Abstract
Purpose
Whilst the treatment paradigm for colorectal cancer has evolved significantly over time, there is still a lack of reliable biomarkers of treatment response. Treatment decisions are based on high-risk features such as advanced TNM stage and histology. The role of the tumour microenvironment, which can influence tumour progression and treatment response, has generated considerable interest. Patient-derived explant cultures allow preservation of native tissue architecture and tumour microenvironment. The aim of the scoping review is to evaluate the utility of patient-derived explant cultures as a preclinical model in colorectal cancer.
Methods
A search was conducted using Ovid MEDLINE, EMBASE, Web of Science, and Cochrane databases from start of database records to September 1, 2022. We included all peer-reviewed human studies in English language which used patient-derived explants as a preclinical model in primary colorectal cancer. Eligible studies were grouped into the following categories: assessing model feasibility; exploring tumour microenvironment; assessing ex vivo drug responses; discovering and validating biomarkers.
Results
A total of 60 studies were eligible. Fourteen studies demonstrated feasibility of using patient-derived explants as a preclinical model. Ten studies explored the tumour microenvironment. Thirty-eight studies assessed ex vivo drug responses of chemotherapy agents and targeted therapies. Twenty-four studies identified potential biomarkers of treatment response.
Conclusions
Given the preservation of tumour microenvironment and tumour heterogeneity, patient-derived explants has the potential to identify reliable biomarkers, treatment resistance mechanisms, and novel therapeutic agents. Further validation studies are required to characterise, refine and standardise this preclinical model before it can become a part of precision medicine in colorectal cancer.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The management of colorectal cancer (CRC) has evolved over time but still involves a combination of surgery, chemotherapy and radiotherapy. More recently, targeted therapies and immunotherapies have become powerful additions to the therapeutic armoury against CRC, with the aim of moving us closer towards precision medicine. However, there are still issues that need to be addressed before it becomes a reality, such as identifying predictive biomarkers of treatment response beyond mutational status (e.g. MMR, RAF/BRAF) to allow patient stratification, understanding mechanisms for treatment response and resistance, and developing novel strategies for poor treatment responders.
For locally advanced rectal cancer, the current recommended treatment is neoadjuvant chemoradiotherapy (NACRT) followed by surgery. Although 10–20% of patients achieve pathological complete response, there remains a significant proportion of patients who will only partially respond or do not respond at all to current NACRT regimens [1]. Currently, all patients undergo the same treatment based on clinical and radiological staging. However, complete responders may benefit from the shift towards organ preservation and “watch and wait" strategy after NACRT to avoid overtreatment [2]. Conversely, poor responders may avoid toxicities and side effects of unnecessary treatment or benefit from alternative treatments such as total neoadjuvant therapy or immunotherapies [3,4,5]. In colon cancer, there is similar uncertainty in the use of adjuvant chemotherapy in patients with early (stage II) disease, with no greater than 5% benefit in 5-year disease-free and overall survival for most patients [6]. Treatment decisions have been based on high-risk features such as T4 primary, evidence of obstruction/perforation, perineural or lymphovascular invasion, and poorly differentiated histology, although the use of circulating tumour DNA has shown promising results in guiding adjuvant therapy [7].
Preclinical models such as cell lines have been used to study tumour cell biology and assess drug efficacy for many years but they lack modelling of the tumour microenvironment (TME). The role of TME, which includes immune cells, stromal cells, extracellular matrix and signalling molecules, has attracted considerable attention in recent times [8]. Indeed, the cellular composition and complex cell–cell interactions within the TME, not just tumour cell intrinsic factors, have a strong influence on tumour progression and treatment response. For example, higher levels of CD3 + and CD8 + tumour-infiltrating lymphocytes are associated with a favourable response to NACRT in rectal cancer [9], whilst regulatory T cells suppress anti-tumour immunity and are associated with poor response to NACRT [10]. In colon cancer, the introduction of the Immunoscore, an immunohistochemistry (IHC) and digital pathology-based assay which quantifies CD3 + and CD8 + T cells in the tumour core and invasive margin, has been shown to predict patients who will benefit from adjuvant chemotherapy [11]. In addition, the characteristics of the TME are altered by tumour cell metabolism that ultimately leads to chemoresistance and tumour progression. Certain conditions, such as hypoxia and acidity, have been shown to impair the function of immune effector cells and promote the accumulation of immunosuppressive cells and cytokines [12].
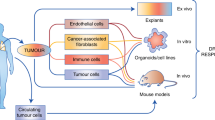
Therefore, a better understanding of the TME, through the use of patient-derived explants (PDEs), may be key to the discovery of more reliable biomarkers of treatment response, potential treatment resistance mechanisms, and novel treatment combinations. PDEs are obtained by cutting fresh human tumour specimens into smaller pieces and are cultured ex vivo over a period of time. The main advantage of PDEs is the preservation of native tissue architecture, TME, heterotypic cell–cell interactions and metabolic crosstalk, without excessive manipulation so they are more likely to mimic the in vivo situation [13]. In addition, PDEs are relatively easy to establish, cost-effective, and provide results in a timely manner, which may make PDEs a more robust preclinical model for translation into clinical practice. This scoping review aims to evaluate the utility of PDEs as a preclinical model in primary CRC to discover and validate potential biomarkers, explore the TME and assess drug responses in order to guide precision medicine.
Methods
Search strategy
This scoping review was designed and performed in accordance with the Preferred Reporting Items for Systematic Reviews and Meta-Analyses extension for Scoping Reviews (PRISMA-ScR) guidelines. A comprehensive search was conducted using the Ovid MEDLINE, EMBASE, Web of Science, and Cochrane Library databases from start of database records to September 1, 2022. The following terms and their variations were used either alone or in combination: “colorectal cancer” and “explants”. The search strategy is supplied in Supplementary Information. Pertinent and electronic links were hand-searched, and cross-referencing was performed for selected studies. The search results were pooled using the Covidence online platform.
