Abstract
Rearrangements of the ROS1 gene occur in 1–2 % of non-small cell lung cancers (NSCLCs). Crizotinib, a highly effective inhibitor of ROS1 kinase activity, is now FDA-approved for the treatment of patients with advanced ROS1-positive NSCLC. Consequently, focus on ROS1 testing is growing. Most laboratories currently rely on fluorescence in situ hybridisation (FISH) assays using a dual-colour break-apart probe to detect ROS1 rearrangements. Given the rarity of these rearrangements in NSCLC, detection of elevated ROS1 protein levels by immunohistochemistry may provide cost-effective screening prior to confirmatory FISH testing. Non-in situ testing approaches also hold potential as stand-alone methods or complementary tests, including multiplex real-time PCR assays and next-generation sequencing (NGS) platforms which include commercial test kits covering a range of fusion genes. In order to ensure high-quality biomarker testing, appropriate tissue handling, adequate control materials and participation in external quality assessment programmes are essential, irrespective of the testing technique employed. ROS1 testing is often only considered after negative tests for EGFR mutation and ALK gene rearrangement, based on the assumption that these oncogenic driver events tend to be exclusive. However, as the use of ROS1 inhibitors becomes routine, accurate and timely detection of ROS1 gene rearrangements will be critical for the optimal treatment of patients with NSCLC. As NGS techniques are introduced into routine diagnostic practice, ROS1 fusion gene testing will be provided as part of the initial testing package.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Lung cancer is the most frequent cause of cancer-related death worldwide and is usually diagnosed in advanced stages [1]. The most common histological lung cancer subgroup is non-small cell lung cancer (NSCLC), which accounts for 80 % of lung cancers [1]. Currently, there are two identified molecular subtypes of NSCLC that have targeted therapies approved for their treatment: mutations in the epidermal growth factor receptor (EGFR) gene and rearrangements in the anaplastic lymphoma kinase (ALK) gene; tumours harbouring these genetic alterations respond well to specific tyrosine kinase inhibitors [2, 3]. In addition to EGFR and ALK, other known oncogenic drivers of NSCLC include hepatocyte growth factor receptor (MET), the GTPase KRAS, human epidermal growth factor receptor 2 (HER2), RET and ROS1 [4, 5].
ROS1 is now recognised as a distinct molecular target in NSCLC [6, 7]. Pre-clinical and clinical studies demonstrate that ROS1 can be efficiently inhibited by the tyrosine kinase inhibitor crizotinib [8, 9], which is approved by the FDA and EMA as a treatment for patients with advanced ALK-positive NSCLC [10, 11]. Crizotinib was recently approved by the FDA for patients with advanced ROS1-positive NSCLC [12]; therefore, detection of ROS1 gene rearrangements is critical for the optimal treatment of ROS1-positive NSCLC patients. In this article, we review the current state of molecular diagnostics for ROS1-positive NSCLC, discuss our experience with the relevant technologies and provide guidance on the detection of ROS1-positive tumours.
Rationale for targeting ROS1 fusions in NSCLC
Although v-ROS1 had already been identified as a unique oncogenic sequence in the avian sarcoma virus (VR2) [13], a chicken retrovirus, it was only in 2003 that the genomic structure of ROS1 was fully characterised [14]. ROS1 belongs to the human receptor tyrosine kinase (RTK) family and is evolutionarily close to the ALK family, forming part of the scientific basis for using inhibitors of ALK as inhibitors of ROS1. The ROS1 gene is located on chromosome 6 (6q22) and encodes a transmembrane receptor protein with unique features. The extracellular N-terminal domain spans more than 1800 amino acids, which makes it one of the largest extracellular domains amongst all human RTKs. Despite this, no human ROS1 ligand has been found to date and the physiological function of this orphan receptor is still unclear. The C-terminal portion of ROS1 contains a kinase domain and a single transmembrane domain [9, 15–17].
Genomic rearrangements involving ROS1 occur in 1–2 % of NSCLCs [9, 18–23]. ROS1 gene rearrangement was initially discovered in the glioblastoma cell line V118MG [24]. In this cell line, an intrachromosomal deletion on chromosome 6 fused the 5′ region of a gene named FIG to the 3′ region of ROS1. Since then, many more novel ROS1 fusion partners have been found. Importantly, the ROS1 kinase domain is retained in all of these fusion events and the expressed fusion genes have been reported to be oncogenic. Known ROS1 fusion partners in lung cancer include FIG, CD74, SLC34A2 and SDC4, and the list is growing. CD74-ROS1 is the most frequently detected ROS1 fusion in this group of patients. With all of the known fusion genes, the ROS1 kinase domain is fully retained and the ROS1 junction point at the messenger RNA (mRNA) level invariably occurs at the 5′ end of exons 32, 34, 35 or 36 (Fig. 1 and Table 1).
a Schematic diagram of ROS1 fusions in NSCLC showing ROS1 tyrosine kinase domain (TKI, dark grey), ROS1 transmembrane domain (TM, mid-grey) and coiled-coil domains (CC, light grey) in ROS1 fusion proteins (KDELR2-ROS1 is not shown). Reproduced from Gainor and Shaw [35]. b Frequencies of different ROS1 fusion partners. Adapted from Gainor and Shaw [35], with additional data from more recent studies as reported in Table 1
Unlike in ALK, where the fusion partner provides a dimerisation domain that induces constitutive oligomerisation and thus activation of the kinase, the mechanism by which ROS1 fusion proteins become constitutively active is not exactly known. In fact, many of the known ROS1 fusion partners do not contain dimerisation domains [22]. What is known is that several signalling pathways are activated by ROS1 fusion proteins. Expression of FIG-ROS1, CD74-ROS1 or SCD4-ROS1 in fibroblasts or Ba/F3 cells has been shown to result in auto-phosphorylation of ROS1 and phosphorylation of SHP-2, MAP-ERK kinase, ERK, STAT3 and AKT, and these effects have been blocked by pharmacological inhibition of ROS1. Subcellular localisation and downstream signalling may differ depending on the fusion partner of ROS1 [19, 20, 25, 26], but in general, the activated pathways seem to involve common growth and survival pathways that are also activated by other RTKs.
Efficacy and safety of ROS1 inhibitor therapy
ROS1 inhibition by crizotinib has been studied in a number of early-phase clinical trials in patients with advanced ROS1-positive NSCLC (Table 2). In the ROS1 expansion cohort of a phase 1 trial of crizotinib, the objective response rate (ORR) was 72 %. Median duration of response was 17.6 months and median progression-free survival (PFS) was 19.2 months. No relationship was observed between ROS1 fusion partner and duration of crizotinib treatment [8]. Furthermore, ORR with crizotinib was 80 % and median PFS was 9.1 months in heavily pre-treated patients in a retrospective study [27]. Consistent with this, in patients with advanced ROS1-positive NSCLC receiving crizotinib in a French phase 2 trial, ORR was 69 % and median PFS was 9.1 months [28]. Finally, ORR was 69 % and median PFS was 12.9 months with crizotinib in a phase 2 trial in East Asian patients with advanced ROS1-positive NSCLC [29]. Across the clinical studies in ROS1-positive NSCLC, crizotinib treatment was well tolerated, with an adverse event profile similar to that seen in ALK-positive NSCLC [3, 30]. Phase 2 trials in ROS1-positive NSCLC are currently ongoing.
Detection of ROS1 gene rearrangements
As mentioned above, ROS1 gene rearrangement is one of several addictive oncogenic events which may drive a proportion of pulmonary adenocarcinomas. Since ROS1-positive tumours are very sensitive to treatment with tyrosine kinase inhibitors such as crizotinib, detecting this rare genetic alteration may be an important step in the diagnostic work-up of a patient with lung adenocarcinoma.
