Abstract
Main Conclusion
14-3-3 isoforms were relatively less conserved at the C-terminal region across plant groups. Both Os 14-3-3f and Os 14-3-3g were inducible with differential gene expression levels under different abiotic stress and developmental stages in sensitive and tolerant indica rice cultivars as confirmed both at transcript and protein level.
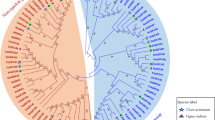
Plant 14-3-3s has been well characterized to function in several signaling pathways, biotic as well as abiotic stress and nutrient metabolism. We attempted comprehensive analysis of 14-3-3 genes in different plant lineages such as green algae (Chlamydomonas reinhardtii), moss (Physcomitrella patens) and lycophyte (Selaginella moellendorffii), dicot Arabidopsis thaliana and monocot Oryza sativa sub sp. japonica at the gene and protein level. Sequence alignment results revealed that 14-3-3 isoforms were evolutionarily conserved across all taxa with variable C-terminal end. Phylogenetic analysis indicated that the majority of 14-3-3 isoforms in rice belong to the non-epsilon group that clustered separately from the dicot group. Segmental duplication event played a significant role in the expansion of both, Arabidopsis and rice, 14-3-3 isoforms as revealed by synteny studies. In silico gene expression using Massive Parallel Signature Sequencing and microarray analysis revealed that 14-3-3 isoforms have variable expression in different tissue types and under different abiotic stress regime in Arabidopsis and japonica rice. Both, semi-quantitative and qPCR results, confirmed that Os14-3-3f and Os14-3-3g were inducible under abiotic stress in lamina and roots of indica rice and relatively higher under salinity and cold stress in Nonabokra, under dehydration stress in N-22 and under exogenous ABA in IR-29 usually after 3–6 h of treatment. Both, 14-3-3f and 14-3-3g, were highly expressed in flag leaves, stems and panicles and mature roots. These results were further confirmed by immunoblot analysis of rice cultivars using Os14-3-3f antibody generated from recombinant Os14-3-3f protein. The results provide the first comprehensive report of Os14-3-3 gene expression in indica rice cultivars which differ in tolerance to abiotic stress that might be useful for further research.














Similar content being viewed by others
Abbreviations
- DAP:
-
Days after pollination
- MPSS:
-
Massive parallel signature sequences
- TPM:
-
Transcript per million
References
Bihn EA, Paul A-L, Wang SW et al (1997) Localization of 14-3-3 proteins in the nuclei of Arabidopsis and maize. Plant J 12:1439–1445. doi:10.1046/j.1365-313x.1997.12061439.x
Blanc G, Wolfe KH (2004) Widespread paleopolyploidy in model plant species inferred from age distributions of duplicate genes. Plant Cell 16:1667–1678. doi:10.1105/tpc.021345
Brenner S, Johnson M, Bridgham J et al (2000) Gene expression analysis by massively parallel signature sequencing (MPSS) on microbead arrays. Nat Biotech 18:630–634
Campo S, Peris-Peris C, Montesinos L et al (2011) Expression of the maize ZmGF14-6 gene in rice confers tolerance to drought stress while enhancing susceptibility to pathogen infection. J Exp Bot 63:983–999
Cao H, Xu Y, Yuan L et al (2016) Molecular characterization of the 14-3-3 gene family in Brachypodium distachyon L. reveals high evolutionary conservation and diverse responses to abiotic stresses. Front Plant Sci 7:1099. doi:10.3389/fpls.2016.01099
Chandler P, Robertson M (1994) Gene expression regulated by abscisic acid and its relation to stress tolerance. Annu Rev Plant Physiol Plant Mol Biol 45:113–141. doi:10.1146/annurev.pp.45.060194.000553
Chen F, Li Q, Sun L, He Z (2006) The rice 14-3-3 gene family and its involvement in responses to biotic and abiotic stress. DNA Res 13:53–63
Daugherty CJ, Rooney MF, Miller PW, Ferl RJ (1996) Molecular organization and tissue-specific expression of an Arabidopsis 14-3-3 gene. Plant Cell 8:1239–1248
