Abstract
Main conclusion 30 expansin genes were identified in the jujube genome. Phylogenetic analysis classified expansins into 17 subgroups. Closely related expansins share a conserved gene structure. ZjEXPs had different expression patterns in different tissues.
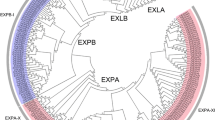
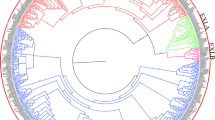
Plant-specific expansins were first discovered as pH-dependent cell-wall-loosening proteins involved in diverse physiological processes. No comprehensive analysis of the expansin gene family has yet been carried out at the whole genome level in Chinese jujube (Ziziphus jujuba Mill.). In this study, 30 expansin genes were identified in the jujube genome. These genes, which were distributed with varying densities across 10 of the 12 jujube chromosomes, could be divided into four subfamilies: 19 ZjEXPAs, 3 ZjEXPBs, 1 ZjEXLA, and 7 ZjEXLBs. Phylogenetic analysis of expansin genes in Arabidopsis, rice, apple, grape, and jujube classified these genes into 17 subgroups. Members of the same subfamily and subgroup shared conserved gene structure and motif compositions. Homology analysis identified 20 homologous gene pairs between jujube and Arabidopsis. Further analysis of ZjEXP gene promoter regions uncovered various growth, development and stress-responsive cis-acting elements. Expression analysis and transcript profiling revealed that ZjEXPs had different expression patterns in different tissues at various developmental stages. ZjEXPA4 and ZjEXPA6 were highly expressed in young fruits, ZjEXPA3 and ZjEXPA5 were significantly expressed in flowers, and ZjEXPA7 was specifically expressed in young leaves. The results of this study, the first systematic analysis of the jujube expansin gene family, can serve as a strong foundation for further elucidation of the physiological functions and biological roles of jujube expansin genes.








Similar content being viewed by others
Abbreviations
- EXLA(B):
-
Expansin-like A(B)
- EXPA(B):
-
α-Expansin (ß-Expansin)
References
Abuqamar S, Ajeb S, Sham A, Enan MR, Iratni R (2013) A mutation in the expansin-like A2 gene enhances resistance to necrotrophic fungi and hypersensitivity to abiotic stress in Arabidopsis thaliana. Mol Plant Pathol 14(8):813–827
Artimo P, Jonnalagedda M, Arnold K, Baratin D, Csardi G, de Castro E, Duvaud S, Flegel V, Fortier A, Gasteiger E, Grosdidier A, Hernandez C, Ioannidis V, Kuznetsov D, Liechti R, Moretti S, Mostaguir K, Redaschi N, Rossier G, Xenarios I, Stockinger H (2012) ExPASy: SIB bioinformatics resource portal. Nucleic Acids Res 40:W597–W603
Bailey TL, Boden M, Buske FA, Frith M, Grant CE, Clementi L, Ren J, Li WW, Noble WS (2009) MEME SUITE: tools for motif discovery and searching. Nucleic Acids Res 37:W202–W208
Boron AK, Van Loock B, Suslov D, Markakis MN, Verbelen JP, Vissenberg K (2015) Over-expression of AtEXLA2 alters etiolated arabidopsis hypocotyl growth. Ann Bot 115(1):67–80
Bu J, Zhao J, Liu M (2016) Expression stabilities of candidate reference genes for RT-qPCR in Chinese jujube (Ziziphus jujuba Mill.) under a variety of conditions. PLoS One 11(4):e0154212
Che J, Yamaji N, Shen RF, Ma JF (2016) An Al-inducible expansin gene, OsEXPA10 is involved in root cell elongation of rice. Plant J 88(1):132–142
Chen Y, Han Y, Zhang M, Zhou S, Kong X, Wang W (2016) Overexpression of the wheat expansin gene TaEXPA2 improved seed production and drought tolerance in transgenic tobacco plants. PLoS One 11(4):e0153494