The traditional approach to detecting ROS1 gene rearrangement is by the use of so-called dual ‘break-apart’ fluorescence in situ hybridisation (FISH) probes, where the rearrangement separates the two ends of the ROS1 gene and thus the two probes. The rearrangement event, when oncogenic, fuses the portion of the ROS1 gene bearing the tyrosine kinase domain with another partner to create a ROS1 fusion gene. An alternative approach to the identification of the abnormal DNA sequence created by the rearrangement event is to use massive parallel ‘next-generation’ sequencing (NGS). A variety of approaches using this technology may be used, and commercial platforms are now available, for use with test kits covering a range of fusion genes, including ROS1. Following transcription, fusion gene mRNA provides another possibility for detection with polymerase chain reaction (PCR) technology using a multiplex platform capable of detecting a range of known ROS1 fusion gene transcripts. For oncogenic activity, the ROS1 fusion gene transcript must be translated into protein with tyrosine kinase activity. Elevation of ROS1 protein levels in tumour cells may be detected by immunohistochemistry (IHC) using antibodies against ROS1 protein. This leaves open a possible scenario, rather like testing for ALK gene rearrangement, where elevations in the protein may be used as a surrogate marker for the presence of a ROS1 gene rearrangement. Furthermore, a positive IHC test is likely to be an indication of a functional rearrangement, since the protein must be present for oncogenic activity and the protein tyrosine kinase is the target of drug therapy. Details of these various testing approaches are discussed below.
As with all current biomarker testing in non-small cell carcinomas, adequate quality and quantity of tissue is required for testing, and this has been extensively discussed elsewhere [31–34]. Issues relating to pre-analytics are discussed below. For many laboratories, ROS1 testing is not yet a routine. Instead, it may be a test considered after several more routine tests, such as EGFR or KRAS mutation and ALK gene rearrangement, prove negative. Consequently, the samples to be used for ROS1 testing may have been exhausted by prior tests, placing the test at risk when pursued in this way. As ROS1 testing becomes more routine, driven by drug approval and availability, or when ROS1 detection is more generally covered within a targeted NGS gene panel, these risks should diminish.
Fluorescence in situ hybridisation
The screening strategy for ROS1 rearrangement was developed based on the experience of ALK testing in lung adenocarcinomas [17, 23, 32]. While ROS1 alterations may be detected with a variety of techniques, most laboratories rely on FISH assays using a dual-colour break-apart probe design. These involve labelling the 3′ (centromeric) part of the fusion breakpoint with one fluorochrome and the 5′ (telomeric) part with another fluorochrome. It is important to choose a 3′ probe colour [16, 22, 35–37] that allows ROS1 (most commonly a green 3′ fluorochrome) and ALK (an orange 3′ fluorochrome) tests to be distinguished [38], particularly if the two tests are to be run together on one slide, in parallel.
There are two positive ROS1 rearrangement patterns. One is the break-apart pattern (‘classic’ pattern) with one fusion signal (native ROS1) and two separated 3′ and 5′ signals. The other positive pattern is an isolated 3′ signal pattern, usually an isolated green signal, (‘atypical’ pattern) with one fusion signal (native ROS1) and one 3′ signal without the corresponding 5′ signal (Fig. 2). Table 3 summarises the criteria for ROS1 FISH interpretation in NSCLC [23, 32, 35, 36].
Examples of different FISH signal patterns using ROS1 break-apart assays. a–d Vysis LSI ROS1 (Cen) SpectrumGreen Probe and Vysis LSI ROS1 (Tel) SpectrumOrange Probe (Abbott Molecular, IL, USA) on histological specimens. a Normal (negative) ROS1 pattern: two fused signals. b Typical ROS1-positive pattern with fused and split signals. c Atypical ROS1-positive pattern with one fusion signal and isolated 3′ green signals. d Increased ROS1 copy number. This pattern should not be interpreted as positive; e–f ZytoLight SPEC ROS1 (Cen) Green Probe and (Tel) Orange Probe (ZytoVision, Bremerhaven, Germany) on cytological specimens. e Split signals. f Isolated 3′ green signals
For optimal FISH results, there are a number of relevant factors. Use of sections older than 6 months may result in poor hybridisation. In the post-analytic phase, it is important that only intact tumour cells with non-overlapping nuclei are scored. Furthermore, the use of an automated software system (e.g. BioView Duet system, Rehovot, Israel) can facilitate FISH scoring. It should be noted that FISH testing for ROS1 (and ALK) is not restricted to histological tissue sections, but is also applicable to cytological specimens [39, 40].
Immunohistochemistry
Given the rarity of ROS1 rearrangements in NSCLC, screening of tumours by IHC may allow unnecessary FISH analysis in ROS1-negative cases to be avoided and thus dramatically reduce the cost of testing. Based on available data, IHC is an effective screening tool to detect ROS1-positive NSCLC, with a sensitivity of 100 % in most studies and a variable specificity ranging from 92 to 100 %, depending on the threshold used to define positivity [20, 36, 37, 41–45]. These results are based on the use of the ROS1 (D4D6) rabbit monoclonal antibody (Cell Signaling Technology, Danvers, MA, USA) applied at dilutions ranging from 1:50 to 1:1000 with various antigen retrieval methods and use of different amplification and detection systems, in automated instruments or manually. Tumour specimens with known ROS1 rearrangement, or a cellblock of the HCC78 cell line harbouring the SLC34A2-ROS1 fusion gene, can serve as positive controls [20] (Figs. 3a and 4a). In contrast to ALK, where the ganglion cells of the appendix serve as an adequate external control, there is currently no good external benign tissue control for ROS1.
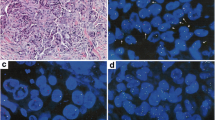
a–f Examples of ROS1 IHC in histological NSCLC specimens (D4D6 antibody, Ventana BenchMark XT; DAB chromogen). a HCC78 cell line (cellblock; ×400). b NSCLC with diffuse, strongly positive staining (×200). c NSCLC with diffuse, granular cytoplasmic staining (×400). d Adenocarcinoma with heterogeneous staining (×200). e Non-neoplastic type II pneumocytes with weak ROS1 staining (×630). f Bone metastasis of a ROS1-negative NSCLC showing strong granular staining of non-neoplastic osteoclastic giant cells (×400). g–h Aberrant immunostaining of ROS1 in a transbronchial biopsy with lung adenocarcinoma. g H&E stain, asterisks show tumour cells. h ROS1 IHC in adjacent hyperplastic type II pneumocytes (arrows) but not in tumour cells (asterisks)
Currently, there is no universally accepted system for how to score IHC results. The thresholds used include either any staining above faint background (if present) or moderate or strong staining (2+/3+). Another option is the use of an H-score with optimal threshold for ROS1 positivity defined as >100 [40] or >150 [43]. Notably, weak and focal staining was found in 31 % of 253 ROS1 wild-type lung carcinomas in one study [44]. However, this had hardly any influence on specificity when appropriate thresholds of positivity were used (i.e. H-score >150). Thus, none of the reported scoring methods have been shown to be clearly superior to the others since all resulted in very good to excellent correlation with FISH results.
Positive ROS1 IHC typically reveals finely granular cytoplasmic staining (Fig. 3). However, the staining pattern may depend on the function and subcellular location of the gene fusion partner [44, 46]. Globular ROS1 immunoreactivity has been described in tumour specimens with the CD74-ROS1 fusion, and membranous staining has been observed in tumours with the EZR-ROS1 fusion [41, 44]. Interestingly, ROS1 expression levels in ROS1-positive lung cancers and cell lines can vary from cell to cell, suggesting dynamic ROS1 protein expression despite homogeneous presence of ROS1 gene rearrangement (Figs. 3 and 4). Detection of ROS1 protein expression in ROS1-positive adenocarcinomas with signet ring cells is challenging since the cytoplasm is largely replaced by non-reactive mucin [44]. The same pitfall has already been shown for ALK IHC [47]. One should be aware that weak ROS1 expression is occasionally detectable in non-neoplastic hyperplastic type II pneumocytes (Fig. 3e, h) and in alveolar macrophages. In bone metastases, there is strong granular cytoplasmic staining of osteoclast-type giant cells (Fig. 3f).
