Abstract
Background
Peripheral neuropathies in mitochondrial disease are caused by mutations in nuclear genes encoding mitochondrial proteins, or in the mitochondrial genome. Whole exome or genome sequencing enable parallel testing of nuclear and mtDNA genes, and it has significantly advanced the genetic diagnosis of inherited diseases. Despite this, approximately 40% of all Charcot-Marie-Tooth (CMT) cases remain undiagnosed.
Methods
The genome-phenome analysis platform (GPAP) in RD-Connect was utilised to create a cohort of 2087 patients with at least one Human Phenotype Ontology (HPO) term suggestive of a peripheral neuropathy, from a total of 10,935 patients. These patients’ genetic data were then analysed and searched for variants in known mitochondrial disease genes.
Results
A total of 1,379 rare variants were identified, 44 of which were included in this study as either reported pathogenic or likely causative in 42 patients from 36 families. The most common genes found to be likely causative for an autosomal dominant neuropathy were GDAP1 and GARS1. We also detected heterozygous likely pathogenic variants in DNA2, MFN2, DNM2, PDHA1, SDHA, and UCHL1. Biallelic variants in SACS, SPG7, GDAP1, C12orf65, UCHL1, NDUFS6, ETFDH and DARS2 and variants in the mitochondrial DNA (mtDNA)-encoded MT-ATP6 and MT-TK were also causative for mitochondrial CMT. Only 50% of these variants were already reported as solved in GPAP.
Conclusion
Variants in mitochondrial disease genes are frequent in patients with inherited peripheral neuropathies. Due to the clinical overlap between mitochondrial disease and CMT, agnostic exome or genome sequencing have better diagnostic yields than targeted gene panels.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Inherited peripheral neuropathies or Charcot-Marie-Tooth disease (CMT) is a group of closely related disorders affecting the motor and sensory neurones of the peripheral nervous system. With an estimated prevalence of 40 individuals per 100,000, CMT is among the most common hereditary neuromuscular disorders. CMT is categorised into demyelinating and axonal subtypes, based on the primary mechanism of degeneration [1, 2].
CMT typically presents in the first or second decade of life, but the age of onset can vary widely, ranging from early infancy to late adulthood. Typical symptoms include symmetrical distal weakness and atrophy, sensory loss, foot deformities (pes cavus or planus, hammer toes, and clawing fingers), and reduced or absent tendon reflexes. Sensorineural hearing loss and phrenic nerve mediated respiratory insufficiency have also been reported. For the vast majority of patients, these symptoms are painless and, although progressively debilitating, not life threatening [3].
CMT exhibits significant genetic heterogeneity, with over 100 causative genes identified to date, and can be inherited through several different modes of inheritance, including autosomal dominant, autosomal recessive, X-linked, and mitochondrial. These genes encompass a wide range of functions, including those involved in peripheral nerve structure, myelin production, axonal transport, and mitochondrial dynamics [4, 5]. This genetic diversity contributes to the significant phenotypic variability observed in CMT, with variations in age of onset, disease progression, severity of symptoms, and specific clinical features. Understanding the underlying genetic subtypes and their associated phenotypic variations is crucial for accurate diagnosis, prognosis, and personalised management strategies. Clinical diagnosis is complex and often elusive, focusing on clinical presentation, neurophysiological tests, genetic and molecular tests, and previously on nerve biopsy. The introduction of next-generation sequencing (NGS) technology has significantly advanced the genetic diagnosis of inherited rare diseases in recent years [6]. Despite this, in approximately 40% of all CMT cases a genetic diagnosis is not identified [7].
The normal functioning of neurons and their long axons is dependent on healthy mitochondria, which play critical roles in axonal transport, energy production, calcium buffering, and other essential cellular functions. The distribution of mitochondria along peripheral axons is regulated by a continuous process associated with mitochondrial fusion and fission, which is collectively known as mitochondrial dynamics [8]. Disruptions to mitochondria have been increasingly recognised as a cause of peripheral nerve dysfunction. In keeping with this, a third of patients with mitochondrial disorders develop peripheral neuropathy [9,10,11]. Peripheral neuropathies in mitochondrial disease are typically the result of mutations in genes that are either nuclear encoded, and whose products are subsequently transported to the mitochondria, or in the mitochondrial genome itself.
