Abstract
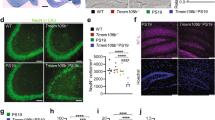
TMEM106B is a risk modifier of multiple neurological conditions, where a single coding variant and multiple non-coding SNPs influence the balance between susceptibility and resilience. Two key questions that emerge from past work are whether the lone T185S coding variant contributes to protection, and if the presence of TMEM106B is helpful or harmful in the context of disease. Here, we address both questions while expanding the scope of TMEM106B study from TDP-43 to models of tauopathy. We generated knockout mice with constitutive deletion of TMEM106B, alongside knock-in mice encoding the T186S knock-in mutation (equivalent to the human T185S variant), and crossed both with a P301S transgenic tau model to study how these manipulations impacted disease phenotypes. We found that TMEM106B deletion accelerated cognitive decline, hind limb paralysis, tau pathology, and neurodegeneration. TMEM106B deletion also increased transcriptional correlation with human AD and the functional pathways enriched in KO:tau mice aligned with those of AD. In contrast, the coding variant protected against tau-associated cognitive decline, synaptic impairment, neurodegeneration, and paralysis without affecting tau pathology. Our findings reveal that TMEM106B is a critical safeguard against tau aggregation, and that loss of this protein has a profound effect on sequelae of tauopathy. Our study further demonstrates that the coding variant is functionally relevant and contributes to neuroprotection downstream of tau pathology to preserve cognitive function.







Similar content being viewed by others
Data availability
RNA-seq data from this study was deposited at Gene Expression Omnibus (GEO) (http://www.ncbi.nlm.nih.gov/geo/) under accession ID GSE223376.
References
Allen M, Carrasquillo MM, Funk C, Heavner BD, Zou F, Younkin CS et al (2016) Human whole genome genotype and transcriptome data for Alzheimer’s and other neurodegenerative diseases. Sci Data 3:160089. https://doi.org/10.1038/sdata.2016.89
Bennett DA, Schneider JA, Arvanitakis Z, Wilson RS (2012) Overview and findings from the religious orders study. Curr Alzheimer Res 9:628–645
Briggs DI, Defensor E, Memar Ardestani P, Yi B, Halpain M, Seabrook G et al (2017) Role of endoplasmic reticulum stress in learning and memory Impairment and Alzheimer’s disease-Like neuropathology in the PS19 and APP(Swe) mouse models of tauopathy and amyloidosis. ENeuro. https://doi.org/10.1523/ENEURO.0025-17.2017
Brody AH, Nies SH, Guan F, Smith LM, Mukherjee B, Salazar SA et al (2022) Alzheimer risk gene product Pyk2 suppresses tau phosphorylation and phenotypic effects of tauopathy. Mol Neurodegener 17:32. https://doi.org/10.1186/s13024-022-00526-y
Cabron AS, Borgmeyer U, Richter J, Peisker H, Gutbrod K, Dormann P et al (2023) Lack of a protective effect of the Tmem106b “protective SNP” in the Grn knockout mouse model for frontotemporal lobar degeneration. Acta Neuropathol Commun 11:21. https://doi.org/10.1186/s40478-023-01510-3
Chang A, Xiang X, Wang J, Lee C, Arakhamia T, Simjanoska M et al (2022) Homotypic fibrillization of TMEM106B across diverse neurodegenerative diseases. Cell 185:1346-1355.e1315. https://doi.org/10.1016/j.cell.2022.02.026
Chen-Plotkin AS, Unger TL, Gallagher MD, Bill E, Kwong LK, Volpicelli-Daley L et al (2012) TMEM106B, the risk gene for frontotemporal dementia, is regulated by the microRNA-132/212 cluster and affects progranulin pathways. J Neurosci 32:11213–11227. https://doi.org/10.1523/JNEUROSCI.0521-12.2012