ROS1 IHC in ethanol-fixed and previously Papanicolaou-stained cytological specimens (D4D6 antibody, Leica BondMax; AEC chromogen, ×400). a HCC78 cell line (positive control; cytospin). b ROS1-positive adenocarcinoma. c Small group of ROS1-positive adenocarcinoma cells surrounded by numerous benign respiratory epithelial cells
Lung cancer is often diagnosed by cytology alone, necessitating ROS1 testing in cytological specimens. For cellblocks, the same IHC protocols can be used as for formalin-fixed paraffin-embedded (FFPE) tissue specimens. In the laboratory of one of the authors (L.B.), ROS1 IHC on the BondMax immunostainer is routinely used on conventional, ethanol-fixed and Papanicolaou-stained cytological specimens, including smears or cytospin preparations (Fig. 4). Notably, in the case of limited cytological material, FISH for confirmation of a positive IHC result can be applied to the immunostained slide if 3-amino-9-ethylcarbazole (AEC) was used as a red chromogen. Although IHC on cytological specimens is common practice in many laboratories, immunocytochemistry performed on smears and/or cytospin slides may be much more influenced by various pre-analytical factors [48, 50]. Thus, it should be performed only in laboratories with experience and appropriate quality assurance in place. The simultaneous use of cell blocks would allow testing sequentially for several biomarkers.
ROS1 IHC has a great advantage over FISH in that it can detect rare positive cells or cell groups within a majority of non-neoplastic reactive cells that would be easily missed by FISH. This can be particularly helpful in cytological specimens where architectural tissue context is lost. Although it has been proposed to score ROS IHC only in specimens containing ≥20 tumour cells [41], a positive result in even only a few clearly neoplastic cells can be considered diagnostic.
In summary, IHC is a cost-effective method that can be used to efficiently screen patients with lung cancer for ROS1 rearrangements. Given the laboratory-dependent variability of specificity, confirmation of positive or doubtful ROS1 IHC by FISH or another method is highly recommended.
Non-in situ technologies
In addition to FISH and IHC, a number of non-in situ approaches based on real-time PCR (RT-PCR) or NGS have been developed for the detection of ROS1 gene rearrangements. RT-PCR assays require multiple specific primer sets to discriminate amongst known fusion variants, which can be confirmed by subsequent sequencing [50]. The breakpoints of ROS1 are located at exons 32, 34, 35 and 36, and the most frequent ROS1 fusion partners include SLC34A2, CD74, TPM3, SDC4, EZR, LRIG3, FIG or GOPC, MSN, KDELR2 and CCDC6 [18, 19, 21, 22, 51]. RT-PCR has been successfully utilised to identify positive cases with a sensitivity of 100 % and a specificity of 85–100 %, using FISH as the reference standard method [37, 42]. Multiplex RT-PCR is easy to perform, rapid and relatively inexpensive but may be challenging using RNA extracted from FFPE samples [52]. In addition, as the list of ROS1 fusion partners is quite large and still growing, RT-PCR is likely to miss rare variants. These reasons have limited the use of the technique in clinical practice. Recently, a very sensitive RT-PCR-based method was devised to detect the overexpression of 3′ regions of fusion transcripts involving tumour genes constitutionally repressed or expressed at very low levels [53]; this approach has been successfully applied to ALK gene fusions in lung cancer [53, 54]. Unfortunately, this method cannot be easily applied to ROS1, since the gene is also expressed in normal and hyperplastic lung tissue [15, 55]. An alternative transcript-based method for detecting ROS1 fusion genes is also available. The NanoString assay, capable of detecting known fusion gene transcripts and employing a dual capture and reporter probe system, provides a convenient and commercially available assay that has shown good concordance with FISH and IHC results for ROS1 [50, 55].
A series of innovative approaches to detect gene fusions in multiple targets has been developed using NGS (Table 4). It is remarkable that some of these comprehensive assays require as little as 10 ng of RNA [56], with relatively low failure rates in paraffin-embedded tissue (5.6 % in the authors’ experience [unpublished data]). A very sensitive NGS technique to assess ROS1 and other gene rearrangements in lung cancer is anchored multiplex PCR that targets only the gene of interest, allowing the detection of the specific alteration irrespective of fusion partner. Validation of a gene rearrangement panel using 319 FFPE samples showed 100 % sensitivity and 100 % specificity compared with reference assays [51].
These promising results suggest potential application of non-in situ methodologies in clinical practice, as stand-alone methods or as complementary tests within algorithms for the selection of patients to be treated with ROS1, RET or NTRK inhibitors [57]. However, published data for these assays are still limited.
Concordance between FISH, IHC and PCR
There is good correlation between FISH and IHC using clone D4D6 with a highly sensitive amplification kit. Although some discrepant cases have been reported, ROS1 testing by IHC seems to be highly sensitive, but less specific, also when compared with ALK IHC for detection of the corresponding gene rearrangement. As suggested by others [41], IHC testing of specimens containing at least 20 tumour cells and application of an H-score cut-off of >100 are highly concordant with ROS1 rearrangement by FISH or RT-PCR.
Currently, there is very limited published information on the concordance of IHC, in situ hybridisation (ISH) and non-in situ tests for the detection of ROS1 gene rearrangements in lung adenocarcinoma [36, 37, 42, 58]; less than 30 cases with gene rearrangements have been subjected to comparative study of the three methods. Four of the ROS1 tests currently hold in vitro diagnostic (IVD) and CE-marked status (Table 5). The general consensus seems to be that IHC, ISH and non-in situ methods all are promising for the detection of ROS1-positive cases, with concordance rates well above 90 %. Preliminary conclusions from limited studies suggest a role for IHC as a screening tool, but so far the lack of an IVD-classified IHC assay is problematic. Advantages of RT-PCR analysis include the highest sensitivity reported and the ability to identify translocation partners. On the other hand, there are potential issues around the quality and quantity of mRNA that may be obtained from routine, FFPE NSCLC diagnostic tissues. Furthermore, familiarity and availability of FISH as a technique in detecting other markers, such as ALK, is also important. Perhaps the use of more than one technique could be of value until further experience of testing and companion diagnostics with IVD status have emerged.
General recommendations for ROS1 testing
Current guidelines either do not refer to ROS1 testing [59] or mention it briefly without making any strong recommendation [33]. With recent changes in the status of crizotinib for the treatment of ROS1-positive NSCLC, the case for recommending ROS1 testing will now increase. Certainly, assuming drug treatment is available, response rates in treated patients whose tumours bear a ROS1 rearrangement are impressive [8].
The group of patients more likely to bear a ROS1 fusion gene is largely the same as that currently recommended for testing for EGFR gene mutation and ALK gene rearrangement. Clinical features such as gender, ethnicity and smoking status are not used to select patients for EGFR or ALK testing [33, 59].
Currently, however, as mentioned above, ROS1 testing is often part of a second phase of testing in a patient whose tumour is negative for more common, routinely tested alterations such as EGFR and KRAS mutation and ALK gene rearrangement and who is a lifelong never or long-time ex-smoker. This is based upon the observations that these various addictive oncogenic driver events tend to be mutually exclusive in occurrence and are almost exclusively found in adenocarcinoma. As mentioned above, as NGS techniques are introduced into routine diagnostic practice, ROS1 fusion gene testing will be provided as part of the testing ‘package’, whether or not this particular test was actively sought by a treating physician.
At present, when ROS1 testing is required, it will be reasonable to test the same tumours currently being selected for EGFR mutation and ALK gene rearrangement. Although this should ideally occur in parallel, this is not possible in all cases. Therefore, in order to save tissue and time, it is wise to cut extra blank sections at the first cutting session [60]. FISH remains the core test for the time being. Laboratories may use IHC as a screening tool, but with our current state of knowledge, a positive IHC test should be confirmed by FISH testing. If laboratories chose to use a multiplex PCR approach instead of FISH, they should be aware of the possible pitfalls of this highly sensitive and specific technique, in terms of sample quality and risks of test failure.