Reanalysis of existing sequencing data provides an opportunity to enhance the diagnostic yield by leveraging improved bioinformatics pipelines and updated literature [12,13,14,15,16,17,18,19,20,21,22,23,24,25,26,27,28,29]. Previous studies have demonstrated that data reanalysis or re-evaluation can significantly increase the diagnostic yield, depending on the timing of the initial analysis and the interval before reanalysis. The American College of Medical Genetics and Genomics (ACMG) has outlined guidelines for variant-level re-evaluation and case-level reanalysis of genomic test results [30]. Keeping phenotypic descriptions comprehensive and up to date is recommended, as it can improve the specificity of the phenotype and contribute to increased diagnostic success for unsolved cases.
The RD-Connect Genome-Phenome Analysis Platform (GPAP) is an online platform (accessible at https://platform.rd-connect.eu/) designed to facilitate the analysis of genome-phenome data for rare disease (RD) diagnosis and gene discovery [31]. The platform includes heterogeneous datasets, contributed by various clinical researchers, generated in different clinical centres and genomic facilities. To ensure consistency and standardisation across all patients and their relatives, the GPAP platform incorporates a mechanism for submitting pseudonymised phenotypic data using recognised ontologies and standards including the Human Phenotype Ontology (HPO) [32], the Orphanet Rare Disease Ontology (ORDO) [33], and the Online Mendelian Inheritance in Man database (OMIM) [34]. By harmonising the information using these ontologies, the GPAP enhances interoperability and facilitates comprehensive analysis by standardised pipelines. WES/WGS raw data (fastq files) are also processed through a standardised pipeline making sequencing results between different facilities, kits, etc., better comparable.
In our study we aimed to identify a genetic diagnosis on a cohort of patients with possible inherited peripheral neuropathy, with particular focus on mitochondrial genes through the RD-Connect Platform.
Methods
Selection criteria
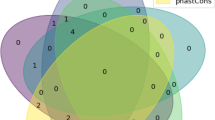
A total of 10,935 individuals (patients and relatives) were visible to all registered and authorised users on the RD-Connect GPAP platform in January 2023. Clinicians uploading patient genomic records on the platform are asked to phenotypically characterise the individuals through HPO terms. We compiled a list of HPO terms commonly associated with inherited neuropathies to create a virtual peripheral neuropathy cohort within GPAP (see Supplemental Materials for the complete list). A search for individuals within the platform with any of these HPO terms resulted in 2087 patients forming our inherited peripheral neuropathy cohort, but not all of these patients have been diagnosed with peripheral neuropathy as a symptom like distal weakness or areflexia could have a different cause (e.g. distal myopathy) (Fig. 1).
Target gene selection
To identify potential genes of interest to target, a comprehensive list of genes associated with mitochondrial disorders was compiled. The list was assembled in two ways. The first component included a high-evidence gene panel for mitochondrial disorders obtained from Genomics England PanelApp (Mitochondrial disorders panel v.4.0). We selected only the high-evidence panel (‘green list’), excluding the medium-evidence (‘amber list’) and low-evidence panels (‘red list’), to ensure the reliability of pathogenicity associations in the included genes. The second component involved a thorough review of the literature, where we included nuclear-encoded genes consistently reported to be involved in mitochondrial pathways. This way the target panel contained all known human disease genes encoding mitochondrial proteins including nuclear genes with involvement of mitochondrial function/dynamics or mutations in the mtDNA. The full list of genes is shown in Supplemental Materials.