Study selection
The study included only full-text English studies including primary CRC PDEs as a preclinical model. The study excluded reviews, commentaries, posters, conference abstracts/proceedings, and studies not performed with humans. Two reviewers (M.Mui and M.C.) performed the search and extracted data independently. Eligible studies were categorized into different groups determined by the use of PDEs for assessing feasibility, exploring TME, drug testing, and biomarker discovery and validation.
Results
The initial database search identified 1641 records, of which 203 studies were assessed for eligibility after eliminating duplicates and screening abstracts. A total of 60 studies were included in the final analysis. The PRISMA flow diagram is shown in Fig. 1. The publication years ranged from 1962 to 2022. We identified 14 studies that assessed the feasibility of using PDEs as a preclinical model for CRC, 10 studies that explored the TME, 38 studies that assessed ex vivo drug responses, and 24 studies that identified potential biomarkers of treatment response or clinical outcome.
CRC PDE culture techniques
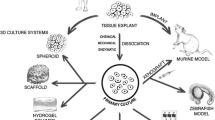
A variety of different culture methods and conditions were used to establish CRC PDEs as shown in Supplementary Table 1. All studies obtained tumour tissue from surgical specimens, except for one which collected biopsies endoscopically before surgery [14] and one which did not specify tissue source [15]. Tumour tissue was either processed by mechanical fragmentation or slicing using an instrument such as vibratome or tissue chopper. Subsequently, the explants were maintained either as free-floating cultures or placed on scaffolds such as gelatin or collagen sponges [14, 16,17,18,19,20,21,22,23,24,25,26,27,28,29,30,31,32,33,34,35], metal grids [36,37,38,39,40,41,42,43], or pore membranes [44,45,46,47,48], with or without agitation. Culture media formulations vary widely among studies but common base media include RPMI 1640 and Dulbecco's Modified Eagle Medium (DMEM), which are often supplemented with foetal bovine serum (FBS) and antibiotics. For prolonged incubation, the majority of studies used a temperature of 37°C, with 5% CO2 and 95% air, although a number of studies opted to use a higher O2 concentration [15, 37,38,39,40,41,42,43, 45, 47, 49, 50]. Culture duration ranged from 4 h to 122 days, depending on the aims of the study and assays performed.
Feasibility studies on CRC PDEs
We identified 14 studies that assessed the feasibility of using PDEs as a preclinical model for CRC (Table 1). Ten studies demonstrated preservation of tissue morphology in short-term cultures as assessed by haematoxylin and eosin (H&E) [36,37,38, 43, 44, 46, 51,52,53,54]. Five studies performed IHC to measure proliferation markers such as Ki-67 [44, 46, 53, 55] and bromodeoxyuridine (BrdU) [52] to confirm tissue viability, although one study reported a slight but significant decrease in proliferation index after 7 days [55]. Two studies analysed gene expression patterns and found that they were relatively stable over time and comparable to the original tumour [44, 56]. One study confirmed tissue viability by demonstrating the ability to incorporate uridine and thymidine during DNA synthesis using autoradiography [49]. One study investigated the feasibility of combining proteomics using reverse-phase protein microarrays with ex vivo cultures and demonstrated the sensitivity and robustness of the system [26]. A more recent study utilised fluorescence-based imaging to assess (i) cell viability, which remained high during culture and (ii) metabolic activity, which showed a 50% decrease in the first week but remained relatively stable during the remaining culture period, suggesting a period of adaptation ex vivo [54].
Exploring TME using CRC PDEs
We identified 10 studies that explored the TME using CRC PDEs (Table 2). Early studies investigated the role of enzymes such as plasminogen activator urokinase and gelatinase in CRC development and metastasis [57, 58]. The majority of subsequent studies focused on the role of different types and levels of cytokines (e.g. IL-6, IL-8, IL-10) or chemokines (e.g. CCL2, CXCL1, CXCL5, CCL5, CXCL10) in the TME that might affect cell function and migration [15, 59,60,61,62,63]. One study found that tumour-infiltrating T cells were able to survive in culture despite the absence of in vivo factors, suggesting that the TME may play a role [39]. A more recent study demonstrated that a high density of tumour-infiltrating lymphocytes with positive T-bet transcription factor (Tbet + TILs) was associated with higher interferon-gamma (IFN-γ) levels both at baseline and following programmed cell death protein 1 (PD-1) blockade [50].
Assessing ex vivo drug responses using CRC PDEs
We identified 38 studies that assessed ex vivo drug responses using CRC PDEs (Supplementary Table 2). Commonly used chemotherapy drugs in CRC (e.g. 5-FU, oxaliplatin, irinotecan) and their combinations were tested. More recent studies included targeted therapies such as cetuximab [29, 32, 53, 64], bevacizumab [29, 32], gefitinib [56], and selumetinib [43]. A large number of studies performed the histoculture drug response assay (HDRA) with 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) as an endpoint to identify effective drugs based on inhibition of tumour viability [14, 16, 17, 19, 20, 23, 24, 27,28,29,30, 32, 33, 44, 65]. Two studies performed a similar colorimetric assay using water-soluble tetrazolium 8 (WST-8) [53, 66] and one study performed a luminometric adenosine triphosphate (ATP) assay [56] to measure cell viability and assess chemosensitivity. Apart from cell proliferation assays, a number of studies compared treated and untreated explants using IHC to measure proliferation markers (e.g. BrdU [21], Ki-67 [31, 44, 46, 47, 53, 55, 64, 67], proliferating cell nuclear antigen (PCNA) [45]) or apoptotic markers (e.g. caspase-3 [31, 53, 64, 67, 68]), while other studies employed H&E staining to assess tissue morphology [38, 43, 46, 53, 54, 64, 67]. Six studies analysed drug responses on a protein and gene expression level [40, 41, 44, 45, 48, 56].