There are a number of general principles that should be observed in order to ensure high-quality laboratory testing for predictive biomarkers in NSCLC. The use of adequate control materials, awareness that test outcomes may be confounded by pre-analytical issues in tissue handling and processing and the need for laboratories to participate in external quality assessment programmes are discussed in the following sections.
Guidance on the use of controls
With ISH techniques, the case for study serves as a control when consistently presenting signals, both in tumour cells and in the accompanying normal cells (lymphocytes, fibroblasts, non-neoplastic lung epithelium). A pre-hybridisation assessment of digestion is useful in difficult samples (e.g. very small biopsies with low tumour content).
With IHC, ROS1 protein expression may also be detected in normal cells, namely histiocytes/giant cells, reactive type II pneumocyte hyperplasia and bronchiolar metaplasia at the tumour periphery or in subpleural areas. In most cases, the expression in these cells is weak to moderate (1+/2+ in intensity), and it is unclear whether protein stability may be affected by pre-analytical variables (e.g. time of fixation) [43]. To control for appropriate analytical conditions of ROS1 testing by IHC, it is mandatory to include a piece of tissue from a ROS1 FISH-positive tumour on the same slide of the neoplasm of interest or on a separate slide to use in the same run.
Pre-analytical variables and factors affecting quality of biopsies and surgical samples
Regarding the ROS1 epitope, the influence of pre-analytical factors has not been investigated systematically, but experiences from our group and several others [36, 61] show that the protein is relatively stable and may be detected reliably by IHC. In addition, the corresponding genomic alterations can be reproducibly detected by ISH [62]. To improve nucleic acid stability, new techniques of fixation may become useful [63]. Nonetheless, attention should be paid to a number of basic requirements in order to avoid false-negative results.
Resection material
Surgical material such as lobectomies should initially be handled macroscopically (documentation and gross sectioning) following a standardised protocol. To greater standardise the work-up of resection material, vacuum preservation might be considered [64]. It is important that these procedures are conducted in a standardised way.
Biopsies
Biopsies almost always are transferred into the fixation solution immediately after removal from the patient. The small tissue fragments should be fixed for no longer than 24 h; as shown for several antigens, a gradual decrease in antigenicity may appear over time. Prior or in parallel to ROS1, several immunohistochemical, ISH and molecular markers may need to be analysed to confirm the subtype of NSCLC and the immunological and molecular profile. This means that a minimum amount of tissue/cells is required for reliable analysis. In the context of personalised medicine, this multi-parameter analysis plays an increasing role, which has led to the suggestion to provide at least three to four endobronchial biopsy specimens for pathology.
External quality assessment
For the successful treatment of patients, it is of great importance that molecular test results are accurate, highly reliable, clearly understandable to the clinician and reported within an acceptable turnaround time [59]. In 2012, the European Society of Pathology (ESP) proposed an external quality assessment (EQA) scheme to promote high-quality biomarker testing in NSCLC for EGFR mutation analysis and ALK rearrangement detection. From 2014 onwards, ROS1 testing was also included [65]. The EQA was performed at the beginning of the development of ROS1 testing. The rate of false negativity for IHC on a limited number of ROS1-positive cases was approximately 15 %. The rate of false positivity was <10 %. For FISH, although the number of evaluable cases was limited, no false-negative scores were present in a ROS1-positive control cell line. Overall, at an early stage of ROS1 testing, the EQA showed promising results, emphasising the need for regular EQA monitoring.
Integration of ROS1 into current testing algorithms
Since clinical trials with crizotinib in ROS1-positive patients have used FISH, this method has been considered the ‘gold standard’ for determining ROS1 positivity by the FDA in the USA. European guidelines currently recommend ROS1 rearrangement testing in patients with advanced NSCLC who have previously tested negative for EGFR mutation and ALK rearrangement, including all stage IIIB/IV histological subtypes in non-smokers and the non-squamous cell carcinoma subtype in current or ex-smokers [33]. As the demand for ROS1 testing increases, it is reasonable that ROS1 rearrangement be considered for testing concurrently with ALK rearrangement and EGFR mutation. Cutting extra blank sections for ROS1 testing (and also for additional tests such as PD-L1) at the first cutting session is good practice to avoid tissue waste, especially when the amount of tumour tissue is scarce.
Although validation with large series is needed, IHC could also become a good preliminary, rapid screening method. An algorithm based on IHC screening with further confirmation by a ROS1 break-apart FISH assay in positive or doubtful cases seems appropriate. Nevertheless, in the near future the possibility of using transcript-based methods in a single-tube assay to detect several oncogenic fusions involving the ALK, RET, ROS1 and NTRK1 genes could drastically limit the use of IHC and FISH tests. The algorithm presented in Fig. 5 is proposed for use in routine clinical practice.
Conclusions
With the recent FDA approval of crizotinib for patients with advanced ROS1-positive NSCLC, ROS1 fusion proteins comprise one of only three oncogenic drivers in NSCLC for which an approved targeted therapy is available. Like ALK, ROS1 gene rearrangements may be detected using a break-apart FISH assay, IHC and a number of non-in situ methods. Although FISH was the methodology used in the clinical trials of crizotinib, IHC can also be used as a screening approach if carefully validated. The real-world possibility of false-positive FISH or IHC results strengthens the case for confirmation of positive cases by a second methodology. Targeted NGS is a valid alternative if cost and turnaround time are reasonable. As testing for ROS1 becomes increasingly important for patients with advanced NSCLC, it will be key to share experience and recommendations on how to accurately implement these diagnostic methodologies into routine practice. Regardless of which testing method(s) is used, it is key that routine testing for ROS1 in the clinical setting be carefully validated, with appropriate controls and participation in EQA schemes. To achieve efficient molecular testing in NSCLC and an optimal turnaround time for test results, we propose that EGFR, ALK and ROS1 are tested for upfront and in parallel in NSCLC specimens.