Cho HT, Cosgrove DJ (2000) Altered expression of expansin modulates leaf growth and pedicel abscission in Arabidopsis thaliana. Proc Natl Acad Sci USA 97(17):9783–9788
Cho HT, Cosgrove DJ (2002) Regulation of root hair initiation and expansin gene expression in Arabidopsis. Plant Cell 14(2):3237–3253
Cosgrove DJ (2000) Loosening of plant cell walls by expansins. Nature 407(6802):321–326
Cosgrove DJ (2005) Growth of the plant cell wall. Nat Rev Mol Cell Biol 6(11):850–861
Cosgrove DJ (2015) Plant expansins: diversity and interactions with plant cell walls. Curr Opin Plant Biol 25:162–172
Cosgrove DJ, Bedinger P, Durachko DM (1997) Group I allergens of grass pollen as cell wall-loosening agents. Proc Natl Acad Sci USA 94(10):6559–6564
Dal Santo S, Vannozzi A, Tornielli GB, Fasoli M, Venturini L, Pezzotti M, Zenoni S (2013) Genome-wide analysis of the expansin gene superfamily reveals grapevine-specific structural and functional characteristics. PLoS One 8(4):e62206
Ding A, Marowa P, Kong Y (2016) Genome-wide identification of the expansin gene family in tobacco (Nicotiana tabacum). Mol Genet Genomics 291(5):1891–1907
Finn RD, Attwood TK, Babbitt PC, Bateman A, Bork P, Bridge AJ, Chang HY, Dosztányi Z, El-Gebali S, Fraser M, Gough J, Haft D, Holliday GL, Huang H, Huang X, Letunic I, Lopez R, Lu S, Marchler-Bauer A, Mi H, Mistry J, Natale DA, Necci M, Nuka G, Orengo CA, Park Y, Pesseat S, Piovesan D, Potter SC, Rawlings ND, Redaschi N, Richardson L, Rivoire C, Sangrador-Vegas A, Sigrist C, Sillitoe I, Smithers B, Squizzato S, Sutton G, Thanki N, Thomas PD, Tosatto SC, Wu CH, Xenarios I, Yeh LS, Young SY, Mitchell AL (2017) InterPro in 2017—beyond protein family and domain annotations. Nucleic Acids Res 45(D1):D190–D199
Gao QH, Wu CS, Wang M (2013) The jujube (Ziziphus jujuba Mill.) fruit: a review of current knowledge of fruit composition and health benefits. J Agric Food Chem 61(14):3351–3363
Georgelis N, Nikolaidis N, Cosgrove DJ (2015) Bacterial expansins and related proteins from the world of microbes. Appl Microbiol Biotechnol 99(9):3807–3823
Guimaraes LA, Mota APZ, Araujo ACG, de Alencar Figueiredo LF, Pereira BM, de Passos Saraiva MA, Silva RB, Danchin EGJ, Guimaraes PM, Brasileiro ACM (2017) Genome-wide analysis of expansin superfamily in wild Arachis discloses a stress-responsive expansin-like B gene. Plant Mol Biol 94(1–2):79–96
Hu B, Jin JP, Guo AY, Zhang H, Luo JH, Gao G (2015) GSDS 2.0: an upgraded gene feature visualization server. Bioinformatics 31(8):1296–1297
Huang J, Zhang C, Zhao X, Fei Z, Wan K, Zhang Z, Pang X, Yin X, Bai Y, Sun X, Gao L, Li R, Zhang J, Li X (2016) The jujube genome provides insights into genome evolution and the domestication of sweetness/acidity taste in fruit trees. PLoS Genet 12(12):e1006433
Kende H, Bradford K, Brummell D, Cho HT, Cosgrove D, Fleming A, Gehring C, Lee Y, McQueen-Mason S, Rose J, Voesenek LA (2004) Nomenclature for members of the expansin superfamily of genes and proteins. Plant Mol Biol 55(3):311–314
Krzywinski M, Schein J, Birol I, Connors J, Gascoyne R, Horsman D, Jones SJ, Marra MA (2009) Circos: an information aesthetic for comparative genomics. Genome Res 19(9):1639–1645
