) WHO Classification of Tumours of Endocrine Organs, 4th edn. IARC, Lyon, France, pp 196–204
Shimada H, Chatten J, Newton WA Jr, Sachs N, Hamoudi AB, Chiba T, Marsden HB, Misugi K (1984) Histopathologic prognostic factors in neuroblastic tumors: definition of subtypes of ganglioneuroblastoma and an age-linked classification of neuroblastomas. J Natl Cancer Inst 73:405–416. https://doi.org/10.1093/jnci/73.2.405
Chang HH, Lee H, Hu MK, Tsao PN, Juan HF, Huang MC, Shih YY, Wang BJ, Jeng YM, Chang CL, Huang SF, Tsay YG, Hsieh FJ, Lin KH, Hsu WM, Liao YF (2010) Notch1 expression predicts an unfavorable prognosis and serves as a therapeutic target of patients with neuroblastoma. Clin Cancer Res 16:4411–4420. https://doi.org/10.1158/1078-0432.CCR-09-3360
Dorneburg C, Goß AV, Fischer M, Roels F, Barth TF, Berthold F, Kappler R, Oswald F, Siveke JT, Molenaar JJ, Debatin KM, Beltinger C (2016) γ-Secretase inhibitor I inhibits neuroblastoma cells, with NOTCH and the proteasome among its targets. Oncotarget 7:62799–62813. https://doi.org/10.18632/oncotarget.11715
Shiraishi T, Sakaitani M, Otsuguro S, Maenaka K, Suzuki T, Nakaya T (2019) Novel Notch signaling inhibitor NSI-1 suppresses nuclear translocation of the Notch intracellular domain. Int J Mol Med 44:1574–1584. https://doi.org/10.3892/ijmm.2019.4280
Ross RA, Walton JD, Han D, Guo HF, Cheung NK (2015) A distinct gene expression signature characterizes human neuroblastoma cancer stem cells. Stem Cell Res 15:419–426. https://doi.org/10.1016/j.scr.2015.08.008
Minami K, Jimbo N, Tanaka Y, Ogawa H, Hokka D, Nishio W, Yoshimura M, Itoh T, Maniwa Y (2020) Insulinoma-associated protein 1 is a prognostic biomarker in pulmonary high-grade neuroendocrine carcinoma. J Surg Oncol 122:243–253. https://doi.org/10.1002/jso.25960
Huang P, Kishida S, Cao D, Murakami-Tonami Y, Mu P, Nakaguro M, Koide N, Takeuchi I, Onishi A, Kadomatsu K (2011) The neuronal differentiation factor NeuroD1 downregulates the neuronal repellent factor Slit2 expression and promotes cell motility and tumor formation of neuroblastoma. Cancer Res 71:2938–2948. https://doi.org/10.1158/0008-5472.CAN-10-3524
Shen X, Xu X, Xie C, Liu H, Yang D, Zhang J, Wu Q, Feng W, Wang L, Du L, Xuan L, Meng C, Zhang H, Wang W, Wang Y, Xie T, Huang Z (2020) YAP promotes the proliferation of neuroblastoma cells through decreasing the nuclear location of p27Kip1 mediated by Akt. Cell Prolif 53:e12734. https://doi.org/10.1111/cpr.12734
Yang C, Tan J, Zhu J, Wang S, Wei G (2017) YAP promotes tumorigenesis and cisplatin resistance in neuroblastoma. Oncotarget 8:37154–37163. https://doi.org/10.18632/oncotarget.16209
Coggins GE, Farrel A, Rathi KS, Hayes CM, Scolaro L, Rokita JL, Maris JM (2019) YAP1 mediates resistance to MEK1/2 inhibition in neuroblastomas with hyperactivated RAS signaling. Cancer Res 79:6204–6214. https://doi.org/10.1158/0008-5472.CAN-19-1415
Liu M, Inoue K, Leng T, Zhou A, Guo S, Xiong ZG (2017) ASIC1 promotes differentiation of neuroblastoma by negatively regulating Notch signaling pathway. Oncotarget 8:8283–8293. https://doi.org/10.18632/oncotarget.14164
Sano R, Krytska K, Tsang M, Erickson SW, Teicher BA, Saunders L, Jones RT, Smith MA, Maris JM, Mosse YP (2018) Pediatric Preclinical Testing Consortium evaluation of a DLL3-targeted antibody drug conjugate rovalpituzumab tesirine, in neuroblastoma. Cancer Res 78 (13_Supplement): LB-136. https://doi.org/10.1158/1538-7445.AM2018-LB-136
Acknowledgements
The authors would like to thank Dr Isabella Morra (Pathology Unit at “Città della Salute e della Scienza” Hospital, Turin) for histological revision of neuroblastoma samples.
Funding
Open access funding provided by Università degli Studi di Torino within the CRUI-CARE Agreement. This work was partially supported by grants from Fondazione Umberto Veronesi (to FF).
Author information
Authors and Affiliations
Contributions
Study design: MP, MV; RNA extraction and RT-PCR analyses: FN, JM; data analyses: MV, MP, JM, FN, LB; database management: JM, FN, CT, GO. Statistical analyses: SOA; clinical and follow up data: FF, MB, DC; writing—original draft preparation: MP, MV, JM, FN; writing—review and editing: all authors; funding acquisition: FF; all authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Ethical approval
Before the study started, all cases were de-identified and coded by a pathology staff member not involved in the study, and all data were accessed anonymously. The study was approved by the Institutional Board review of Citta della Salute e della Scienza, Turin, Italy (n° 966/2011). The study was conducted in accordance with The Code of Ethics of the World Medical Association (Declaration of Helsinki).
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Supplementary file 1
(DOCX 26 kb)

Supplementary file 2
(PNG 469 kb)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Metovic, J., Napoli, F., Osella-Abate, S. et al. Overexpression of INSM1, NOTCH1, NEUROD1, and YAP1 genes is associated with adverse clinical outcome in pediatric neuroblastoma. Virchows Arch 481, 925–933 (2022). https://doi.org/10.1007/s00428-022-03406-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00428-022-03406-4