Biomarker discovery and validation using CRC PDEs
We identified 24 studies using CRC PDEs to discover and validate potential biomarkers of treatment response or clinical outcome (Supplementary Table 3). In about half of these studies, they were correlated with treatment response using ex vivo HDRA results as a surrogate marker. Response to chemotherapeutic agents as assessed by HDRA was shown to be predictive of in vivo treatment response with moderate specificity and sensitivity [14, 17, 29, 32, 33]. One study showed that chemosensitivity to 5-FU and cisplatin depended on the presence of serum p53 antibody [18]. Another study investigated the enzyme activity of dihydropyrimidine dehydrogenase (DPD) & orotate phosphoribosyltransferase (OPRT) and found that the combination of these levels was predictive of 5-FU positive sensitivity [20]. Various immunohistochemical markers including p53 [19], p21 [19], and ATP binding cassette subfamily G member 2 (ABCG2) [65] have been correlated with response to various chemotherapeutic agents. Similarly, there were differential gene expressions of thymidylate synthase (TS) [22, 30], DPD & OPRT [25], amphiregulin (AREG) & epiregulin (EREG) [64], microRNA 34a (miR-34a) [34], and checkpoint-with-forkhead-and-ring-finger-domains (CHFR) [35] between treatment responders and non-responders. Tumours with mismatch repair (MMR) defects were closely correlated with chemosensitivities to combined regimens of PDX101 with 5-FU + leucovorin + oxaliplatin (FLOX) and 5-FU + leucovorin + irinotecan (FLIRI) [27]. A more recent study incorporated readouts from different assays (H&E, IHC and CCK-8 assay) to generate a score to predict clinical response with 91.67% specificity and 100% sensitivity [53].
Discussion
To our knowledge, this is the first scoping review evaluating the utility of PDEs as a preclinical model in primary CRC to discover and validate potential biomarkers, explore the TME and assess drug responses. The use of PDEs is not a novel concept and has been around since the 1960s in various formats [36, 37, 49, 69,70,71]. Early studies investigated the feasibility of such “tissue cultures” and assessed ex vivo response to different chemotherapeutic agents. Many of these studies had small sample sizes and included different types of cancers apart from CRC, which made comparisons challenging and conclusions difficult to draw. In the 1990s, Hoffmann et al. successfully optimised and popularised the platform for drug testing, with the assay now commonly known as HDRA [72]. In addition to CRC [14, 17, 29, 32, 33], previous studies demonstrated that HDRA results correlated well with in vivo drug responses and clinical outcomes in various types of solid tumours [73,74,75].
Despite its initial success, PDEs have not been widely used in basic research and have had minimal impact in terms of incorporating them into cancer drug development pipeline or precision medicine approach to guide clinical decision-making. This may be due to their relatively short-term viability, low throughput due to finite amount of tissue, lack of standardised response readouts, and challenges in genetic manipulation using technologies such as siRNA or CRISPR. Preference for established preclinical models, such as cell lines and xenografts, as well as the surging popularity of organoids in the early 2000s [76] may also have played a role. Organoids are 3D, stem-cell-derived structures that resemble their in vivo tissue counterparts and can be maintained in culture, with passaging, for a potentially indefinite period. Consequently, it provides useful insights into the effects of tumour heterogeneity and allows testing of different drugs and cellular therapies, making it an attractive preclinical tool to predict therapy response. However, some of the potential disadvantages of organoid cultures include in vitro selection of specific clones that are able to grow in those conditions, lack of heterotypic cell types (e.g. stromal and immune cells), low success rates, and longer times to establish successful cultures which may limit clinical applications.
With a better appreciation of the significance of the TME in determining cancer progression and treatment response, there is renewed interest in the use of PDEs to maintain tissue architecture and reflect tumour-stromal interactions and metabolic crosstalk. In addition, they are relatively easy to establish, cost-effective, and provide results within a clinically acceptable timeframe. CRC PDEs can be generated from tumour tissue obtained from either endoscopic biopsies or surgical specimens. Because of intra-tumour heterogeneity, obtaining tissue from biopsies increases the risk of working on a sample that is not representative of the overall tumour. The risk still exists in surgical specimens, albeit lower than in endoscopic biopsies. On the other hand, obtaining tissue from surgical specimens can have the limitation of not being treatment-naïve in the case of locally advanced rectal cancer where most patients undergo NACRT before proceeding to surgery. This may alter characteristics of the original tumour and TME but also increase the risk of sampling error due to tumour regression, necrosis and fibrosis.
Different tissue processing methods have been described but mechanical fragmentation into smaller tumour pieces was most commonly performed, particularly for HDRA. Alternatively, tissue slices of varying thickness were used in some studies for better consistency and reproducibility. In contrast to the methods used in establishing cell lines and organoids, tumour fragments or slices do not need to be enzymatically digested to obtain a single cell suspension. This allows preservation of the TME containing immune cells, stromal cells and extracellular matrix that provides physical scaffolding for cellular components, as well as biochemical signals that are essential for tissue differentiation and homeostasis. Although free-floating cultures may be the simplest to establish and maintain, cells tend to migrate out of the explants to form monolayer cultures with loss of tissue architecture [77]. In order to improve nutrient and waste exchange and tissue viability, some studies have successfully incorporated agitation or adopted the use of scaffolds such as gelatin or collagen sponges into their culturing techniques.