Kuluev BR, Knyazev AB, Lebedev YP, Chemeris AV (2012) Morphological and physiological characteristics of transgenic tobacco plants expressing expansin genes: AtEXP10 from Arabidopsis and PnEXPA1 from poplar. Russ J Plant Physiol 59(1):97–104
Kuluev BR, Knyazev AV, Mikhaylova EV, Chemeris AV (2017) The role of expansin genes PtrEXPA3, and PnEXPA3, in the regulation of leaf growth in poplar. Russ J Plant Physiol 53(6):651–660
Kwon YR, Lee HJ, Kim KH, Hong SW, Lee SJ, Lee H (2008) Ectopic expression of Expansin3 or Expansinβ1 causes enhanced hormone and salt stress sensitivity in Arabidopsis. Biotechnol Lett 30(7):1281–1288
Lee Y, Choi DS, Kende H (2001) Expansins: ever-expanding numbers and functions. Curr Opin Plant Biol 4(6):527–532
Lescot M, Déhais P, Thijs G, Marchal K, Moreau Y, Van de Peer Y, Rouzé P, Rombauts S (2002) PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res 30(1):325–327
Li B, Dewey CN (2011) RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinformatics 12:323
Li L, Stoeckert CJ, Roos DS (2003) OrthoMCL: identification of ortholog groups for eukaryotic genomes. Genome Res 13(9):2178–2189
Li N, Pu Y, Gong Y, Yu Y, Ding H (2016) Genomic location and expression analysis of expansin gene family reveals the evolutionary and functional significance in Triticum aestivum. Genes Genom 38(11):1021–1030
Lin C, Choi HS, Cho HT (2011) Root hair-specific EXPANSIN A7 is required for root hair elongation in Arabidopsis. Mol Cells 31(4):393–397
Liu MJ (2003) Genetic diversity of Chinese jujube (Ziziphus jujuba Mill.). Acta Hort 623(623):351–355
Liu MJ (2006) Chinese jujube: botany and horticulture. Hortic Rev 32:229–298
Liu M, Wang M (2009) Chinese jujube germsplasm resources. Forestry Publishing House, Beijing
Liu M, Zhao J, Cai Q, Liu G, Wang J, Zhao Z, Liu P, Dai L, Yan G, Wang W, Li X, Chen Y, Sun Y, Liu Z, Lin M, Xiao J, Chen Y, Li X, Wu B, Ma Y, Jian J, Yang W, Yuan Z, Sun X, Wei Y, Yu L, Zhang C, Liao S, He R, Guang X, Wang Z, Zhang Y, Luo L (2014) The complex jujube genome provides insights into fruit tree biology. Nat Commun 5:5315
Liu H, Li H, Zhang H, Li J, Xie B, Xu J (2016) The expansin gene PttEXPA8 from poplar (Populus tomentosa) confers heat resistance in transgenic tobacco. Plant Tiss Org Cult 126(2):353–359
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using realtime quantitative PCR and the 2 −ΔΔCT method. Methods 25(4):402–408
Lu ZM, Liu K, Yan ZX, Li XG (2010) Research status of nutrient component and health functions of Ziziphus jujuba Mill. Acta Hortic Sin 37(12):2017–2024
Lu YG, Liu LF, Wang X, Han ZH, Ouyang B, Zhang JH, Li HX (2016) Genome-wide identification and expression analysis of the expansin gene family in tomato. Mol Genet Genomics 291(2):597–608
Palapol Y, Kunyamee S, Thongkhum M, Ketsa S, Ferguson IB, van Doorn WG (2015) Expression of expansin genes in the pulp and the dehiscence zone of ripening durian (Durio zibethinus) fruit. J Plant Physiol 182:33–39
Park CH, Kim TW, Son SH, Hwang JY, Lee SC, Chang SC, Kim SH, Kim SW, Kim SK (2011) Brassinosteroids control AtEXPA5 gene expression in Arabidopsis thaliana. Phytochemistry 71(4):380–387
Perini MA, Sin IN, Villarreal NM, Marina M, Powell AL, Martínez GA, Civello PM (2017) Overexpression of the carbohydrate binding module from Solanum lycopersicum expansin 1 (Sl-EXP1) modifies tomato fruit firmness and Botrytis cinerea susceptibility. Plant Physiol Biochem 113:122–132