Debernardi JM, Rodriguez RE, Mecchia MA, Palatnik JF (2012) Functional specialization of the plant miR396 regulatory network through distinct microRNA–target interactions. PLoS Genet 8:e1002419
Degenkolbe T, Do PT, Zuther E et al (2009) Expression profiling of rice cultivars differing in their tolerance to long-term drought stress. Plant Mol Biol 69:133–153. doi:10.1007/s11103-008-9412-7
DeLille JM, Sehnke PC, Ferl RJ (2001) The Arabidopsis 14-3-3 family of signaling regulators. Plant Physiol 126:35–38. doi:10.1104/pp.126.1.35
Denison FC, Paul A-L, Zupanska AK, Ferl RJ (2011) 14-3-3 proteins in plant physiology. Semin Cell Dev Biol 22:720–727. doi:10.1016/j.semcdb.2011.08.006
Eisen MB, Spellman PT, Brown PO, Botstein D (1998) Cluster analysis and display of genome-wide expression patterns. Proc Natl Acad Sci USA 95:14863–14868
Ferl RJ (2004) 14-3-3 proteins: regulation of signal-induced events. Physiol Plant 120:173–178. doi:10.1111/j.0031-9317.2004.0239.x
Ferl RJ, Manak MS, Reyes MF (2002) The 14-3-3s. Genome Biol 3:reviews3010.1–reviews3010.7
Fu H, Subramanian RR, Masters SC (2000) 14-3-3 Proteins: structure, function, and regulation. Annu Rev Pharmacol Toxicol 40:617–647. doi:10.1146/annurev.pharmtox.40.1.617
Fukuda M, Asano S, Nakamura T et al (1997) CRM1 is responsible for intracellular transport mediated by the nuclear export signal. Nature 390:308–311
Ganguly M, Datta K, Roychoudhury A et al (2012) Overexpression of Rab16A gene in indica rice variety for generating enhanced salt tolerance. Plant Signal Behav 7:502–509. doi:10.4161/psb.19646
Gaut BS, Morton BR, McCaig BC, Clegg MT (1996) Substitution rate comparisons between grasses and palms: synonymous rate differences at the nuclear gene Adh parallel rate differences at the plastid gene rbcL. Proc Natl Acad Sci USA 93:10274–10279
He Y, Zhang Y, Chen L et al (2017) A member of the 14-3-3 gene family in Brachypodium distachyon, BdGF14d, confers salt tolerance in transgenic tobacco plants. Front Plant Sci 8:340
Ho S-L, Huang L-F, Lu C-A et al (2013) Sugar starvation-and GA-inducible calcium-dependent protein kinase 1 feedback regulates GA biosynthesis and activates a 14-3-3 protein to confer drought tolerance in rice seedlings. Plant Mol Biol 81:347–361
Jaspert N, Throm C, Oecking C (2011) Arabidopsis 14-3-3 proteins: fascinating and less fascinating aspects. Front Plant Sci 2:96
Kelley LA, Mezulis S, Yates CM et al (2015) The Phyre2 web portal for protein modeling, prediction and analysis. Nat Protoc 10:845–858
Krzywinski M, Schein J, Birol İ et al (2009) Circos: an information aesthetic for comparative genomics. Genome Res 19:1639–1645. doi:10.1101/gr.092759.109
Li R, Jiang X, Jin D et al (2015) Identification of 14-3-3 family in common bean and their response to abiotic stress. PLoS ONE 10:e0143280
Liu D, Bienkowska J, Petosa C et al (1995) Crystal structure of the zeta isoform of the 14-3-3 protein. Nature 376:191–194
Liu Q, Zhang S, Liu B (2016) 14-3-3 proteins: macro-regulators with great potential for improving abiotic stress tolerance in plants. Biochem Biophys Res Commun 477:9–13. doi:10.1016/j.bbrc.2016.05.120
Lovell SC, Davis IW, Arendall III WB et al (2003) Structure validation by Cα geometry: ϕ,ψ and Cβ deviation. Proteins Str Func Bioinfo 50(3):437–450. doi:10.1002/prot.10286
Muthuramalingam P, Krishnan SR, Pothiraj R, Ramesh M (2017) Global transcriptome analysis of combined abiotic stress signaling genes unravels key players in Oryza sativa L.: an in silico approach. Front Plant Sci 8:759
Nakano M, Nobuta K, Vemaraju K et al (2006) Plant MPSS databases: signature-based transcriptional resources for analyses of mRNA and small RNA. Nucleic Acids Res 34:D731–D735. doi:10.1093/nar/gkj077
Nakashima K, Yamaguchi-Shinozaki K (2013) ABA signaling in stress-response and seed development. Plant Cell Rep 32:959–970. doi:10.1007/s00299-013-1418-1
