Abstract
Medulloblastoma comprises four distinct molecular variants with distinct genetics, transcriptomes, and outcomes. Subgroup affiliation has been previously shown to remain stable at the time of recurrence, which likely reflects their distinct cells of origin. However, a therapeutically relevant question that remains unanswered is subgroup stability in the metastatic compartment. We assembled a cohort of 12-paired primary-metastatic tumors collected in the MAGIC consortium, and established their molecular subgroup affiliation by performing integrative gene expression and DNA methylation analysis. Frozen tissues were collected and profiled using Affymetrix gene expression arrays and Illumina methylation arrays. Class prediction and hierarchical clustering were performed using existing published datasets. Our molecular analysis, using consensus integrative genomic data, establishes the unequivocal maintenance of molecular subgroup affiliation in metastatic medulloblastoma. We further validated these findings by interrogating a non-overlapping cohort of 19 pairs of primary-metastatic tumors from the Burdenko Neurosurgical Institute using an orthogonal technique of immunohistochemical staining. This investigation represents the largest reported primary-metastatic paired cohort profiled to date and provides a unique opportunity to evaluate subgroup-specific molecular aberrations within the metastatic compartment. Our findings further support the hypothesis that medulloblastoma subgroups arise from distinct cells of origin, which are carried forward from ontogeny to oncology.



Similar content being viewed by others
References
Brunet J-P, Tamayo P, Golub TR, Mesirov JP (2004) Metagenes and molecular pattern discovery using matrix factorization. Proc Natl Acad Sci 101:4164–4169. doi:10.1073/pnas.0308531101
Cho YJ, Tsherniak A, Tamayo P, Santagata S, Ligon A, Greulich H, Berhoukim R, Amani V, Goumnerova L, Eberhart CG et al (2011) Integrative genomic analysis of medulloblastoma identifies a molecular subgroup that drives poor clinical outcome. J Clin Oncol 29:1424–1430
Dolecek TA, Propp JM, Stroup NE, Kruchko C (2012) CBTRUS statistical report: primary brain and central nervous system tumors diagnosed in the United States in 2005–2009. Neuro Oncol 14(Suppl 5):v1–v49. doi:10.1093/neuonc/nos218
Gandola L, Massimino M, Cefalo G et al (2009) Hyperfractionated accelerated radiotherapy in the Milan strategy for metastatic medulloblastoma. J Clin Oncol 27:566–571. doi:10.1200/JCO.2008.18.4176
Gibson P, Tong Y, Robinson G et al (2010) Subtypes of medulloblastoma have distinct developmental origins. Nature 468:1095–1099. doi:10.1038/nature09587
Hovestadt V, Jones DTW, Picelli S et al (2014) Decoding the regulatory landscape of medulloblastoma using DNA methylation sequencing. Nature 510:537–541. doi:10.1038/nature13268
Hovestadt V, Remke M, Kool M et al (2013) Robust molecular subgrouping and copy-number profiling of medulloblastoma from small amounts of archival tumour material using high-density DNA methylation arrays. Acta Neuropathol 125:913–916. doi:10.1007/s00401-013-1126-5
Irizarry RA (2003) Summaries of Affymetrix GeneChip probe level data. Nucleic Acids Res 31:15e. doi:10.1093/nar/gng015
Jakacki RI, Burger PC, Zhou T et al (2012) Outcome of children with metastatic medulloblastoma treated with carboplatin during craniospinal radiotherapy: a Children’s Oncology Group Phase I/II study. J Clin Oncol 30:2648–2653. doi:10.1200/JCO.2011.40.2792
Johnson BE, Mazor T, Hong C et al (2014) Mutational analysis reveals the origin and therapy-driven evolution of recurrent glioma. Science 343:189–193. doi:10.1126/science.1239947
Kawauchi D, Robinson G, Uziel T et al (2012) A mouse model of the most aggressive subgroup of human medulloblastoma. Cancer Cell 21:168–180. doi:10.1016/j.ccr.2011.12.023
Kool M, Korshunov A, Remke M et al (2012) Molecular subgroups of medulloblastoma: an international meta-analysis of transcriptome, genetic aberrations, and clinical data of WNT, SHH, Group 3, and Group 4 medulloblastomas. Acta Neuropathol. doi:10.1007/s00401-012-0958-8
Kool M, Koster J, Bunt J et al (2008) Integrated genomics identifies five medulloblastoma subtypes with distinct genetic profiles, pathway signatures and clinicopathological features. PLoS One 3:e3088. doi:10.1371/journal.pone.0003088
Northcott PA, Shih DJH, Remke M et al (2012) Rapid, reliable, and reproducible molecular sub-grouping of clinical medulloblastoma samples. Acta Neuropathol 123:615–626. doi:10.1007/s00401-011-0899-7
Northcott PA, Jones DTW, Kool M et al (2012) Medulloblastomics: the end of the beginning. Nat Rev Cancer 12:818–834. doi:10.1038/nrc3410
Northcott PA, Korshunov A, Witt H et al (2011) Medulloblastoma comprises four distinct molecular variants. J Clin Oncol 29:1408–1414. doi:10.1200/JCO.2009.27.4324
Packer RJ, Gajjar A, Vezina G et al (2006) Phase III study of craniospinal radiation therapy followed by adjuvant chemotherapy for newly diagnosed average-risk medulloblastoma. J Clin Oncol 24:4202–4208. doi:10.1200/JCO.2006.06.4980