Colonna M (2023) The biology of TREM receptors. Nat Rev Immunol. https://doi.org/10.1038/s41577-023-00837-1
De Jager PL, Ma Y, McCabe C, Xu J, Vardarajan BN, Felsky D et al (2018) A multi-omic atlas of the human frontal cortex for aging and Alzheimer’s disease research. Sci Data 5:180142. https://doi.org/10.1038/sdata.2018.142
Etelainen TS, Silva MC, Uhari-Vaananen JK, De Lorenzo F, Jantti MH, Cui H et al (2023) A prolyl oligopeptidase inhibitor reduces tau pathology in cellular models and in mice with tauopathy. Sci Transl Med. https://doi.org/10.1126/scitranslmed.abq2915
Feng T, Lacrampe A, Hu F (2021) Physiological and pathological functions of TMEM106B: a gene associated with brain aging and multiple brain disorders. Acta Neuropathol 141:327–339. https://doi.org/10.1007/s00401-020-02246-3
Feng T, Luan L, Katz II, Ullah M, Van Deerlin VM, Trojanowski JQ et al (2022) TMEM106B deficiency impairs cerebellar myelination and synaptic integrity with Purkinje cell loss. Acta Neuropathol Commun 10:33. https://doi.org/10.1186/s40478-022-01334-7
Feng T, Mai S, Roscoe JM, Sheng RR, Ullah M, Zhang J et al (2020) Loss of TMEM106B and PGRN leads to severe lysosomal abnormalities and neurodegeneration in mice. EMBO Rep 21:e50219. https://doi.org/10.15252/embr.202050219
Feng T, Sheng RR, Solé-Domènech S, Ullah M, Zhou X, Mendoza CS et al (2020) A role of the frontotemporal lobar degeneration risk factor TMEM106B in myelination. Brain. https://doi.org/10.1093/brain/awaa154
Fowler SW, Chiang AC, Savjani RR, Larson ME, Sherman MA, Schuler DR et al (2014) Genetic modulation of soluble abeta rescues cognitive and synaptic impairment in a mouse model of Alzheimer’s disease. J Neurosci 34:7871–7885. https://doi.org/10.1523/JNEUROSCI.0572-14.2014
Gallagher MD, Posavi M, Huang P, Unger TL, Berlyand Y, Gruenewald AL et al (2017) A dementia-associated risk variant near TMEM106B alters chromatin architecture and gene expression. Am J Hum Genet 101:643–663. https://doi.org/10.1016/j.ajhg.2017.09.004
Gotzl JK, Mori K, Damme M, Fellerer K, Tahirovic S, Kleinberger G et al (2014) Common pathobiochemical hallmarks of progranulin-associated frontotemporal lobar degeneration and neuronal ceroid lipofuscinosis. Acta Neuropathol 127:845–860. https://doi.org/10.1007/s00401-014-1262-6
Jiang YX, Cao Q, Sawaya MR, Abskharon R, Ge P, DeTure M et al (2022) Amyloid fibrils in FTLD-TDP are composed of TMEM106B and not TDP-43. Nature 605:304–309. https://doi.org/10.1038/s41586-022-04670-9
Jiao HS, Yuan P, Yu JT (2023) TMEM106B aggregation in neurodegenerative diseases: linking genetics to function. Mol Neurodegener 18:54. https://doi.org/10.1186/s13024-023-00644-1
Jun MH, Han JH, Lee YK, Jang DJ, Kaang BK, Lee JA (2015) TMEM106B, a frontotemporal lobar dementia (FTLD) modifier, associates with FTD-3-linked CHMP2B, a complex of ESCRT-III. Mol Brain 8:85. https://doi.org/10.1186/s13041-015-0177-z
Keren-Shaul H, Spinrad A, Weiner A, Matcovitch-Natan O, Dvir-Szternfeld R, Ulland TK et al (2017) A unique microglia type associated with restricting development of Alzheimer’s disease. Cell 169:1276-1290.e1217. https://doi.org/10.1016/j.cell.2017.05.018
Klein ZA, Takahashi H, Ma M, Stagi M, Zhou M, Lam TT et al (2017) Loss of TMEM106B ameliorates lysosomal and frontotemporal dementia-related phenotypes in progranulin-deficient mice. Neuron 95(281–296):e286. https://doi.org/10.1016/j.neuron.2017.06.026