Whole exome sequencing (WES) reanalysis and reinterpretation
The cohort was created virtually, as previously described [35,36,37]. The data were reanalysed using a centralised, automated analysis, and filtering approach developed within the RD-Connect GPAP. Variants were filtered for those (a) located in genes associated with mitochondrial diseases (see list in Supplemental Materials), (b) Genome Aggregation Database (gnomAD) allele frequency of < 0.005, (c) internal RD-Connect GPAP allele frequency of < 0.02, (d) a HIGH or MODERATE putative effect prediction, as defined by SnpEFF, a genetic variant annotation and functional effect prediction toolbox, and (e) having zygosity compatible with the inheritance patterns of the gene (based on OMIM database) in which the variants were observed. Variants classified as benign or likely benign in ClinVar were excluded from further analysis. For single nucleotide variants (SNVs), a Combined Annotation Dependent Depletion (CADD) score above 20 was used as an additional criterion for pathogenicity assessment. It is important to note that base insertions, deletions, and duplications did not generate CADD scores and were evaluated using literature search and online tools such as ClinVar. In addition to consulting ClinVar and the ACMG guidelines, we also considered relevant literature surrounding each variant, when available, to further inform our assessment. In total, 1379 variants of interest were identified in 959 patients, of which 1326 variants were excluded for the following reasons: (a) previously reported as benign; (b) low pathogenicity prediction, as indicated by a CADD score of less than 20; (c) annotated in GPAP as solved with alternative genetic diagnosis; (d) incompatible inheritance pattern; (e) incompatible phenotype or non-neuropathic phenotype on further phenotype analysis; (f) incomplete HPO list resulting in ambiguous clinical diagnosis; and (g) conflicting reports of pathogenicity of variant in the literature. Additionally, we excluded variants of uncertain significance, where interpretation was not possible due to insufficient information. With this approach we aimed to maximise the chance that the included variants were likely to be causative.
Results
Cohort description
Of the 10,935 individuals on GPAP, a cohort of 2087 patients was created for those that had at least one HPO term suggestive of a possible neuropathic phenotype. Within this cohort, 1379 variants were identified in 959 patients. Of the 959 patients, 507 were male (52.87%), 415 were female (43.27%), and for 37 (3.86%) their gender was not reported.
Of the 1379 variants, 984 were heterozygous, 262 homozygous, 125 putative compound heterozygous, and 8 were located in the mitochondrial genome. Forty-four variants in 42 patients were found to be either previously reported as causative or identified here as likely to be causative of the patient’s disease. Approximately half of these cases (50%) had already been marked as solved in GPAP. Heterozygous variants were only considered to be potentially causative in genes with known autosomal dominant or X-linked dominant inheritance. Variants in genes with autosomal recessive inheritance were only considered causative if homozygous or compound heterozygous. Contact has been attempted with the involved centres to establish whether a causative variant has been identified, or to discuss the variant of interest.
Variants identified in mitochondrial disease genes
The 44 included variants comprised 18 heterozygous, 14 homozygous, 4 putative compound heterozygous, and 8 mitochondrial variants (Tables 1, 2, 3). The genes responsible for each inheritance mode are shown in Fig. 2. These variants were present in 42 patients from 36 families, contributed by a total of 14 different GPAP submitter groups; a total of 322 HPO terms were registered, with an average of 6 terms per patient. The number of registered terms ranged from a minimum of 1 to a maximum of 16. Of the 42 patients included, 21 were male (50%), 20 were female (48%), and 1 patient’s gender was unknown (2%).
We also identified 7 patients with HPO terms representing a peripheral neuropathy and heterozygous pathogenic or likely pathogenic SPG7 mutations. The role of SPG7 heterozygous variants in the aetiology of neuropathies is currently disputed. To gain a better insight whether this association is real, we tested the frequency of heterozygous SPG7 variants in our neuropathy cohort, compared to the wider GPAP. We found that 1.097% (120 of 10,935) of GPAP participants harbour pathogenic or likely pathogenic heterozygous SPG7 variants, while only 0.240% (5 out of 2,087) of our neuropathy cohort. Consequently, this finding does not support the role of heterozygous SPG7 mutations in the aetiology of mitochondrial neuropathies. It is worth noting that RD-Connect is made up of mostly affected individuals and not healthy controls; therefore, it is not accurately demonstrating the prevalence of reportedly pathogenic heterozygous SPG7 mutations in a healthy population.
Discussion
In this study, our aim was to investigate the occurrence of mitochondrial disease genes in patients with CMT. “Mitochondrial CMT” has been previously reported to be caused by a number of mitochondrial disease genes [5], suggesting a link between mitochondrial dysfunction and neuropathies. While certain forms of mitochondrial CMT, such as MFN2, SANDO, and MNGIE, are well documented, and have been detected by candidate gene sequencing in the past, the mechanism leading to a predominant neuropathy in some individuals remains unclear.