Most contemporary studies used a commercially available cell culture medium such as RPMI 1640 or DMEM, supplemented with FBS and antibiotics. FBS has been used in human cell cultures for decades and contains a variety of growth factors that are important in overall cell proliferation and differentiation, while antibiotics reduce bacterial contamination of cultures by intestinal flora from tissue specimens. In some studies that omitted FBS, other growth-promoting supplements such as B-27 [54, 67, 68] and epidermal growth factor (EGF) [54, 56, 67, 68] were added to improve viability, although evidence is scarce and inconclusive in studies on non-CRC explants [78, 79]. With the recent development of more physiological cell culture media which mimics the composition of human plasma (e.g. Plasmax, Human Plasma-Like Medium) [80], it will be interesting to know if they will better support tumour niche and prolong viability of PDEs. Most PDEs were maintained at a temperature of 37°C and a CO2 level of 5%, which are similar to physiological parameters in the human body. There were variations in O2 level, with a number of studies opting for a hyperoxic environment presumably due to the fact that hypoxia reduces tissue viability, although it is difficult to measure the amount of O2 that will diffuse into the tissue in this experimental setup [78, 81].
With the almost endless combinations of culture methods and conditions, it is not surprising that the most optimal system for CRC explant cultures has not been established. In addition, different assays and quality standards were used to characterise tissue viability at various time points. Most studies reported on tissue morphology based on H&E stain, although careful quantification was sometimes lacking. In early studies, cell proliferation was assessed using autoradiography, which measured the ability to incorporate uridine and thymidine during DNA synthesis [49, 70]. Subsequently, this was replaced by immunohistochemical staining of proliferation markers such as Ki-67 [44, 46, 53, 55], which is a nuclear protein expressed in all phases of the cell cycle, except the resting phase (G0). More recently, newer techniques such as fluorescence-based viability assays [53, 54] and gene expression analysis [44, 56] have been utilised to further support the use of explants as a preclinical model. Regardless of culture methods and conditions, most studies demonstrated that CRC PDEs can be maintained ex vivo for at least 3–7 days without significant loss of viability. Remarkably, da Mata et al. was able to maintain their PDE cultures for a maximum of 122 days (median, 28 days), which may be partly explained by the use of a similar culture medium which has successfully established CRC organoids, as well as an agitation-based system which promotes diffusion of oxygen and soluble factors [54].
Given the main advantage of the explant model is the preservation of the TME, a better characterisation of this environment is crucial to understand the mechanisms for treatment response and provide new insight into potential resistance mechanisms. To this end, previous studies have used PDEs to investigate the roles of different enzymes [57, 58] and cytokines [15, 50, 59,60,61,62,63] which may contribute to cancer progression and metastasis. As expected, the cytokine profiles differed significantly between normal and tumour microenvironment. Consequently, the cytokine profile influences the immune cell populations present in the TME. Disappointingly, there has been a lack of studies that looked at immune cell subtypes and spatial relationships between tumour and immune cells [39, 50], despite these factors being predictive of chemotherapeutic drug responses and patient outcomes in CRC [9,10,11]. Recent results from pancreatic and ovarian PDEs have demonstrated that immune cells, such as macrophages, cytotoxic T lymphocytes and regulatory T cells, can be retained in cultures, ensuring that this platform will provide functional relevance and redefine evaluation of current and novel therapies [82,83,84].
Over the years, the use of explant cultures has predominantly focused on assessing ex vivo response to commonly used chemotherapeutic agents, and increasingly, targeted therapies and immunotherapies used in the clinical setting. Many studies have utilised the HDRA which can identify effective treatments based on inhibition of tumour viability [14, 16, 17, 19, 20, 23, 24, 27,28,29,30, 32, 33, 44, 65]. Briefly, it involves incubating tumour fragments on collagen or gelatin sponge, with or without drugs, for a specific time period and then adding MTT, which viable cells convert into purple formazan that is measured by spectrophotometry. Whilst this metabolic activity assay is easy to use and rapid and allows high throughput, the end results may be affected by different factors (e.g. pH, cellular ion concentration, cell types and numbers). In addition, MTT is insoluble in cell culture media and needs to be dissolved in dimethyl sulfoxide (DMSO) or isopropanol, thus it is mainly used as an endpoint detection method. IHC staining has also been commonly used to compare drug-treated and untreated PDEs. Treatment effects are quantified by calculating the percentage of cells expressing proliferation markers (e.g. Ki-67, PCNA) and apoptotic markers (e.g. cleaved-caspase-3), although the scoring may be subjective.
At present, only 7.5% of all potential anti-cancer drugs that enter phase I clinical trials are eventually approved for clinical use [85]. Undoubtedly, a major obstacle in new drug development and subsequent approval is the ability to predict clinical efficacy in preclinical models [86]. The potential ability to differentiate between treatment responders and non-responders can greatly assist in selection of patients for clinical trials. Potential biomarkers of treatment response, such as gene expression (TS, miRNA34, CHFR) and protein markers (e.g. p53, p21, ABCG2), were investigated but none of them have been validated in other studies. Moreover, many of them were identified from HDRA, instead of correlating with in vivo clinical response, so they must be interpreted with caution. A recent notable advance is the development of the CANScript platform by Majumder et al. which incorporated results from a combination of ex vivo assays (H&E; Ki-67 and cleaved caspase-3 on IHC; CCK-8 assay) to generate a score to predict clinical response with 91.67% specificity and 100% sensitivity [53].