References
Torre LA, Bray F, Siegel RL, Ferlay J, Lortet-Tieulent J, Jemal A (2015) Global cancer statistics, 2012. CA Cancer J Clin 65(2):87–108. doi:10.3322/caac.21262
Rosell R, Carcereny E, Gervais R, Vergnenegre A, Massuti B, Felip E, Palmero R, Garcia-Gomez R, Pallares C, Sanchez JM, Porta R, Cobo M, Garrido P, Longo F, Moran T, Insa A, De Marinis F, Corre R, Bover I, Illiano A, Dansin E, de Castro J, Milella M, Reguart N, Altavilla G, Jimenez U, Provencio M, Moreno MA, Terrasa J, Munoz-Langa J, Valdivia J, Isla D, Domine M, Molinier O, Mazieres J, Baize N, Garcia-Campelo R, Robinet G, Rodriguez-Abreu D, Lopez-Vivanco G, Gebbia V, Ferrera-Delgado L, Bombaron P, Bernabe R, Bearz A, Artal A, Cortesi E, Rolfo C, Sanchez-Ronco M, Drozdowskyj A, Queralt C, de Aguirre I, Ramirez JL, Sanchez JJ, Molina MA, Taron M, Paz-Ares L (2012) Spanish Lung Cancer Group in collaboration with Groupe Francais de P-C, Associazione Italiana Oncologia T Erlotinib versus standard chemotherapy as first-line treatment for European patients with advanced EGFR mutation-positive non-small-cell lung cancer (EURTAC): a multicentre, open-label, randomised phase 3 trial. Lancet Oncol 13(3):239–246. doi:10.1016/S1470-2045(11)70393-X
Solomon BJ, Mok T, Kim DW, Wu YL, Nakagawa K, Mekhail T, Felip E, Cappuzzo F, Paolini J, Usari T, Iyer S, Reisman A, Wilner KD, Tursi J, Blackhall F, Investigators P (2014) First-line crizotinib versus chemotherapy in ALK-positive lung cancer. N Engl J Med 371(23):2167–2177. doi:10.1056/NEJMoa1408440
Savas P, Hughes B, Solomon B (2013) Targeted therapy in lung cancer: IPASS and beyond, keeping abreast of the explosion of targeted therapies for lung cancer. J Thorac Dis 5(Suppl 5):S579–S592. doi:10.3978/j.issn.2072-1439.2013.08.52
Shaw AT, Forcione DG, Digumarthy SR, Iafrate AJ (2011) Case records of the Massachusetts General Hospital. Case 21-2011. A 31-year-old man with ALK-positive adenocarcinoma of the lung. N Engl J Med 365(2):158–167. doi:10.1056/NEJMcpc1102202
Kris MG, Johnson BE, Berry LD, Kwiatkowski DJ, Iafrate AJ, Wistuba II, Varella-Garcia M, Franklin WA, Aronson SL, Su P-F (2014) Using multiplexed assays of oncogenic drivers in lung cancers to select targeted drugs. Jama 311(19):1998–2006
Network TCGAR (2014) Comprehensive molecular profiling of lung adenocarcinoma. Nature 511 (7511):543-550. doi: 10.1038/nature13385 http://www.nature.com/nature/journal/v511/n7511/abs/nature13385.html#supplementary-information
Shaw AT, Ou SH, Bang YJ, Camidge DR, Solomon BJ, Salgia R, Riely GJ, Varella-Garcia M, Shapiro GI, Costa DB, Doebele RC, Le LP, Zheng Z, Tan W, Stephenson P, Shreeve SM, Tye LM, Christensen JG, Wilner KD, Clark JW, Iafrate AJ (2014) Crizotinib in ROS1-rearranged non-small-cell lung cancer. N Engl J Med 371(21):1963–1971. doi:10.1056/NEJMoa1406766
Bergethon K, Shaw AT, Ou SH, Katayama R, Lovly CM, McDonald NT, Massion PP, Siwak-Tapp C, Gonzalez A, Fang R, Mark EJ, Batten JM, Chen H, Wilner KD, Kwak EL, Clark JW, Carbone DP, Ji H, Engelman JA, Mino-Kenudson M, Pao W, Iafrate AJ (2012) ROS1 rearrangements define a unique molecular class of lung cancers. J Clin Oncol 30(8):863–870. doi:10.1200/JCO.2011.35.6345
Pfizer. (2016) Crizotinib Summary of Product Characteristics.
Pfizer. (2016) Crizotinib Prescribing Information.
FDA (2016) FDA expands use of Xalkori to treat rare form of advanced non-small cell lung cancer. http://www.fda.gov/newsevents/newsroom/pressannouncements/ucm490329.htm.
Shibuya M, Hanafusa H, Balduzzi PC (1982) Cellular sequences related to three new onc genes of avian sarcoma virus (fps, yes, and ros) and their expression in normal and transformed cells. J Virol 42(1):143–152
Charest A, Lane K, McMahon K, Park J, Preisinger E, Conroy H, Housman D (2003) Fusion of FIG to the receptor tyrosine kinase ROS in a glioblastoma with an interstitial del (6) (q21q21). Genes Chromosom Cancer 37(1):58–71. doi:10.1002/gcc.10207
Acquaviva J, Wong R, Charest A (2009) The multifaceted roles of the receptor tyrosine kinase ROS in development and cancer. Biochim Biophys Acta 1795(1):37–52. doi:10.1016/j.bbcan.2008.07.006
Davies KD, Doebele RC (2013) Molecular pathways: ROS1 fusion proteins in cancer. Clin Cancer Res 19(15):4040–4045. doi:10.1158/1078-0432.CCR-12-2851
Kim MH, Shim HS, Kang DR, Jung JY, Lee CY, Kim DJ, Lee JG, Bae MK, Kim HR, Lim SM, Kim EY, Park JS, Chung KY, Kim HJ, Kim JH, Cho BC (2014) Clinical and prognostic implications of ALK and ROS1 rearrangements in never-smokers with surgically resected lung adenocarcinoma. Lung Cancer 83(3):389–395. doi:10.1016/j.lungcan.2014.01.003
Govindan R, Ding L, Griffith M, Subramanian J, Dees ND, Kanchi KL, Maher CA, Fulton R, Fulton L, Wallis J, Chen K, Walker J, McDonald S, Bose R, Ornitz D, Xiong D, You M, Dooling DJ, Watson M, Mardis ER, Wilson RK (2012) Genomic landscape of non-small cell lung cancer in smokers and never-smokers. Cell 150(6):1121–1134. doi:10.1016/j.cell.2012.08.024
Rikova K, Guo A, Zeng Q, Possemato A, Yu J, Haack H, Nardone J, Lee K, Reeves C, Li Y, Hu Y, Tan Z, Stokes M, Sullivan L, Mitchell J, Wetzel R, Macneill J, Ren JM, Yuan J, Bakalarski CE, Villen J, Kornhauser JM, Smith B, Li D, Zhou X, Gygi SP, Gu TL, Polakiewicz RD, Rush J, Comb MJ (2007) Global survey of phosphotyrosine signaling identifies oncogenic kinases in lung cancer. Cell 131(6):1190–1203. doi:10.1016/j.cell.2007.11.025
Rimkunas VM, Crosby KE, Li D, Hu Y, Kelly ME, Gu TL, Mack JS, Silver MR, Zhou X, Haack H (2012) Analysis of receptor tyrosine kinase ROS1-positive tumors in non-small cell lung cancer: identification of a FIG-ROS1 fusion. Clin Cancer Res 18(16):4449–4457. doi:10.1158/1078-0432.CCR-11-3351
Seo JS, Ju YS, Lee WC, Shin JY, Lee JK, Bleazard T, Lee J, Jung YJ, Kim JO, Shin JY, Yu SB, Kim J, Lee ER, Kang CH, Park IK, Rhee H, Lee SH, Kim JI, Kang JH, Kim YT (2012) The transcriptional landscape and mutational profile of lung adenocarcinoma. Genome Res 22(11):2109–2119. doi:10.1101/gr.145144.112
Takeuchi K, Soda M, Togashi Y, Suzuki R, Sakata S, Hatano S, Asaka R, Hamanaka W, Ninomiya H, Uehara H, Lim Choi Y, Satoh Y, Okumura S, Nakagawa K, Mano H, Ishikawa Y (2012) RET, ROS1 and ALK fusions in lung cancer. Nat Med 18(3):378–381. doi:10.1038/nm.2658
Yoshida A, Kohno T, Tsuta K, Wakai S, Arai Y, Shimada Y, Asamura H, Furuta K, Shibata T, Tsuda H (2013) ROS1-rearranged lung cancer: a clinicopathologic and molecular study of 15 surgical cases. Am J Surg Pathol 37(4):554–562. doi:10.1097/PAS.0b013e3182758fe6
