Abstract
We reconstructed the natural history and temporal evolution of the most common childhood brain malignancy, medulloblastoma, by single-cell whole-genome sequencing (sc-WGS) of tumours representing its major molecular sub-classes and clinical risk groups. Favourable-risk disease sub-types assessed (MBWNT and infant desmoplastic/nodular MBSHH) typically comprised a single clone with no evidence of further evolution. In contrast, highest risk sub-classes (MYC-amplified MBGroup3 and TP53-mutated MBSHH) were most clonally diverse and displayed gradual evolutionary trajectories. Clinically adopted biomarkers (e.g. chromosome 6/17 aberrations; CTNNB1/TP53 mutations) were typically early-clonal/initiating events, exploitable as targets for early-disease detection; in analyses of spatially distinct tumour regions, a single biopsy was sufficient to assess their status. Importantly, sc-WGS revealed novel events which arise later and/or sub-clonally and more commonly display spatial diversity; their clinical significance and role in disease evolution post-diagnosis now require establishment. These findings reveal diverse modes of tumour initiation and evolution in the major medulloblastoma sub-classes, with pathogenic relevance and clinical potential.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Medulloblastoma (MB) is the most common malignant brain tumour of childhood. Approximately 30% of patients die from their disease and survivors are left with life-long disease and treatment-associated morbidities [31]. Diversity between individual MBs has long been recognised, encompassing established disease molecular groups (i.e. MBWNT, MBSHH, MBGroup3 and MBGroup4) and subgroups (i.e. MBGroup3-4 subgroups I–VIII) with distinct clinico-molecular characteristics, alongside well-defined genomic features at the bulk-tumour level [3, 14, 22, 30, 31, 42, 43]. Critically, this inter-tumoural diversity has been exploited clinically; specific molecular sub-classes define risk-associated therapy stratification in current clinical trials. MBWNT tumours, and infant MBSHH tumours with desmoplastic/nodular (DN) pathology, are associated with high survival rates (i.e. favourable-risk) and receive reduced-intensity therapies, while MYC-amplified MBGroup3 and TP53-mutated MBSHH carry the highest risk and gravest prognoses [14, 31].
The potential presence of clinically significant cellular heterogeneity within individual MBs was first suggested by studies which demonstrated intra-tumoural variation of specific genetic defects at the single-cell level (e.g. MYC amplification [8, 11, 41]). Moreover, multiple sampling of bulk MB tumours has suggested spatial heterogeneity of genetic markers within individual tumours, questioning whether a single diagnostic biopsy captures all necessary genetic information for use in clinical management [28]. Systematic single-cell studies of MB are essential to further explore these issues. Studies to date have, however, focussed on intra-tumoural transcriptional variability and its relationship to cellular origins and cerebellar development [15, 32, 46, 49], with only one specific study analysing genetic heterogeneity within two Li–Fraumeni-associated MBs [35]. The cellular genetic constituency of the major MB sub-classes, and its relationship to disease initiation, clonal and temporal evolution, thus remains poorly understood [8].
Here, we report the application of sc-WGS to reconstruct the development and clonal evolution of 14 MBs, encompassing sampling of two spatially distinct regions of each tumour. Unlike previous single-cell MB transcriptomic studies predicated on fresh tumour material [15, 32, 35, 46, 49], our use of fresh-frozen tumours enabled specific selection of molecularly and clinically phenotyped tumours representative of the major MB sub-classes and risk groups. Using these data, we derive a clinically focussed understanding of MB intra-tumoural heterogeneity, providing critical insights into differences in initiation, drivers, clonal evolution trajectories and spatial impact. Moreover, our findings highlight how understanding MB at the single-cell level offers significant potential to improve the clinical management of MB, through informing biomarkers for early disease detection, strategies for diagnostic sampling, and therapeutic targeting.
Methods
Sample selection
Fresh-frozen tumour material from 14 clinically annotated human primary MBs was selected for analysis, using extant molecular pathology data (Fig. 1; TP53/CTNNB1 mutation, MYC/N amplification and DNA methylation array (molecular subgroup and copy number profiles)) [42]. Samples were collected with written, informed consent from UK Children’s Cancer and Leukaemia Group (CCLG, https://www.cclg.org.uk/) institutions and collaborating centres (study approval reference BS-2008–12). Tumour investigations were performed with approval from Newcastle/North Tyneside Research Ethics Committee (study reference 07/Q0905/71). Central review of histological variants was performed according to 2016 World Health Organization (WHO) criteria [22]. Metastatic status was determined according to Chang's criteria [4]. Patients < 4.0 years of age at diagnosis were regarded as infants. Tumours were assigned to the four consensus disease molecular groups (MBWNT, MBSHH, MBGrp3 and MBGrp4) [22] using established DNA methylation array-based methods [42]. Second-generation MBSHH subgroups were assigned according to Schwalbe et al. 2017 [42]. Second-generation MBGrp3/4 subgroups [43] were assigned using the ‘Grp3 and Grp4 Classifier’ (https://www.molecularneuropathology.org/mnp/classifier/7). A cerebellar sample was obtained post-mortem from the Newcastle Brain Tissue Resource (https://nbtr.ncl.ac.uk/).
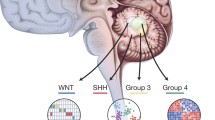
Single-cell analysis of childhood medulloblastoma: patient cohort and study design. a Selected cohort of 14 patients representing key medulloblastoma clinico-molecular sub-classes. Cohort encompasses all 4 molecular groups WNT (blue), SHH (light red—infant SHH and dark red—childhood SHH), Group3 (yellow) and Group4 (green), multiple subtypes and risk groups. MYC-amplified tumours with low A and high B percentages of amplified cells were selected for comparative analysis. b Study pipeline showing the main milestones. Scale bar on the tumour photographs equals 10 mm. Abbreviations: LCA large cell and anaplastic, CLA classic, DN desmoplastic/nodular, amp amplification, mut mutation, FACS fluorescence-activated single-cell sorting, WGA whole-genome amplification, WGS whole-genome sequencing, CNV copy number variant, Grp group, r1/2 region 1/2, FCS forward scatter, DOD dead of disease, ADF alive disease-free, DOOC dead of other cause
Two separate tissue pieces were selected from each tumour (received as two distinct surgical pieces), ranging from 2.4 × 2.6 mm (smallest) to 5.4 × 4.9 mm (largest) (Fig. 1) in size, to represent tissue fragment sizes typically used in diagnostic genetic analysis. For each region, histological sections confirmed material quality and 30 µm cryosections were produced for single-cell analyses; the remainder was used for comparative bulk tumour assessment.