Paul A-L, Liu L, McClung S et al (2009) Comparative interactomics: analysis of Arabidopsis 14-3-3 complexes reveals highly conserved 14-3-3 interactions between humans and plants. J Proteome Res 8:1913–1924. doi:10.1021/pr8008644
Purwestri YA, Ogaki Y, Tamaki S et al (2009) The 14-3-3 protein GF14c acts as a negative regulator of flowering in rice by interacting with the florigen Hd3a. Plant Cell Physiol 50:429–438
Raghavendra AS, Gonugunta VK, Christmann A, Grill E (2017) ABA perception and signalling. Trends Plant Sci 15:395–401. doi:10.1016/j.tplants.2010.04.006
Reddy INBL, Kim B-K, Yoon I-S et al (2017) Salt tolerance in rice: focus on mechanisms and approaches. Rice Sci 24:123–144. doi:10.1016/j.rsci.2016.09.004
Roberts MR (2017) 14-3-3 Proteins find new partners in plant cell signalling. Trends Plant Sci 8:218–223. doi:10.1016/S1360-1385(03)00056-6
Roberts MR, Salinas J, Collinge DB (2002) 14-3-3 proteins and the response to abiotic and biotic stress. Plant Mol Biol 50:1031–1039. doi:10.1023/A:1021261614491
Rosenquist M, Alsterfjord M, Larsson C, Sommarin M (2001) Data mining the Arabidopsis genome reveals fifteen 14-3-3 genes expression is demonstrated for two out of five novel genes. Plant Physiol 127:142–149. doi:10.1104/pp.127.1.142
Schmid M, Davison TS, Henz SR et al (2005) A gene expression map of Arabidopsis thaliana development. Nat Genet 37:501–506
Schoonheim PJ, Sinnige MP, Casaretto JA et al (2007) 14-3-3 adaptor proteins are intermediates in ABA signal transduction during barley seed germination. Plant J 49:289–301. doi:10.1111/j.1365-313X.2006.02955.x
Sehnke PC, Ferl RJ (2000) Plant 14-3-3s: omnipotent metabolic phosphopartners? Sci STKE 2000(56):pe1
Sehnke PC, Henry R, Cline K, Ferl RJ (2000) Interaction of a plant 14-3-3 protein with the signal peptide of a thylakoid-targeted chloroplast precursor protein and the presence of 14-3-3 isoforms in the chloroplast stroma. Plant Physiol 122:235–242
Shankar R, Bhattacharjee A, Jain M (2016) Transcriptome analysis in different rice cultivars provides novel insights into desiccation and salinity stress responses. Sci Rep 6:23719. doi:10.1038/srep23719
Shanko AV, Mesenko MM, Klychnikov OI et al (2003) Proton pumping in growing part of maize root: its correlation with 14-3-3 protein content and changes in response to osmotic stress. Biochemistry 68:1320–1326
Shin R, Alvarez S, Burch AY et al (2007) Phosphoproteomic identification of targets of the Arabidopsis sucrose nonfermenting-like kinase SnRK2.8 reveals a connection to metabolic processes. Proc Natl Acad Sci USA 104:6460–6465. doi:10.1073/pnas.0610208104
Shin R, Jez JM, Basra A et al (2011) 14-3-3 proteins fine-tune plant nutrient metabolism. FEBS Lett 585:143–147. doi:10.1016/j.febslet.2010.11.025
Sirichandra C, Davanture M, Turk BE et al (2010) The Arabidopsis ABA-activated kinase OST1 phosphorylates the bZIP transcription factor ABF3 and creates a 14-3-3 binding site involved in its turnover. PLoS ONE 5:e13935
Sun G, Xie F, Zhang B (2011) Transcriptome-wide identification and stress properties of the 14-3-3 gene family in cotton (Gossypium hirsutum L.). Funct Integr Genomics 11:627–636. doi:10.1007/s10142-011-0242-3
Sun X, Luo X, Sun M et al (2013) A Glycine soja 14-3-3 protein GsGF14o participates in stomatal and root hair development and drought tolerance in Arabidopsis thaliana. Plant Cell Physiol 55:99–118
Sun X, Sun M, Jia B et al (2016) A 14-3-3 family protein from wild soybean (Glycine Soja) regulates ABA sensitivity in Arabidopsis. PLoS ONE 10:e0146163
Takahashi Y, Kinoshita T, Shimazaki K (2007) Protein phosphorylation and binding of a 14-3-3 protein in Vicia guard cells in response to ABA. Plant Cell Physiol 48:1182–1191
Tamura K, Peterson D, Peterson N et al (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739. doi:10.1093/molbev/msr121
Taoka K, Ohki I, Tsuji H et al (2011) 14-3-3 proteins act as intracellular receptors for rice Hd3a florigen. Nature 476:332–335