Patel AP, Tirosh I, Trombetta JJ et al (2014) Single-cell RNA-seq highlights intratumoral heterogeneity in primary glioblastoma. Science 344:1396–1401. doi:10.1126/science.1254257
Pei Y, Moore CE, Wang J et al (2012) An animal model of MYC-driven medulloblastoma. Cancer Cell 21:155–167. doi:10.1016/j.ccr.2011.12.021
Polkinghorn WR, Tarbell NJ (2007) Medulloblastoma: tumorigenesis, current clinical paradigm, and efforts to improve risk stratification. Nat Clin Pract Oncol 4:295–304. doi:10.1038/ncponc0794
Ramaswamy V, Northcott PA, Taylor MD (2011) FISH and chips: the recipe for improved prognostication and outcomes for children with medulloblastoma. Cancer Genet 204:577–588. doi:10.1016/j.cancergen.2011.11.001
Ramaswamy V, Remke M, Bouffet E et al (2013) Recurrence patterns across medulloblastoma subgroups: an integrated clinical and molecular analysis. Lancet Oncol 14:1200–1207. doi:10.1016/S1470-2045(13)70449-2
Remke M, Hielscher T, Korshunov A et al (2011) FSTL5 is a marker of poor prognosis in non-WNT/Non-SHH medulloblastoma. J Clin Oncol 29:3852–3861. doi:10.1200/JCO.2011.36.2798
Remke M, Hielscher T, Northcott PA et al (2011) Adult medulloblastoma comprises three major molecular variants. J Clin Oncol 29:2717–2723. doi:10.1200/JCO.2011.34.9373
Sottoriva A, Spiteri I, Piccirillo SGM et al (2013) Intratumor heterogeneity in human glioblastoma reflects cancer evolutionary dynamics. Proc Natl Acad Sci 110:4009–4014. doi:10.1073/pnas.1219747110
Tarbell NJ, Friedman H, Polkinghorn WR et al (2013) High-risk medulloblastoma: a pediatric oncology group randomized trial of chemotherapy before or after radiation therapy (POG 9031). J Clin Oncol 31:2936–2941. doi:10.1200/JCO.2012.43.9984
Taylor MD, Northcott PA, Korshunov A et al (2012) Molecular subgroups of medulloblastoma: the current consensus. Acta Neuropathol 123:465–472. doi:10.1007/s00401-011-0922-z
Thompson MC, Fuller C, Hogg TL et al (2006) Genomics identifies medulloblastoma subgroups that are enriched for specific genetic alterations. J Clin Oncol 24:1924–1931. doi:10.1200/JCO.2005.04.4974
Tibshirani R, Hastie T, Narasimhan B, Chu G (2002) Diagnosis of multiple cancer types by shrunken centroids of gene expression. Proc Natl Acad Sci 99:6567–6572. doi:10.1073/pnas.082099299
Wang X, Ramaswamy V, Remke M et al (2013) Intertumoral and intratumoral heterogeneity as a barrier for effective treatment of medulloblastoma. Neurosurgery 60(Suppl 1):57–63. doi:10.1227/01.neu.0000430318.01821.6f
Wu X, Northcott PA, Dubuc A et al (2012) Clonal selection drives genetic divergence of metastatic medulloblastoma. Nature 482:529–533. doi:10.1038/nature10825
Acknowledgments
XW is supported by a CIHR Vanier Canada Graduate Scholarship, McLaughlin Centre for Molecular Medicine MD/PhD Scholarship, and the Ruggles MD/PhD Innovation Award. MDT is supported by a CIHR Clinician Scientist Phase II award, funds from the Garron Family Chair in Childhood Cancer Research at The Hospital for Sick Children and The University of Toronto, and operating funds from the Canadian Institutes of Health Research, the National Institutes of Health (R01CA159859 and R01CA148699) and the Pediatric Brain Tumor Foundation. VR is supported by a CIHR fellowship, an Alberta Innovates-Health Solutions Clinical Fellowship and a Young Investigator Award from Alex’s Lemonade Stand Foundation. BHK is supported by the Gattuso-Slaight Personalized Cancer Medicine Fund at the Princess Margaret Cancer Centre. We wish to acknowledge the Labatt Brain Tumour Research Centre and Tumour Tissue Repository, which are supported by b.r.a.i.n.child and Meagan’s Walk.
Author information
Authors and Affiliations
Corresponding author
Additional information
X. Wang and A. M. Dubuc contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
401_2015_1389_MOESM1_ESM.eps
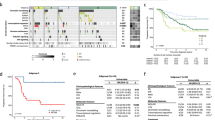
Supplementary material 1 (EPS 1542 kb) Supplementary Fig. 1 PCA of the primary and metastatic medulloblastoma samples described in using the 1,000 most differentially expressed genes (a) and 10,000 most differentially methylated probes (b). Coloured ellipsoids (red = SHH, yellow = Group 3, green = Group 4) represent 1.5 SDs of the data distribution for each subgroup. Individual patients are indicated with a unique colour
401_2015_1389_MOESM2_ESM.docx
Supplementary material 2 (DOCX 25 kb) Supplementary Table 1. Medulloblastoma subgroup predictions and consensus using integrative genomics analysis based on gene expression and 450k DNA methylation
401_2015_1389_MOESM3_ESM.xlsx
Supplementary material 3 (XLSX 60 kb) Supplementary Table 2. Detailed demographic, clinical, and histological characteristics of patients
Rights and permissions
About this article
Cite this article
Wang, X., Dubuc, A.M., Ramaswamy, V. et al. Medulloblastoma subgroups remain stable across primary and metastatic compartments. Acta Neuropathol 129, 449–457 (2015). https://doi.org/10.1007/s00401-015-1389-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00401-015-1389-0