Bulk tumour analysis
For each tumour, DNA was isolated (DNeasy Blood and Tissue Kit (Qiagen, Cat No. 69504) from the two bulk tumour regions, combined and subjected to deep (180x) whole-exome sequencing (WES). Sequencing libraries (Agilent SureSelect Human All Exon kit (Agilent Technologies; Santa Clara, CA, USA)), fragmentation (hydrodynamic shearing system (Covaris, Massachusetts, USA)), product purification (AMPure XP system (Beckman Coulter, Beverly, USA)) and quantification (Agilent Bioanalyzer 2100 system), followed by Illumina paired-end sequencing (Illumina; San Diego, CA, USA), were performed according to the manufacturer’s instructions. Raw reads were mapped to the GRCh37/hg19 reference genome.
Chromosome arm-level and focal copy number variations (CNVs) were assessed using Nexus Copy Number 10.0 (BioDiscovery, El Segundo, CA, USA) and validated by visual inspection. Copy number segments were classified as balanced, gained (log2 ratio > 0.2) or lost (log2 ratio < − 0.2) [30]. Whole arm chromosomal changes were considered gained/lost when ≥ 90% of the probe signals exceeded defined thresholds [14].
Mutational analysis interrogated 192 genes previously either reported as frequently mutated driver genes in medulloblastoma or recognised to be acquired in relapsed disease [25, 29, 30, 38]. Next-generation sequencing datasets were analysed using Genome Analysis Toolkit (GATK) version 3.8, according to Broad Institute’s best practices (Burrows-Wheeler alignment, Samtools, Picard, Haplotype Caller, MuTect, ANNOVAR) [5, 10, 20, 21, 47]. Variants were predicted pathogenic if their consequence included coding or splice donor/acceptor mutations, ExAC and 1000 genomes frequency < 0.01, included stop codon or frameshift variants, were predicted to be deleterious by SIFT/Polyphen and FATHMM or were known mutations in MB [44]. Where germline data were available only somatic variants were reported. Variants with a bidirectional read depth ≥ 5 were further curated by visual inspection in Integrative Genomics Viewer (IGV) [39].
Single-cell isolation and DNA sequencing
Single nuclei were isolated using standard methods and FACS sorting [2], followed by DNA extraction and whole-genome amplification (WGA) using the GenomePlex WGA4 kit (Sigma-Aldrich) according to the manufacturer’s protocol. DNA purification was performed according to Macaulay et al. [23]. An additional single-cell DNA quality control of WGA DNA fidelity by multiplex PCR-based analysis of multiple differentially sized amplification products was included as an adaptation to the protocol (Supplementary Methods Fig. 1). Cells with at least 4 products in multiplex PCRs were taken forward for library preparation.
Library preparation was performed using the Nextera XT 96-Index Kit (Illumina, cat. no. FC-131-1002) [23]. Shallow WGS was performed on NextSeq (Illumina), using 150 bp pair-end reads at the Sequencing Core Facility at Newcastle University to obtain target coverage of 0.15X for each single-cell library. Sequencing depth was determined from preliminary down-sampling experiments, as the optimal depth which produced findings consistent with both copy number aberrations detected at higher read-depths, and findings from parallel bulk tumour analysis (Supplementary Figs. 1 and 2). Data were processed using the CASAVA 1.8.1 pipeline (Illumina), and sequence reads were converted to fastq files. Sequencing reads from each single cell were demultiplexed using an in-house Perl script (demultiplex.pl) into independent fastq files, where each file represented the sequencing reads from one cell (https://emea.support.illumina.com/content/dam/illuminasupport/documents/documentation/ /bcl2fastq/bcl2fastq_letterbooklet_15038058brpmi.pdf). DNAs from 32 normal post-mortem cerebellar cells were sequenced as diploid controls, with 8 normal cells being sequenced per single-cell batch run.
Single-cell data analysis
Eight hundred sixty-eight tumour nuclei were sequenced. Raw reads in the fastq format [6] were processed and quality-checked using FastQC (Babraham Bioinformatics). 865 tumour nuclei passed the initial QC step. Illumina adapters were trimmed and reads with Phred-scale base qualities of 10 and below were filtered out using Cutadapt [24]. Raw reads were mapped to human reference genome version GRCh37/hg19 using BWA-MEM algorithm of Burrows–Wheeler Aligner (BWA) [20] version 0.7.17. The output SAM files were converted to compressed BAM files using Genome Analysis Toolkit [26]. Alignment summary reports were generated using SAMtools [21] flagstat function (detailed data quality scores used for the final selection of cells are shown in Supplementary table 1). Base quality recalibration was performed according to GATK guidelines.
Copy number aberrations were assessed from processed*. Bam files using the CNVKIT copy number analysis pipeline [45]; an on-target bin size of 50 kb was used for optimal resolution and segmentation. Coverages were estimated by calculating the mean read depth for each genomic bin. The MAD (median absolute deviation) quality metric was used to assess the noisiness of sequencing data. Genomes with two standard deviations below or above the mean bin count were excluded from further analyses, leaving 430 nuclei for further assessment of copy number calls.
A panel of normal cells was compiled as a copy number reference from 72 high-quality diploid (normal) genome sequences. Copy number regions with segmented log2 ratio estimates of − 1 and 1 (circular binary segmentation, p < 0.00001 [33]) were assigned as single copy gain and loss, respectively. Copy number region smoothing was performed by merging adjacent segments with the same absolute copy numbers or regions with non-significant difference in segment ratios. Copy number outputs were validated by visual assessment in IGV browser. CNVs were taken forward for analysis if detected in > 2 cells and with consistent genomic coordinates. Putative driver genes within focal and subchromosomal CNV regions identified were assessed using the Cancermine database [19].