Tian F, Wang T, Xie Y et al (2015) Genome-wide identification, classification, and expression analysis of 14-3-3 gene family in Populus. PLoS ONE 10:e0123225
Walia H, Wilson C, Condamine P et al (2005) Comparative transcriptional profiling of two contrasting rice genotypes under salinity stress during the vegetative growth stage. Plant Physiol 139:822–835. doi:10.1104/pp.105.065961
Wang X, Yang P, Zhang X et al (2009) Proteomic analysis of the cold stress response in the moss, Physcomitrella patens. Proteomics 9:4529–4538
Witkos TM, Koscianska E, Krzyzosiak WJ (2011) Practical aspects of microRNA target prediction. Curr Mol Med 11:93–109. doi:10.2174/156652411794859250
Wu K, Rooney MF, Ferl RJ (1997) The Arabidopsis 14-3-3 multigene family. Plant Physiol 114:1421–1431
Xu WF, Shi WM (2006) Expression profiling of the 14-3-3 gene family in response to salt stress and potassium and iron deficiencies in young tomato (Solanum lycopersicum) roots: analysis by real-time RT–PCR. Ann Bot 98:965–974
Xu W, Jia L, Shi W et al (2012) Smart role of plant 14-3-3 proteins in response to phosphate deficiency. Plant Signal Behav 7:1047–1048. doi:10.4161/psb.20997
Xu W, Jia L, Shi W et al (2013) The tomato 14-3-3 protein TFT4 modulates H+ efflux, basipetal auxin transport, and the PKS5-J3 pathway in the root growth response to alkaline stress. Plant Physiol 163:1817–1828
Yamaguchi-Shinozaki K, Shinozaki K (2006) Transcriptional regulatory networks in cellular responses and tolerance to dehydration and cold stresses. Annu Rev Plant Biol 57:781–803. doi:10.1146/annurev.arplant.57.032905.105444
Yan J, He C, Wang J et al (2004) Overexpression of the Arabidopsis 14-3-3 protein GF14λ in cotton leads to a “stay-green” phenotype and improves stress tolerance under moderate drought conditions. Plant Cell Physiol 45:1007–1014
Yang Z-P, Li H-L, Guo D et al (2014) Identification and characterization of the 14-3-3 gene family in Hevea brasiliensis. Plant Physiol Biochem 80:121–127. doi:10.1016/j.plaphy.2014.03.034
Yang L, You J, Wang Y et al (2017) Systematic analysis of the G-box factor 14-3-3 gene family and functional characterization of GF14a in Brachypodium distachyon. Plant Physiol Biochem 117:1–11. doi:10.1016/j.plaphy.2017.05.013
Yao Y, Du Y, Jiang L, Liu J-Y (2007) Molecular analysis and expression patterns of the 14-3-3 gene family from Oryza Sativa. J Biochem Mol Biol 40(3):349–357
Zeng L, Shannon MC, Lesch SM (2001) Timing of salinity stress affects rice growth and yield components. Agric Water Manag 48:191–206. doi:10.1016/S0378-3774(00)00146-3
Zhang Z, Li J, Zhao X-Q et al (2006) KaKs_Calculator: calculating Ka and Ks through model selection and model averaging. Genomics Proteomics Bioinform 4:259–263. doi:10.1016/S1672-0229(07)60007-2
Zhang Z, Yu J, Li D et al (2010) PMRD: plant microRNA database. Nucleic Acids Res 38:D806–D813. doi:10.1093/nar/gkp818
Acknowledgements
We gratefully acknowledge the Director, Bose Institute, for providing fellowship to NY and for infrastructural support. We are also thankful to Mr. Jadab Ghosh, Mrs. Kaberi Ghosh and Mr. Mrinal Das, Bose Institute, for their technical help. We also thank Dr. Subarna Thakur (Bioinformatics Centre, Bose Institute) for helping in synteny map preparation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Ethical statement regarding use of plant material
The authors state that rice seeds, leaves and other planting materials were procured as per local/national regulations. All experimental materials were grown in a contained facility without causing any harm to the natural resource and biodiversity.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Yashvardhini, N., Bhattacharya, S., Chaudhuri, S. et al. Molecular characterization of the 14-3-3 gene family in rice and its expression studies under abiotic stress. Planta 247, 229–253 (2018). https://doi.org/10.1007/s00425-017-2779-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-017-2779-4