As with any other preclinical models, the PDE model has its own challenges include obtaining sufficient tumour tissue, timely and careful processing of tissue, optimising culture method and conditions, and establishing the optimal window of time for drug testing or co-cultures. With rapid developments in biomolecular technologies (e.g. tumour-on-a-chip microfluidic platforms to screen multiple drugs on a single PDE) and endpoint analysis (e.g. multiplex immunofluorescence staining to differentiate cell types in the TME, non-destructive assays to permit readouts at multiple timepoints), it is hoped that some of these challenges can be overcome to bring back PDEs as a powerful preclinical model in cancer research.
At present, our scoping review on its utility in translational research remains inconclusive, largely because of the limited number and heterogeneity of previous studies. Further validation studies are required to characterise, refine and standardise this preclinical model before it can become a part of precision medicine. This will require close collaboration between clinicians and scientists to support ongoing research efforts.
Conclusion
In summary, given the preservation of the TME and tumour heterogeneity, PDEs represent a more clinically relevant model for CRC than other preclinical models. In addition, they are relatively easy to establish, cost-effective, and provide results within a clinically acceptable timeframe. With the introduction of immunotherapies such as anti-PD1, which has shown excellent clinical efficacy in a subset of CRC patients, there has never been a stronger incentive to incorporate PDEs in future preclinical studies. We believe that the use of PDEs for functional assays, in combination with molecular profiling, has the potential to identify more reliable biomarkers, potential treatment resistance mechanisms and novel therapeutic agents to improve patient outcomes in CRC.
Data Availability
All data supporting the findings of this study are available within the paper and its Supplementary Information.
References
Maas M et al (2010) Long-term outcome in patients with a pathological complete response after chemoradiation for rectal cancer: a pooled analysis of individual patient data. Lancet Oncol 11(9):835–844. https://doi.org/10.1016/S1470-2045(10)70172-8
Garcia-Aguilar J et al (2022) Organ preservation in patients with rectal adenocarcinoma treated with Total neoadjuvant therapy. J Clin Oncol 40(23):2546–2556. https://doi.org/10.1200/JCO.22.00032
Bahadoer RR et al (2021) Short-course radiotherapy followed by chemotherapy before total mesorectal excision (TME) versus preoperative chemoradiotherapy, TME, and optional adjuvant chemotherapy in locally advanced rectal cancer (RAPIDO): a randomised, open-label, phase 3 trial. Lancet Oncol 22(1):29–42. https://doi.org/10.1016/S1470-2045(20)30555-6
Conroy T et al (2021) Neoadjuvant chemotherapy with FOLFIRINOX and preoperative chemoradiotherapy for patients with locally advanced rectal cancer (UNICANCER-PRODIGE 23): a multicentre, randomised, open-label, phase 3 trial. Lancet Oncol 22(5):702–715. https://doi.org/10.1016/S1470-2045(21)00079-6
Cercek A et al (2022) PD-1 blockade in mismatch repair-deficient, locally advanced rectal cancer. N Engl J Med 386(25):2363–2376. https://doi.org/10.1056/NEJMoa2201445
Sargent D et al (2009) Evidence for cure by adjuvant therapy in colon cancer: observations based on individual patient data from 20,898 patients on 18 randomized trials. J Clin Oncol 27(6):872–877. https://doi.org/10.1200/JCO.2008.19.5362
Tie J et al (2022) Circulating tumor DNA analysis guiding adjuvant therapy in stage II colon cancer. N Engl J Med 386(24):2261–2272. https://doi.org/10.1056/NEJMoa2200075
Mackenzie NJ et al (2022) Modelling the tumor immune microenvironment for precision immunotherapy. Clin Transl Immunology 11(6):e1400. https://doi.org/10.1002/cti2.1400
Anitei MG et al (2014) Prognostic and predictive values of the immunoscore in patients with rectal cancer. Clin Cancer Res 20(7):1891–1899. https://doi.org/10.1158/1078-0432.CCR-13-2830
McCoy MJ et al (2015) Low stromal Foxp3+ regulatory T-cell density is associated with complete response to neoadjuvant chemoradiotherapy in rectal cancer. Br J Cancer 113(12):1677–1686. https://doi.org/10.1038/bjc.2015.427
Galon J, Lanzi A (2020) Immunoscore and its introduction in clinical practice. Q J Nucl Med Mol Imaging 64(2):152–161. https://doi.org/10.23736/S1824-4785.20.03249-5
Hatfield SM et al (2015) Immunological mechanisms of the antitumor effects of supplemental oxygenation. Sci Transl Med 7(277):27730. https://doi.org/10.1126/scitranslmed.aaa1260
Kenerson HL et al (2020) Tumor slice culture as a biologic surrogate of human cancer. Ann Transl Med 8(4):114. https://doi.org/10.21037/atm.2019.12.88
Kanamori N, Fujii M, Iwai S, Nemoto N, Sakurai I (1999) In vitro and in vivo sensitivity tests for anticancer drugs: comparative study of histoculture drug response assay (HDRA) utilizing biopsy specimens and histopathological effects produced by preoperative chemotherapy using UFT in colo-rectal cancer patients. Nihon Univ J Med 41(5):239–250
Mutala LB et al (2021) The caspase-1/IL-18 axis of the inflammasome in tumor cells: a modulator of the Th1/Tc1 response of tumor-infiltrating T lymphocytes in colorectal cancer. Cancers 13(2). https://doi.org/10.3390/cancers13020189