Lewandoski M, Meyers EN, Martin GR (1997) Analysis of Fgf8 gene function in vertebrate development. Cold Spring Harb Symp Quant Biol 62:159–168
Luningschror P, Werner G, Stroobants S, Kakuta S, Dombert B, Sinske D et al (2020) The FTLD risk factor TMEM106B Regulates the transport of lysosomes at the axon initial segment of motoneurons. Cell Rep 30:3506-3519.e3506. https://doi.org/10.1016/j.celrep.2020.02.060
Mathys H, Peng Z, Boix CA, Victor MB, Leary N, Babu S et al (2023) Single-cell atlas reveals correlates of high cognitive function, dementia, and resilience to Alzheimer’s disease pathology. Cell 186:4365-4385.e4327. https://doi.org/10.1016/j.cell.2023.08.039
Min SW, Chen X, Tracy TE, Li Y, Zhou Y, Wang C et al (2015) Critical role of acetylation in tau-mediated neurodegeneration and cognitive deficits. Nat Med 21:1154–1162. https://doi.org/10.1038/nm.3951
Mootha VK, Lindgren CM, Eriksson KF, Subramanian A, Sihag S, Lehar J et al (2003) PGC-1alpha-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat Genet 34:267–273. https://doi.org/10.1038/ng1180
Mortazavi A, Williams BA, McCue K, Schaeffer L, Wold B (2008) Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat Methods 5:621–628. https://doi.org/10.1038/nmeth.1226
Mostafavi S, Gaiteri C, Sullivan SE, White CC, Tasaki S, Xu J et al (2018) A molecular network of the aging human brain provides insights into the pathology and cognitive decline of Alzheimer’s disease. Nat Neurosci 21:811–819. https://doi.org/10.1038/s41593-018-0154-9
Nicholson AM, Finch NA, Wojtas A, Baker MC, Perkerson RB 3rd, Castanedes-Casey M et al (2013) TMEM106B p. T185S regulates TMEM106B protein levels: implications for frontotemporal dementia. J Neurochem 126:781–791. https://doi.org/10.1111/jnc.12329
Nicholson AM, Rademakers R (2016) What we know about TMEM106B in neurodegeneration. Acta Neuropathol 132:639–651. https://doi.org/10.1007/s00401-016-1610-9
Nicholson AM, Zhou X, Perkerson RB, Parsons TM, Chew J, Brooks M et al (2018) Loss of Tmem106b is unable to ameliorate frontotemporal dementia-like phenotypes in an AAV mouse model of C9ORF72-repeat induced toxicity. Acta Neuropathol Commun 6:42. https://doi.org/10.1186/s40478-018-0545-x
Perneel J, Rademakers R (2022) Identification of TMEM106B amyloid fibrils provides an updated view of TMEM106B biology in health and disease. Acta Neuropathol 144:807–819. https://doi.org/10.1007/s00401-022-02486-5
Rademakers R, Nicholson AM, Ren Y, Koga S, Nguyen HP, Brooks M et al (2021) Loss of Tmem106b leads to cerebellum Purkinje cell death and motor deficits. Brain Pathol 31:e12945. https://doi.org/10.1111/bpa.12945
Robinson MD, McCarthy DJ, Smyth GK (2010) edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26:139–140. https://doi.org/10.1093/bioinformatics/btp616
Rodriguez CI, Buchholz F, Galloway J, Sequerra R, Kasper J, Ayala R et al (2000) High-efficiency deleter mice show that FLPe is an alternative to Cre-loxP. Nat Genet 25:139–140. https://doi.org/10.1038/75973
Schweighauser M, Arseni D, Bacioglu M, Huang M, Lovestam S, Shi Y et al (2022) Age-dependent formation of TMEM106B amyloid filaments in human brains. Nature 605:310–314. https://doi.org/10.1038/s41586-022-04650-z