Tumour ploidy was inferred using copy number relative ratios [2]. For tumour purity assessment, cells derived from aneuploid tumours (from bulk analysis), but with entirely balanced diploid single-cell copy number states, were considered normal/non-tumour.
Mutation analysis
Fifteen deleterious mutations observed in bulk tumour analysis were selected for further investigation in the single-cell DNA from their corresponding tumours using targeted Sanger sequencing, for inclusion in analyses of clonal composition and genomic evolution.
Clone identification
Numeric matrixes consisting of copy number segments were derived from the final copy number calls for each normal and tumour single-cell genome (https://bioconductor.org/packages/release/bioc/html/CNTools.html). Matrices were clustered using Euclidean distances for each genomic bin, hierarchical clustering and Ward linkage. Optimal clusters were calculated using multiple methods as Elbow, Silhouette and Gap statistics. Clustering heatmaps were visualised using heatmap.2 function of the gplot R package [48]. Clusters were further investigated using principal component analysis (PCA) using k = 2 dimensions and visualised applying the multidimensional scaling (MDS) R utility [7].
(Sub)clonality measures
CNVs and mutations were assigned as clonal when detected in each identified CNV sub-clone within a tumour, and subclonal when only detected in certain subclones. Mutational analyses assumed dropout of one allele at random following single-cell WGA [16].
Phylogenetic tree reconstruction
Neighbour-joining (NJ) trees were estimated from the phylo object constructed by computing pairwise Euclidean distances of the copy number segment numeric matrixes using nj function of ape R package [34]. All trees were re-rooted to a normal diploid cell matrix using the interactive root.phylo function. The final trees were plotted as downward-directed using plot.phylo function, and the edges were coloured according to the clone colour affiliations.
Modelling tumour genomic evolution
Tree formation (hierarchical clustering) and optimal clone numbers (multidimensional analysis) were imputed to infer evolutionary models using the SCICoNE package [18]. Individual tumour data were visualised using the “fishplot” R package [27] (https://www.rstudio.com/). Tumours were assigned to three modes of evolution: (i) no evolution, characterised by acquisition of established driver events and structural chromosomal alterations at initiation, leading to the development of a single, stable rapidly expanding clone, (ii) punctuated evolution, established features and structural chromosomal aberrations are acquired early and further copy number events accumulate in a punctuated clonal burst, consistent with clone formation and gradual stable expansion, (iii) gradual (branching) evolution, where established features and structural chromosomal aberrations are acquired at initiation, followed by the parallel formation of clones and their subsequent branching [9, 12].
Statistical analysis
Categorical variables were compared using Fisher’s exact tests, with a significance threshold of p < 0.05. p values were adjusted for multiple testing using “fdr” method (https://www.rdocumentation.org/packages/fda.usc/versions/2.0.2/topics/FDR). Analyses were performed using R.
Results
scDNA sequencing of medulloblastoma: patient selection and study design
To assess intra-tumoural genetic heterogeneity in the major sub-classes of MB, 14 primary tumours were selected, based on their bulk tumour profiles (Fig. 1a, Supplementary Figs. 1 and 2), to represent all four disease molecular groups and their characteristic genetic features: MBWNT, MBSHH (infant and childhood subgroups), Group3 (subgroups II and V) and Group 4 (subgroups VI, VII and VIII). Tumours further represented the major molecular risk-groups in current clinical use [22, 31], encompassing favourable (MBWNT and infant DN MBSHH), standard, high- and very high-risk tumours (MYC-amplified MBGroup3 and TP53-mutated MBSHH) [42, 43] (Fig. 1a), and recently described whole-chromosome aberration phenotypes (i.e. Chr 7 + . Chr8- and/or Chr 11-; Goschzik et al. [14]). The study design and workflow are summarised in Fig. 1b.
Tumour purity and risk-associated diversity in clonal substructure
We sequenced 868 nuclei from the selected cohort (62 cells per tumour). After performing quality controls (see Methods section), 430 high quality single-nuclei datasets were taken forward for analysis. All tumours had a high tumour cell content (i.e. aneuploid; mean, 95%; range, 82–100% of cells), with low proportions of normal diploid cells evident (Fig. 2a). To dissect genetic clonal substructure within each tumour, PCA/k-means (Fig. 2b) and Euclidean distance (Fig. 2c; Supplementary Fig. 3) clustering were independently applied to the single-cell CNV data. The number and composition of dissected clusters were concordant across both approaches; in each case, normal diploid cells formed a separate cluster, while cancer cell CNV profiles revealed diverse structures between tumours, ranging from 1 to 4 distinct clusters or subclones (Fig. 2b–c).
Tumour purity and clonal substructures in medulloblastoma. a Percentage of tumour (white; aneuploid) and normal (green; diploid) cells within each tumour. b Clonal sub-populations dissected by PCA/k-means clustering. Identified aneuploid clusters are labelled in grey, orange, blue and pink. Groups with a normal/diploid copy number profile are shown in green. c Clonal substructures dissected by hierarchical clustering (clustering dendrograms are presented in Supplementary Fig. 3). Three heatmaps represent the aneuploid copy number profiles for three patient groups with no, moderate or extensive clonal substructure. Single cells are plotted along the y-axis, and CNAs are plotted in genomic order along the x-axis. Clonal subpopulation identity (clone A, B, C or D), patient numbers and subgroup identifiers are shown on the left. Patient subtypes and risk profiles are indicated on the right. d Graph showing percentage of cells in each identified subpopulation. Abbreviations: PC principal component
Tumours could be assigned to 3 broad categories according to the number of sub-populations observed (Fig. 2b–d). First, low-complexity tumours with no evidence of genetic clonal substructure (n = 5), where all cells represented a single clone; notably, all were favourable-risk tumours (3 of 4 MBWNT tumours examined and both infant MBSHH tumours). Second, tumours with extensive clonal substructure (i.e. ≥ 3 sub-populations; n = 4); these were completely aligned with the very high-risk (VHR) tumours assessed (i.e. both MYC-amplified MBGroup3 and both TP53-mutated/MYCN-amplified MBSHH). All tumours examined with high-risk LCA pathology (3/3) also fell within this group. Finally, a group of tumours with 2 sub-populations, which represented multiple sub-classes (WNT, Grp3 and Grp4) and different non-VHR risk profiles. Statistically, low-complexity, wholly clonal tumours were significantly associated with favourable-risk disease (5/6 vs. 0/8, p = 0.003, Fisher’s exact test), while the highest risk tumour sub-classes assessed were associated with the most extensive sub-clonality (4/4 vs. 0/10, p = 0.001, Fisher’s exact test). Importantly, cell numbers analysed were not associated with the number of subclones we identified (data not shown). Based on the mean number of 31 cells analysed per tumour, and the criteria we set for subclone detection (i.e. reproducible copy number patterns in 3 independent cells), we estimate that our approach was sensitive to detect subclones present at a frequency greater than approximately 10%.