Birchmeier C, Sharma S, Wigler M (1987) Expression and rearrangement of the ROS1 gene in human glioblastoma cells. Proc Natl Acad Sci U S A 84(24):9270–9274
Jun HJ, Johnson H, Bronson RT, de Feraudy S, White F, Charest A (2012) The oncogenic lung cancer fusion kinase CD74-ROS activates a novel invasiveness pathway through E-Syt1 phosphorylation. Cancer Res 72(15):3764–3774. doi:10.1158/0008-5472.CAN-11-3990
Charest A, Kheifets V, Park J, Lane K, McMahon K, Nutt CL, Housman D (2003) Oncogenic targeting of an activated tyrosine kinase to the Golgi apparatus in a glioblastoma. Proc Natl Acad Sci U S A 100(3):916–921. doi:10.1073/pnas.242741799
Mazieres J, Zalcman G, Crino L, Biondani P, Barlesi F, Filleron T, Dingemans AM, Lena H, Monnet I, Rothschild SI, Cappuzzo F, Besse B, Thiberville L, Rouviere D, Dziadziuszko R, Smit EF, Wolf J, Spirig C, Pecuchet N, Leenders F, Heuckmann JM, Diebold J, Milia JD, Thomas RK, Gautschi O (2015) Crizotinib therapy for advanced lung adenocarcinoma and a ROS1 rearrangement: results from the EUROS1 cohort. J Clin Oncol 33(9):992–999. doi:10.1200/JCO.2014.58.3302
Moro-Sibilot D, Faivre L, Zalcman G, Perol M, Mazières J, Barlesi F, Otto J, Monnet I, Cortot A, Wislez M, Léna H, Souquet PJ, Lantuejoul S, Rouquette I, Mcleer-Florin A, Ferretti G, Hoog-Labouret N, Nowak F, Jimenez M, Vassal G (2015) Crizotinib in patients with ROS1 NSCLC. Preliminary results of the AcSé trial. J Thorac Oncol 10(9):S261–S406. doi:10.1016/S1556-0864(16)30011-9
Goto K, Yang JCH, Kim D-W, Lu S, Seto T, Yang J-J, Yamamoto N, Ahn M-J, Takahashi T, Yamanaka T, Kemner A, Roychowdhury D, Paolini J, Wilner K, Wu Y-L (2016) Phase II study of crizotinib in East Asian patients (pts) with ROS1-positive advanced non-small cell lung cancer (NSCLC). J Clin Oncol 34(15 suppl.):Abstr. 9022
Shaw AT, Kim DW, Nakagawa K, Seto T, Crino L, Ahn MJ, De Pas T, Besse B, Solomon BJ, Blackhall F, Wu YL, Thomas M, O'Byrne KJ, Moro-Sibilot D, Camidge DR, Mok T, Hirsh V, Riely GJ, Iyer S, Tassell V, Polli A, Wilner KD, Janne PA (2013) Crizotinib versus chemotherapy in advanced ALK-positive lung cancer. N Engl J Med 368(25):2385–2394. doi:10.1056/NEJMoa1214886
Dietel M, Bubendorf L, Dingemans AM, Dooms C, Elmberger G, Garcia RC, Kerr KM, Lim E, Lopez-Rios F, Thunnissen E, Van Schil PE, von Laffert M (2016) Diagnostic procedures for non-small-cell lung cancer (NSCLC): recommendations of the European Expert Group. Thorax 71(2):177–184. doi:10.1136/thoraxjnl-2014-206677
Thunnissen E, Bubendorf L, Dietel M, Elmberger G, Kerr K, Lopez-Rios F, Moch H, Olszewski W, Pauwels P, Penault-Llorca F, Rossi G (2012) EML4-ALK testing in non-small cell carcinomas of the lung: a review with recommendations. Virchows Arch 461(3):245–257. doi:10.1007/s00428-012-1281-4
Kerr KM, Bubendorf L, Edelman MJ, Marchetti A, Mok T, Novello S, O'Byrne K, Stahel R, Peters S, Felip E, Panel M, Panel M (2014) Second ESMO consensus conference on lung cancer: pathology and molecular biomarkers for non-small-cell lung cancer. Ann Oncol 25(9):1681–1690. doi:10.1093/annonc/mdu145
Thunnissen E, van der Oord K, den Bakker M (2014) Prognostic and predictive biomarkers in lung cancer. A review. Virchows Arch 464(3):347–358. doi:10.1007/s00428-014-1535-4
Gainor JF, Shaw AT (2013) Novel targets in non-small cell lung cancer: ROS1 and RET fusions. Oncologist 18(7):865–875. doi:10.1634/theoncologist.2013-0095
Mescam-Mancini L, Lantuejoul S, Moro-Sibilot D, Rouquette I, Souquet PJ, Audigier-Valette C, Sabourin JC, Decroisette C, Sakhri L, Brambilla E, McLeer-Florin A (2014) On the relevance of a testing algorithm for the detection of ROS1-rearranged lung adenocarcinomas. Lung Cancer 83(2):168–173. doi:10.1016/j.lungcan.2013.11.019
Shan L, Lian F, Guo L, Qiu T, Ling Y, Ying J, Lin D (2015) Detection of ROS1 gene rearrangement in lung adenocarcinoma: comparison of IHC, FISH and real-time RT-PCR. PLoS One 10(3):e0120422. doi:10.1371/journal.pone.0120422
Jin Y, Sun PL, Kim H, Park E, Shim HS, Jheon S, Kim K, Lee CT, Chung JH (2015) Erratum to: ROS1 gene rearrangement and copy number gain in non-small cell lung cancer. Virchows Arch 467(1):123. doi:10.1007/s00428-015-1775-y
Bozzetti C, Nizzoli R, Tiseo M, Squadrilli A, Lagrasta C, Buti S, Gasparro D, Zanoni D, Majori M, De Filippo M, Mazzoni F, Maddau C, Naldi N, Sammarelli G, Frati C, Pinto C, Ardizzoni A (2015) ALK and ROS1 rearrangements tested by fluorescence in situ hybridization in cytological smears from advanced non-small cell lung cancer patients. Diagn Cytopathol 43(11):941–946. doi:10.1002/dc.23318
Savic S, Bubendorf L (2012) Role of fluorescence in situ hybridization in lung cancer cytology. Acta Cytol 56(6):611–621. doi:10.1159/000339792
Boyle TA, Masago K, Ellison KE, Yatabe Y, Hirsch FR (2015) ROS1 immunohistochemistry among major genotypes of non-small-cell lung cancer. Clin Lung Cancer 16(2):106–111. doi:10.1016/j.cllc.2014.10.003
Cao B, Wei P, Liu Z, Bi R, Lu Y, Zhang L, Zhang J, Yang Y, Shen C, Du X, Zhou X (2016) Detection of lung adenocarcinoma with ROS1 rearrangement by IHC, FISH, and RT-PCR and analysis of its clinicopathologic features. Onco Targets Ther 9:131–138. doi:10.2147/OTT.S94997
Sholl LM, Sun H, Butaney M, Zhang C, Lee C, Janne PA, Rodig SJ (2013) ROS1 immunohistochemistry for detection of ROS1-rearranged lung adenocarcinomas. Am J Surg Pathol 37(9):1441–1449. doi:10.1097/PAS.0b013e3182960fa7
Yoshida A, Tsuta K, Wakai S, Arai Y, Asamura H, Shibata T, Furuta K, Kohno T, Kushima R (2014) Immunohistochemical detection of ROS1 is useful for identifying ROS1 rearrangements in lung cancers. Mod Pathol 27(5):711–720. doi:10.1038/modpathol.2013.192
Rogers TM, Russell PA, Wright G, Wainer Z, Pang JM, Henricksen LA, Singh S, Stanislaw S, Grille J, Roberts E, Solomon B, Fox SB (2015) Comparison of methods in the detection of ALK and ROS1 rearrangements in lung cancer. J Thorac Oncol 10(4):611–618. doi:10.1097/JTO.0000000000000465
Bruce B, Khanna G, Ren L, Landberg G, Jirstrom K, Powell C, Borczuk A, Keller ET, Wojno KJ, Meltzer P, Baird K, McClatchey A, Bretscher A, Hewitt SM, Khanna C (2007) Expression of the cytoskeleton linker protein ezrin in human cancers. Clin Exp Metastasis 24(2):69–78. doi:10.1007/s10585-006-9050-x
Murakami Y, Mitsudomi T, Yatabe Y (2012) A Screening Method for the ALK Fusion Gene in NSCLC. Front Oncol 2:24. doi:10.3389/fonc.2012.00024