To explore the potential presence of additional mitochondrial forms of CMT, we conducted a search using RD-Connect GPAP. We created a virtual cohort of patients classified by HPO terms typical for CMT and we searched for causative variants in all known mitochondrial disease genes. Our search identified 1,379 variants in 93 genes affecting a total of 959 patients. Further analysis showed that only 44 of these variants, in 16 disease genes, were potentially causative affecting 42 patients in 36 families. In addition to some known variants in genes already known to be associated with neuropathy such as GDAP1, GARS1, DNM2, MFN2, SACS, C12orf65, SPG7, SDHA, PDHA1, and MT-ATP6, we detected some variants in unexpected genes such as DNA2, NDUFS6, ETFDH, SDHA, DARS2 and UCHL1, and MT-TK highlighting that neuropathy presents more widely in mitochondrial disease.
Mutations of SPG7 are predominantly autosomal recessive; however, heterozygous SPG7 mutations have also been reported in association with diverse clinical manifestations such as optic atrophy, spastic paraplegia, and peripheral neuropathy [38]. The putative role of these heterozygous SPG7 variants in the aetiology of neuropathies remains controversial. In an attempt to elucidate this association further, we assessed the frequency of heterozygous SPG7 variants within our neuropathy cohort. We observed a reduced frequency of heterozygous SPG7 mutation carriers within our cohort compared to the rest of the samples we could access in GPAP. This decreased prevalence in our neuropathy cohort does not support the pathogenic role of heterozygous SPG7 mutations in the aetiology of mitochondrial neuropathies. However, we detected biallelic SPG7 variants in 3 patients suggest the association of SPG7 with autosomal recessive peripheral neuropathy.
By exploring the genetic landscape of peripheral neuropathies and identifying potential causative variants in genes with mitochondrial mechanisms, we contribute to a deeper understanding of the molecular mechanisms underlying this complex disease. Our findings underscore the utility of large-scale data analysis and the value of reanalysing exome/genome data over time as new genetic associations are reported and NGS technologies evolve. We also emphasise the importance of comprehensive phenotypic characterisation of patients by clinicians to assist future cohort studies and increase the yield of a genetic diagnosis.
Whole exome sequencing (WES) has emerged as a valuable diagnostic tool [18], but even with its application, a substantial number of patients remain undiagnosed. The diagnostic yield from WES varies depending on the setting and inclusion criteria, ranging from 15 to 60% [39]. To address this diagnostic gap, before performing new genomic testing (whole genome sequencing, RNA sequencing, proteomics, etc.), expert reanalysis of WES data holds promise for identifying missed or newly discovered genetic variants [18, 40].
This study has important limitations that should be acknowledged. Firstly, there is a possibility of missed patients due to incomplete or incorrect HPO term entry by the uploading clinician. Although efforts were made to ensure accurate phenotypic characterisation, variations in data entry may have resulted in some patients being inadvertently excluded from the analysis. Secondly, gene segregation could not be performed in cases where only index patient data were available, as opposed to a trio. Attempts were made to contact all centres whose variants were included in this study to ascertain if the patient had been previously diagnosed or to discuss the variant identified. Moreover, the platform does not currently enable the efficient cohort-search feature for mitochondrial encoded genes, meaning it was necessary to select individual variants within the mitochondrial genome, which could lead to potential omissions.
Additionally, despite the potential benefits of reanalysis, several challenges and limitations need to be considered. The interpretation of variants can be complex and may vary between different laboratories or databases, leading to discrepancies in variant classification. Updates in variant databases and evolving guidelines for variant interpretation also demand regular re-evaluation of previously reported variants. Furthermore, the identification of disease-causing genes or variants may require functional studies or additional evidence beyond bioinformatic analysis. Lastly, the dynamic nature of genetic research requires continuous data monitoring and re-evaluation as new genetic associations, and disease mechanisms emerge. Despite these challenges, reanalysis provides an opportunity to enhance diagnostic yield and contribute to the evolving understanding of rare diseases, ultimately improving patient care and management.