Uncovering clonal genetic composition and phylogenetic relationships
We dissected the CNV and mutational composition of each identified clone and their inter-relationships (Fig. 3, Supplementary Fig. 4, Supplementary Tables 2 and 3). As expected, the five tumours without evidence of substructure harboured a single homogeneous clone with shared genetic features. Marked clonal variation was observed within the remaining tumours; all displayed admixtures of clones with unique CNV and mutational content (Fig. 3). Applying neighbour-joining clustering, we determined the phylogenetic relationships between individual clones. In all cases, each identified subclone formed a distinct taxon but shared a common ancestor with other subclones in the same tumour (Fig. 3). Thus, identified clones were phylogenetically related, but the degree of relationship (i.e. the distances between different pairs of taxa) differed between tumours.
Content of clones and their phylogenetic inter-relationships. Clonal admixture plots for the three subclonal landscape categories are shown. Each distinct clone is represented in a different colour and proportional to the number of cells in which it was identified. Small, shaded squares and shaded labels on the phylogenetic trees represent the clone colour affiliation. CNVs and mutations defining more than one clone are listed beneath the shaded squares. Alterations unique to a single clone are labelled on the corresponding branches of phylogenetic trees. Established diagnostic and prognostic genomic features of medulloblastoma are shown in bold. GLI2 amplification was detected in a single cell, but is illustrated as a key biomarker, A—GLI2 amplification. B—MYC amplification was detected in 8/62 cells, but these did not pass QC and were excluded from analyses. C—MYC amplification in all clones. Abbreviations: FR favourable risk, SR standard risk, HR high risk, VHR very high risk, foc focal, sub subchromosomal
Medulloblastoma evolutionary trajectories
Modelling clonal evolution in our cohort using SCICoNE [18] revealed the three previously identified groups of tumours with different clonal substructures (Figs. 2c, 3) were associated with distinct evolutionary trajectories (p = 7.9 × 10–7; Fisher’s exact test) (Fig. 4). Favourable-risk tumours without clonal substructure showed no evidence of clonal genomic evolution (i.e. single-clone expansion), acquiring established driver events (e.g. CTNNB1 mutation in MBWNT; PTCH1 mutation in infant MBSHH) [31] and structural chromosomal alterations at initiation, associated with the development of a single, stable, rapidly expanding clone.
Evolutionary trajectories in medulloblastoma. Fish plots for groups of tumours associated with three different evolutionary trajectories. Plots display changes in clonal structure over pseudo-time, involving two time points, “Earlier” and “Later”. Consistent with previous figures, clonal admixtures are represented in the corresponding colours. Tumour initiating events, preceding subclone formation, are listed on the left (bracketed). Labelled shaded circles represent the events leading to subclone formation, and display their estimated time of origin. Acquired and lost alterations are shown at the bottom and top of the plot, respectively. The well-defined genomic biomarkers of medulloblastoma are shown in bold. See Figs. 1 and 3 for abbreviations
Tumours with a moderate clonal substructure followed a route of punctuated evolution. They obtained a number of established features (e.g. a combination of Chr7 + , Chr8– and Chr11− in the absence of gene-specific mutations) and structural chromosomal aberrations early in tumourigenesis [14, 31, 42]. The accumulation of further copy number events is then associated with a punctuated clonal burst, consistent with formation of both clones at similar time points, followed by gradual expansion and clonal stability over time.
The VHR tumours examined, with most extensive clonal substructures, best fitted a model of gradual (branching) evolution (Fig. 4). These tumours acquired high-risk drivers (e.g. TP53 mutation, MYC and/or MYCN amplifications), alongside multiple structural CNVs at medulloblastoma initiation. Further accumulation and loss of CNVs is consistent with ongoing instability and formation of clones, followed by their subsequent branching. These findings are consistent with the diminution of some clones over time, while others gradually expand. This evolutionary trajectory shows the highest complexity and magnitude of events, consistent with their aggressive clinical behaviour and low survival rates [14, 31, 42].
Clonality, detectability and timing of origin of individual events
We next assessed individual CNV and SNV mutations (Fig. 3), their clonality, and its relationship to their detection at the sc-WGS and bulk tumour levels (WES and DNA-methylation array data) (Fig. 5). First, we investigated diagnostic and prognostic disease biomarkers in current clinical use [31], or supported by trials-based studies [14]; these were assessed strictly within their relevant subgroup-specific context (Fig. 3a). The overwhelming majority (20/23; 87%) of such biomarkers observed were clonal, emerged early at initiation and were detected in all three datasets examined. The remaining three instances were sub-clonal and occurred later (MYCN and GLI2 amplifications in MBSHH patient 7 and Chr8- in MBGrp4 patient 14). Other clonal whole-arm/chromosome CNVs detected are summarised on Fig. 5b. Importantly, a series of sub-clonal CNVs were detected at the single-cell level; the majority were observed in small proportions of cells (mean 31% of cells, range from 12 to 85% of cells) and occurred later in tumourigenesis. They were predominantly undetectable (22/37) in both bulk tumour datasets (WES and DNA methylation array; Fig. 5c). Subchromosomal (> 15 Mb) and focal (< 15 Mb) CNVs detected at the single-cell level, and associated with putative driver genes, are summarised in Supplementary table 4; 15/19 of these were detectable at the bulk tumour level. Notably, the resolution of our analysis allowed identification of smaller focal CNVs. The smallest focal gains/losses reproducibly detected at our sequencing resolution (i.e. 0.15x) were, respectively, a 2.5 Mb GLI2 amplification (patient 7) and a 6.5 Mb deletion on chr8 (patient 8); the presence of both was confirmed in our bulk data sets. To conclude, 74% of the assessed CNVs (115/155) were clonal early events, thus, rather than occurring gradually over time, polyploidy is typically associated with medulloblastoma initiation.