Fischer AH, Schwartz MR, Moriarty AT, Wilbur DC, Souers R, Fatheree L, Booth CN, Clayton AC, Kurtyz DF, Padmanabhan V, Crothers BA (2014) Immunohistochemistry practices of cytopathology laboratories: a survey of participants in the College of American Pathologists Nongynecologic Cytopathology Education Program. Arch Pathol Lab Med 138(9):1167–1172. doi:10.5858/arpa.2013-0259-CP
Kirbis IS, Maxwell P, Flezar MS, Miller K, Ibrahim M (2011) External quality control for immunocytochemistry on cytology samples: a review of UK NEQAS ICC (cytology module) results. Cytopathology 22(4):230–237. doi:10.1111/j.1365-2303.2011.00867.x
Lee SE, Lee B, Hong M, Song JY, Jung K, Lira ME, Mao M, Han J, Kim J, Choi YL (2015) Comprehensive analysis of RET and ROS1 rearrangement in lung adenocarcinoma. Mod Pathol 28(4):468–479. doi:10.1038/modpathol.2014.107
Zheng Z, Liebers M, Zhelyazkova B, Cao Y, Panditi D, Lynch KD, Chen J, Robinson HE, Shim HS, Chmielecki J, Pao W, Engelman JA, Iafrate AJ, Le LP (2014) Anchored multiplex PCR for targeted next-generation sequencing. Nat Med 20(12):1479–1484. doi:10.1038/nm.3729
Ribeiro-Silva A, Zhang H, Jeffrey SS (2007) RNA extraction from ten year old formalin-fixed paraffin-embedded breast cancer samples: a comparison of column purification and magnetic bead-based technologies. BMC Mol Biol 8:118. doi:10.1186/1471-2199-8-118
Gruber K, Horn H, Kalla J, Fritz P, Rosenwald A, Kohlhaufl M, Friedel G, Schwab M, Ott G, Kalla C (2014) Detection of rearrangements and transcriptional up-regulation of ALK in FFPE lung cancer specimens using a novel, sensitive, quantitative reverse transcription polymerase chain reaction assay. J Thorac Oncol 9(3):307–315. doi:10.1097/JTO.0000000000000068
Moskalev EA, Frohnauer J, Merkelbach-Bruse S, Schildhaus HU, Dimmler A, Schubert T, Boltze C, Konig H, Fuchs F, Sirbu H, Rieker RJ, Agaimy A, Hartmann A, Haller F (2014) Sensitive and specific detection of EML4-ALK rearrangements in non-small cell lung cancer (NSCLC) specimens by multiplex amplicon RNA massive parallel sequencing. Lung Cancer 84(3):215–221. doi:10.1016/j.lungcan.2014.03.002
Lira ME, Choi YL, Lim SM, Deng S, Huang D, Ozeck M, Han J, Jeong JY, Shim HS, Cho BC, Kim J, Ahn MJ, Mao M (2014) A single-tube multiplexed assay for detecting ALK, ROS1, and RET fusions in lung cancer. J Mol Diagn 16(2):229–243. doi:10.1016/j.jmoldx.2013.11.007
Thermo Fisher Scientific. Oncomine™ Solid Tumour Fusion Transcript Kit Product Information. Available at https://www.thermofisher.com/uk/en/home/clinical/molecular-diagnostics/oncology-diagnostics.html
Swanton C, Govindan R (2016) Clinical implications of genomic discoveries in lung cancer. N Engl J Med 374(19):1864–1873. doi:10.1056/NEJMra1504688
Cha YJ, Lee JS, Kim HR, Lim SM, Cho BC, Lee CY, Shim HS (2014) Screening of ROS1 rearrangements in lung adenocarcinoma by immunohistochemistry and comparison with ALK rearrangements. PLoS One 9(7):e103333. doi:10.1371/journal.pone.0103333
Lindeman NI, Cagle PT, Beasley MB, Chitale DA, Dacic S, Giaccone G, Jenkins RB, Kwiatkowski DJ, Saldivar JS, Squire J, Thunnissen E, Ladanyi M, College of American Pathologists International Association for the Study of Lung C, Association for Molecular P (2013) Molecular testing guideline for selection of lung cancer patients for EGFR and ALK tyrosine kinase inhibitors: guideline from the College of American Pathologists, International Association for the Study of Lung Cancer, and Association for Molecular Pathology. J Mol Diagn 15(4):415–453. doi:10.1016/j.jmoldx.2013.03.001
Felip E, Concha Á, de Castro J, Gómez-Román J, Garrido P, Ramírez J, Isla D, Sanz J, Paz-Ares L, López-Ríos F (2015) Biomarker testing in advanced non-small-cell lung cancer: a National Consensus of the Spanish Society of Pathology and the Spanish Society of Medical Oncology. Clin Transl Oncol 17(2):103–112. doi:10.1007/s12094-014-1248-9
Zhang N, Yang JJ, Zhang XC, Xie Z, Wang BC, Tu HY, Jiang BY, Wu YL (2014) Responses to crizotinib in a patient with c-ros oncogene 1, receptor tyrosine kinase-positive advanced lung adenocarcinoma: A case report. Oncol Lett 8(6):2624–2626. doi:10.3892/ol.2014.2571
Jurmeister P, Lenze D, Berg E, Mende S, Schaper F, Kellner U, Herbst H, Sers C, Budczies J, Dietel M, Hummel M, von Laffert M (2015) Parallel screening for ALK, MET and ROS1 alterations in non-small cell lung cancer with implications for daily routine testing. Lung Cancer 87(2):122–129. doi:10.1016/j.lungcan.2014.11.018
Bussolati G, Annaratone L, Medico E, D'Armento G, Sapino A (2011) Formalin fixation at low temperature better preserves nucleic acid integrity. PLoS One 6(6):e21043. doi:10.1371/journal.pone.0021043
Bussolati G, Annaratone L, Maletta F (2015) The pre-analytical phase in surgical pathology. Recent Results Cancer Res 199:1–13. doi:10.1007/978-3-319-13957-9_1
Tembuyser L, Tack V, Zwaenepoel K, Pauwels P, Miller K, Bubendorf L, Kerr K, Schuuring E, Thunnissen E, Dequeker EM (2014) The relevance of external quality assessment for molecular testing for ALK positive non-small cell lung cancer: results from two pilot rounds show room for optimization. PLoS One 9(11):e112159. doi:10.1371/journal.pone.0112159
Arai Y, Totoki Y, Takahashi H, Nakamura H, Hama N, Kohno T, Tsuta K, Yoshida A, Asamura H, Mutoh M, Hosoda F, Tsuda H, Shibata T (2013) Mouse model for ROS1-rearranged lung cancer. PLoS One 8(2):e56010. doi:10.1371/journal.pone.0056010
Cai W, Li X, Su C, Fan L, Zheng L, Fei K, Zhou C, Manegold C, Schmid-Bindert G (2013) ROS1 fusions in Chinese patients with non-small-cell lung cancer. Ann Oncol 24(7):1822–1827. doi:10.1093/annonc/mdt071
Cheng H, Ye L, Xue L (2014) Detection of ROS1 gene rearrangement by FISH and analysis of its clinical features in non-small cell lung cancer patients. Zhonghua Zhong Liu Za Zhi 36(10):751–754
Davies KD, Le AT, Theodoro MF, Skokan MC, Aisner DL, Berge EM, Terracciano LM, Cappuzzo F, Incarbone M, Roncalli M, Alloisio M, Santoro A, Camidge DR, Varella-Garcia M, Doebele RC (2012) Identifying and targeting ROS1 gene fusions in non-small cell lung cancer. Clin Cancer Res 18(17):4570–4579. doi:10.1158/1078-0432.CCR-12-0550
Fu S, Liang Y, Lin YB, Wang F, Huang MY, Zhang ZC, Wang J, Cen WJ, Shao JY (2015) The Frequency and Clinical Implication of ROS1 and RET Rearrangements in Resected Stage IIIA-N2 Non-Small Cell Lung Cancer Patients. PLoS One 10(4):e0124354. doi:10.1371/journal.pone.0124354