In conclusion, inherited neuropathies are rare and complex disorders affecting the peripheral nerves, and mitochondria play a crucial role in the disease. This study shows how frequently is CMT caused by mutations in primary mitochondrial disease genes. This study demonstrates the potential of RD-Connect as a powerful tool for unravelling the underlying causes of rare diseases, including inherited neuropathies, and highlights the importance of international collaboration to improve the diagnosis and treatment of rare diseases.
Data availability
All genomic data are deposited in the RD-Connect Genome Phenome Analysis Database (GPAP).
References
Hicks CW, Selvin E (2019) Epidemiology of peripheral neuropathy and lower extremity disease in diabetes. Curr DiabRep 19:1–8
Martyn C, Hughes R (1997) Epidemiology of peripheral neuropathy. J Neurol Neurosurg Psychiatry 62(4):310
Eggermann K, Gess B, Häusler M, Weis J, Hahn A, Kurth I (2018) Hereditary neuropathies: clinical presentation and genetic panel diagnosis. Dtsch Arztebl Int 115(6):91
Morena J, Gupta A, Hoyle JC (2019) Charcot-Marie-Tooth: from molecules to therapy. Int J Mol Sci 20(14):3419
Horvath R, Medina J, Reilly MM, Shy ME, Zuchner S (2023) Peripheral neuropathy in mitochondrial disease. Handbook of clinical neurology. 194: Elsevier, p. 99–116.
Cortese A, Wilcox JE, Polke JM, Poh R, Skorupinska M, Rossor AM et al (2020) Targeted next-generation sequencing panels in the diagnosis of Charcot-Marie-tooth disease. Neurology 94(1):e51–e61
Fridman V, Bundy B, Reilly M, Pareyson D, Bacon C, Burns J et al (2015) CMT subtypes and disease burden in patients enrolled in the inherited neuropathies consortium natural history study: a cross-sectional analysis. J Neurol Neurosurg Psychiatry 86(8):873–878
Mandal A, Drerup CM (2019) Axonal transport and mitochondrial function in neurons. Front Cell Neurosci 13:373
Pareyson D, Piscosquito G, Moroni I, Salsano E, Zeviani M (2013) Peripheral neuropathy in mitochondrial disorders. Lancet Neurol 12(10):1011–1024
Finsterer J (2005) Mitochondrial neuropathy. Clin Neurol Neurosurg 107(3):181–186
McFarland R, Taylor RW, Turnbull DM (2010) A neurological perspective on mitochondrial disease. Lancet Neurol 9(8):829–840
Wenger AM, Guturu H, Bernstein JA, Bejerano G (2017) Systematic reanalysis of clinical exome data yields additional diagnoses: implications for providers. Genet Med 19(2):209–214
Al-Nabhani M, Al-Rashdi S, Al-Murshedi F, Al-Kindi A, Al-Thihli K, Al-Saegh A et al (2018) Reanalysis of exome sequencing data of intellectual disability samples: yields and benefits. Clin Genet 94(6):495–501
Initiative EG, Berkovic SF, Goldstein DB, Heinzen EL, Laughlin BL, Lowenstein DH et al (2019) The epilepsy genetics initiative: systematic reanalysis of diagnostic exomes increases yield. Epilepsia 60(5):797–806
Al-Murshedi F, Meftah D, Scott P (2019) Underdiagnoses resulting from variant misinterpretation: time for systematic reanalysis of whole exome data? Eur J Med Genet 62(1):39–43
Baker SW, Murrell JR, Nesbitt AI, Pechter KB, Balciuniene J, Zhao X et al (2019) Automated clinical exome reanalysis reveals novel diagnoses. J Mol Diagn 21(1):38–48
Li J, Gao K, Yan H, Xiangwei W, Liu N, Wang T et al (2019) Reanalysis of whole exome sequencing data in patients with epilepsy and intellectual disability/mental retardation. Gene 700:168–175
Liu P, Meng L, Normand EA, Xia F, Song X, Ghazi A et al (2019) Reanalysis of clinical exome sequencing data. N Engl J Med 380(25):2478–2480
Schmitz-Abe K, Li Q, Rosen SM, Nori N, Madden JA, Genetti CA et al (2019) Unique bioinformatic approach and comprehensive reanalysis improve diagnostic yield of clinical exomes. Eur J Hum Genet 27(9):1398–1405