Clonality, detectability and timing of established medulloblastoma biomarkers (a) diagnostic and prognostic disease biomarkers in current clinical use, or supported by trials-based studies; shown in bold) [14, 31]. Summaries of other whole-arm/chromosome and mutational clonal (b) and subclonal (c) medulloblastoma biomarkers. Specific biomarkers were assessed only in their defined subgroup-specific context. See Fig. 1 for abbreviations
Spatial heterogeneity at the cellular level
To assess any spatial differences in clonal substructure, we created clonal admixture plots reflecting the percentage content of each clone within the two discrete areas of each tumour assessed (Fig. 6a). Of the 9 tumours with clonal substructure, 5 had a detectable clone which was only present in a single sampled region of the tumour, and clonal content was significantly different between regions in two tumours (Fig. 6a and Supplementary Table 5a, b). Region-specific clones formed distinct taxa within their tumour region on phylogenetic reconstruction (Fig. 6a). As expected, the five tumours with no evidence of clonal substructure did not display any spatial clonal variation (data not shown).
Spatial heterogeneity in medulloblastoma. a Clonal admixture plots and phylogenetic trees, reconstructed for two independent regions per tumour. Tumours with region-specific clones are framed in red. Significant differences in overall clonal composition between two regions are marked by black asterisks (p < 0.05 Fisher’s exact test), ns no significant difference; significant differences in the abundance of individual clones between two tumour regions is marked by clone colour-matching asterisks (p < 0.05, Fisher’s exact test). b Table summarises individual CNVs observed at significantly different frequencies between tumour regions (p < 0.05, Fisher’s exact test). Shaded colour blocks in each row represent the clones (coloured as on Figs. 3 and 6a) in which each CNV was detected. c Table summarises the regional distribution of established medulloblastoma biomarkers
We further tested the distribution of individual genetic events across the two regions assessed within each tumour. The majority of CNVs (141/155) did not significantly differ in cellular frequency between the two tumour regions (Supplementary table 5C); 13 CNVs showed significant spatial variation (Fig. 6b). All of these were CNVs of unknown clinical significance. 14/15 tested SNV mutations were detected in both tumour regions (Supplementary Fig. 4). All assessed established medulloblastoma biomarkers were observed in both tumour regions, and at equivalent frequencies (Fig. 6c). PTCH1 mutation and GLI2 amplification were observed below our frequency threshold (i.e. ≥ 3 cells) in Patients 5 and 7, respectively, and thus were excluded from spatial analyses.
Discussion
This study reveals diverse modes of initiation, clonal evolution and intra-tumoural genetic complexity between childhood MBs sampled from its major disease sub-classes and risk-groups, and provides a first understanding of their relationships to disease biology and clinical behaviour.
Our discoveries have enabled definition of three distinct trajectories of disease initiation, development and clonal evolution. First, genetically homogeneous tumours with no clonal substructure, which acquire events at initiation (e.g. CTNNB1 mutation and chromosome 6 loss in MBWNT) followed by expansion of a single clone (i.e. no evolution); such trajectories were only observed in favourable-risk disease groups examined (e.g. MBWNT, Infant MBSHH). Second, MBs with an extensive clonal landscape, characterised by multiple events at initiation, followed by the gradual evolution of subclones with unique genetic composition, resulting in the most diverse clonal substructures observed. These trajectories were only found in very high-risk tumours—their clonal diversity and clinical behaviour being consistent with the involvement of initiating events previously associated with genomic instability (TP53 mutation and/or MYC(N) amplification [31, 35, 37]); and the presence of aggressive large-cell/anaplastic histology [11]. Independent inspection of single-cell WGS data reported for 2 Li–Fraumeni-associated primary MBSHH tumours with TP53 mutations [35] revealed evolutionary trajectories closely similar to the gradual trajectories we observed for our MBSHH-TP53 mutated tumours, providing important validation of our findings for this tumour group. Thirdly, we also characterised an intermediate group of medulloblastomas with a moderate clonal composition, which were sampled from multiple non-VHR risk-groups. These typically acquired multiple events at initiation, followed by a punctuated evolutionary trajectory. From our findings, we proffer a model of genetic initiation, evolution, and clonal substructure for childhood medulloblastoma (Fig. 7). This model defines clearly distinct evolutionary paths along which disease evolution may proceed and which, while not absolute, provide a critical foundation for future investigations. Of note, we included a rare relapsing WNT patient (#1) in our investigation; this patient had clonal substructure equivalent to the other WNT tumours analysed, suggesting other factors play a role in WNT relapse.
