Abstract
Main conclusion
We identified 66 melon SAUR genes by bioinformatic analyses. CmSAUR19, 38, 58, 62 genes are specifically expressed in different stages of fruit growth, suggesting their participation in regulating fruit development.
Abstract
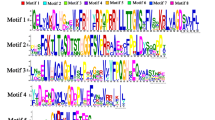
Auxin plays a crucial role in plant growth by regulating the multiple auxin response genes. However, in melon (Cucumis melo L.), the functions of the auxin early response gene family SAUR (Small auxin up RNA) genes in fruit development are still poorly understood. Through genome-wide characterization of CmSAUR family in melon, we identified a total of 66 CmSAUR genes. The open reading frames of the CmSAUR genes ranged from 234 to 525 bp, containing only one exon and lacking introns. Chromosomal position and phylogenetic tree analyses found that the two gene clusters in the melon chromosome are highly homologous in the Cucurbitaceae plants. Among the four conserved motifs in CmSAUR proteins, motif 1, motif 2, and motif 3 located in most of the family protein sequences, and motif 4 showed a close correlation with the two gene clusters. The CmSAUR28 and CmSAUR58 genes have auxin response elements located in the promoters, suggesting they may be involved in the auxin signaling pathway to regulate fruit development. Through transcriptomic profiling in the four developmental stages of fruit and different lateral organs, we selected 16 differentially-expressed SAUR genes for performing further expression analyses. qRT-PCR results showed that five SAUR genes are specifically expressed in flower organs and ovaries. CmSAUR19 and CmSAUR58 were significantly accumulated in the early developmental stage of the fruit. CmSAUR38 and CmAUR62 showed high expression in the climacteric and post-climacteric stages, suggesting their specific role in controlling fruit ripening. This work provides a foundation for further exploring the function of the SAUR gene in fruit development.






Similar content being viewed by others
Data availability
All data generated or analyzed during this study are included in this published article. Raw reads for RNA-Seq data generated in this study have been deposited to the NCBI SRA (https://www.ncbi.nlm.nih.gov/sra) and are available under the BioProject accession numbers PRJNA543288 and PRJNA803327.
Abbreviations
- ARF:
-
Auxin response factors
- Aux/IAA:
-
Auxin/indole-3-acetic acid
- BR:
-
Brassinosteroids
- cDNA:
-
Complementary DNA
- DAP:
-
Days after pollination
- GAPDH:
-
Glyceraldehyde-3-phosphate dehydrogenase
- GH3:
-
Gretchen Hagen 3
- ORF:
-
Open reading frame
- qRT-PCR:
-
Quantitative real-time PCR
- SAUR:
-
Small auxin-up RNA
References
Abel S, Theologis A (1996) Early genes and auxin action. Plant Physiol 111:9–17. https://doi.org/10.1104/pp.111.1.9
Abel S, Oeller PW, Theologis A (1994) Early auxin-induced genes encode short-lived nuclear proteins. Proc Natl Acad Sci USA 91:326–330. https://doi.org/10.1073/pnas.91.1.326
Bai MY, Shang JX, Oh E, Fan M, Bai Y, Zentella R, Sun TP, Wang ZY (2012) Brassinosteroid, gibberellin and phytochrome impinge on a common transcription module in Arabidopsis. Nat Cell Biol 14:810–817. https://doi.org/10.1038/ncb2546
Chae K, Isaacs CG, Reeves PH, Maloney GS, Muday GK, Nagpal P, Reed JW (2012) Arabidopsis SMALL AUXIN UP RNA63 promotes hypocotyl and stamen filament elongation. Plant J 71:684–697. https://doi.org/10.1111/j.1365-313X.2012.05024.x
Deng G, Huang X, Xie L, Tan SB, Gbokie T, Bao YN, Xie ZL, Yi KX (2019) Identification and expression of SAUR genes in the CAM Plant Agave. Genes 10:555. https://doi.org/10.3390/genes10070555
Favero DS, Le KN, Neff MM (2017) Brassinosteroid signaling converges with SUPPRESSOR OF PHYTOCHROME B4-#3 to influence the expression of SMALL AUXIN UP RNA genes and hypocotyl growth. The Plant Journal: for Cell and Molecular Biology 89(6):1133–1145. https://doi.org/10.1111/tpj.13451