Jalkh N, Corbani S, Haidar Z, Hamdan N, Farah E, Abou Ghoch J et al (2019) The added value of WES reanalysis in the field of genetic diagnosis: lessons learned from 200 exomes in the Lebanese population. BMC Med Genom 12(1):1–7
Salfati EL, Spencer EG, Topol SE, Muse ED, Rueda M, Lucas JR et al (2019) Re-analysis of whole-exome sequencing data uncovers novel diagnostic variants and improves molecular diagnostic yields for sudden death and idiopathic diseases. Genome Med 11(1):1–8
Tan NB, Stapleton R, Stark Z, Delatycki MB, Yeung A, Hunter MF et al (2020) Evaluating systematic reanalysis of clinical genomic data in rare disease from single center experience and literature review. Mol Genet Genom Med 8(11):e1508
Denommé-Pichon A-S, Matalonga L, de Boer E, Jackson A, Benetti E, Banka S et al (2023) A Solve-RD ClinVar-based reanalysis of 1522 index cases from ERN-ITHACA reveals common pitfalls and misinterpretations in exome sequencing. Genet Med 25(4):100018
James KN, Clark MM, Camp B, Kint C, Schols P, Batalov S et al (2020) Partially automated whole-genome sequencing reanalysis of previously undiagnosed pediatric patients can efficiently yield new diagnoses. NPJ Genom Med 5(1):33
Matalonga L, Hernández-Ferrer C, Piscia D, Schüle R, Synofzik M, Töpf A et al (2021) Solving patients with rare diseases through programmatic reanalysis of genome-phenome data. Eur J Hum Genet 29(9):1337–1347
Ji J, Leung ML, Baker S, Deignan JL, Santani A (2021) Clinical exome reanalysis: current practice and beyond. Mol Diagn Ther 25:529–536
Wright CF, McRae JF, Clayton S, Gallone G, Aitken S, FitzGerald TW et al (2018) Making new genetic diagnoses with old data: iterative reanalysis and reporting from genome-wide data in 1133 families with developmental disorders. Genet Med 20(10):1216–1223
Nambot S, Thevenon J, Kuentz P, Duffourd Y, Tisserant E, Bruel A-L et al (2018) Clinical whole-exome sequencing for the diagnosis of rare disorders with congenital anomalies and/or intellectual disability: substantial interest of prospective annual reanalysis. Genet Med 20(6):645–654
Ewans LJ, Schofield D, Shrestha R, Zhu Y, Gayevskiy V, Ying K et al (2018) Whole-exome sequencing reanalysis at 12 months boosts diagnosis and is cost-effective when applied early in Mendelian disorders. Genet Med 20(12):1564–1574
Deignan JL, Chung WK, Kearney HM, Monaghan KG, Rehder CW, Chao EC et al (2019) Points to consider in the reevaluation and reanalysis of genomic test results: a statement of the American college of medical genetics and genomics (ACMG). Genet Med 21(6):1267–1270
Laurie S, Piscia D, Matalonga L, Corvó A, Fernández-Callejo M, Garcia-Linares C et al (2022) The RD-connect genome-phenome analysis platform: accelerating diagnosis, research, and gene discovery for rare diseases. Hum Mutat 43(6):717–733
Köhler S, Carmody L, Vasilevsky N, Jacobsen JOB, Danis D, Gourdine J-P et al (2019) Expansion of the human phenotype ontology (HPO) knowledge base and resources. Nucleic Acids Res 47(D1):D1018–D1027
Nguengang Wakap S, Lambert DM, Olry A, Rodwell C, Gueydan C, Lanneau V et al (2020) Estimating cumulative point prevalence of rare diseases: analysis of the orphanet database. Eur J Hum Genet 28(2):165–173
Amberger JS, Bocchini CA, Schiettecatte F, Scott AF, Hamosh A (2015) OMIM. org: online mendelian inheritance in man (OMIM®), an online catalog of human genes and genetic disorders. Nucleic Acids Res 43(D1):D789–D798
De Ligt J, Willemsen MH, Van Bon BW, Kleefstra T, Yntema HG, Kroes T et al (2012) Diagnostic exome sequencing in persons with severe intellectual disability. N Engl J Med 367(20):1921–1929