Our findings have immediate clinical relevance. The majority of established medulloblastoma biomarkers examined (e.g. CTNNB1 in MBWNT; PTCH1/TP53/MYCN in MBSHH; MYC in MBGroup3; chr7 + /ch11- and i17q in MBGroup3/4) [14, 31, 42] were revealed as early clonal/initiating events in tumour development, consistent with their driver roles in experimental models of spontaneous tumourigenesis, where tested [13, 17, 36, 40]. Their ubiquitous spatial presence highlights their potential importance for therapeutic targeting and informs requirements for tumour sampling in diagnostics and therapeutic stratification. Our data support the sufficiency of a single biopsy for the diagnostic assessment of established biomarkers, challenging previous recommendations that actionable targets found in a single biopsy are seldom clonal across the entire tumour [28]. Moreover, this finding highlights the potential of these key biomarkers of medulloblastoma initiation for use in cancer early detection and prevention strategies. For instance, clonal early events could be further investigated as potential early detection biomarkers in cerebrospinal fluid (CSF) or plasma circulating free DNA (cfDNA) screening. The early occurrence of multiple whole chromosome events suggests polyploidy commonly occurs as an early event in medulloblastoma development. Against this background, we additionally observed a series of previously unappreciated subclonal and/or region-specific events by sc-WGS, which were commonly undetectable in our WES analysis of the corresponding bulk tumour samples. Their biological and clinical significance now require investigation; to determine whether they play any role in further temporal evolution and resistance to therapy, or simply represent uninvolved ‘passenger’ events, will be of particular interest. In contrast to the aforementioned established biomarkers, multi-regional and/or single-cell sampling is essential for their detection.
Cellular transcriptional diversity has been previously observed within individual medulloblastomas [15, 46]; our data indicate this can arise against simpler genetic architectures. For instance, WNT tumours revealed here as genetically homogeneous, are associated with more diverse transcriptionally defined cellular sub-populations: four transcriptional meta-programs with variable underlying gene signatures were previously identified by single-cell transcriptome analyses of five WNT medulloblastomas [15]. Together, these findings suggest intra-tumoural differences in gene expression are influenced by non-genetic factors.
Our focus on tumours representing the major medulloblastoma disease sub-classes and molecular risk-groups is a significant strength of our study. However, we acknowledge, that future studies involving larger patient cohorts will be necessary to fully establish the relationship of clonal states to specific clinical and molecular disease features, and their translational relevance. The number of single cells investigated per tumour represents a potential limitation and it is possible that additional rare clones may be detected in deeper analyses; however, and importantly, cell number analysed was not associated with the number of subclones we identified in this study. Further studies will now be essential to explore these findings in larger clinically annotated cohorts, and with increased cell numbers. Furthermore, single-cell co-studies of genetic status and gene expression, ideally with longitudinal sampling over disease-course including at relapse, and in disease-relevant models, will be critical to discern their inter-relationships and relative contributions to tumour development, clinical behaviour and evolution.
In clinical therapeutics, our findings support a model whereby highest risk tumours may display greatest clonal diversity, which supports the hypothesis that clonal diversity increases potential for adaptation, evolution and escape under therapy, while favourable-risk tumour groups are least complex. Our findings are further supported by observations from a multi-regional bulk tumour sampling study, which demonstrated high-risk paediatric neuroblastomas, Wilms tumours, and rhabdomyosarcoma subtypes exhibit more frequent and extensive clonal branching [1]. Thus, in concert with morphological, molecular and cellular estimates currently used as prognostic markers, measures of clonality have potential as a common marker of prognosis in childhood cancers, which must now be considered and further investigated.
Data availability
Data are available from the European Genome-Phenome Archive (EGA). Accession number is EGAS00001006392.
References
Andersson N, Bakker B, Karlsson J, Valind A, Holmquist Mengelbier L, Spierings DCJ et al (2020) Extensive clonal branching shapes the evolutionary history of high-risk pediatric cancers. Cancer Res 80:1512–1523. https://doi.org/10.1158/0008-5472.Can-19-3468
Baslan T, Kendall J, Rodgers L, Cox H, Riggs M, Stepansky A et al (2012) Genome-wide copy number analysis of single cells. Nat Protoc 7:1024–1041. https://doi.org/10.1038/nprot.2012.039
Cavalli FMG, Remke M, Rampasek L, Peacock J, Shih DJH, Luu B et al (2017) Intertumoral heterogeneity within medulloblastoma subgroups. Cancer Cell 31:737-754.e736. https://doi.org/10.1016/j.ccell.2017.05.005
Chang CH, Housepian EM, Herbert C Jr (1969) An operative staging system and a megavoltage radiotherapeutic technic for cerebellar medulloblastomas. Radiology 93:1351–1359. https://doi.org/10.1148/93.6.1351
Cibulskis K, Lawrence MS, Carter SL, Sivachenko A, Jaffe D, Sougnez C et al (2013) Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat Biotechnol 31:213–219. https://doi.org/10.1038/nbt.2514
Cock PJ, Fields CJ, Goto N, Heuer ML, Rice PM (2010) The Sanger FASTQ file format for sequences with quality scores, and the Solexa/Illumina FASTQ variants. Nucleic Acids Res 38:1767–1771. https://doi.org/10.1093/nar/gkp1137