Furukawa T et al (1992) Chemosensitivity testing of clinical gastrointestinal cancers using histoculture and the MTT end-point. Anticancer Res 12(5):1377–1382
Furukawa T, Kubota T, Hoffman RM (1995) Clinical applications of the histoculture drug response assay. Clin Cancer Res 1(3):305–311
Takeda A et al (1999) Clinical significance of serum p53 antibody detection on chemosensitivity assay in human colorectal cancer. J Surg Oncol 71(2):112–116. https://doi.org/10.1002/(sici)1096-9098(199906)71:2%3c112::aid-jso10%3e3.0.co;2-p
Hosaka N et al (2001) Correlation of immunohistochemical p53 labeling index with inhibition rate in chemosensitivity test in gastric and colon cancer. Anticancer Res 21(1A):229–235
Isshi K et al (2002) Predicting 5-FU sensitivity using human colorectal cancer specimens: comparison of tumor dihydropyrimidine dehydrogenase and orotate phosphoribosyl transferase activities with in vitro chemosensitivity to 5-FU. Int J Clin Oncol 7(6):335–342. https://doi.org/10.1007/s101470200051
Yanqun L, Eu KW, Seow-Choen F, Cheah PY (2002) Differential cytostatic effect of sodium salicylate in human colorectal cancers using an individualized histoculture system. Cancer Chemother Pharmacol 49(6):473–478. https://doi.org/10.1007/s00280-002-0441-7
Inoue T et al (2005) Expression level of thymidylate synthase is a good predictor of chemosensitivity to 5-fluorouracil in colorectal cancer. J Gastroenterol 40(2):143–147. https://doi.org/10.1007/s00535-004-1512-9
Kwon YE, Kim KH (2006) Octahedral PT(IV) complex K101 induces apoptosis via ERK1/2 activation and the p53 pathway in human colon cancer cells. Anticancer Drugs 17(5):553–558. https://doi.org/10.1097/00001813-200606000-00009
Kwon YE, Park JY, Kim WK (2007) In vitro histoculture drug response assay and in vivo blood chemistry of a novel Pt(IV) compound, K104. Anticancer Res 27(1A):321–326
Kinoshita M et al (2007) Gene expression profile of 5-fluorouracil metabolic enzymes in primary colorectal cancer: potential as predictive parameters for response to fluorouracil-based chemotherapy. Anticancer Res 27(2):851–856
Pirnia F et al (2009) Novel functional profiling approach combining reverse phase protein microarrays and human 3-D ex vivo tissue cultures: expression of apoptosis-related proteins in human colon cancer. Proteomics 9(13):3535–3548. https://doi.org/10.1002/pmic.200800159
Kim JC et al (2009) Evaluation of novel histone deacetylase inhibitors as therapeutic agents for colorectal adenocarcinomas compared to established regimens with the histoculture drug response assay. Int J Colorectal Dis 24(2):209–218. https://doi.org/10.1007/s00384-008-0590-1
Um JW, Lee JH, Kim SH, Lee ES, Kim YS (2012) Can immunohistochemistry of multidrug-resistant proteins replace the histoculture drug response assay in colorectal adenocarcinomas? Hepatogastroenterology 59(116):1075–1078. https://doi.org/10.5754/hge09411
Yoon YS et al (2012) Applicability of histoculture drug response assays in colorectal cancer chemotherapy. Anticancer Res 32(8):3581–3586. http://ar.iiarjournals.org/content/32/8/3581.full.pdf+html. Accessed 28 Sep 2022
Zhang Q et al (2014) TS mRNA levels can predict pemetrexed and raltitrexed sensitivity in colorectal cancer. Cancer Chemother Pharmacol 73(2):325–333. https://doi.org/10.1007/s00280-013-2354-z
Flanagan L et al (2016) Low levels of Caspase-3 predict favourable response to 5FU-based chemotherapy in advanced colorectal cancer: Caspase-3 inhibition as a therapeutic approach. Cell Death Dis 7(101524092):e2087. https://doi.org/10.1038/cddis.2016.7
Yoon YS et al (2017) Development and applicability of integrative tumor response assays for metastatic colorectal cancer. Anticancer Res 37(3):1297–1303. https://doi.org/10.21873/anticanres.11447
Ji WB et al (2017) Clinical significance of 5-fluorouracil chemosensitivity testing in patients with colorectal cancer. Anticancer Res 37(5):2679–2682. https://doi.org/10.21873/anticanres.11616
Li et al Y (2018) miR-34a regulates multidrug resistance via positively modulating OAZ2 signaling in colon cancer cells. J Immunol Res 2018. https://doi.org/10.1155/2018/7198514
Hagiwara T et al (2022) CHFR-promoter-methylation status is predictive of response to irinotecan-based systemic chemotherapy in advanced colorectal cancer. Anticancer Res 42(2):697–707. https://doi.org/10.21873/anticanres.15528
Rovin S (1962) The influence of carbon dioxide on the cultivation of human neoplastic explants in Vitro*. Can Res 22(3):384–387
Röller M-R, Owen SP, Heidelberger C (1966) Studies on the Organ Culture of Human Tumors1. Cancer Research 26(4_Part_1):626–637
Pritchett CJ, Senior PV, Sunter JP, Watson AJ, Appleton DR, Wilson RG (1982) Human colorectal tumours in short-term organ culture. a stathmokinetic study. Cell Tissue Kinet 15(5):555–564. https://doi.org/10.1111/j.1365-2184.1982.tb01577.x
Golby SJC, Chinyama C, Spencer J (2002) Proliferation of t-cell subsets that contact tumour cells in colorectal cancer. Clin Exp Immunol 127(1):85–91. https://doi.org/10.1046/j.1365-2249.2002.01730.x
Monteleone G et al (2006) Silencing of SH-PTP2 defines a crucial role in the inactivation of epidermal growth factor receptor by 5-aminosalicylic acid in colon cancer cells. Cell Death Differ 13(2):202–211. https://doi.org/10.1038/sj.cdd.4401733