Go H, Kim DW, Kim D, Keam B, Kim TM, Lee SH, Heo DS, Bang YJ, Chung DH (2013) Clinicopathologic analysis of ROS1-rearranged non-small-cell lung cancer and proposal of a diagnostic algorithm. J Thorac Oncol 8(11):1445–1450. doi:10.1097/JTO.0b013e3182a4dd6e
Jin Y, Sun PL, Kim H, Park E, Shim HS, Jheon S, Kim K, Lee CT, Chung JH (2015) ROS1 gene rearrangement and copy number gain in non-small cell lung cancer. Virchows Arch 466(1):45–52. doi:10.1007/s00428-014-1679-2
Karlsson A, Brunnstrom H, Lindquist KE, Jirstrom K, Jonsson M, Rosengren F, Reutersward C, Cirenajwis H, Borg A, Jonsson P, Planck M, Jonsson G, Staaf J (2015) Mutational and gene fusion analyses of primary large cell and large cell neuroendocrine lung cancer. Oncotarget 6(26):22028–22037. doi:10.18632/oncotarget.4314
Kim HR, Lim SM, Kim HJ, Hwang SK, Park JK, Shin E, Bae MK, Ou SH, Wang J, Jewell SS, Kang DR, Soo RA, Haack H, Kim JH, Shim HS, Cho BC (2013) The frequency and impact of ROS1 rearrangement on clinical outcomes in never smokers with lung adenocarcinoma. Ann Oncol 24(9):2364–2370. doi:10.1093/annonc/mdt220
Kirita K, Izumo T, Matsumoto Y, Hiraishi Y, Tsuchida T (2016) Bronchoscopic Re-biopsy for Mutational Analysis of Non-small Cell Lung Cancer. Lung. doi:10.1007/s00408-016-9864-5
Li C, Fang R, Sun Y, Han X, Li F, Gao B, Iafrate AJ, Liu XY, Pao W, Chen H, Ji H (2011) Spectrum of oncogenic driver mutations in lung adenocarcinomas from East Asian never smokers. PLoS One 6(11):e28204. doi:10.1371/journal.pone.0028204
Matsuura S, Shinmura K, Kamo T, Igarashi H, Maruyama K, Tajima M, Ogawa H, Tanahashi M, Niwa H, Funai K, Kohno T, Suda T, Sugimura H (2013) CD74-ROS1 fusion transcripts in resected non-small cell lung carcinoma. Oncol Rep 30(4):1675–1680. doi:10.3892/or.2013.2630
Okamoto I, Sakai K, Morita S, Yoshioka H, Kaneda H, Takeda K, Hirashima T, Kogure Y, Kimura T, Takahashi T, Atagi S, Seto T, Sawa T, Yamamoto M, Satouchi M, Okuno M, Nagase S, Takayama K, Tomii K, Maeda T, Oizumi S, Fujii S, Akashi Y, Nishino K, Ebi N, Nakagawa K, Nakanishi Y, Nishio K (2014) Multiplex genomic profiling of non-small cell lung cancers from the LETS phase III trial of first-line S-1/carboplatin versus paclitaxel/carboplatin: results of a West Japan Oncology Group study. Oncotarget 5(8):2293–2304. doi:10.18632/oncotarget.1906
Scheffler M, Schultheis A, Teixido C, Michels S, Morales-Espinosa D, Viteri S, Hartmann W, Merkelbach-Bruse S, Fischer R, Schildhaus HU, Fassunke J, Sebastian M, Serke M, Kaminsky B, Randerath W, Gerigk U, Ko YD, Kruger S, Schnell R, Rothe A, Kropf-Sanchen C, Heukamp L, Rosell R, Buttner R, Wolf J (2015) ROS1 rearrangements in lung adenocarcinoma: prognostic impact, therapeutic options and genetic variability. Oncotarget 6(12):10577–10585. doi:10.18632/oncotarget.3387
Suehara Y, Arcila M, Wang L, Hasanovic A, Ang D, Ito T, Kimura Y, Drilon A, Guha U, Rusch V, Kris MG, Zakowski MF, Rizvi N, Khanin R, Ladanyi M (2012) Identification of KIF5B-RET and GOPC-ROS1 fusions in lung adenocarcinomas through a comprehensive mRNA-based screen for tyrosine kinase fusions. Clin Cancer Res 18(24):6599–6608. doi:10.1158/1078-0432.CCR-12-0838
Wang R, Zhang Y, Pan Y, Li Y, Hu H, Cai D, Li H, Ye T, Luo X, Zhang Y, Li B, Shen L, Sun Y, Chen H (2015) Comprehensive investigation of oncogenic driver mutations in Chinese non-small cell lung cancer patients. Oncotarget 6(33):34300–34308. doi:10.18632/oncotarget.5549
Warth A, Muley T, Dienemann H, Goeppert B, Stenzinger A, Schnabel PA, Schirmacher P, Penzel R, Weichert W (2014) ROS1 expression and translocations in non-small-cell lung cancer: clinicopathological analysis of 1478 cases. Histopathology 65(2):187–194. doi:10.1111/his.12379
Zhang Q, Sun T, Kang P, Qian K, Deng B, Zhou J, Wang R, Jiang B, Li K, Liu F, Wu S, Tan Q (2016) Combined analysis of rearrangement of ALK, ROS1, somatic mutation of EGFR, KRAS, BRAF, PIK3CA, and mRNA expression of ERCC1, TYMS, RRM1, TUBB3, EGFR in patients with non-small cell lung cancer and their clinical significance. Cancer Chemother Pharmacol 77(3):583–593. doi:10.1007/s00280-016-2969-y
Zhao C, Li X, Li J, Zhang Y, Ren S, Chen X, Zhou C (2014) Detecting ALK, ROS1 and RET Fusion Genes in Cell Block Samples. Transl Oncol 7(3):363–367. doi:10.1016/j.tranon.2014.04.013
Zhong S, Zhang H, Bai D, Gao D, Zheng J, Ding Y (2015) Detection of ALK, ROS1 and RET fusion genes in non-small cell lung cancer patients and its clinicopathologic correlation. Zhonghua Bing Li Xue Za Zhi 44(9):639–643
Drilon A, Wang L, Arcila ME, Balasubramanian S, Greenbowe JR, Ross JS, Stephens P, Lipson D, Miller VA, Kris MG, Ladanyi M, Rizvi NA (2015) Broad, Hybrid Capture-Based Next-Generation Sequencing Identifies Actionable Genomic Alterations in Lung Adenocarcinomas Otherwise Negative for Such Alterations by Other Genomic Testing Approaches. Clin Cancer Res 21(16):3631–3639. doi:10.1158/1078-0432.CCR-14-2683
Leary RJ, Kinde I, Diehl F, Schmidt K, Clouser C, Duncan C, Antipova A, Lee C, McKernan K, De La Vega FM, Kinzler KW, Vogelstein B, Diaz LA Jr, Velculescu VE (2010) Development of personalized tumor biomarkers using massively parallel sequencing. Sci Transl Med 2(20):20ra14. doi:10.1126/scitranslmed.3000702
Acknowledgments
This paper was produced following a consensus meeting on 25 November 2015, Berlin, Germany, organised and sponsored by Pfizer. Medical writing support was provided by ACUMED® (Tytherington, UK), an Ashfield Company, part of UDG Healthcare plc, and was funded by Pfizer.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
AR and GR have participated in advisory boards on behalf of Pfizer. FLR has received honoraria and research funding from Abbott, Pfizer and Roche. GR has participated in advisory boards on behalf of Pfizer and Qiagen. KK and LB have received honoraria from Pfizer. PP has received research grants and honoraria from Pfizer. The remaining authors have no conflicts of interests to declare.
Additional information
This paper reflects the consensus of an expert working group of pathologists that met in Berlin on 25 November 2015. Authors are listed in alphabetical order and contributed equally to this manuscript
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Bubendorf, L., Büttner, R., Al-Dayel, F. et al. Testing for ROS1 in non-small cell lung cancer: a review with recommendations. Virchows Arch 469, 489–503 (2016). https://doi.org/10.1007/s00428-016-2000-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00428-016-2000-3