Cox M. CT (2008) Multidimensional scaling. Handbook of data visualization. Springer, Berlin, Heidelberg
Danilenko M, Clifford SC, Schwalbe EC (2021) Inter and intra-tumoral heterogeneity as a platform for personalized therapies in medulloblastoma. Pharmacol Ther 228:107828. https://doi.org/10.1016/j.pharmthera.2021.107828
Davis A, Gao R, Navin N (2017) Tumor evolution: linear, branching, neutral or punctuated? Biochim Biophys Acta Rev Cancer 1867:151–161. https://doi.org/10.1016/j.bbcan.2017.01.003
DePristo MA, Banks E, Poplin R, Garimella KV, Maguire JR, Hartl C et al (2011) A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat Genet 43:491–498. https://doi.org/10.1038/ng.806
Ellison DW, Kocak M, Dalton J, Megahed H, Lusher ME, Ryan SL et al (2011) Definition of disease-risk stratification groups in childhood medulloblastoma using combined clinical, pathologic, and molecular variables. J Clin Oncol 29:1400–1407. https://doi.org/10.1200/jco.2010.30.2810
Gao R, Davis A, McDonald TO, Sei E, Shi X, Wang Y et al (2016) Punctuated copy number evolution and clonal stasis in triple-negative breast cancer. Nat Genet 48:1119–1130. https://doi.org/10.1038/ng.3641
Gibson P, Tong Y, Robinson G, Thompson MC, Currle DS, Eden C et al (2010) Subtypes of medulloblastoma have distinct developmental origins. Nature 468:1095–1099. https://doi.org/10.1038/nature09587
Goschzik T, Schwalbe EC, Hicks D, Smith A, Zur Muehlen A, Figarella-Branger D et al (2018) Prognostic effect of whole chromosomal aberration signatures in standard-risk, non-WNT/non-SHH medulloblastoma: a retrospective, molecular analysis of the HIT-SIOP PNET 4 trial. Lancet Oncol 19:1602–1616. https://doi.org/10.1016/s1470-2045(18)30532-1
Hovestadt V, Smith KS, Bihannic L, Filbin MG, Shaw ML, Baumgartner A et al (2019) Resolving medulloblastoma cellular architecture by single-cell genomics. Nature 572:74–79. https://doi.org/10.1038/s41586-019-1434-6
Huang L, Ma F, Chapman A, Lu S, Xie XS (2015) Single-cell whole-genome amplification and sequencing: methodology and applications. Annu Rev Genomics Hum Genet 16:79–102. https://doi.org/10.1146/annurev-genom-090413-025352
Kawauchi D, Robinson G, Uziel T, Gibson P, Rehg J, Gao C et al (2012) A mouse model of the most aggressive subgroup of human medulloblastoma. Cancer Cell 21:168–180. https://doi.org/10.1016/j.ccr.2011.12.023
Kuipers J, Tuncel MA, Ferreira P, Jahn K, Beerenwinkel N (2020) Single-cell copy number calling and event history reconstruction. bioRxiv. https://doi.org/10.1101/2020.04.28.065755
Lever J, Zhao EY, Grewal J, Jones MR, Jones SJM (2019) CancerMine: a literature-mined resource for drivers, oncogenes and tumor suppressors in cancer. Nat Methods 16:505–507. https://doi.org/10.1038/s41592-019-0422-y
Li H, Durbin R (2010) Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 26:589–595. https://doi.org/10.1093/bioinformatics/btp698
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N et al (2009) The sequence alignment/map format and SAMtools. Bioinformatics 25:2078–2079. https://doi.org/10.1093/bioinformatics/btp352
Louis DN, Perry A, Reifenberger G, von Deimling A, Figarella-Branger D, Cavenee WK et al (2016) The 2016 World Health Organization classification of tumors of the central nervous system: a summary. Acta Neuropathol 131:803–820. https://doi.org/10.1007/s00401-016-1545-1
Macaulay IC, Haerty W, Kumar P, Li YI, Hu TX, Teng MJ et al (2015) G&T-seq: parallel sequencing of single-cell genomes and transcriptomes. Nat Methods 12:519–522. https://doi.org/10.1038/nmeth.3370
Martin M (2011) Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J 17:10–12. https://doi.org/10.14806/ej.17.1.200
Martínez-Jiménez F, Muiños F, Sentís I, Deu-Pons J, Reyes-Salazar I, Arnedo-Pac C et al (2020) A compendium of mutational cancer driver genes. Nat Rev Cancer 20:555–572. https://doi.org/10.1038/s41568-020-0290-x
McKenna A, Hanna M, Banks E, Sivachenko A, Cibulskis K, Kernytsky A et al (2010) The Genome analysis toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res 20:1297–1303. https://doi.org/10.1101/gr.107524.110
Miller CA, McMichael J, Dang HX, Maher CA, Ding L, Ley TJ et al (2016) Visualizing tumor evolution with the fishplot package for R. BMC Genom 17:880. https://doi.org/10.1186/s12864-016-3195-z
Morrissy AS, Cavalli FMG, Remke M, Ramaswamy V, Shih DJH, Holgado BL et al (2017) Spatial heterogeneity in medulloblastoma. Nat Genet 49:780–788. https://doi.org/10.1038/ng.3838
Morrissy AS, Garzia L, Shih DJ, Zuyderduyn S, Huang X, Skowron P et al (2016) Divergent clonal selection dominates medulloblastoma at recurrence. Nature 529:351–357. https://doi.org/10.1038/nature16478
Northcott PA, Buchhalter I, Morrissy AS, Hovestadt V, Weischenfeldt J, Ehrenberger T et al (2017) The whole-genome landscape of medulloblastoma subtypes. Nature 547:311–317. https://doi.org/10.1038/nature22973
Northcott PA, Robinson GW, Kratz CP, Mabbott DJ, Pomeroy SL, Clifford SC et al (2019) Medulloblastoma. Nat Rev Dis Primers 5:11. https://doi.org/10.1038/s41572-019-0063-6
Ocasio J, Babcock B, Malawsky D, Weir SJ, Loo L, Simon JM et al (2019) scRNA-seq in medulloblastoma shows cellular heterogeneity and lineage expansion support resistance to SHH inhibitor therapy. Nat Commun 10:5829. https://doi.org/10.1038/s41467-019-13657-6
Olshen AB, Venkatraman ES, Lucito R, Wigler M (2004) Circular binary segmentation for the analysis of array-based DNA copy number data. Biostatistics 5:557–572. https://doi.org/10.1093/biostatistics/kxh008
Paradis E, Schliep K (2019) ape 5.0: an environment for modern phylogenetics and evolutionary analyses in R. Bioinformatics 35:526–528. https://doi.org/10.1093/bioinformatics/bty633
Parra RG, Przybilla MJ, Simovic M, Susak H, Ratnaparkhe M, Wong JK et al (2021) Single cell multi-omics analysis of chromothriptic medulloblastoma highlights genomic and transcriptomic consequences of genome instability. bioRxiv. https://doi.org/10.1101/2021.06.25.449944