Stroobants S, D’Hooge R, Damme M (2021) Aged Tmem106b knockout mice display gait deficits in coincidence with Purkinje cell loss and only limited signs of non-motor dysfunction. Brain Pathol 31:223–238. https://doi.org/10.1111/bpa.12903
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA et al (2005) Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci U S A 102:15545–15550. https://doi.org/10.1073/pnas.0506580102
Takeuchi H, Iba M, Inoue H, Higuchi M, Takao K, Tsukita K et al (2011) P301S mutant human tau transgenic mice manifest early symptoms of human tauopathies with dementia and altered sensorimotor gating. PLoS ONE 6:e21050. https://doi.org/10.1371/journal.pone.0021050
Van Deerlin VM, Sleiman PM, Martinez-Lage M, Chen-Plotkin A, Wang LS, Graff-Radford NR et al (2010) Common variants at 7p21 are associated with frontotemporal lobar degeneration with TDP-43 inclusions. Nat Genet 42:234–239. https://doi.org/10.1038/ng.536
van der Touw W, Chen HM, Pan PY, Chen SH (2017) LILRB receptor-mediated regulation of myeloid cell maturation and function. Cancer Immunol Immunother 66:1079–1087. https://doi.org/10.1007/s00262-017-2023-x
van der Zee J, Van Langenhove T, Kleinberger G, Sleegers K, Engelborghs S, Vandenberghe R et al (2011) TMEM106B is associated with frontotemporal lobar degeneration in a clinically diagnosed patient cohort. Brain 134:808–815. https://doi.org/10.1093/brain/awr007
Wan YW, Al-Ouran R, Mangleburg CG, Perumal TM, Lee TV, Allison K et al (2020) Meta-analysis of the Alzheimer’s disease human brain transcriptome and functional dissection in mouse models. Cell Rep 32:107908. https://doi.org/10.1016/j.celrep.2020.107908
Wang M, Beckmann ND, Roussos P, Wang E, Zhou X, Wang Q et al (2018) The Mount Sinai cohort of large-scale genomic, transcriptomic and proteomic data in Alzheimer’s disease. Sci Data 5:180185. https://doi.org/10.1038/sdata.2018.185
Werner G, Damme M, Schludi M, Gnorich J, Wind K, Fellerer K et al (2020) Loss of TMEM106B potentiates lysosomal and FTLD-like pathology in progranulin-deficient mice. EMBO Rep 21:e50241. https://doi.org/10.15252/embr.202050241
Yanamandra K, Kfoury N, Jiang H, Mahan TE, Ma S, Maloney SE et al (2013) Anti-tau antibodies that block tau aggregate seeding in vitro markedly decrease pathology and improve cognition in vivo. Neuron 80:402–414. https://doi.org/10.1016/j.neuron.2013.07.046
Yoshiyama Y, Higuchi M, Zhang B, Huang SM, Iwata N, Saido TC et al (2007) Synapse loss and microglial activation precede tangles in a P301S tauopathy mouse model. Neuron 53:337–351. https://doi.org/10.1016/j.neuron.2007.01.010
Zhang B, Carroll J, Trojanowski JQ, Yao Y, Iba M, Potuzak JS et al (2012) The microtubule-stabilizing agent, epothilone D, reduces axonal dysfunction, neurotoxicity, cognitive deficits, and Alzheimer-like pathology in an interventional study with aged tau transgenic mice. J Neurosci 32:3601–3611. https://doi.org/10.1523/JNEUROSCI.4922-11.2012
Zhang T, Pang W, Feng T, Guo J, Wu K, Nunez Santos M et al (2023) TMEM106B regulates microglial proliferation and survival in response to demyelination. Sci Adv. https://doi.org/10.1126/sciadv.add2676
Zhang Y, Parmigiani G, Johnson WE (2020) ComBat-seq: batch effect adjustment for RNA-seq count data. NAR Genom Bioinform. https://doi.org/10.1093/nargab/lqaa078
Zhang Z, Song M, Liu X, Kang SS, Kwon IS, Duong DM et al (2014) Cleavage of tau by asparagine endopeptidase mediates the neurofibrillary pathology in Alzheimer’s disease. Nat Med 20:1254–1262. https://doi.org/10.1038/nm.3700