Stolfi C et al (2008) Mesalazine negatively regulates CDC25A protein expression and promotes accumulation of colon cancer cells in S phase. Carcinogenesis 29(6):1258–1266. https://doi.org/10.1093/carcin/bgn122
Stolfi C et al (2012) 2-Methoxy-5-amino-N-hydroxybenzamide, a derivative of mesalamine, inhibits colon cancer cell growth through cyclo-oxygenase-2-dependent and -independent mechanisms. Clin Sci 123(5–6):295–306. https://doi.org/10.1042/CS20110556
Gavert N et al (2022) Ex vivo organotypic cultures for synergistic therapy prioritization identify patient-specific responses to combined MEK and Src inhibition in colorectal cancer. Nat Cancer 3(2):219–231. https://doi.org/10.1038/s43018-021-00325-2
Vaira V et al (2010) Preclinical model of organotypic culture for pharmacodynamic profiling of human tumors. Proc Natl Acad Sci 107(18):8352–8356. https://doi.org/10.1073/pnas.0907676107
Stolfi C et al (2014) A functional role for Smad7 in sustaining colon cancer cell growth and survival. Cell Death Dis 5. https://doi.org/10.1038/cddis.2014.49
Sonnichsen R et al (2018) Individual susceptibility analysis using patient-derived slice cultures of colorectal carcinoma. Clin Colorectal Cancer 17(2):e189–e199. https://doi.org/10.1016/j.clcc.2017.11.002
Laudisi F et al (2019) Progranulin sustains STAT3 hyper-activation and oncogenic function in colorectal cancer cells. Mol Oncol 13(10):2142–2159. https://doi.org/10.1002/1878-0261.12552
Hewitt SL et al (2020) Intratumoral IL12 mRNA therapy promotes TH1 transformation of the tumor microenvironment. Clin Cancer Res 26(23):6284–6298. https://doi.org/10.1158/1078-0432.CCR-20-0472
Wolberg WH, Brown RR (1962) Autoradiographic studies of in vitro incorporation of uridine and thymidine by human tumor Tissue*. Can Res 22(9):1113–1119
Ott E et al (2019) The density of Tbet+ tumor-infiltrating T lymphocytes reflects an effective and druggable preexisting adaptive antitumor immune response in colorectal cancer, irrespective of the microsatellite status. Oncoimmunology 8(4):e1562834. https://doi.org/10.1080/2162402X.2018.1562834
Kalus M (1972) Carcinoma and adenomatous polyps of the colon and rectum in biopsy and organ tissue culture. Cancer 30(4):972–982. https://doi.org/10.1002/1097-0142(197210)30:4%3c972::AID-CNCR2820300418%3e3.0.CO;2-A
Hood CJ, Parham DM (1998) A simple method of tumour culture. Pathol Res Pract 194(3):177–181
Majumder B et al (2015) Predicting clinical response to anticancer drugs using an ex vivo platform that captures tumour heterogeneity. Nat Commun 6(1):6169. https://doi.org/10.1038/ncomms7169
da Mata S et al (2021) Patient-derived explants of colorectal cancer: histopathological and molecular analysis of long-term cultures. Cancers 13(18). https://doi.org/10.3390/cancers13184695
Brouquet A et al (2011) A model of primary culture of colorectal cancer and liver metastasis to predict chemosensitivity. J Surg Res 166(2):247–254. https://doi.org/10.1016/j.jss.2009.04.039
Unger F et al (2015) Precision cut cancer tissue slices in anti-cancer drug testing. J Mol Pathophysiol 4:108. https://doi.org/10.5455/jmp.20151023055556
Harvey SR, Lawrence DD, Madeja JM, Abbey SJ, Markus G (1988) Secretion of plasminogen activators by human colorectal and gastric tumor explants. Clin Exp Metas 6(6):431–450. https://doi.org/10.1007/BF01784375
Yamagata S, Yoshii Y, Suh JG, Tanaka R, Shimizu S (1991) Occurrence of an active form of gelatinase in human gastric and colorectal-carcinoma tissues. Cancer Lett 59(1):51–55. https://doi.org/10.1016/0304-3835(91)90135-5
Michielsen AJ et al (2011) Tumour tissue microenvironment can inhibit dendritic cell maturation in colorectal cancer. PLoS ONE 6(11):e27944. https://doi.org/10.1371/journal.pone.0027944
Muthuswamy R et al (2012) NF-kappaB hyperactivation in tumor tissues allows tumor-selective reprogramming of the chemokine microenvironment to enhance the recruitment of cytolytic T effector cells. Can Res 72(15):3735–3743. https://doi.org/10.1158/0008-5472.CAN-11-4136
O’Toole A et al (2014) Tumour microenvironment of both early- and late-stage colorectal cancer is equally immunosuppressive. Br J Cancer 111(5):927–932. https://doi.org/10.1038/bjc.2014.367
Kistner L, Doll D, Holtorf A, Nitsche U, Janssen KP (2017) Interferon-inducible CXC-chemokines are crucial immune modulators and survival predictors in colorectal cancer. Oncotarget 8(52):89998–90012. https://doi.org/10.18632/oncotarget.21286
Benkhelifa S et al (2019) Aberrant up-regulation ofiNOS/NO system is correlated with an increased abundance of Foxp3(+)cells and reduced effector/memory cell markers expression during colorectal cancer: immunomodulatory effects of cetuximab combined with chemotherapy. Inflammopharmacology 27(4):685–700. https://doi.org/10.1007/s10787-019-00566-9
Brijwani N et al (2017) Rationally co-targeting divergent pathways in KRAS wild-type colorectal cancers by CANscript technology reveals tumor dependence on Notch and Erbb2. Sci Rep 7(1):1502. https://doi.org/10.1038/s41598-017-01566-x
Yuan SQ et al (2009) Correlation of chemosensitivity tested using histoculture drug response assay to expression of multidrug resistance genes and proteins in colorectal cancer tissues. Chin J Cancer 28(9):932–938. https://doi.org/10.5732/cjc.008.10787