Saito T, Tuan PA, Katsumi-Horigane A, Bai S, Ito A, Sekiyama Y, Ono H, Moriguchi T (2015) Development of flower buds in the Japanese pear (Pyrus pyrifolia) from late autumn to early spring. Tree Physiol 35(6):653–662
Sampedro J, Cosgrove DJ (2005) The expansin superfamily. Genome Biol 6(12):242. https://doi.org/10.1186/gb-2005-6-12-242
Sarkar P, Bosneaga E, Auer M (2009) Plant cell walls throughout evolution: towards a molecular understanding of their design principles. J Exp Bot 60(13):3615–3635
Song S, Zhou H, Sheng S, Cao M, Li Y, Pang X (2017) Genome-wide organization and expression profiling of the SBP-box gene family in Chinese jujube (Ziziphus jujuba Mill.). Int J Mol Sci 18(8):1734
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30(12):2725–2729
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specifc gap penalties and weight matrix choice. Nucleic Acids Res 22(22):4673–4680
Tovar-Herrera OE, Batista-Garcia RA, Sanchez-Carbente Mdel R, Iracheta-Cardenas MM, Arevalo-Nino K, Folch-Mallol JL (2015) A novel expansin protein from the white-rot fungus Schizophyllum commune. PLoS One 10(3):e0122296
Trapnell C, Roberts A, Goff L, Pertea G, Kim D, Kelley DR, Pimentel H, Salzberg SL, Rinn JL, Pachter L (2012) Differential gene and transcript expression analysis of RNA-Seq experiments with Hophat and Cufflinks. Nat Protoc 7(3):562–578
Tsuchiya M, Satoh S, Iwai H (2015) Distribution of XTH, expansin, and secondary-wall-related CesA in floral and fruit abscission zones during fruit development in tomato (Solanum lycopersicum). Front Plant Sci 6:323
Valdivia ER, Sampedro J, Lamb JC, Chopra S, Cosgrove DJ (2007) Recent proliferation and translocation of pollen group 1 allergen genes in the maize genome. Plant Physiol 143(3):1269–1281
Wakasa Y, Hatsuyama Y, Takahashi A, Sato T, Niizeki M, Harada T (2003) Divergent expression of six expansin genes during apple fruit ontogeny. Eur J Hort Sci 68(6):253–259
Yan A, Wu M, Yan L, Hu R, Ali I, Gan Y (2014) AtEXP2 is involved in seed germination and abiotic stress response in Arabidopsis. PLoS One 9(1):e85208
Yennawar NH, Li LC, Dudzinski DM, Tabuchi A, Cosgrove DJ (2006) Crystal structure and activities of EXPB1 (Zea m 1), a β-expansin and group-1 pollen allergen from maize. Proc Natl Acad Sci USA 103(40):14664–14671
Zhang S, Xu R, Gao Z, Chen C, Jiang Z, Shu H (2014a) A genome-wide analysis of the expansin genes in Malus × Domestica. Mol Genet Genom 289(2):225–236
Zhang W, Yan H, Chen W, Liu J, Jiang C, Jiang H, Zhu S (2014b) Genome-wide identification and characterization of maize expansin genes expressed in endosperm. Mol Genet Genomics 289(6):1061–1074
Zhu Y, Wu N, Song W, Yin G, Qin Y, Yan Y, Hu Y (2014) Soybean (Glycine max) expansin gene superfamily origins: segmental and tandem duplication events followed by divergent selection among subfamilies. BMC Plant Biol 14:93
Acknowledgements
This work was supported by grants from the Fundamental Research Funds for the Central Universities (2016ZCQ05) and Major Science and Technology Special Project of Xuchang, Henan province (20170112006).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Hou, L., Zhang, Z., Dou, S. et al. Genome-wide identification, characterization, and expression analysis of the expansin gene family in Chinese jujube (Ziziphus jujuba Mill.). Planta 249, 815–829 (2019). https://doi.org/10.1007/s00425-018-3020-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-018-3020-9