Goda H, Sawa S, Asami T, Fujioka S, Shimada Y, Yoshida S (2004) Comprehensive comparison of auxin-regulated and brassinosteroid-regulated genes in Arabidopsis. Plant Physiol 134:1555–1573. https://doi.org/10.1104/pp.103.034736
Gretchen H (2015) Auxin signal transduction. Essays Biochem 58:1–12
Guo Y, Jiang QY, Hu Z, Sun XJ, Fan SJ, Zhang H (2018) Function of the auxin-responsive gene TaSAUR75 under salt and drought stress. Crop Journal 6(2):181–190
Hagen G, Guilfoyle TJ (2002) Auxin-responsive gene expression: genes, promoters and regulatory factors. Plant Mol Biol 49:373–385. https://doi.org/10.1007/978-94-010-0377-3_9
Hagen G, Kleinschmidt AJ, Guilfoyle TJ (1984) Auxin regulated gene expression in intact soybean hypocotyl and excised hypocotyl sections. Planta 16:147–153. https://doi.org/10.1007/BF00410211
Jain M, Tyagi AK, Khurana JP (2006) Genome-wide analysis, evolutionary expansion, and expression of early auxin-responsive SAUR gene family in rice (Oryza sativa). Genomics 88:360–371. https://doi.org/10.1016/j.ygeno.2006.04.008
Kathare KP, Dharmasiri S, Dharmasiri N (2018) SAUR53 regulates organ elongation and apical hook development in Arabidopsis. Plant Signal Behav 13(10):e1514896. https://doi.org/10.1080/15592324.2018.1514896
Knauss S, Rohrmeier T, Lehle L (2003) The auxin-induced maize gene ZmSAUR2 encodes a short-lived nuclear protein expressed in elongating tissues. J Biol Chem 278:23936–23943. https://doi.org/10.1074/jbc.M212585200
Kong Y, Zhu Y, Gao C, She W, Lin W, Chen Y, Han N, Bian H, Zhu M, Wang J (2013) Tissue-specific expression of SMALL AUXIN UP RNA41 differentially regulates cell expansion and root meristem patterning in Arabidopsis. Plant Cell Physiol 54:609–621. https://doi.org/10.1093/pcp/pct028
Lim PO, Lee IC, Kim J, Kim HJ, Ryu JS, Woo HR, Hong GH (2010) Auxin response factor 2 (ARF2) plays a major role in regulating auxin-mediated leaf longevity. J Exp Bot 61(5):1419–1430. https://doi.org/10.1093/jxb/erq010
Markakis MN, Boron AK, Van Loock B, Saini K, Cirera S, Verbelen JP, Vissenberg K (2013) Characterization of a small auxin-up RNA (SAUR)-like gene involved in Arabidopsis thaliana development. PLoS ONE 8:e82596. https://doi.org/10.1371/journal.pone.0082596
Mascarenhas D, Mettler IJ, Pierce D, Lowe HW (1990) Intron-mediated enhancement of heterologous gene expression in maize. Plant Mol Biol 15:913–920
McClure BA, Guilfoyle T (1987) Characterization of a class of small auxin-inducible soybean polyadenylated RNAs. Plant Mol Biol 9:611–623. https://doi.org/10.1007/BF00020537
Nemhauser JL, Mockler TC, Chory J (2004) Interdependency of brassinosteroid and auxin signaling in Arabidopsis. PLoS Biol 2:e258. https://doi.org/10.1371/journal.pbio.0020258
Oh E, Zhu JY Wang ZY (2012) Interaction between BZR1 and PIF4 integrates brassinosteroid and environmental responses. Nat. Cell Biol 14, 802–809. http://dx.doi.org/https://doi.org/10.1038/ncb2545
Paponov IA, Paponov M, Teale W, Menges M, Chakrabortee S, Murray JA, Palme K (2008) Comprehensive transcriptome analysis of auxin responses in Arabidopsis. Mol Plant 1:321–337. https://doi.org/10.1093/mp/ssm021
Parra G, Bradna K, Rose AB, Korf I (2011) Comparative and functional analysis of intron-mediated enhancement signals reveals conserved features among plants. Nucleic Acids Res 39(13):5328–5337. https://doi.org/10.1093/nar/gkr043
Ren H, Gray MW (2015) SAUR proteins as effectors of hormonal and environmental signals in plant growth. Mol Plant 8(8):1153–1164. https://doi.org/10.1016/j.molp.2015.05.003
Ren H, Park MY, Spartz AK, Wong JH, Gray WM, Muday GK (2018) A subset of plasma membrane-localized PP2C.D phosphatases negatively regulate SAUR-mediated cell expansion in Arabidopsis. PLoS Genet 2:1–27. https://doi.org/10.1371/journal.pgen.1007455
Shin JH, Mila I, Liu M, Rodrigues MA, Vernoux T, Pirrello J, Bouzayen M (2019) The RIN-regulated small Auxin-up RNA SAUR69 is involved in the unripe-to-ripe phase transition of tomato fruit via enhancement of the sensitivity to ethylene. New Phytol 222(2):820–836