Matsuoka H, Nagaya M, Tsukikawa S, Yanagi Y, Isogai A, Kubota S (2006) Repeated hepatic intra-arterial chemotherapy based on results of anticancer drug sensitivity test in patients with synchronous hepatic metastases from colorectal cancer. Surgery 140(3):387–395. https://doi.org/10.1016/j.surg.2006.03.001
Ahmed M et al (2020) Repurposing antibacterial AM404 as a potential anticancer drug for targeting colorectal cancer stem-like cells. Cancers 12(1):106. https://doi.org/10.3390/cancers12010106
Lo Re D et al (2018) Increased immune cell infiltration in patient-derived tumor explants treated with Traniplatin: An original Pt(iv) pro-drug based on Cisplatin and Tranilast. Chem Commun 54(60):8324–8327. https://doi.org/10.1039/c8cc02071j
Wright JC, Cobb JP, Gumport SL, Safadi D, Walker DG, Golomb FM (1962) Further investigation of the relation between the clinical and tissue culture response to chemotherapeutic agents on human cancer. Cancer 15(2):284–293. https://doi.org/10.1002/1097-0142(196203/04)15:2%3c284::AID-CNCR2820150212%3e3.0.CO;2-I
Wolberg WH (1964) Studies on the mechanism of human tumor resistance to the fluorinated Pyrimidines*. Can Res 24(8):1437–1447
Hurley JD, Yount LJ (1965) Selection of anticancer drug for palliation using tissue culture sensitivity studies. Am J Surg 109:39–42. https://doi.org/10.1016/s0002-9610(65)80100-3
Hoffman RM (1991) In vitro sensitivity assays in cancer: a review, analysis, and prognosis. J Clin Lab Anal 5(2):133–143. https://doi.org/10.1002/jcla.1860050211
Hasegawa Y et al (2007) Evaluation of optimal drug concentration in histoculture drug response assay in association with clinical efficacy for head and neck cancer. Oral Oncol 43(8):749–756. https://doi.org/10.1016/j.oraloncology.2006.09.003
Jung PS et al (2013) Progression-free survival is accurately predicted in patients treated with chemotherapy for epithelial ovarian cancer by the histoculture drug response assay in a prospective correlative clinical trial at a single institution. Anticancer Res 33(3):1029–34. https://www.ncbi.nlm.nih.gov/pubmed/23482777. Accessed 28 Sep 2022
Shinden Y et al (2016) Clinical significance of the histoculture drug response assay in breast cancer. Anticancer Res 36(11):6173–6178. https://doi.org/10.21873/anticanres.11210
Sato T et al (2011) Long-term expansion of epithelial organoids from human colon, adenoma, adenocarcinoma, and Barrett’s epithelium. Gastroenterology 141(5):1762–1772. https://doi.org/10.1053/j.gastro.2011.07.050
Centenera MM, Raj GV, Knudsen KE, Tilley WD, Butler LM (2013) Ex vivo culture of human prostate tissue and drug development. Nat Rev Urol 10(8):483–487. https://doi.org/10.1038/nrurol.2013.126
Misra S et al (2019) Ex vivo organotypic culture system of precision-cut slices of human pancreatic ductal adenocarcinoma. Sci Rep 9(1):2133. https://doi.org/10.1038/s41598-019-38603-w
Naipal KA et al (2016) Tumor slice culture system to assess drug response of primary breast cancer. BMC Cancer 16:78. https://doi.org/10.1186/s12885-016-2119-2
Vande Voorde J et al (2019) Improving the metabolic fidelity of cancer models with a physiological cell culture medium. Sci Adv 5(1):eaau7314. https://doi.org/10.1126/sciadv.aau7314
Davies EJ et al (2015) Capturing complex tumour biology in vitro: histological and molecular characterisation of precision cut slices. Sci Rep 5:17187. https://doi.org/10.1038/srep17187
Lim CY et al (2018) Organotypic slice cultures of pancreatic ductal adenocarcinoma preserve the tumor microenvironment and provide a platform for drug response. Pancreatology 18(8):913–927. https://doi.org/10.1016/j.pan.2018.09.009
Abreu S et al (2020) Patient-derived ovarian cancer explants: preserved viability and histopathological features in long-term agitation-based cultures. Sci Rep 10(1):19462. https://doi.org/10.1038/s41598-020-76291-z
Kokkinos J et al (2021) Ex vivo culture of intact human patient derived pancreatic tumour tissue. Sci Rep 11(1):1944. https://doi.org/10.1038/s41598-021-81299-0
Toniatti C, Jones P, Graham H, Pagliara B, Draetta G (2014) Oncology drug discovery: planning a turnaround. Cancer Discov 4(4):397–404. https://doi.org/10.1158/2159-8290.CD-13-0452
Kamb A, Wee S, Lengauer C (2007) Why is cancer drug discovery so difficult? Nat Rev Drug Discov 6(2):115–120. https://doi.org/10.1038/nrd2155
Funding
Open Access funding enabled and organized by CAUL and its Member Institutions No funding was obtained for this study.
Author information
Authors and Affiliations
Contributions
Milton Mui, Alexander G. Heriot and Joseph Kong conceived and designed the study. Milton Mui and Molly Clark collected the data. All authors contributed to the analysis and interpretation of the results. Milton Mui took the lead in writing the manuscript. All authors provided critical feedback and contributed to the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Mui, M., Clark, M., Vu, T.M.S.H. et al. Use of patient-derived explants as a preclinical model for precision medicine in colorectal cancer: A scoping review. Langenbecks Arch Surg 408, 392 (2023). https://doi.org/10.1007/s00423-023-03133-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00423-023-03133-7