Zhou X, Brooks M, Jiang P, Koga S, Zuberi AR, Baker MC et al (2020) Loss of Tmem106b exacerbates FTLD pathologies and causes motor deficits in progranulin-deficient mice. EMBO Rep 21:e50197. https://doi.org/10.15252/embr.202050197
Zhou X, Nicholson AM, Ren Y, Brooks M, Jiang P, Zuberi A et al (2020) Loss of TMEM106B leads to myelination deficits: implications for frontotemporal dementia treatment strategies. Brain 143:1905–1919. https://doi.org/10.1093/brain/awaa141
Zhu XC, Yu JT, Jiang T, Wang P, Cao L, Tan L (2015) CR1 in Alzheimer’s disease. Mol Neurobiol 51:753–765. https://doi.org/10.1007/s12035-014-8723-8
Acknowledgements
We thank Philip De Jager for sharing then-unpublished data on the link between TMEM106B and cognitive resilience that sparked our original interest in this protein. We thank Chelsea Jiaqi Zong, Zoe Lai, Jennifer Saldana, and Melissa Comstock for support with the mouse colonies; Gabriella Perez for help with imaging, Cecilia Ljungberg for help with slide scanning, I-Chih Tan for equipment support and repair, and Denise Lanza, Lan Liao, and Jason Heaney for help with mouse creation and cryorecovery. TMEM106B knockout germplasm was provided by the KnockOut Mouse Project (KOMP) Repository and the Mouse Biology Program at the University of California Davis from ES cells generated by the Wellcome Trust Sanger Institute. The results published here are in whole or in part based on data obtained from the AD Knowledge Portal (https://adknowledgeportal.org). The specific human AD RNAseq datasets used here were generated by: 1) the Rush Alzheimer’s Disease Center, Rush University Medical Center, Chicago. 2) Dr. Allan Levey from Emory University based on postmortem brain tissue collected by Dr. Eric Schadt from the Mount Sinai School of Medicine through the Mount Sinai VA Medical Center Brain Bank, and 3) the Mayo RNAseq study led by Dr. Nilüfer Ertekin-Taner, Mayo Clinic, Jacksonville, FL.
Funding
This work was supported by National Institutes of Health grants R21AG056028, RF1AG058188, and P01AG066606-01A1 and -S1 (JLJ), R01AG074009 (IAR and JLJ), F31AG067676-01A1 (CAW), and T32NS043124 (support for GAE), Alzheimer’s Association grant ZEN-19–591129 (JLJ), and by Federal Work Study funds to BCM (support for QN and PJK). This work was assisted by several advanced technology core laboratories at BCM, including the RNA In Situ Hybridization Core, Bioengineering Core, and Genetically Engineered Rodent Models Core. BCM core laboratories were funded in part by National Institutes of Health grants S10 OD016167 and P50 HD103555 (RNA ISH), P30 CA125123 (GERM), and P30 EY002520 (Bioeng).
Author information
Authors and Affiliations
Contributions
JLJ and IAR conceptualized the study; GAE, YH, QN, KWP, RGG, CZ, and PJK performed experimental investigation; GAE, CAW, QN, and IAR contributed to formal analysis; JLJ, CAW, and IAR wrote the paper; JLJ, CAW, YX, and IAR obtained funding for the work.
Corresponding author
Ethics declarations
Conflict of interest
CZ is a paid employee of NeuroScience Associates. The remaining authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Edwards, G.A., Wood, C.A., He, Y. et al. TMEM106B coding variant is protective and deletion detrimental in a mouse model of tauopathy. Acta Neuropathol 147, 61 (2024). https://doi.org/10.1007/s00401-024-02701-5
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00401-024-02701-5