Spartz AK, Lee SH, Wenger JP, Gonzalez N, Itoh H, Inzé D, Peer WA, Murphy AS, Overvoorde PJ, Gray WM (2012) The SAUR19 subfamily of SMALL AUXIN UP RNA genes promote cell expansion. Plant J 70:978–990. https://doi.org/10.1111/j.1365-313X.2012.04946.x
Stamm P, Kumar PP (2013) Auxin and gibberellin responsive Arabidopsis SMALL AUXIN UP RNA36 regulates hypocotyl elongation in the light. Plant Cell Rep 32:759–769. https://doi.org/10.1007/s00299-013-1406-5
Taniguchi M, Nakamura M, Tasaka M, Morita M (2014) Identification of gravitropic response indicator genes in Arabidopsis inflorescence stems. Plant Signal Behav 9:e29570. https://doi.org/10.4161/psb.29570
Tian YY, Bai SLG, Dang ZH, Hao JF, Zhang J, Hasi A (2019) Genome-wide identification and characterization of long non-coding RNAs involved in fruit ripening and the climacteric in Cucumis melo. BMC Plant Biol 19:369. https://doi.org/10.1186/s12870-019-1942-4
Vert G, Chory J (2011) Crosstalk in cellular signaling: Background noise or the real thing? Dev. Cell 21:985–991. https://doi.org/10.1016/j.devcel.2011.11.006
Wang SK, Bai YH, Shen CJ, Wu YR, Zhang SN, Jiang DA, Guilfoyle TJ, Chen M, Qi YH (2010) Auxin-related gene families in abiotic stress response in Sorghum bicolor. Funct Integr Genomics 10:533–546. https://doi.org/10.1007/s10142-010-0174-3
Wang ZY, Bai MY, Oh E, Zhu JY (2012) Brassinosteroid signaling network and regulation of photomorphogenesis. Annu Rev Genet 46:701–724. https://doi.org/10.1146/annurev-genet-102209-163450
Wang HF, Shang QM (2019) Genome-wide Identification and Expression Analysis of the SAUR Gene Family in Cucumis sativus. Acta Horticulturae Sinica 46(6): 1093–1111 (in Chinese). https://doi.org/10.16420/j.issn.0513-353x.2018-0873
Wu J, Liu S, He Y, Guan X, Zhu X, Cheng L, Wang J, Lu G (2012) Genome-wide analysis of SAUR gene family in Solanaceae species. Gene 509:38–50. https://doi.org/10.1016/j.gene.2012.08.002
Wu B, Wang L, Pan GY, Li T, Li X, Hao JH (2020) Genome-wide characterization and expression analysis of the auxin response factor (ARF) gene family during melon (Cucumis melo L.) fruit development. Protoplasma 257:979–992. https://doi.org/10.1007/s00709-020-01484-2
Xie DS, Xu YC, Wang JP, Liu WR, Zhou Q, Luo SB, Huang W, He XM, Li Q, Peng QW, Yang XY, Yuan JQ, Yu JG, Wang XY, Lucas WJ, Huang SW, Jiang B, Zhang ZH (2019) The wax gourd genomes offer insights into the genetic diversity and ancestral cucurbit karyotype. Nat Commun 10:5158. https://doi.org/10.1038/s41467-019-13185-3
Zhang H, Yu ZJ, Yao XD, Chen JL, Chen X, Zhou HW, Lou YX, Ming F, Jin Y (2021) Genome-wide identification and characterization of small auxin-up RNA (SAUR) gene family in plants: evolution and expression profiles during normal growth and stress response. BMC Plant Biol 21(1):4. https://doi.org/10.1186/s12870-020-02781-x
Zhang N, Huang X, Bao YN, Wang B, Zeng HX, Cheng WS, Tang M, Li YH, Ren J, Sun YH (2017) Genome-wide identification of SAUR genes in watermelon (Citrullus lanatus). Physiology and molecular biology of plants 23(3). http://dx.doi.org/https://doi.org/10.1007/s12298-017-0442-y
Zhou J, Wen ZW, Mei YY (2018) The mechanism of SAUR72 in the regulation of leaf senescence in Arabidopsis. Chinese Journal of Plant Physiology 54(3): 379–385 (in Chinese). http://dx.doi.org/https://doi.org/10.13592/j.cnki.ppj.2018.1001
Acknowledgements
This work was supported by the National Natural Science Foundation of China (31860563), Natural Science Foundation of Inner Mongolia Autonomous Region (2021BS03002), and the Inner Mongolia University High-Level Talent Research Program (10000-21311201/056).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by Fuminori Takahashi.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Tian, Z., Han, J., Che, G. et al. Genome-wide characterization and expression analysis of SAUR gene family in Melon (Cucumis melo L.). Planta 255, 123 (2022). https://doi.org/10.1007/s00425-022-03908-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00425-022-03908-0