Pei Y, Moore CE, Wang J, Tewari AK, Eroshkin A, Cho YJ et al (2012) An animal model of MYC-driven medulloblastoma. Cancer Cell 21:155–167. https://doi.org/10.1016/j.ccr.2011.12.021
Rausch T, Jones DT, Zapatka M, Stütz AM, Zichner T, Weischenfeldt J et al (2012) Genome sequencing of pediatric medulloblastoma links catastrophic DNA rearrangements with TP53 mutations. Cell 148:59–71. https://doi.org/10.1016/j.cell.2011.12.013
Richardson S, Hill RM, Kui C, Lindsey JC, Grabovksa Y, Keeling C et al (2021) Emergence and maintenance of actionable genetic drivers at medulloblastoma relapse. Neuro Oncol. https://doi.org/10.1093/neuonc/noab178
Robinson JT, Thorvaldsdóttir H, Winckler W, Guttman M, Lander ES, Getz G et al (2011) Integrative genomics viewer. Nat Biotechnol 29:24–26. https://doi.org/10.1038/nbt.1754
Roussel MF, Stripay JL (2020) Modeling pediatric medulloblastoma. Brain Pathol 30:703–712. https://doi.org/10.1111/bpa.12803
Ryan SL, Schwalbe EC, Cole M, Lu Y, Lusher ME, Megahed H et al (2012) MYC family amplification and clinical risk-factors interact to predict an extremely poor prognosis in childhood medulloblastoma. Acta Neuropathol 123:501–513. https://doi.org/10.1007/s00401-011-0923-y
Schwalbe EC, Lindsey JC, Nakjang S, Crosier S, Smith AJ, Hicks D et al (2017) Novel molecular subgroups for clinical classification and outcome prediction in childhood medulloblastoma: a cohort study. Lancet Oncol 18:958–971. https://doi.org/10.1016/s1470-2045(17)30243-7
Sharma T, Schwalbe EC, Williamson D, Sill M, Hovestadt V, Mynarek M et al (2019) Second-generation molecular subgrouping of medulloblastoma: an international meta-analysis of Group 3 and Group 4 subtypes. Acta Neuropathol 138:309–326. https://doi.org/10.1007/s00401-019-02020-0
Sherry ST, Ward MH, Kholodov M, Baker J, Phan L, Smigielski EM et al (2001) dbSNP: the NCBI database of genetic variation. Nucleic Acids Res 29:308–311. https://doi.org/10.1093/nar/29.1.308
Talevich E, Shain AH, Botton T, Bastian BC (2016) CNVkit: genome-wide copy number detection and visualization from targeted DNA sequencing. PLoS Comput Biol 12:e1004873. https://doi.org/10.1371/journal.pcbi.1004873
Vladoiu MC, El-Hamamy I, Donovan LK, Farooq H, Holgado BL, Sundaravadanam Y et al (2019) Childhood cerebellar tumours mirror conserved fetal transcriptional programs. Nature 572:67–73. https://doi.org/10.1038/s41586-019-1158-7
Wang K, Li M, Hakonarson H (2010) ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res 38:e164. https://doi.org/10.1093/nar/gkq603
Warnes GR, Bolker B, Bonebakker L, Gentleman R, Huber W, Liaw A, et al (2020) Various R Programming Tools for Plotting Data [R package gplots version 3.1.0]. City
Zhang L, He X, Liu X, Zhang F, Huang LF, Potter AS et al (2019) Single-cell transcriptomics in medulloblastoma reveals tumor-initiating progenitors and oncogenic cascades during tumorigenesis and relapse. Cancer Cell 36:302-318.e307. https://doi.org/10.1016/j.ccell.2019.07.009
Acknowledgements
This study was funded by Cancer Research UK (Grants C8464/A13457 and C8464/A23391). We thank members of the Newcastle University Flow Cytometry Core facility, especially Dr Andew Filby and Ms Carly Knill for their advice on FACS experiments.
Author information
Authors and Affiliations
Contributions
MD, DW and SCC designed the study. MD prepared the figures. MD and SCC wrote the manuscript. MD, CK, SC, RH and PZ performed the laboratory experiments. MD and MF optimised single-cell extraction and DNA amplification protocols. MD, RH and JC optimised single-cell sequencing-related protocols. MZ prepared the bioinformatics pipelines. MZ and MD did the bioinformatics analyses. MD and MZ performed statistical analyses. MD performed other data analyses and data interpretation. SL contributed to data analyses. ECS, DH, VR and SCC provided help with data interpretation. AJ, RMH and SB provided patient data and clinical interpretation. All authors are accountable for all aspects of the work and approved the final copy.
Corresponding author
Ethics declarations
Conflict of interest
All authors declare no conflict of interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
401_2022_2464_MOESM3_ESM.xlsx
Supplementary Tables 2 CNV calls from single-cell and bulk sequencing data. Tables summarize: (a) CNVs in the single-cell data; (b) Whole arm/chromosome CNVs identified in bulk WES data (XLSX 29 KB)
401_2022_2464_MOESM4_ESM.xlsx
Supplementary Tables 3 A list of key mutations identified in the bulk WES and selected for validation in single-cell DNA. (a) Table summarises 16 mutations selected for validation (b–e) Summaries of Sanger sequencing results listed separately for (b) WNT, (c) SHH, (d) Grp3 and (e) Grp4 patients (XLSX 29 KB)
401_2022_2464_MOESM5_ESM.xlsx
Supplementary Table 4 A summary of medulloblastoma driver genes covered by focal and sub-chromosomal CNV loci in the single-cell data. Table provides a list of focal and subchromosomal CNVs with identified driver gene loci. The functions of the genes are highlighted. The size, coordinates, and clonal status of CNVs are recorded (XLSX 21 KB)
401_2022_2464_MOESM6_ESM.xlsx
Supplementary Tables 5 Statistical significance scores for the spatially resolved findings. Tables provide statistical significance scores: (a) for the proportions of cells contained in the clones of region 1 (R1) versus the proportions of cells in the clones of region 2 (R2) (b) comparing proportions of cells in certain individual clones in region 1 vs region 2 (c) for the individual CNVs, when comparing proportions of cells containing CNVs in region 1 versus region 2 (XLSX 36 KB)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Danilenko, M., Zaka, M., Keeling, C. et al. Single-cell DNA sequencing identifies risk-associated clonal complexity and evolutionary trajectories in childhood medulloblastoma development. Acta Neuropathol 144, 565–578 (2022). https://doi.org/10.1007/s00401-022-02464-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00401-022-02464-x