Thevenon J, Duffourd Y, Masurel-Paulet A, Lefebvre M, Feillet F, El Chehadeh-Djebbar S et al (2016) Diagnostic odyssey in severe neurodevelopmental disorders: toward clinical whole-exome sequencing as a first-line diagnostic test. Clin Genet 89(6):700–707
Laurie S, Fernandez-Callejo M, Marco-Sola S, Trotta JR, Camps J, Chacón A et al (2016) From wet-lab to variations: concordance and speed of bioinformatics pipelines for whole genome and whole exome sequencing. Hum Mutat 37(12):1263–1271
Sánchez-Ferrero E, Coto E, Beetz C, Gamez J, Corao A, Diaz M et al (2013) SPG7 mutational screening in spastic paraplegia patients supports a dominant effect for some mutations and a pathogenic role for p. A510V. Clin Genet 83(3):257–262
Wise AL, Manolio TA, Mensah GA, Peterson JF, Roden DM, Tamburro C et al (2019) Genomic medicine for undiagnosed diseases. The Lancet 394(10197):533–540
Kaplanis J, Samocha KE, Wiel L, Zhang Z, Arvai KJ, Eberhardt RY et al (2020) Evidence for 28 genetic disorders discovered by combining healthcare and research data. Nature 586(7831):757–762
Acknowledgements
T.F. received support from Queens’ College, University of Cambridge, the Wolfson Foundation, and the Royal College of Physicians. R.H. is a Wellcome Trust Investigator (109915/Z/15/Z, and 226653/Z/22/Z), who receives support from the Medical Research Council (UK) (MR/V009346/1), the Addenbrooke’s Charitable Trust (G100142), the Evelyn Trust, the Stoneygate Trust, the Lily Foundation, Action for AT, Muscular Dystrophy UK, and an MRC strategic award to establish an International Centre for Genomic Medicine in Neuromuscular Diseases (ICGNMD) MR/S005021/1. This research was supported by the NIHR Cambridge Biomedical Research Centre (BRC-1215-20014). The views expressed are those of the authors and not necessarily those of the NIHR or the Department of Health and Social Care. This study was further supported by the Horizon 2020 research and innovation programme via grant 779257 “Solve-RD”. Data were analysed using the RD-Connect Genome-Phenome Analysis Platform developed under FP7/2007-2013 funded project (grant agreement nº 305444) and funding from European Joint Programme in Rare Disease (EJP-RD) and INB/ELIXIR-ES. HL receives support from the Canadian Institutes of Health Research (CIHR) for Foundation Grant FDN-167281 (Precision Health for Neuromuscular Diseases), Transnational Team Grant ERT-174211 (ProDGNE), and Network Grant OR2-189333 (NMD4C), from the Canada Foundation for Innovation (CFI-JELF 38412), the Canada Research Chairs Program (Canada Research Chair in Neuromuscular Genomics and Health, 950-232279), the European Commission (Grant # 101080249), and the Canada Research Coordinating Committee New Frontiers in Research Fund (NFRFG-2022-00033) for SIMPATHIC, and from the Government of Canada, Canada First Research Excellence Fund (CFREF) for the Brain-Heart Interconnectome (CFREF-2022-00007). KP holds a CIHR postdoctoral fellowship award under grant no. MFE-491707. This study makes use of data and tools shared/provided through the RD-Connect GPAP, which received funding originally from the European Union Seventh Framework Programme (FP7/2007–2013) under grant agreement No. 305444.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
The authors have no financial or non-financial interest that are directly or indirectly related to the work submitted for publication.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ferreira, T., Polavarapu, K., Olimpio, C. et al. Variants in mitochondrial disease genes are common causes of inherited peripheral neuropathies. J Neurol 271, 3546–3553 (2024). https://doi.org/10.1007/s00415-024-12319-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00415-024-12319-y